

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
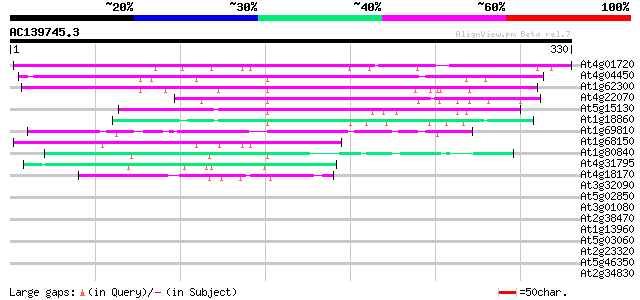
Query= AC139745.3 - phase: 0 /pseudo
(330 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At4g01720 putative DNA-binding protein 206 1e-53
At4g04450 putative DNA-binding protein 149 2e-36
At1g62300 transcription factor WRKY6 145 2e-35
At4g22070 WRKY transcription factor 31 (WRKY31) 133 1e-31
At5g15130 unknown protein 88 7e-18
At1g18860 WRKY transcription factor 61 (WRKY61) 85 6e-17
At1g69810 unknown protein 74 1e-13
At1g68150 putative DNA binding protein 73 2e-13
At1g80840 transcription factor like protein 55 4e-08
At4g31795 WRKY like transcription factor 47 2e-05
At4g18170 DNA binding like protein 41 8e-04
At3g32090 unknown protein 40 0.001
At5g02850 unknown protein 40 0.002
At3g01080 WRKY transcription factor 58 (WRKY58) 38 0.007
At2g38470 putative WRKY-type DNA binding protein 38 0.007
At1g13960 WRKY DNA-binding protein 4 (WRKY4) 38 0.007
At5g03060 putative protein 38 0.009
At2g23320 putative WRKY-type DNA-binding protein 38 0.009
At5g46350 putative protein 37 0.011
At2g34830 WRKY-type DNA binding protein-like 37 0.011
>At4g01720 putative DNA-binding protein
Length = 489
Score = 206 bits (524), Expect = 1e-53
Identities = 163/415 (39%), Positives = 210/415 (50%), Gaps = 95/415 (22%)
Query: 3 SSAAVSKEENLENSETEMSILESELRRVQEENHKLRIMLEQITKSYSQLQAQLFITLQKQ 62
S S + + ++T++S L+ EL R+ EENHKL+ +L+++++SY+ LQ ++ + Q Q
Sbjct: 82 SCHGTSSNDGDDKTKTQISRLKLELERLHEENHKLKHLLDEVSESYNDLQRRVLLARQTQ 141
Query: 63 KPN-HGQNMEENHGMVSEQIFLN------NNNASVSDGKQACPHD------------HPA 103
H + E+ S Q N N+ + K+ P D P
Sbjct: 142 VEGLHHKQHEDVPQAGSSQALENRRPKDMNHETPATTLKRRSPDDVDGRDMHRGSPKTPR 201
Query: 104 EDSSHSSKLEEPTQ--DLIPFKKARVSIRARSEA--------------------PLPTI- 140
D + S+ EE D +P++KARVS+RARS+A P P
Sbjct: 202 IDQNKSTNHEEQQNPHDQLPYRKARVSVRARSDATTVNDGCQWRKYGQKMAKGNPCPRAY 261
Query: 141 ----VALWL*DVQFANRCAEDKTILITTYEGNHNHPLPPAATAIAHTTSAAAAMLLSSST 196
+A+ + RCAED TIL TTYEGNHNHPLPP+ATA+A TTSAAAAMLLS S+
Sbjct: 262 YRCTMAVGCPVRKQVQRCAEDTTILTTTYEGNHNHPLPPSATAMAATTSAAAAMLLSGSS 321
Query: 197 SS----TLRKESATGYLS--NSFPYATMATSTLSASQPFPTITLDFTQ------------ 238
SS TL SAT S ++FPY T +TLSAS PFPTITLD T
Sbjct: 322 SSNLHQTLSSPSATSSSSFYHNFPY-TSTIATLSASAPFPTITLDLTNPPRPLQPPPQFL 380
Query: 239 ------------NHNLSMHHNRVPLPLFFSHKLPPLLQLGQPPPSSMVESVSAAISSDPN 286
N SM++N L L P L Q PP MV+SV AAI+ DPN
Sbjct: 381 SQYGPAAFLPNANQIRSMNNNNQQL-------LIPNLFGPQAPPREMVDSVRAAIAMDPN 433
Query: 287 FTTALAAAISSIIGPQRSGDGNN-----NLAGVVPG------SPQLPQSCTTFST 330
FT ALAAAIS+IIG + + NN N G SPQLPQSCTTFST
Sbjct: 434 FTAALAAAISNIIGGGNNDNNNNTDINDNKVDAKSGGSSNGDSPQLPQSCTTFST 488
>At4g04450 putative DNA-binding protein
Length = 528
Score = 149 bits (376), Expect = 2e-36
Identities = 135/424 (31%), Positives = 188/424 (43%), Gaps = 120/424 (28%)
Query: 6 AVSKEENLENSETEMSILESELRRVQEENHKLRIMLEQITKSYSQLQAQLFITLQKQKPN 65
+V EE + ++ E + L EL++ E+N +L+ ML Q T +++ LQ QL +++Q+ +
Sbjct: 101 SVDMEE--KRTKCENAQLREELKKASEDNQRLKQMLSQTTNNFNSLQMQLVAVMRQQEDH 158
Query: 66 HGQNMEENHG----------MVSEQIF------------------------LNNNNASVS 91
H EN+ MV Q L ++S
Sbjct: 159 HHLATTENNDNVKNRHEVPEMVPRQFIDLGPHSDEVSSEERTTVRSGSPPSLLEKSSSRQ 218
Query: 92 DGKQACPHDHPAEDSSH--------------------------SSKLEEPTQDLIPFKKA 125
+GK+ + E S+ SSK+ E +KA
Sbjct: 219 NGKRVLVREESPETESNGWRNPNKVPKHHASSSICGGNGSENASSKVIEQAAAEATMRKA 278
Query: 126 RVSIRARSEAPLPTIVALWL*DVQF-------------------------ANRCAEDKTI 160
RVS+RARSEAP+ + W Q RCAED+TI
Sbjct: 279 RVSVRARSEAPMLSDGCQWRKYGQKMAKGNPCPRAYYRCTMAVGCPVRKQVQRCAEDRTI 338
Query: 161 LITTYEGNHNHPLPPAATAIAHTTSAAAAMLLSSSTSSTL-RKESATGYLSNSFPYATMA 219
LITTYEGNHNHPLPPAA +A TT+AAA+MLLS ST S + T L+ + + +
Sbjct: 339 LITTYEGNHNHPLPPAAMNMASTTTAAASMLLSGSTMSNQDGLMNPTNLLARTILPCSSS 398
Query: 220 TSTLSASQPFPTITLDFTQNHNLSMHHNRVPLPLFFSHKLPPLLQLGQ------------ 267
+T+SAS PFPTITLD T++ N +N PL + L++L Q
Sbjct: 399 MATISASAPFPTITLDLTESPN---GNNPTNNPLMQFSQRSGLVELNQSVLPHMMGQALY 455
Query: 268 -------------PPPSSMVESVS---AAISSDPNFTTALAAAISSII-GPQRSGDGNNN 310
P + ESVS AAI+S+PNF ALAAAI+SII G +GNNN
Sbjct: 456 YNQQSKFSGLHMPSQPLNAGESVSAATAAIASNPNFAAALAAAITSIINGSNNQQNGNNN 515
Query: 311 LAGV 314
+ V
Sbjct: 516 NSNV 519
>At1g62300 transcription factor WRKY6
Length = 553
Score = 145 bits (367), Expect = 2e-35
Identities = 127/405 (31%), Positives = 178/405 (43%), Gaps = 102/405 (25%)
Query: 8 SKEENLENSETEMSILESELRRVQEENHKLRIMLEQITKSYSQLQAQLFITLQKQKPNHG 67
S E + ++ E+ L+ EL+++ +N KLR +L Q++ SY+ LQ L +Q+Q+ +
Sbjct: 145 SSEMEDKRAKNELVKLQDELKKMTMDNQKLRELLTQVSNSYTSLQMHLVSLMQQQQQQNN 204
Query: 68 QNMEENHG-----------------MVSEQIFLNNNNASV-------------SDGKQAC 97
+ +E V E ++N+++ S+GK+
Sbjct: 205 KVIEAAEKPEETIVPRQFIDLGPTRAVGEAEDVSNSSSEDRTRSGGSSAAERRSNGKRLG 264
Query: 98 PHDHPAEDSSHSSKLEEPTQDLIP------FKKARVSIRARSEAPLPTIVALWL*DVQF- 150
+ P +S+ K+ T +KARVS+RARSEAP+ + W Q
Sbjct: 265 REESPETESNKIQKVNSTTPTTFDQTAEATMRKARVSVRARSEAPMISDGCQWRKYGQKM 324
Query: 151 ------------------------ANRCAEDKTILITTYEGNHNHPLPPAATAIAHTTSA 186
RCAED++ILITTYEGNHNHPLPPAA A+A TT+A
Sbjct: 325 AKGNPCPRAYYRCTMATGCPVRKQVQRCAEDRSILITTYEGNHNHPLPPAAVAMASTTTA 384
Query: 187 AAAMLLSSSTSSTLRKESATGYLSNSFPYATMATSTLSASQPFPTITLDFT--------- 237
AA MLLS S SS + T L+ + + + +T+SAS PFPT+TLD T
Sbjct: 385 AANMLLSGSMSSHDGMMNPTNLLARAVLPCSTSMATISASAPFPTVTLDLTHSPPPPNGS 444
Query: 238 ---------QNHNLSMHH---------NRVP--LP------LFFSHKLPPLLQLGQPP-- 269
NHN M N P LP L+ K L G P
Sbjct: 445 NPSSSAATNNNHNSLMQRPQQQQQQMTNLPPGMLPHVIGQALYNQSKFSGLQFSGGSPST 504
Query: 270 ----PSSMVESVSAAISSDPNFTTALAAAISSIIGPQRSGDGNNN 310
S V A+++DPNFT ALAA ISS+I DG N
Sbjct: 505 AAFSQSHAVADTITALTADPNFTAALAAVISSMINGTNHHDGEGN 549
>At4g22070 WRKY transcription factor 31 (WRKY31)
Length = 538
Score = 133 bits (335), Expect = 1e-31
Identities = 109/279 (39%), Positives = 144/279 (51%), Gaps = 65/279 (23%)
Query: 98 PHDHPAEDSSHSSK---LEEPTQDLIPFKKARVSIRARSEAPLPTIVALWL*DVQF---- 150
P +P+ +S+ ++ + + + +KARVS+RARSEA + + W Q
Sbjct: 253 PKHNPSSSNSNGNRNGNVIDQSAAEATMRKARVSVRARSEAAMISDGCQWRKYGQKMAKG 312
Query: 151 ---------------------ANRCAEDKTILITTYEGNHNHPLPPAATAIAHTTSAAAA 189
RCAED++ILITTYEGNHNHPLPPAATA+A TT+AAA+
Sbjct: 313 NPCPRAYYRCTMAGGCPVRKQVQRCAEDRSILITTYEGNHNHPLPPAATAMASTTTAAAS 372
Query: 190 MLLSSSTSSTLRKESATGYLSNSFPYATMATSTLSASQPFPTITLDFTQN---HNLSMHH 246
MLLS S SS + T L+ + + + +T+SAS PFPTITLD T + +N +M
Sbjct: 373 MLLSGSMSSQDGLMNPTNLLARAILPCSSSMATISASAPFPTITLDLTNSPNGNNPNMTT 432
Query: 247 NRVPL------PLFFSHKLPPL----------------LQLGQPP-----PSSMVESVSA 279
N PL P F LP + LQL P SS+ ESVSA
Sbjct: 433 NN-PLMQFAQRPGFNPAVLPQVVGQAMYNNQQQSKFSGLQLPAQPLQIAATSSVAESVSA 491
Query: 280 A---ISSDPNFTTALAAAISSII---GPQRSGDGNNNLA 312
A I+SDPNF ALAAAI+SI+ Q + NNN+A
Sbjct: 492 ASAAIASDPNFAAALAAAITSIMNGSSHQNNNTNNNNVA 530
Score = 40.0 bits (92), Expect = 0.002
Identities = 22/71 (30%), Positives = 42/71 (58%), Gaps = 8/71 (11%)
Query: 9 KEENLENSETEMSILESELRRVQEENHKLRIMLEQITKSYSQLQAQLFITLQKQKPNHGQ 68
K +EN++ L+ EL++++ EN +LR ML Q T +++ LQ QL +++Q+ +
Sbjct: 105 KRAKIENAQ-----LQEELKKMKIENQRLRDMLSQATTNFNALQMQLVAVMRQQEQ---R 156
Query: 69 NMEENHGMVSE 79
N ++H + E
Sbjct: 157 NSSQDHLLAQE 167
>At5g15130 unknown protein
Length = 548
Score = 87.8 bits (216), Expect = 7e-18
Identities = 84/312 (26%), Positives = 131/312 (41%), Gaps = 78/312 (25%)
Query: 65 NHGQNMEENHGMVSEQIFLNNNNASVSDGKQACPHDHPAEDSSHSSKLEEPTQDLIPFKK 124
N+G E G+ E + + + + GK PA S + E Q+ + K+
Sbjct: 155 NNGNGGEPKEGLSMENRANSGSEEAWAPGKVTGKRSSPAPASGGDADGEAGQQNHV--KR 212
Query: 125 ARVSIRARSEAPLPTIVALWL*DVQF-------------------------ANRCAEDKT 159
ARV +RAR + P W Q RCA+D +
Sbjct: 213 ARVCVRARCDTPTMNDGCQWRKYGQKIAKGNPCPRAYYRCTVAPGCPVRKQVQRCADDMS 272
Query: 160 ILITTYEGNHNHPLPPAATAIAHTTSAAAAMLLSSSTSSTLRKESATGYLSNS------- 212
ILITTYEG H+H LP +AT +A TTSAAA+MLLS S+SS + NS
Sbjct: 273 ILITTYEGTHSHSLPLSATTMASTTSAAASMLLSGSSSSPAAEMIGNNLYDNSRFNNNNK 332
Query: 213 -----------FPYATM-----------ATSTLSAS--------QPFPTITLDFTQNHNL 242
P T+ ++S LS + Q FP+ +L+F+ +
Sbjct: 333 SFYSPTLHSPLHPTVTLDLTAPQHSSSSSSSLLSLNFNKFSNSFQRFPSTSLNFSSTSST 392
Query: 243 SMHHNRVPLPLFFSHKLPPL-------LQLGQ-------PPPSSMVESVSAAISSDPNFT 288
S + + + LP + + +Q G S+ E+++ A++SDP+F
Sbjct: 393 SSNPSTLNLPAIWGNGYSSYTPYPYNNVQFGTSNLGKTVQNSQSLTETLTKALTSDPSFH 452
Query: 289 TALAAAISSIIG 300
+ +AAAIS+++G
Sbjct: 453 SVIAAAISTMVG 464
Score = 39.3 bits (90), Expect = 0.003
Identities = 21/73 (28%), Positives = 38/73 (51%), Gaps = 3/73 (4%)
Query: 4 SAAVSKEENLENSETEMSILESELRRVQEENHKLRIMLEQITKSYSQLQAQLFITLQKQK 63
S + + + E+ ++E+ V+EEN KL+ MLE+I Y L+ + F +Q++
Sbjct: 20 SVQIHEANKGDGDHQELESAKAEMSEVKEENEKLKGMLERIESDYKSLKLRFFDIIQQEP 79
Query: 64 PN---HGQNMEEN 73
N QNM ++
Sbjct: 80 SNTATKNQNMVDH 92
>At1g18860 WRKY transcription factor 61 (WRKY61)
Length = 480
Score = 84.7 bits (208), Expect = 6e-17
Identities = 94/359 (26%), Positives = 139/359 (38%), Gaps = 118/359 (32%)
Query: 61 KQKPNHGQNMEENHGMVSEQIFLNNNNASVSDGKQACPHDHPAEDSSHSSKLEEPTQDLI 120
K N + +E +H + + ++NNN S +D H + E Q+L+
Sbjct: 119 KALSNPNEKLEIDHNQETMSLEISNNNKIRSQNSFGFKND----GDDHEDEDEILPQNLV 174
Query: 121 PFKKARVSIRARSEAPLPTIVALWL*DVQF-------------------------ANRCA 155
KK RVS+R+R E P W Q RC+
Sbjct: 175 --KKTRVSVRSRCETPTMNDGCQWRKYGQKIAKGNPCPRAYYRCTIAASCPVRKQVQRCS 232
Query: 156 EDKTILITTYEGNHNHPLPPAATAIAHTTSAAAAMLLSSSTSST---------------- 199
ED +ILI+TYEG HNHPLP +ATA+A TSAAA+MLLS ++SS+
Sbjct: 233 EDMSILISTYEGTHNHPLPMSATAMASATSAAASMLLSGASSSSSAAADLHGLNFSLSGN 292
Query: 200 ----------LRKESATGY----------LSNSFPYATMAT------STLSASQPFPTIT 233
L+ S++G+ S+ P+ +M S +S S +P+
Sbjct: 293 NITPKPKTHFLQSPSSSGHPTVTLDLTTSSSSQQPFLSMLNRFSSPPSNVSRSNSYPSTN 352
Query: 234 LDFTQNHNLSMH--------------------HNRVPLPLFF---------------SHK 258
L+F+ N N M+ H + P S
Sbjct: 353 LNFSNNTNTLMNWGGGGNPSDQYRAAYGNINTHQQSPYHKIIQTRTAGSSFDPFGRSSSS 412
Query: 259 LPPLLQL---------GQPPPSSMVESVSAAISSDPNFTTALAAAISSIIGPQRSGDGN 308
P + L PS E++ A I++DP+F +ALA A+SSI+G D N
Sbjct: 413 HSPQINLDHIGIKNIISHQVPSLPAETIKA-ITTDPSFQSALATALSSIMGGDLKIDHN 470
>At1g69810 unknown protein
Length = 387
Score = 73.6 bits (179), Expect = 1e-13
Identities = 79/270 (29%), Positives = 121/270 (44%), Gaps = 40/270 (14%)
Query: 11 ENLENSETEMSILESELRRVQEENHKLRIMLEQITKSYSQLQAQLFITLQK-----QKPN 65
+N+E S+ + S L + E KL + Q+ + ++ + ++ +K + N
Sbjct: 117 KNVEESKDKRSALGFGFQIQSYEASKLDDLCRQVKLANAENKC---VSSRKDVKSVRNEN 173
Query: 66 HGQNMEENHGMVSEQIFLNNNNASVSDGKQACPHDHPAEDSSHSSKLEEPTQDLIPFKKA 125
H +EE+ EQ L V K +C D D K + T P +A
Sbjct: 174 HQDVLEEH-----EQTGLKKTRVCV---KASC-EDPSINDGCQWRKYGQKTAKTNPLPRA 224
Query: 126 RVSIRARSEAPLPTIVALWL*DVQFANRCAEDKT-ILITTYEGNHNHPLPPAATAIAHTT 184
S P+ V RC E++T +TTYEGNH+HPLP A+ +A T
Sbjct: 225 YYRCSMSSNCPVRKQV----------QRCGEEETSAFMTTYEGNHDHPLPMEASHMAAGT 274
Query: 185 SAAAAMLLSSSTSSTLRKESATGYLSNSFPYATMATSTLSASQPFPTITLDFTQNHNLSM 244
SAAA++L S S+SS+ S + LS FP+ + ST ++ PT+TLD T+ +
Sbjct: 275 SAAASLLQSGSSSSS---SSTSASLSYFFPFHHFSISTTNS---HPTVTLDLTRPN---- 324
Query: 245 HHNRVP--LPLFFSHKLPPLLQLGQPPPSS 272
+ N++P PL S PPPSS
Sbjct: 325 YPNQLPDDYPLSSSSFSLNFSSPDPPPPSS 354
Score = 38.1 bits (87), Expect = 0.007
Identities = 19/60 (31%), Positives = 37/60 (61%), Gaps = 2/60 (3%)
Query: 15 NSETEMSILESELRRVQEENHKLRIMLEQITKSYSQLQAQLFITLQKQKPNHGQNMEENH 74
+ E E+ ++++ +V+EEN KL+++L I +Y+ LQ Q+ L +Q+ +ME +H
Sbjct: 21 DKEEELDATKAKVEKVREENEKLKLLLSTILNNYNSLQMQVSKVLGQQQ--GASSMELDH 78
>At1g68150 putative DNA binding protein
Length = 374
Score = 72.8 bits (177), Expect = 2e-13
Identities = 71/240 (29%), Positives = 101/240 (41%), Gaps = 47/240 (19%)
Query: 3 SSAAVSKEENLENSETEMSILESELRRVQEENHKLRIMLEQITKSYSQLQA--------- 53
SS+ + EN E+ L+ ++ V+EEN +LR ++EQ + Y L+
Sbjct: 77 SSSLGLRTREEENEREELLQLQIQMESVKEENTRLRKLVEQTLEDYRHLEMKFPVIDKTK 136
Query: 54 ----QLFITLQKQKPNHGQNMEENHGMVSEQIFLNNNNASVS-DGKQACPHDHPAEDSSH 108
++F+ +Q ++ + G S+S + KQ A S H
Sbjct: 137 KMDLEMFLGVQGKRCVDITSKARKRGAERSPSMEREIGLSLSLEKKQKQEESKEAVQSHH 196
Query: 109 ----SSKLEEPTQDLIPF----KKARVSIRARSEA--------------------PLPTI 140
SS L+ +I +KARVS+RAR E P P
Sbjct: 197 QRYNSSSLDMNMPRIISSSQGNRKARVSVRARCETATMNDGCQWRKYGQKTAKGNPCPRA 256
Query: 141 -----VALWL*DVQFANRCAEDKTILITTYEGNHNHPLPPAATAIAHTTSAAAAMLLSSS 195
VA + RC ED +ILITTYEG HNHPLP ATA+A T S + +LL SS
Sbjct: 257 YYRCTVAPGCPVRKQVQRCLEDMSILITTYEGTHNHPLPVGATAMASTASTSPFLLLDSS 316
>At1g80840 transcription factor like protein
Length = 302
Score = 55.5 bits (132), Expect = 4e-08
Identities = 67/298 (22%), Positives = 112/298 (37%), Gaps = 59/298 (19%)
Query: 21 SILESELRRVQEENHKLRIMLEQITKSYSQLQAQLFITLQKQKPNHGQNM---EENHGMV 77
S L EL RV EN KL ML + +Y+ L+ QL + K + ++
Sbjct: 31 SALVEELNRVSAENKKLSEMLTLMCDNYNVLRKQLMEYVNKSNITERDQISPPKKRKSPA 90
Query: 78 SEQIFLNNNNASVSDGKQACPHDHPAEDSSHSSKLEEPT----------------QDLIP 121
E F VS+ ++ + + ++E +D
Sbjct: 91 REDAFSCAVIGGVSESSSTDQDEYLCKKQREETVVKEKVSRVYYKTEASDTTLVVKDGYQ 150
Query: 122 FKKARVSIRARSEAPLPTIVALWL*DVQF---ANRCAEDKTILITTYEGNHNHPLPPAAT 178
++K + + +P R ED+++L+ TYEG HNHP+P
Sbjct: 151 WRKYGQKVTRDNPSPRAYFKCACAPSCSVKKKVQRSVEDQSVLVATYEGEHNHPMPSQI- 209
Query: 179 AIAHTTSAAAAMLLSSSTSSTLRKESATGYLSNSFPYATMATSTLSASQPFPTITLDFTQ 238
S + R S G S S P A S+L+ P T+D +
Sbjct: 210 ---------------DSNNGLNRHISHGG--SASTPVAANRRSSLTV----PVTTVDMIE 248
Query: 239 NHNLSMHHNRVPLPLFFSHKLPPLLQLGQPPPSSMVESVSAAISSDPNFTTALAAAIS 296
+ ++ +R+ P KL +VE ++++++ DPNFT ALAAA++
Sbjct: 249 SKKVTSPTSRIDFPQV--QKL-------------LVEQMASSLTKDPNFTAALAAAVT 291
>At4g31795 WRKY like transcription factor
Length = 310
Score = 46.6 bits (109), Expect = 2e-05
Identities = 53/220 (24%), Positives = 86/220 (39%), Gaps = 37/220 (16%)
Query: 9 KEENLENSETEMSILESELRRVQEENHKLRIMLEQITKSYSQLQAQLFITLQKQKPNHGQ 68
K + LE E+ S L EL RV EN KL ML ++ +SY++L L +Q P Q
Sbjct: 36 KRKWLEQDESA-SELREELNRVNSENKKLTEMLARVCESYNELHNHLEKLQSRQSPEIEQ 94
Query: 69 -------NMEENHGMVSEQIFLNNNNASVSDGKQACPHDH--------------PAEDSS 107
++ + I L++ S + H H P DS
Sbjct: 95 TDIPIKKRKQDPDEFLGFPIGLSSGKTENSSSNEDHHHHHQQHEQKNQLLSCKRPVTDSF 154
Query: 108 HSSKLEE---PTQ---------DLIPFKKARVSIRARSEAPLPTIVALWL*DV---QFAN 152
+ +K+ PT+ D ++K + + +P + +
Sbjct: 155 NKAKVSTVYVPTETSDTSLTVKDGFQWRKYGQKVTRDNPSPRAYFRCSFAPSCPVKKKVQ 214
Query: 153 RCAEDKTILITTYEGNHNHPLPPAATAIAHTTSAAAAMLL 192
R AED ++L+ TYEG HNH P A+ A + ++ + L
Sbjct: 215 RSAEDPSLLVATYEGTHNHLGPNASEGDATSQGGSSTVTL 254
>At4g18170 DNA binding like protein
Length = 318
Score = 41.2 bits (95), Expect = 8e-04
Identities = 55/193 (28%), Positives = 79/193 (40%), Gaps = 55/193 (28%)
Query: 41 LEQITKSYSQLQAQLFITLQKQKPNHG-QNMEENHGMVSE-QIFLNNNNASVSDGKQACP 98
L Q T S +++F + Q+PN N N G +E + ++ +N+S S+
Sbjct: 63 LLQKTFGLSPSSSEVFNSSIDQEPNRDVTNDVINGGACNETETRVSPSNSSSSEA----- 117
Query: 99 HDHPAEDSSHSSKLEEPT--QDLIPFK-------------KARVSIRARSE--------- 134
DHP EDS S + E +D I K + RVS +SE
Sbjct: 118 -DHPGEDSGKSRRKRELVGEEDQISKKVGKTKKTEVKKQREPRVSFMTKSEVDHLEDGYR 176
Query: 135 -----------APLPTIVALWL*DVQFAN------RCAEDKTILITTYEGNHNHPLPPAA 177
+P P + + Q N R +D T++ITTYEG HNHP+P
Sbjct: 177 WRKYGQKAVKNSPYPR--SYYRCTTQKCNVKKRVERSFQDPTVVITTYEGQHNHPIPTNL 234
Query: 178 TAIAHTTSAAAAM 190
+SAAAAM
Sbjct: 235 RG----SSAAAAM 243
>At3g32090 unknown protein
Length = 182
Score = 40.4 bits (93), Expect = 0.001
Identities = 29/99 (29%), Positives = 44/99 (44%), Gaps = 9/99 (9%)
Query: 109 SSKLEEPTQDLIPFKKARV-SIRARSEAPLPTIVALWL*DVQFANRCAEDKTILITTYEG 167
S K + T LIP V ++R + LW R ED+++L+ TYEG
Sbjct: 53 SRKRKTLTTSLIPLLVITVCALRCIEVVEKVKVHPLWQ-ATPMVQRSVEDQSVLVATYEG 111
Query: 168 NHNHPLPPAATA-------IAHTTSAAAAMLLSSSTSST 199
HNHP+P + I+H SA+ + + +S T
Sbjct: 112 EHNHPMPSQMDSNNGLNRYISHGGSASTPVAANKRSSLT 150
>At5g02850 unknown protein
Length = 426
Score = 40.0 bits (92), Expect = 0.002
Identities = 34/116 (29%), Positives = 47/116 (40%), Gaps = 6/116 (5%)
Query: 169 HNHPLPPAATAIAHTTSAAAAMLLSSSTSSTLRKESATGYLSNS-----FPYATMATSTL 223
H HP +++ H LL SST + SA+ +S+S P A + L
Sbjct: 31 HGHPTSSSSSQSQHQQIQQQPNLLPSSTVAAASSASASAAVSSSALLSLLPPLPRAQALL 90
Query: 224 SASQPFPTITLDFTQNHNLSMHHNRVPLPLFFS-HKLPPLLQLGQPPPSSMVESVS 278
+ D + N + + R LP F S H LPP L P PSS E +S
Sbjct: 91 QQMAVLTSKLFDVSPNRAIWLSAFRGSLPSFLSSHSLPPPPPLENPSPSSTKEILS 146
>At3g01080 WRKY transcription factor 58 (WRKY58)
Length = 423
Score = 38.1 bits (87), Expect = 0.007
Identities = 17/35 (48%), Positives = 21/35 (59%)
Query: 153 RCAEDKTILITTYEGNHNHPLPPAATAIAHTTSAA 187
R + D +ITTYEG HNH +P A A T+AA
Sbjct: 344 RASTDAKAVITTYEGKHNHDVPAARNGTAAATAAA 378
>At2g38470 putative WRKY-type DNA binding protein
Length = 512
Score = 38.1 bits (87), Expect = 0.007
Identities = 31/102 (30%), Positives = 45/102 (43%), Gaps = 10/102 (9%)
Query: 153 RCAEDKTILITTYEGNHNHPLPPAATAIAHTTSAAAAMLLSSSTSSTLRKESATGY---- 208
R + D +ITTYEG HNH + PAA + T+ A S+S +R + G+
Sbjct: 393 RASHDMRAVITTYEGKHNHDV-PAARGSGYATNRAP----QDSSSVPIRPAAIAGHSNYT 447
Query: 209 LSNSFPYA-TMATSTLSASQPFPTITLDFTQNHNLSMHHNRV 249
S+ PY M + + + PF + N NL N V
Sbjct: 448 TSSQAPYTLQMLHNNNTNTGPFGYAMNNNNNNSNLQTQQNFV 489
Score = 28.1 bits (61), Expect = 6.9
Identities = 13/49 (26%), Positives = 25/49 (50%)
Query: 165 YEGNHNHPLPPAATAIAHTTSAAAAMLLSSSTSSTLRKESATGYLSNSF 213
Y+G+HNHP P + + ++S + + ++S + S +NSF
Sbjct: 226 YKGSHNHPKPQSTRRSSSSSSTFHSAVYNASLDHNRQASSDQPNSNNSF 274
>At1g13960 WRKY DNA-binding protein 4 (WRKY4)
Length = 514
Score = 38.1 bits (87), Expect = 0.007
Identities = 18/35 (51%), Positives = 23/35 (65%), Gaps = 1/35 (2%)
Query: 153 RCAEDKTILITTYEGNHNHPLPPAATAIAHTTSAA 187
R A D ++TTYEG HNH L PAA + +H +AA
Sbjct: 447 RAATDPKAVVTTYEGKHNHDL-PAAKSSSHAAAAA 480
>At5g03060 putative protein
Length = 292
Score = 37.7 bits (86), Expect = 0.009
Identities = 19/77 (24%), Positives = 39/77 (49%)
Query: 10 EENLENSETEMSILESELRRVQEENHKLRIMLEQITKSYSQLQAQLFITLQKQKPNHGQN 69
+EN EN E + +E +++ E N ++ ++ K + L L+K++ +
Sbjct: 43 KENYENLEKDYKSIEESFKQMNEMNEIMKFQYQKQIKELEEKILSLLKDLEKERSEKEEY 102
Query: 70 MEENHGMVSEQIFLNNN 86
M+E GM+SE+ + N+
Sbjct: 103 MKEMKGMISEKEAIIND 119
>At2g23320 putative WRKY-type DNA-binding protein
Length = 317
Score = 37.7 bits (86), Expect = 0.009
Identities = 21/43 (48%), Positives = 27/43 (61%), Gaps = 4/43 (9%)
Query: 153 RCAEDKTILITTYEGNHNHPLPPAATAIAHTTSAAAAMLLSSS 195
R A+D ++LI TYEG+HNH L A A A A A ++L SS
Sbjct: 279 RAADDSSMLIVTYEGDHNHSLSAADLAGA----AVADLILESS 317
>At5g46350 putative protein
Length = 326
Score = 37.4 bits (85), Expect = 0.011
Identities = 24/70 (34%), Positives = 37/70 (52%), Gaps = 4/70 (5%)
Query: 153 RCAEDKTILITTYEGNHNHPLPP-AATAIAHTTSAAAAMLLSSSTSSTLRKESATGYLSN 211
R +D T++ITTYE HNHP+P TA+ T+A+ SS S L + + ++
Sbjct: 221 RSYQDPTVVITTYESQHNHPIPTNRRTAMFSGTTASDYNPSSSPIFSDLIINTPRSFSND 280
Query: 212 SF---PYATM 218
PYA++
Sbjct: 281 DLFRVPYASV 290
>At2g34830 WRKY-type DNA binding protein-like
Length = 427
Score = 37.4 bits (85), Expect = 0.011
Identities = 20/48 (41%), Positives = 28/48 (57%), Gaps = 1/48 (2%)
Query: 153 RCAEDKTILITTYEGNHNHPLPPAATAIAHTT-SAAAAMLLSSSTSST 199
R D +L+ TY HNHP P A+A +T S++++ L SS SST
Sbjct: 254 RSRTDPNMLVITYTSEHNHPWPTQRNALAGSTRSSSSSSLNPSSKSST 301
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.313 0.126 0.354
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,072,322
Number of Sequences: 26719
Number of extensions: 287076
Number of successful extensions: 1400
Number of sequences better than 10.0: 104
Number of HSP's better than 10.0 without gapping: 68
Number of HSP's successfully gapped in prelim test: 36
Number of HSP's that attempted gapping in prelim test: 1283
Number of HSP's gapped (non-prelim): 145
length of query: 330
length of database: 11,318,596
effective HSP length: 100
effective length of query: 230
effective length of database: 8,646,696
effective search space: 1988740080
effective search space used: 1988740080
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 60 (27.7 bits)
Medicago: description of AC139745.3


