

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139743.2 - phase: 0 /pseudo
(461 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
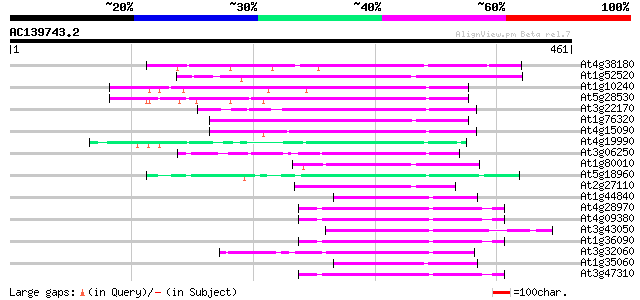
Score E
Sequences producing significant alignments: (bits) Value
At4g38180 unknown protein (At4g38180) 87 2e-17
At1g52520 F6D8.26 69 4e-12
At1g10240 unknown protein 69 4e-12
At5g28530 far-red impaired response protein (FAR1) - like 69 7e-12
At3g22170 far-red impaired response protein, putative 67 2e-11
At1g76320 putative phytochrome A signaling protein 67 2e-11
At4g15090 unknown protein 66 4e-11
At4g19990 putative protein 64 2e-10
At3g06250 unknown protein 64 2e-10
At1g80010 hypothetical protein 64 2e-10
At5g18960 FAR1 - like protein 62 7e-10
At2g27110 Mutator-like transposase 62 9e-10
At1g44840 hypothetical protein 59 6e-09
At4g28970 putative protein 59 7e-09
At4g09380 putative protein 57 2e-08
At3g43050 putative protein 57 2e-08
At1g36090 hypothetical protein 57 2e-08
At3g32060 hypothetical protein 57 2e-08
At1g35060 hypothetical protein 55 6e-08
At3g47310 putative protein 55 8e-08
>At4g38180 unknown protein (At4g38180)
Length = 788
Score = 87.0 bits (214), Expect = 2e-17
Identities = 79/324 (24%), Positives = 142/324 (43%), Gaps = 23/324 (7%)
Query: 113 RLVCERSGSHIVPEKKPKHANTGS----RKCGCLFMISGYQSKQTKEWGLNILNGVHNHP 168
+ VC + G + EK+ K + GC +S + + + +W ++ HNH
Sbjct: 118 QFVCAKEGFRNMNEKRTKDREIKRPRTITRVGCKASLS-VKMQDSGKWLVSGFVKDHNHE 176
Query: 169 MEPALEGHILAG--RLKEDDKKIVRDLTKSKMLPRNILIHLKNQRPHC----MTNVKQVY 222
+ P + H L ++ K ++ L + M PR I+ L + T V
Sbjct: 177 LVPPDQVHCLRSHRQISGPAKTLIDTLQAAGMGPRRIMSALIKEYGGISKVGFTEVDCRN 236
Query: 223 NERQQIWKANRGDKKSLQFLISKLEEHNYT----YYSRTQLESNTIEDIFWAHPTSIKLF 278
R K+ G+ +Q L+ L + N +YS E ++ ++FWA P +I F
Sbjct: 237 YMRNNRQKSIEGE---IQLLLDYLRQMNADNPNFFYSVQGSEDQSVGNVFWADPKAIMDF 293
Query: 279 NNFPTVLVMDSTYKTNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKL 338
+F + D+TY++N YR+P GV G F+ +E E +FVW+
Sbjct: 294 THFGDTVTFDTTYRSNRYRLPFAPFTGVNHHGQPILFGCAFIINETEASFVWLFNTWLAA 353
Query: 339 LSSKMNMPKVIVTDRDMSLMKAVAHVFPESYALNCFFHVQANVKQRCV-LNCKYPLGFKK 397
+S+ + P I TD D + A+ HVFP + C +H+ +++ + K+P F+
Sbjct: 354 MSA--HPPVSITTDHDAVIRAAIMHVFPGARHRFCKWHILKKCQEKLSHVFLKHP-SFES 410
Query: 398 D-GKEVSNRDVVKKIMNAWKAMVE 420
D K V+ + V+ W ++++
Sbjct: 411 DFHKCVNLTESVEDFERCWFSLLD 434
>At1g52520 F6D8.26
Length = 703
Score = 69.3 bits (168), Expect = 4e-12
Identities = 70/289 (24%), Positives = 118/289 (40%), Gaps = 14/289 (4%)
Query: 138 KCGCLFMISGYQSKQTKEWGLNILNGVHNHPMEPALEGHILAGRLKEDDKKI---VRDLT 194
+ GC MI Q +K W + + HNH L G L +K K + V D
Sbjct: 152 RTGCPAMIRMRQV-DSKRWRVVEVTLDHNH-----LLGCKLYKSVKRKRKCVSSPVSDAK 205
Query: 195 KSKMLPRNILIHLKNQRPHCMTNVK-QVYNERQQIWKANRGDKKSLQFLISKLEEHNYTY 253
K+ ++ + N P+ N K Q + RGD ++ +++ N +
Sbjct: 206 TIKLYRACVVDNGSNVNPNSTLNKKFQNSTGSPDLLNLKRGDSAAIYNYFCRMQLTNPNF 265
Query: 254 YSRTQL-ESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNMYRMPMFEVVGVTSTDLT 312
+ + + + ++FWA S + F V+ +DS+Y + + +P+ GV T
Sbjct: 266 FYLMDVNDEGQLRNVFWADAFSKVSCSYFGDVIFIDSSYISGKFEIPLVTFTGVNHHGKT 325
Query: 313 YSVGFGFMTHEKEENFVWVLTMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYALN 372
+ GF+ E E++ W +L+ LS P+ IVTDR L A++ VFP S+
Sbjct: 326 TLLSCGFLAGETMESYHW---LLKVWLSVMKRSPQTIVTDRCKPLEAAISQVFPRSHQRF 382
Query: 373 CFFHVQANVKQRCVLNCKYPLGFKKDGKEVSNRDVVKKIMNAWKAMVES 421
H+ + ++ Y K K V V + AW MV +
Sbjct: 383 SLTHIMRKIPEKLGGLHNYDAVRKAFTKAVYETLKVVEFEAAWGFMVHN 431
>At1g10240 unknown protein
Length = 680
Score = 69.3 bits (168), Expect = 4e-12
Identities = 72/314 (22%), Positives = 141/314 (43%), Gaps = 22/314 (7%)
Query: 83 DELLEWVRRQANKAGFTIVTQRSSLINPMFR------LVCERSGS---HIVPEKKPKHAN 133
D E+ A + GF+I R+ + + + VC R+G+ + E KP+ N
Sbjct: 58 DTAYEFYSTFAKRCGFSIRRHRTEGKDGVGKGLTRRYFVCHRAGNTPIKTLSEGKPQR-N 116
Query: 134 TGSRKCGC--LFMISGYQSKQTKEWGLNILNGVHNHPM-EPALEGHILAGR-LKEDDKKI 189
S +CGC IS + EW + HNH + EP + A R + + DK
Sbjct: 117 RRSSRCGCQAYLRISKLTELGSTEWRVTGFANHHNHELLEPNQVRFLPAYRSISDADKSR 176
Query: 190 VRDLTKSKMLPRNILIHLKNQR---PHCMTNVKQVYNERQQIWKANRGDKKSLQFL--IS 244
+ +K+ + + ++ L+ ++ P + ++ Q +K + +++ FL
Sbjct: 177 ILMFSKTGISVQQMMRLLELEKCVEPGFLPFTEKDVRNLLQSFKKLDPEDENIDFLRMCQ 236
Query: 245 KLEEHNYTYYSRTQLESNT-IEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNMYRMPMFEV 303
++E + + L++N +E+I W++ +SI+ + F +V D+T++ + MP+
Sbjct: 237 SIKEKDPNFKFEFTLDANDKLENIAWSYASSIQSYELFGDAVVFDTTHRLSAVEMPLGIW 296
Query: 304 VGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSKMNMPKVIVTDRDMSLMKAVAH 363
VGV + + G + E ++ W L ++ K P+ I+TD +M L +A+A
Sbjct: 297 VGVNNYGVPCFFGCVLLRDENLRSWSWALQAFTGFMNGK--APQTILTDHNMCLKEAIAG 354
Query: 364 VFPESYALNCFFHV 377
P + C + V
Sbjct: 355 EMPATKHALCIWMV 368
>At5g28530 far-red impaired response protein (FAR1) - like
Length = 700
Score = 68.6 bits (166), Expect = 7e-12
Identities = 75/325 (23%), Positives = 137/325 (42%), Gaps = 36/325 (11%)
Query: 83 DELLEWVRRQANKAGFTIVTQRSSLINPM--FR--LVCERSGSHIVPEKKPKHANTGSRK 138
DE E+ A K+GF+I RS+ + +R VC RSG + P KK + RK
Sbjct: 80 DEAFEYYSTFARKSGFSIRKARSTESQNLGVYRRDFVCYRSGFN-QPRKKANVEHPRERK 138
Query: 139 ---CGCLFMISGYQSKQ----TKEWGLNILNGVHNHPMEPALEGHILAG--RLKEDDKKI 189
CGC + Y +K+ W ++ + VHNH + + +L ++++ D++
Sbjct: 139 SVRCGCDGKL--YLTKEVVDGVSHWYVSQFSNVHNHELLEDDQVRLLPAYRKIQQSDQER 196
Query: 190 VRDLTKSKMLPRNILIHL----------------KNQRPHCMTNVKQVYNERQQIWKANR 233
+ L+K+ P N ++ L K+ R K V + +
Sbjct: 197 ILLLSKAG-FPVNRIVKLLELEKGVVSGQLPFIEKDVRNFVRACKKSVQENDAFMTEKRE 255
Query: 234 GDKKSLQFLISKLEEHNYTY-YSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYK 292
D L L E + + Y T E+ +E+I WA+ S++ ++ F V+V D++Y+
Sbjct: 256 SDTLELLECCKGLAERDMDFVYDCTSDENQKVENIAWAYGDSVRGYSLFGDVVVFDTSYR 315
Query: 293 TNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSKMNMPKVIVTD 352
+ Y + + G+ + +G + E +F W L + + + P+ I+TD
Sbjct: 316 SVPYGLLLGVFFGIDNNGKAMLLGCVLLQDESCRSFTWALQTFVRFMRGR--HPQTILTD 373
Query: 353 RDMSLMKAVAHVFPESYALNCFFHV 377
D L A+ P + + H+
Sbjct: 374 IDTGLKDAIGREMPNTNHVVFMSHI 398
>At3g22170 far-red impaired response protein, putative
Length = 814
Score = 67.4 bits (163), Expect = 2e-11
Identities = 50/230 (21%), Positives = 98/230 (41%), Gaps = 19/230 (8%)
Query: 155 EWGLNILNGVHNHPMEPALEGHILAGRLKEDDKKIVRDLTKSKMLPRNILIHLKNQRPHC 214
+W ++ HNH + PA + E +KI + K + +I LK+
Sbjct: 170 KWVIHSFVREHNHELLPAQA-------VSEQTRKIYAAMAK-QFAEYKTVISLKSDSKSS 221
Query: 215 MTNVKQVYNERQQIWKANRGDKKSLQFLISKLEEHNYTYYSRTQL-ESNTIEDIFWAHPT 273
E+ + GD K L +S+++ N ++ L + ++++FW
Sbjct: 222 F--------EKGRTLSVETGDFKILLDFLSRMQSLNSNFFYAVDLGDDQRVKNVFWVDAK 273
Query: 274 SIKLFNNFPTVLVMDSTYKTNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLT 333
S + +F V+ +D+TY N Y+MP+ VGV +G ++ E + W++
Sbjct: 274 SRHNYGSFCDVVSLDTTYVRNKYKMPLAIFVGVNQHYQYMVLGCALISDESAATYSWLME 333
Query: 334 MLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYALNCFFHVQANVKQ 383
+ + + PKV++T+ D+ + V +FP + +HV V +
Sbjct: 334 TWLRAIGGQ--APKVLITELDVVMNSIVPEIFPNTRHCLFLWHVLMKVSE 381
>At1g76320 putative phytochrome A signaling protein
Length = 670
Score = 67.4 bits (163), Expect = 2e-11
Identities = 48/215 (22%), Positives = 97/215 (44%), Gaps = 4/215 (1%)
Query: 165 HNHPMEPALEGHILAGRLKEDDKKIVRDLTKSKMLPRNILIHLKNQRP-HCMTNVKQVYN 223
HNH + P + + R E K L + K P HL + + +
Sbjct: 91 HNHDLLPEQAHYFRSHRNTELVKSNDSRLRRKKNTPLTDCKHLSAYHDLDFIDGYMRNQH 150
Query: 224 ERQQIWKANRGDKKSLQFLISKLEEHNYTYYSRTQL-ESNTIEDIFWAHPTSIKLFNNFP 282
++ + + GD + L + +++E N ++ E + + ++FW I+ + +F
Sbjct: 151 DKGRRLVLDTGDAEILLEFLMRMQEENPKFFFAVDFSEDHLLRNVFWVDAKGIEDYKSFS 210
Query: 283 TVLVMDSTYKTNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSK 342
V+ +++Y + Y++P+ VGV +G G + + +VW+ M L++
Sbjct: 211 DVVSFETSYFVSKYKVPLVLFVGVNHHVQPVLLGCGLLADDTVYTYVWL--MQSWLVAMG 268
Query: 343 MNMPKVIVTDRDMSLMKAVAHVFPESYALNCFFHV 377
PKV++TD++ ++ A+A V PE+ C +HV
Sbjct: 269 GQKPKVMLTDQNNAIKAAIAAVLPETRHCYCLWHV 303
>At4g15090 unknown protein
Length = 768
Score = 66.2 bits (160), Expect = 4e-11
Identities = 53/228 (23%), Positives = 99/228 (43%), Gaps = 12/228 (5%)
Query: 165 HNHPMEPALEGHILAGR-LKEDDKKIVRDLTKSKMLPRNILIHL-------KNQRPHCMT 216
HNH + PAL H R +K +K + L + + + + KN T
Sbjct: 89 HNHELLPALAYHFRIQRNVKLAEKNNIDILHAVSERTKKMYVEMSRQSGGYKNIGSLLQT 148
Query: 217 NVKQVYNERQQIWKANRGDKKSLQFLISKLEEHNYTYYSRTQL-ESNTIEDIFWAHPTSI 275
+V ++ + + GD + L ++++ N ++ L E + ++FWA S
Sbjct: 149 DVSSQVDKGRYL-ALEEGDSQVLLEYFKRIKKENPKFFYAIDLNEDQRLRNLFWADAKSR 207
Query: 276 KLFNNFPTVLVMDSTYKTNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTML 335
+ +F V+ D+TY ++P+ +GV +G + E E FVW++
Sbjct: 208 DDYLSFNDVVSFDTTYVKFNDKLPLALFIGVNHHSQPMLLGCALVADESMETFVWLIKTW 267
Query: 336 RKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYALNCFFHVQANVKQ 383
+ + + PKVI+TD+D LM AV+ + P + +HV + +
Sbjct: 268 LRAMGGR--APKVILTDQDKFLMSAVSELLPNTRHCFALWHVLEKIPE 313
>At4g19990 putative protein
Length = 672
Score = 63.5 bits (153), Expect = 2e-10
Identities = 76/330 (23%), Positives = 127/330 (38%), Gaps = 75/330 (22%)
Query: 66 EVDTAKFFSDDVKSKNRDELLEWVRRQANKAGFTIVTQ--RSSLINPMF---RLVCERSG 120
E+D + F ++++E E+ + AN GFT + + R S + F + VC R G
Sbjct: 20 EIDEGREF------ESKEEAFEFYKEYANSVGFTTIIKASRRSRMTGKFIDAKFVCTRYG 73
Query: 121 S--------------HIVPEKKPKHANTGSRKCGCLFMISGYQSKQTKEWGLNILNGVHN 166
S +I +K N S K C + + +Q W + L HN
Sbjct: 74 SKKEDIDTGLGTDGFNIPQARKRGRINRSSSKTDCKAFLH-VKRRQDGRWVVRSLVKEHN 132
Query: 167 HPMEPALEGHILAGRLKEDDKKIVRDLTKSKMLPRNILIHLKNQRPHCMTNVKQVYNERQ 226
H I G+ +R+L+ R+
Sbjct: 133 H--------EIFTGQADS-----LRELSG-----------------------------RR 150
Query: 227 QIWKANRGDKKSLQFLISKLEEHNYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLV 286
++ K N K ++ KLE+ + + ++ +IFW + F V+
Sbjct: 151 KLEKLNGAIVKEVKS--RKLEDGDVERLLNFFTDMQSLRNIFWVDAKGRFDYTCFSDVVS 208
Query: 287 MDSTYKTNMYRMPMFEVVGVTSTDLTYSVGFG-FMTHEKEENFVWVLTMLRKLLSSKMNM 345
+D+T+ N Y++P+ GV +GFG +T E + FVW+ K +
Sbjct: 209 IDTTFIKNEYKLPLVAFTGVNHHGQFLLLGFGLLLTDESKSGFVWLFRAWLKAMHG--CR 266
Query: 346 PKVIVTDRDMSLMKAVAHVFPESYALNCFF 375
P+VI+T D L +AV VFP S +CF+
Sbjct: 267 PRVILTKHDQMLKEAVLEVFPSS--RHCFY 294
>At3g06250 unknown protein
Length = 764
Score = 63.5 bits (153), Expect = 2e-10
Identities = 59/232 (25%), Positives = 99/232 (42%), Gaps = 25/232 (10%)
Query: 139 CGCLFMISGYQSKQTKEWGLNILNGVHNHPMEPALEGHILAGRLKEDDKKIVRDLTKSKM 198
CG I + + + W ++ LN HNH +EP G+ KKI D+T
Sbjct: 251 CGAYMRI---KRQDSGGWIVDRLNKDHNHDLEP--------GKKNAGMKKITDDVTGG-- 297
Query: 199 LPRNILIHLKNQRPHCMTNVKQVYNERQQIWKANRGDKKSLQFLISKLEEHNYTYYSRTQ 258
L LI L + H + + + W L + SK E +Y+ +
Sbjct: 298 LDSVDLIELNDLSNHISSTRENTIGKE---WYP-----VLLDYFQSKQAEDMGFFYA-IE 348
Query: 259 LESN-TIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNMYRMPMFEVVGVTSTDLTYSVGF 317
L+SN + IFWA S + F +V D++Y+ Y +P +G +G
Sbjct: 349 LDSNGSCMSIFWADSRSRFACSQFGDAVVFDTSYRKGDYSVPFATFIGFNHHRQPVLLGG 408
Query: 318 GFMTHEKEENFVWVLTMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESY 369
+ E +E F W+ + +S + P+ +V D+D+ + +AVA VFP ++
Sbjct: 409 ALVADESKEAFSWLFQTWLRAMSGR--RPRSMVADQDLPIQQAVAQVFPGTH 458
>At1g80010 hypothetical protein
Length = 696
Score = 63.5 bits (153), Expect = 2e-10
Identities = 40/157 (25%), Positives = 77/157 (48%), Gaps = 7/157 (4%)
Query: 233 RGDKKSLQ--FLISKLEEHNYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDST 290
RG ++LQ F +L N+ Y + ++ ++FW + +++F VL+ D+T
Sbjct: 240 RGGFRALQDFFFQIQLSSPNFLYLMDLA-DDGSLRNVFWIDARARAAYSHFGDVLLFDTT 298
Query: 291 YKTNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSKMNM-PKVI 349
+N Y +P+ VG+ T +G G + + E +VW + R L+ + P++
Sbjct: 299 CLSNAYELPLVAFVGINHHGDTILLGCGLLADQSFETYVW---LFRAWLTCMLGRPPQIF 355
Query: 350 VTDRDMSLMKAVAHVFPESYALNCFFHVQANVKQRCV 386
+T++ ++ AV+ VFP ++ HV N+ Q V
Sbjct: 356 ITEQCKAMRTAVSEVFPRAHHRLSLTHVLHNICQSVV 392
>At5g18960 FAR1 - like protein
Length = 788
Score = 62.0 bits (149), Expect = 7e-10
Identities = 61/310 (19%), Positives = 121/310 (38%), Gaps = 38/310 (12%)
Query: 113 RLVCERSGSHIVPEKKPKHANTGSRKCGCLFMISGYQSKQTKEWGLNILNGVHNHPMEPA 172
R VC R G + CG I + + + W ++ LN HNH +EP
Sbjct: 256 RFVCSREGFQ----------HPSRMGCGAYMRI---KRQDSGGWIVDRLNKDHNHDLEPG 302
Query: 173 LEGHILAGRLKEDDKKIVR--DLTKSKMLPRNILIHLKNQRPHCMTNVKQVYNERQQIWK 230
+ ++ +D + DL + N H+K R N + W
Sbjct: 303 KKNDAGMKKIPDDGTGGLDSVDLIELNDFGNN---HIKKTRE----------NRIGKEWY 349
Query: 231 ANRGDKKSLQFLISKLEEHNYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDST 290
D F + E+ + Y + + + IFWA + + F +V D++
Sbjct: 350 PLLLD----YFQSRQTEDMGFFYAVELDVNNGSCMSIFWADSRARFACSQFGDSVVFDTS 405
Query: 291 YKTNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSKMNMPKVIV 350
Y+ Y +P ++G +G + E +E F+W+ + +S + P+ IV
Sbjct: 406 YRKGSYSVPFATIIGFNHHRQPVLLGCAMVADESKEAFLWLFQTWLRAMSGR--RPRSIV 463
Query: 351 TDRDMSLMKAVAHVFPESYALNCFFHVQANVKQRCVLNCKYPLGFKKD-GKEVSNRDVVK 409
D+D+ + +A+ VFP ++ + ++ ++ + +P FK + K + +
Sbjct: 464 ADQDLPIQQALVQVFPGAHHRYSAWQIREKERENLI---PFPSEFKYEYEKCIYQTQTIV 520
Query: 410 KIMNAWKAMV 419
+ + W A++
Sbjct: 521 EFDSVWSALI 530
Score = 31.2 bits (69), Expect = 1.3
Identities = 31/120 (25%), Positives = 45/120 (36%), Gaps = 30/120 (25%)
Query: 55 GGSGVLPLQPHEVDTAKFFSDDVKSKNRDELLEWVRRQANKAGFTIVT-----QRSSLIN 109
GGSGV P E DTA +E E+ A + GF + T R+
Sbjct: 37 GGSGVEPYVGLEFDTA------------EEAREFYNAYAARTGFKVRTGQLYRSRTDGTV 84
Query: 110 PMFRLVCERSGSHIVPEKKPKHANTGSRKCGCLFMISGYQSKQTKEWGLNILNGVHNHPM 169
R VC + G + + + GC I Q + T +W L+ + HNH +
Sbjct: 85 SSRRFVCSKEGFQL------------NSRTGCTAFIR-VQRRDTGKWVLDQIQKEHNHEL 131
>At2g27110 Mutator-like transposase
Length = 851
Score = 61.6 bits (148), Expect = 9e-10
Identities = 34/134 (25%), Positives = 64/134 (47%), Gaps = 5/134 (3%)
Query: 235 DKKSLQFLISKLEEHNYTYYSRTQL-ESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKT 293
D +L +++ N ++ QL E N + ++FWA S + +F + +D+ Y+
Sbjct: 196 DAHNLLEYFKRMQAENPGFFYAVQLDEDNQMSNVFWADSRSRVAYTHFGDTVTLDTRYRC 255
Query: 294 NMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSKMNMPKV-IVTD 352
N +R+P GV G + E + +F+W + + L++ + P V +VTD
Sbjct: 256 NQFRVPFAPFTGVNHHGQAILFGCALILDESDTSFIW---LFKTFLTAMRDQPPVSLVTD 312
Query: 353 RDMSLMKAVAHVFP 366
+D ++ A VFP
Sbjct: 313 QDRAIQIAAGQVFP 326
>At1g44840 hypothetical protein
Length = 926
Score = 58.9 bits (141), Expect = 6e-09
Identities = 31/118 (26%), Positives = 59/118 (49%), Gaps = 2/118 (1%)
Query: 267 IFWAHPTSIKLFNNFPTVLVMDSTYKTNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEE 326
+F A SI F + +LV+D T+ Y+ + G + Y +GF + E +E
Sbjct: 557 MFLAFGASIAGFRHLRRILVVDGTHLKGKYKGVLLTSSGQDANFQVYPLGFAVVDSENDE 616
Query: 327 NFVWVLTMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYALNCFFHVQANVKQR 384
++ W T L ++++ + I++DR S++ AV VFP++ C H+ N++ +
Sbjct: 617 SWTWFFTKLERIIADSKTL--TILSDRHSSILVAVKRVFPQANHGACIIHLCRNIQTK 672
>At4g28970 putative protein
Length = 914
Score = 58.5 bits (140), Expect = 7e-09
Identities = 43/170 (25%), Positives = 77/170 (45%), Gaps = 14/170 (8%)
Query: 238 SLQFLISKLEEHNYTYYSRTQLESNT-IEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNMY 296
S +++ K+ TY +LE + +F A I+ F V+V+D+T+ +Y
Sbjct: 366 SYLYMLEKVNPGTVTY---VELEGEKKFKYLFIALGACIEGFRAMRKVIVVDATHLKTVY 422
Query: 297 RMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSKMNMPKVIVTDRDMS 356
+ Y + FG + EK+ +++W L L+ + S + V ++DR S
Sbjct: 423 GGMLVIATAQDPNHHHYPLAFGIIDSEKDVSWIWFLEKLKTVYSDVPGL--VFISDRHQS 480
Query: 357 LMKAVAHVFPESYALNCFFHVQANVKQRCVLNCKYPLGFKKDGKEVSNRD 406
+ KAV V+P + C +H+ N++ R ++ KDG V RD
Sbjct: 481 IKKAVKTVYPNALHAACIWHLCQNMRDRVTID--------KDGAAVKFRD 522
>At4g09380 putative protein
Length = 960
Score = 57.4 bits (137), Expect = 2e-08
Identities = 43/170 (25%), Positives = 77/170 (45%), Gaps = 14/170 (8%)
Query: 238 SLQFLISKLEEHNYTYYSRTQLESNT-IEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNMY 296
S +++ K+ TY +LE + +F A I+ F V+V+D+T+ +Y
Sbjct: 366 SYLYMLEKVNPGTVTY---VELEGEKKFKYLFIALGACIEGFRAMRKVIVVDATHLKTVY 422
Query: 297 RMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSKMNMPKVIVTDRDMS 356
+ Y + FG + EK+ +++W L L+ + S + V ++DR S
Sbjct: 423 GGMLVIATAQDPNHHHYPLAFGIIDSEKDVSWIWFLENLKTVYSDVPGL--VFISDRHQS 480
Query: 357 LMKAVAHVFPESYALNCFFHVQANVKQRCVLNCKYPLGFKKDGKEVSNRD 406
+ KAV V+P + C +H+ N++ R ++ KDG V RD
Sbjct: 481 IKKAVKTVYPNALHAACIWHLCQNMRDRVKID--------KDGAAVKFRD 522
>At3g43050 putative protein
Length = 608
Score = 57.4 bits (137), Expect = 2e-08
Identities = 45/187 (24%), Positives = 81/187 (43%), Gaps = 20/187 (10%)
Query: 260 ESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNMYRMPMFEVVGVTSTDLTYSVGFGF 319
E + +F+A I+ FN V+++D+T+ + + Y + FG
Sbjct: 310 EKKNFQYLFFALGACIEGFNAMRKVIIVDATHLKIAQGGVLVVALAQDLEHHHYPIDFGV 369
Query: 320 MTHEKEENFVWVLTMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYALNCFFHVQA 379
+ EK+ ++ W L+ ++ + V V+DR+ SLMK+ ++P S C +H+
Sbjct: 370 LDGEKDVSWSWFFEKLKSVIPDSSEL--VFVSDRNQSLMKSQRELYPLSQHGCCIYHLCQ 427
Query: 380 NVKQRCVLNCKYPLGFKKDGKEVSNRDVVKKIMNAWKAMVESPNQQLYANALVEFKDSCS 439
NVK C + K +G KK M KA E+ +LY +FK+
Sbjct: 428 NVKGACNYSKKKVVG--------------KKFMECAKAYTEAEFLKLYG----DFKERYP 469
Query: 440 DFSIFVD 446
+++ D
Sbjct: 470 AAAVYQD 476
>At1g36090 hypothetical protein
Length = 645
Score = 57.4 bits (137), Expect = 2e-08
Identities = 43/170 (25%), Positives = 77/170 (45%), Gaps = 14/170 (8%)
Query: 238 SLQFLISKLEEHNYTYYSRTQLESNT-IEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNMY 296
S +++ K+ TY +LE + +F A I+ F V+V+D+T+ +Y
Sbjct: 326 SYLYMLEKVNPGTVTY---VELEGEKKFKYLFIALGACIEGFRAMRKVIVVDATHLKTVY 382
Query: 297 RMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSKMNMPKVIVTDRDMS 356
+ Y + FG + EK+ +++W L L+ + S + V ++DR S
Sbjct: 383 GGMLVIATAHDPNHHHYPLAFGIIDSEKDVSWIWFLEKLKTVYSDVPRL--VFISDRHQS 440
Query: 357 LMKAVAHVFPESYALNCFFHVQANVKQRCVLNCKYPLGFKKDGKEVSNRD 406
+ KAV V+P + C +H+ N++ R ++ KDG V RD
Sbjct: 441 IKKAVKTVYPNALHAACIWHLCQNMRDRVKID--------KDGAAVKFRD 482
>At3g32060 hypothetical protein
Length = 487
Score = 57.0 bits (136), Expect = 2e-08
Identities = 46/210 (21%), Positives = 98/210 (45%), Gaps = 12/210 (5%)
Query: 173 LEGHILAGRLKEDDKKIVRDLTKSKMLPRNILIHLKNQRPHCMTNVKQVYNERQQIWKAN 232
L H L G+++ +I+ +L ++K+ + + + H + ++K R++ +K
Sbjct: 165 LHDHFL-GQVETPVPRIIMELVQTKLGVKVSYSTVLRGKYHAIYDLK---GSREESYK-- 218
Query: 233 RGDKKSLQFLISKLEEHNYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYK 292
D +++ K+ + TY E++ + +F A SI+ F VL++D+T+
Sbjct: 219 --DINCYLYMLKKVNDGTVTYLKLD--ENDKFQYVFVALGASIEGFRVMRKVLIVDATHL 274
Query: 293 TNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSKMNMPKVIVTD 352
N Y + Y + F + E + ++ W L+ ++ + V +TD
Sbjct: 275 KNGYGGVLVFASAQDPNRHHYIIAFAVLDGENDASWEWFFEKLKTVVPDTSEL--VFMTD 332
Query: 353 RDMSLMKAVAHVFPESYALNCFFHVQANVK 382
R+ SL+KA+ +V+ ++ C +H+ NVK
Sbjct: 333 RNASLIKAIRNVYTAAHHGYCIWHLSQNVK 362
>At1g35060 hypothetical protein
Length = 873
Score = 55.5 bits (132), Expect = 6e-08
Identities = 31/118 (26%), Positives = 57/118 (48%), Gaps = 2/118 (1%)
Query: 267 IFWAHPTSIKLFNNFPTVLVMDSTYKTNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEE 326
+F A SI+ F + VLV+D T+ Y+ + G + Y + F + E ++
Sbjct: 508 MFLAFGASIQGFKHLRRVLVVDGTHLKGKYKGVLLTASGQDANFQVYPLAFAVVDSENDD 567
Query: 327 NFVWVLTMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYALNCFFHVQANVKQR 384
+ W T L ++++ N I++DR S+ V VFP+++ C H+ N++ R
Sbjct: 568 AWTWFFTKLERIIAD--NNTLTILSDRHESIKVGVKKVFPQAHHGACIIHLCRNIQAR 623
>At3g47310 putative protein
Length = 735
Score = 55.1 bits (131), Expect = 8e-08
Identities = 41/170 (24%), Positives = 75/170 (44%), Gaps = 14/170 (8%)
Query: 238 SLQFLISKLEEHNYTYYSRTQLESNT-IEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNMY 296
S +++ K+ TY +LE + +F A I+ F V+V+D+T+ +Y
Sbjct: 315 SYLYMLEKVNPGTVTY---VELEGEKKFKYLFIALGACIEGFRTMRKVIVVDATHLKTVY 371
Query: 297 RMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSKMNMPKVIVTDRDMS 356
+ Y + FG + E + +++W L L+ + S + V ++DR S
Sbjct: 372 GGMLVIATAQDPNHHHYPLAFGIIDSENDVSWIWFLEKLKTVYSDVPGL--VFISDRHQS 429
Query: 357 LMKAVAHVFPESYALNCFFHVQANVKQRCVLNCKYPLGFKKDGKEVSNRD 406
+ K V V+P + C +H+ N++ R ++ KDG V RD
Sbjct: 430 IKKVVKTVYPNALHAACIWHLCQNMRDRVKID--------KDGAAVKFRD 471
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.134 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,421,904
Number of Sequences: 26719
Number of extensions: 436649
Number of successful extensions: 1261
Number of sequences better than 10.0: 77
Number of HSP's better than 10.0 without gapping: 49
Number of HSP's successfully gapped in prelim test: 28
Number of HSP's that attempted gapping in prelim test: 1174
Number of HSP's gapped (non-prelim): 85
length of query: 461
length of database: 11,318,596
effective HSP length: 103
effective length of query: 358
effective length of database: 8,566,539
effective search space: 3066820962
effective search space used: 3066820962
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC139743.2


