

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139748.4 + phase: 0 /pseudo
(36 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
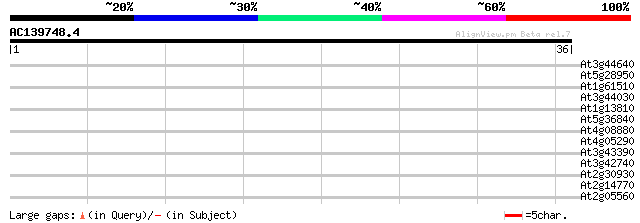
Score E
Sequences producing significant alignments: (bits) Value
At3g44640 putative protein 37 0.002
At5g28950 putative protein 34 0.013
At1g61510 hypothetical protein 29 0.41
At3g44030 putative protein 29 0.54
At1g13810 hypothetical protein 25 5.9
At5g36840 putative protein 25 7.7
At4g08880 hypothetical protein 25 7.7
At4g05290 putative protein 25 7.7
At3g43390 putative protein 25 7.7
At3g42740 putative protein 25 7.7
At2g30930 unknown protein 25 7.7
At2g14770 hypothetical protein 25 7.7
At2g05560 hypothetical protein 25 7.7
>At3g44640 putative protein
Length = 377
Score = 37.4 bits (85), Expect = 0.002
Identities = 13/26 (50%), Positives = 21/26 (80%)
Query: 1 MELPNRIKNNPKYYPWFKNCIGAIDG 26
+ +P +I+ + +YP+FK+CIGAIDG
Sbjct: 127 VRVPAKIRESTSFYPYFKDCIGAIDG 152
>At5g28950 putative protein
Length = 148
Score = 34.3 bits (77), Expect = 0.013
Identities = 11/23 (47%), Positives = 19/23 (81%)
Query: 3 LPNRIKNNPKYYPWFKNCIGAID 25
+P +I+ + + YP+FK+C+GAID
Sbjct: 8 VPRKIRESTRLYPYFKDCVGAID 30
>At1g61510 hypothetical protein
Length = 608
Score = 29.3 bits (64), Expect = 0.41
Identities = 10/18 (55%), Positives = 15/18 (82%)
Query: 9 NNPKYYPWFKNCIGAIDG 26
N+ +Y P+F +CIGA+DG
Sbjct: 370 NDKRYMPYFIDCIGALDG 387
>At3g44030 putative protein
Length = 775
Score = 28.9 bits (63), Expect = 0.54
Identities = 9/24 (37%), Positives = 18/24 (74%)
Query: 3 LPNRIKNNPKYYPWFKNCIGAIDG 26
+P R++ + +Y+P+F +GA+DG
Sbjct: 619 IPERLQVDRRYWPYFSGFVGAMDG 642
>At1g13810 hypothetical protein
Length = 303
Score = 25.4 bits (54), Expect = 5.9
Identities = 10/24 (41%), Positives = 15/24 (61%), Gaps = 3/24 (12%)
Query: 8 KNNPKYYPWFK---NCIGAIDGLL 28
++N K YPW K NC+ + GL+
Sbjct: 177 RDNSKVYPWKKVPYNCVPQLQGLM 200
>At5g36840 putative protein
Length = 434
Score = 25.0 bits (53), Expect = 7.7
Identities = 9/20 (45%), Positives = 13/20 (65%)
Query: 4 PNRIKNNPKYYPWFKNCIGA 23
PN I + +Y P+FK C+ A
Sbjct: 68 PNDINSEGRYRPFFKRCLQA 87
>At4g08880 hypothetical protein
Length = 1175
Score = 25.0 bits (53), Expect = 7.7
Identities = 9/20 (45%), Positives = 13/20 (65%)
Query: 4 PNRIKNNPKYYPWFKNCIGA 23
PN I + +Y P+FK C+ A
Sbjct: 1010 PNDINSEGRYRPFFKRCLQA 1029
>At4g05290 putative protein
Length = 213
Score = 25.0 bits (53), Expect = 7.7
Identities = 9/20 (45%), Positives = 13/20 (65%)
Query: 4 PNRIKNNPKYYPWFKNCIGA 23
PN I + +Y P+FK C+ A
Sbjct: 51 PNDINSEGRYRPFFKRCLQA 70
>At3g43390 putative protein
Length = 1113
Score = 25.0 bits (53), Expect = 7.7
Identities = 9/20 (45%), Positives = 13/20 (65%)
Query: 4 PNRIKNNPKYYPWFKNCIGA 23
PN I + +Y P+FK C+ A
Sbjct: 951 PNDINSEGRYRPFFKRCLQA 970
>At3g42740 putative protein
Length = 230
Score = 25.0 bits (53), Expect = 7.7
Identities = 9/20 (45%), Positives = 13/20 (65%)
Query: 4 PNRIKNNPKYYPWFKNCIGA 23
PN I + +Y P+FK C+ A
Sbjct: 68 PNDINSEGRYRPFFKRCLEA 87
>At2g30930 unknown protein
Length = 164
Score = 25.0 bits (53), Expect = 7.7
Identities = 11/29 (37%), Positives = 15/29 (50%), Gaps = 5/29 (17%)
Query: 2 ELPNRIKNNP-----KYYPWFKNCIGAID 25
+LP N P + PW+KNC G +D
Sbjct: 130 DLPVSTDNQPLLASGEKTPWWKNCCGVLD 158
>At2g14770 hypothetical protein
Length = 1756
Score = 25.0 bits (53), Expect = 7.7
Identities = 9/20 (45%), Positives = 13/20 (65%)
Query: 4 PNRIKNNPKYYPWFKNCIGA 23
PN I + +Y P+FK C+ A
Sbjct: 1412 PNDINSEGRYRPFFKRCLQA 1431
>At2g05560 hypothetical protein
Length = 1471
Score = 25.0 bits (53), Expect = 7.7
Identities = 9/20 (45%), Positives = 13/20 (65%)
Query: 4 PNRIKNNPKYYPWFKNCIGA 23
PN I + +Y P+FK C+ A
Sbjct: 1281 PNDINSEGRYRPFFKRCLQA 1300
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.146 0.491
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 945,784
Number of Sequences: 26719
Number of extensions: 20223
Number of successful extensions: 59
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 48
Number of HSP's gapped (non-prelim): 13
length of query: 36
length of database: 11,318,596
effective HSP length: 12
effective length of query: 24
effective length of database: 10,997,968
effective search space: 263951232
effective search space used: 263951232
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 53 (25.0 bits)
Medicago: description of AC139748.4


