

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
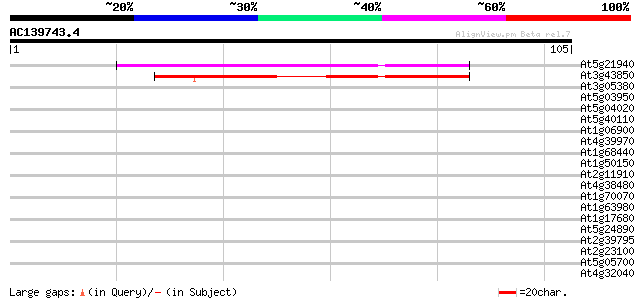
Query= AC139743.4 - phase: 0 /pseudo
(105 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At5g21940 unknown protein 54 2e-08
At3g43850 putative protein 52 5e-08
At3g05380 unknown protein 36 0.003
At5g03950 putative protein 35 0.006
At5g04020 pathogen-induced calmodulin-binding protein (PICBP) 35 0.008
At5g40110 putative protein 33 0.024
At1g06900 unknown protein 33 0.031
At4g39970 unknown protein 32 0.040
At1g68440 unknown protein 32 0.053
At1g50150 hypothetical protein 32 0.053
At2g11910 unknown protein 32 0.069
At4g38480 unknown protein 31 0.090
At1g70070 Similar to Synechocystis antiviral protein (At1g70070) 31 0.090
At1g63980 unknown protein 31 0.090
At1g17680 unknown protein 31 0.090
At5g24890 unknown protein 31 0.12
At2g39795 unknown protein 30 0.15
At2g23100 hypothetical protein 30 0.15
At5g05700 arginine-tRNA-protein transferase 1 homolog 30 0.20
At4g32040 homeodomain containing protein 1 30 0.20
>At5g21940 unknown protein
Length = 264
Score = 53.5 bits (127), Expect = 2e-08
Identities = 29/66 (43%), Positives = 38/66 (56%), Gaps = 1/66 (1%)
Query: 21 APRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGGALDCMEALE 80
+P D S+++SSSIGRNSDD + ENE ES Y G L+ ME+LE
Sbjct: 24 SPSPSPSDSSSSPSSSASSSIGRNSDDGEKSSEDGGDDAGENEVESPYK-GPLEMMESLE 82
Query: 81 EVLPIR 86
+VLP+R
Sbjct: 83 QVLPVR 88
>At3g43850 putative protein
Length = 270
Score = 52.0 bits (123), Expect = 5e-08
Identities = 33/61 (54%), Positives = 37/61 (60%), Gaps = 12/61 (19%)
Query: 28 DQGKVC--STTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGGALDCMEALEEVLPI 85
DQ C S+TSS SIG NSDDD+ ENE ES YN G LD ME+LEE LPI
Sbjct: 15 DQEFACLSSSTSSDSIGENSDDDEG---------GENEIESSYN-GPLDMMESLEEALPI 64
Query: 86 R 86
+
Sbjct: 65 K 65
>At3g05380 unknown protein
Length = 1055
Score = 36.2 bits (82), Expect = 0.003
Identities = 30/96 (31%), Positives = 45/96 (46%), Gaps = 9/96 (9%)
Query: 9 KRVGSRTSVLFDAPRDHDDDQGKVCSTTS---SSSIGRNSDDDDDDEVSSERSMDENEAE 65
KRV + + +A + DD G+ CS T S S R + + E S RS + +
Sbjct: 315 KRVYKKRVKVEEAECNDSDDNGEACSATQGLRSKSQRRKAAIEASREKYSPRS--PKKRD 372
Query: 66 SKYNGGALDCMEALEE----VLPIRLFQSFMVMQEK 97
K+ GA D ++AL E +LP L +S + Q K
Sbjct: 373 DKHTSGAFDALQALAELSASMLPANLMESELSAQLK 408
>At5g03950 putative protein
Length = 252
Score = 35.0 bits (79), Expect = 0.006
Identities = 16/51 (31%), Positives = 27/51 (52%), Gaps = 2/51 (3%)
Query: 23 RDHDDDQG--KVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGG 71
R H DD +VC + S+ + D+ DD+ + + DE E +++Y GG
Sbjct: 95 RTHGDDAESYRVCDSVSNVDENNEAVDEQDDDEDDKTNEDEEEGDNEYRGG 145
>At5g04020 pathogen-induced calmodulin-binding protein (PICBP)
Length = 1495
Score = 34.7 bits (78), Expect = 0.008
Identities = 25/90 (27%), Positives = 38/90 (41%), Gaps = 9/90 (10%)
Query: 3 LETVAFKRVGSRT---------SVLFDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEV 53
+ETVA K S+T ++ HD D GKV TTS ++G N + +
Sbjct: 354 METVASKVQESKTETVGATLWRAICEQTVTGHDHDDGKVDGTTSDGTVGDNEEVCREGSS 413
Query: 54 SSERSMDENEAESKYNGGALDCMEALEEVL 83
R D + E +N +A +E+L
Sbjct: 414 GELREEDGKKTEYVWNETVTLVKQAFDEIL 443
Score = 25.0 bits (53), Expect = 6.4
Identities = 14/42 (33%), Positives = 19/42 (44%), Gaps = 6/42 (14%)
Query: 30 GKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGG 71
G+V + S + N D D D+EV E E S +GG
Sbjct: 291 GRVVTDQESKEVDDNVDGDSDEEVF------EEEVSSSVDGG 326
>At5g40110 putative protein
Length = 280
Score = 33.1 bits (74), Expect = 0.024
Identities = 16/51 (31%), Positives = 26/51 (50%), Gaps = 2/51 (3%)
Query: 23 RDHDDD--QGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGG 71
R H DD +VC + S+ + D+ DD+ + + DE E ++K GG
Sbjct: 95 RTHGDDAESDRVCDSVSNVDENNEAVDEKDDDEDDKTNEDEEEGDNKDGGG 145
>At1g06900 unknown protein
Length = 1023
Score = 32.7 bits (73), Expect = 0.031
Identities = 16/47 (34%), Positives = 24/47 (51%)
Query: 24 DHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNG 70
D DD+ G+ + SS + +DD++D E DE+E E K G
Sbjct: 54 DEDDEDGEEEDSDGSSEDDDDDEDDEEDGEGDEEDEDEDEDEVKGKG 100
>At4g39970 unknown protein
Length = 316
Score = 32.3 bits (72), Expect = 0.040
Identities = 21/64 (32%), Positives = 28/64 (42%), Gaps = 4/64 (6%)
Query: 8 FKRVGSRTSVLFDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSE----RSMDENE 63
FK G TS +FD+P +DDD+ K+ T R + V R MDE +
Sbjct: 129 FKENGWPTSTIFDSPPQNDDDRAKLIDTLQDWKTERYKEIIKSGSVEPRPGVIRLMDEAK 188
Query: 64 AESK 67
A K
Sbjct: 189 AAGK 192
>At1g68440 unknown protein
Length = 306
Score = 32.0 bits (71), Expect = 0.053
Identities = 20/61 (32%), Positives = 30/61 (48%), Gaps = 6/61 (9%)
Query: 11 VGSRTSVLFDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNG 70
V + LFD D+D D CS+T SS ++ +D+ DDE + +D + NG
Sbjct: 76 VSVSSPTLFD---DYDSDSDVSCSSTVSSDDEKDEEDEADDE---DEDVDSIFNRRRVNG 129
Query: 71 G 71
G
Sbjct: 130 G 130
>At1g50150 hypothetical protein
Length = 358
Score = 32.0 bits (71), Expect = 0.053
Identities = 15/50 (30%), Positives = 25/50 (50%)
Query: 19 FDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKY 68
++ + DDD + TS S +D D+DD SSE ++ + E +Y
Sbjct: 241 YNEESESDDDIAESDQCTSQSEAEEETDGDNDDTSSSETKIEGTDDEERY 290
>At2g11910 unknown protein
Length = 168
Score = 31.6 bits (70), Expect = 0.069
Identities = 19/42 (45%), Positives = 26/42 (61%), Gaps = 3/42 (7%)
Query: 44 NSDDDDDDEVSSERSMDENEA-ESKYNGGALDCMEALEEVLP 84
+SDDDDDDE + E DE++A + ++GG D E EE P
Sbjct: 74 DSDDDDDDEDADEDDDDEDDANDEDFSGGEGD--EGEEEADP 113
Score = 25.0 bits (53), Expect = 6.4
Identities = 9/26 (34%), Positives = 17/26 (64%)
Query: 42 GRNSDDDDDDEVSSERSMDENEAESK 67
G + +DDDD+E ++ ++NE E +
Sbjct: 124 GSDDEDDDDEEGDNDDEDEDNEDEEE 149
>At4g38480 unknown protein
Length = 471
Score = 31.2 bits (69), Expect = 0.090
Identities = 13/27 (48%), Positives = 18/27 (66%)
Query: 44 NSDDDDDDEVSSERSMDENEAESKYNG 70
+ DDD DDE S E S D++ +E + NG
Sbjct: 411 DDDDDSDDESSEESSDDDDSSEEEENG 437
Score = 31.2 bits (69), Expect = 0.090
Identities = 12/29 (41%), Positives = 19/29 (65%)
Query: 46 DDDDDDEVSSERSMDENEAESKYNGGALD 74
DDDD D+ SSE S D++++ + G +D
Sbjct: 412 DDDDSDDESSEESSDDDDSSEEEENGEVD 440
Score = 24.6 bits (52), Expect = 8.4
Identities = 11/44 (25%), Positives = 20/44 (45%)
Query: 20 DAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENE 63
D +++ G+V + + DDDDDD+ + D +E
Sbjct: 427 DDDSSEEEENGEVDVEITKDDDNDHGDDDDDDDEDDDGDGDGDE 470
>At1g70070 Similar to Synechocystis antiviral protein (At1g70070)
Length = 1171
Score = 31.2 bits (69), Expect = 0.090
Identities = 18/37 (48%), Positives = 21/37 (56%), Gaps = 1/37 (2%)
Query: 24 DHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMD 60
D DDD + S RNSDDDDDDE +E S+D
Sbjct: 75 DEDDDDEAADEYDNISDEIRNSDDDDDDE-ETEFSVD 110
>At1g63980 unknown protein
Length = 391
Score = 31.2 bits (69), Expect = 0.090
Identities = 14/31 (45%), Positives = 19/31 (61%)
Query: 35 TTSSSSIGRNSDDDDDDEVSSERSMDENEAE 65
T+ +S + DDDDDDE E DE+EA+
Sbjct: 261 TSFDNSDDDDDDDDDDDEEDEEEDEDESEAD 291
Score = 30.4 bits (67), Expect = 0.15
Identities = 16/48 (33%), Positives = 25/48 (51%), Gaps = 2/48 (4%)
Query: 39 SSIGRNSDDDDDDEVSSERSMDENEAESKYNGGALDCMEALEEVLPIR 86
+S + DDDDDD+ E +E+E ES+ + D +E LP +
Sbjct: 261 TSFDNSDDDDDDDDDDDEEDEEEDEDESEADDDDKD--SVIESSLPAK 306
>At1g17680 unknown protein
Length = 896
Score = 31.2 bits (69), Expect = 0.090
Identities = 19/67 (28%), Positives = 32/67 (47%), Gaps = 5/67 (7%)
Query: 12 GSRTSVLFDAPRDHDDDQGKVCSTTSSSSIGRNSD-----DDDDDEVSSERSMDENEAES 66
G+ S L + P + + D+ + T+ NSD DDDDD+ + DE+E E
Sbjct: 14 GNLISELEEGPSNMECDKQVLGGDTNYDDKDLNSDEEGLVDDDDDDSDDDDEGDESEEED 73
Query: 67 KYNGGAL 73
+ G++
Sbjct: 74 DFEAGSV 80
>At5g24890 unknown protein
Length = 240
Score = 30.8 bits (68), Expect = 0.12
Identities = 18/50 (36%), Positives = 27/50 (54%), Gaps = 2/50 (4%)
Query: 35 TTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGGALDCMEALEEVLP 84
++ SSSIG D ++D+E S + D + E G L M +LE+ LP
Sbjct: 59 SSDSSSIGTPGDSEEDEEESENENDDVSSKELGLRG--LASMSSLEDSLP 106
Score = 27.7 bits (60), Expect = 0.99
Identities = 11/27 (40%), Positives = 16/27 (58%)
Query: 44 NSDDDDDDEVSSERSMDENEAESKYNG 70
+ DD+DDDE + DEN++ S G
Sbjct: 177 DDDDEDDDEEDLKSGFDENKSSSDEEG 203
>At2g39795 unknown protein
Length = 250
Score = 30.4 bits (67), Expect = 0.15
Identities = 22/63 (34%), Positives = 33/63 (51%), Gaps = 3/63 (4%)
Query: 23 RDHDDDQGKVCSTTSSSSIGRNSDDDDDDE-VSSERSMDENEAESKYNGGALD--CMEAL 79
RD++ + KV + S N DDDDDDE S+E S+ +K +G L+ CM
Sbjct: 110 RDYNGEHIKVVVSMPSLVSDENDDDDDDDEGPSNESSIPLVVTVTKKSGLTLEFSCMAFP 169
Query: 80 EEV 82
+E+
Sbjct: 170 DEI 172
>At2g23100 hypothetical protein
Length = 1036
Score = 30.4 bits (67), Expect = 0.15
Identities = 19/67 (28%), Positives = 27/67 (39%), Gaps = 19/67 (28%)
Query: 16 SVLFDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGGALDC 75
S FD D DDD + + DDDDDD+ D+N+ Y G C
Sbjct: 171 SPFFDVEDDDDDD------------VDLDDDDDDDDD-------DDNDLSDIYYGKCKCC 211
Query: 76 MEALEEV 82
+ E++
Sbjct: 212 QKRFEDI 218
>At5g05700 arginine-tRNA-protein transferase 1 homolog
Length = 632
Score = 30.0 bits (66), Expect = 0.20
Identities = 18/62 (29%), Positives = 30/62 (48%), Gaps = 7/62 (11%)
Query: 12 GSRTSVLFDAPRDHDD-DQGKV------CSTTSSSSIGRNSDDDDDDEVSSERSMDENEA 64
G+ +++ A +H+D +QG+ CS + DDDDDDE E +++
Sbjct: 509 GASETLVEPAASEHEDMEQGETNDNFMGCSDEDEDEDEDDDDDDDDDEEMYETESEDSHI 568
Query: 65 ES 66
ES
Sbjct: 569 ES 570
>At4g32040 homeodomain containing protein 1
Length = 383
Score = 30.0 bits (66), Expect = 0.20
Identities = 20/64 (31%), Positives = 32/64 (49%), Gaps = 7/64 (10%)
Query: 38 SSSIGRN-SDDDDDDEVSSERSMDENEAESKYNGGALDCMEALEEVLPIRLFQSFMVMQE 96
S S G+ SDD+DD++V SE +M + + DC+ ++P +S M +
Sbjct: 226 SESNGKTMSDDEDDNQVESEVNMFDGSLDGS------DCLMGFGPLVPTERERSLMERVK 279
Query: 97 KYLK 100
K LK
Sbjct: 280 KELK 283
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.312 0.129 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,567,059
Number of Sequences: 26719
Number of extensions: 111586
Number of successful extensions: 1737
Number of sequences better than 10.0: 263
Number of HSP's better than 10.0 without gapping: 181
Number of HSP's successfully gapped in prelim test: 85
Number of HSP's that attempted gapping in prelim test: 1026
Number of HSP's gapped (non-prelim): 548
length of query: 105
length of database: 11,318,596
effective HSP length: 81
effective length of query: 24
effective length of database: 9,154,357
effective search space: 219704568
effective search space used: 219704568
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 52 (24.6 bits)
Medicago: description of AC139743.4


