

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
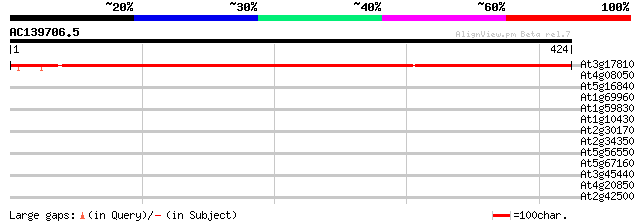
Query= AC139706.5 - phase: 0
(424 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At3g17810 dihydroorotate dehydrogenase like protein 674 0.0
At4g08050 34 0.13
At5g16840 unknown protein 34 0.17
At1g69960 serine/threonine protein phosphatase (type 2A) 32 0.66
At1g59830 unknown protein 31 1.1
At1g10430 serine/threonine protein phosphatase, PP2A, catalytic ... 31 1.1
At2g30170 unknown protein 30 2.5
At2g34350 nodulin-like protein 30 3.3
At5g56550 putative protein 29 5.6
At5g67160 anthranilate N-hydroxycinnamoyl/benzoyltransferase-lik... 28 7.3
At3g45440 receptor like protein kinase 28 7.3
At4g20850 putative protein 28 9.6
At2g42500 putative serine/threonine protein phosphatase catalyti... 28 9.6
>At3g17810 dihydroorotate dehydrogenase like protein
Length = 426
Score = 674 bits (1739), Expect = 0.0
Identities = 338/429 (78%), Positives = 381/429 (88%), Gaps = 8/429 (1%)
Query: 1 MANLS--MTQLKTRNSASRFSFNF---SKKVYPRPSRVDFKVFASEGQVSEPDLSVKVNG 55
MA++S + + +S + S +F S++ + P+RV K+ S SEPDLSV VNG
Sbjct: 1 MASMSFALNRFSGLSSKTTLSADFDPSSRRSFLPPTRVGLKI--SSAAESEPDLSVTVNG 58
Query: 56 LHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYARLRANANG 115
L MPNPFVIGSGPPGTNYTVMKRAFDEGWG VIAKTVSLDA+KVINVTPRYARLR +NG
Sbjct: 59 LKMPNPFVIGSGPPGTNYTVMKRAFDEGWGAVIAKTVSLDASKVINVTPRYARLRTGSNG 118
Query: 116 SAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAWEELIDRVE 175
SAK ++IGWQNIELISDRPLETMLKEF++LK+EYPDRILIAS+MEEYNK AWEELIDRVE
Sbjct: 119 SAKTDVIGWQNIELISDRPLETMLKEFERLKKEYPDRILIASVMEEYNKTAWEELIDRVE 178
Query: 176 QTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKMTPNITDIS 235
QTG+DA+EINFSCPHGMPER+MGAAVGQDCALL+EVCGWINAKATVPVWAKMTPNITDI+
Sbjct: 179 QTGVDALEINFSCPHGMPERRMGAAVGQDCALLDEVCGWINAKATVPVWAKMTPNITDIT 238
Query: 236 QPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAVHPIALGKVM 295
+PARV+L SGCEG+AAINTIMSVMGI++ TLRPEPCVEGYSTPGGYS KAV PIAL KVM
Sbjct: 239 EPARVSLKSGCEGIAAINTIMSVMGIDMKTLRPEPCVEGYSTPGGYSYKAVRPIALAKVM 298
Query: 296 SIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYGLVKKLSLE 355
+IAKMMKSEF SE+ SLS IGGVETG DAAEFILLG+NTVQVCTGVMMHGYG VK L E
Sbjct: 299 NIAKMMKSEF-SEDRSLSGIGGVETGYDAAEFILLGSNTVQVCTGVMMHGYGHVKTLCAE 357
Query: 356 LKDFMKKHNFTSIEDFRGASLQYFTTHTDLVHRQQEAIKQRKAIKKGLQSDKDWTGDGFV 415
LKDFMK+HNF++IE+FRG SLQYFTTHTDLV RQ+EA++QRKA K+GL+SDKDWTGDGFV
Sbjct: 358 LKDFMKQHNFSTIEEFRGHSLQYFTTHTDLVKRQKEAVEQRKAEKRGLKSDKDWTGDGFV 417
Query: 416 KESESMVSN 424
KE+ESMVSN
Sbjct: 418 KETESMVSN 426
>At4g08050
Length = 1428
Score = 34.3 bits (77), Expect = 0.13
Identities = 39/146 (26%), Positives = 65/146 (43%), Gaps = 10/146 (6%)
Query: 48 DLSVKVNGLHMPNPFVI-GSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRY 106
DL V +NG+ +P FV+ + ++ R F G VI + K+ ++
Sbjct: 638 DLPVMINGVEVPTDFVVLDMEVEHKDPLILGRPFLASVGAVI----DVKEGKIGLNLGKH 693
Query: 107 ARLRANANGSAKGEIIGWQ---NIELISDRPLET-MLKEFKQLKEEYPDRI-LIASIMEE 161
+L+ N N + +G + N +IS ET +KEFK+ ++ + I +A +EE
Sbjct: 694 IKLQFNINRTPQGSTEDGRTSGNDRVISREEYETERVKEFKKRYDKQDETIEKLAHTVEE 753
Query: 162 YNKAAWEELIDRVEQTGIDAIEINFS 187
EE E+ GI+ NFS
Sbjct: 754 RRNYPLEEKEAYFEEKGIEYSAANFS 779
>At5g16840 unknown protein
Length = 259
Score = 33.9 bits (76), Expect = 0.17
Identities = 31/104 (29%), Positives = 46/104 (43%), Gaps = 9/104 (8%)
Query: 309 NYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYGLVKKLSLELKDFMKKHNFTSI 368
NYS A ET AE ++ A V + ++ G+ L K + K F +KH FTS
Sbjct: 77 NYSPPAAPHAETQSSGAESVVQKAE--DVVSSMLAKGFILGKDAVGKAKAFDEKHGFTST 134
Query: 369 EDFRGASL-------QYFTTHTDLVHRQQEAIKQRKAIKKGLQS 405
ASL Q T T LV+ + +A+ Q + + +S
Sbjct: 135 ATAGVASLDQKIGLSQKLTAGTSLVNEKIKAVDQNFQVTERTKS 178
>At1g69960 serine/threonine protein phosphatase (type 2A)
Length = 307
Score = 32.0 bits (71), Expect = 0.66
Identities = 29/96 (30%), Positives = 39/96 (40%), Gaps = 3/96 (3%)
Query: 62 FVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYARLRANANGSAKGEI 121
F IG P TNY M D G+ V +TVSL A + R LR N ++
Sbjct: 67 FRIGGSSPDTNYLFMGDYVDRGYYSV--ETVSLLVALKVRYRDRLTILRGNHESRQITQV 124
Query: 122 IGWQNIELISDRPLETMLKEFKQLKEEYPDRILIAS 157
G+ + E + + K F L + P LI S
Sbjct: 125 YGFYD-ECLRKYGNANVWKHFTDLFDYLPLTALIES 159
>At1g59830 unknown protein
Length = 306
Score = 31.2 bits (69), Expect = 1.1
Identities = 29/96 (30%), Positives = 39/96 (40%), Gaps = 3/96 (3%)
Query: 62 FVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYARLRANANGSAKGEI 121
F IG P TNY M D G+ V +TVSL A + R LR N ++
Sbjct: 66 FRIGGNAPDTNYLFMGDYVDRGYYSV--ETVSLLVALKVRYRDRLTILRGNHESRQITQV 123
Query: 122 IGWQNIELISDRPLETMLKEFKQLKEEYPDRILIAS 157
G+ + E + + K F L + P LI S
Sbjct: 124 YGFYD-ECLRKYGNANVWKYFTDLFDYLPLTALIES 158
>At1g10430 serine/threonine protein phosphatase, PP2A, catalytic
subunit
Length = 306
Score = 31.2 bits (69), Expect = 1.1
Identities = 29/96 (30%), Positives = 39/96 (40%), Gaps = 3/96 (3%)
Query: 62 FVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYARLRANANGSAKGEI 121
F IG P TNY M D G+ V +TVSL A + R LR N ++
Sbjct: 66 FRIGGNAPDTNYLFMGDYVDRGYYSV--ETVSLLVALKVRYRDRLTILRGNHESRQITQV 123
Query: 122 IGWQNIELISDRPLETMLKEFKQLKEEYPDRILIAS 157
G+ + E + + K F L + P LI S
Sbjct: 124 YGFYD-ECLRKYGNANVWKYFTDLFDYLPLTALIES 158
>At2g30170 unknown protein
Length = 298
Score = 30.0 bits (66), Expect = 2.5
Identities = 29/93 (31%), Positives = 41/93 (43%), Gaps = 10/93 (10%)
Query: 27 YPRPSRVDF--KVFASEGQVSEPDLSVKVNGLH-MPNPFVIGSGPPGTNYTVMKRAFDEG 83
+P PSRVDF + SE Q P+LS+ V G+H +P+P + G + R
Sbjct: 21 HPNPSRVDFLCRCAPSEIQPLRPELSLSV-GIHAIPHPDKVEKGGEDAFFVSSYR----- 74
Query: 84 WGGVIAKTVSLDAAKVINVTPRYARLRANANGS 116
GGV+A + +V P AN S
Sbjct: 75 -GGVMAVADGVSGWAEQDVDPSLFSKELMANAS 106
>At2g34350 nodulin-like protein
Length = 2301
Score = 29.6 bits (65), Expect = 3.3
Identities = 16/62 (25%), Positives = 32/62 (50%), Gaps = 2/62 (3%)
Query: 331 GANTVQVCTGVMMHGYGLVKKLSLE-LKDFMKK-HNFTSIEDFRGASLQYFTTHTDLVHR 388
G+++ + + H G V+KL+ L+D ++K H + + GA + F + DL H+
Sbjct: 1221 GSDSYNILLNFVTHSDGKVRKLASSCLRDVLQKSHGTKAWQSVSGAITEMFQNYLDLAHK 1280
Query: 389 QQ 390
+
Sbjct: 1281 SE 1282
>At5g56550 putative protein
Length = 172
Score = 28.9 bits (63), Expect = 5.6
Identities = 15/42 (35%), Positives = 19/42 (44%)
Query: 281 YSAKAVHPIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGG 322
Y K+ +LG V S+ +MK F S NY TGG
Sbjct: 95 YEGKSQSFTSLGNVKSLEDLMKRGFKSRNYGAKRKSSRSTGG 136
>At5g67160 anthranilate N-hydroxycinnamoyl/benzoyltransferase-like
protein
Length = 434
Score = 28.5 bits (62), Expect = 7.3
Identities = 25/96 (26%), Positives = 41/96 (42%), Gaps = 4/96 (4%)
Query: 98 KVINVTPRYA-RLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIA 156
K+ +VT +L A AN A +I +I+ + +M+K +EE L
Sbjct: 229 KMFHVTKENVLKLDAKANDEADQKI---SSIQAVLAYIWRSMVKHSGMSREEETHCRLPI 285
Query: 157 SIMEEYNKAAWEELIDRVEQTGIDAIEINFSCPHGM 192
++ + N EE V QTGI + + HG+
Sbjct: 286 NMRQRLNPPLEEECFGNVSQTGIATVTVGELLDHGL 321
>At3g45440 receptor like protein kinase
Length = 669
Score = 28.5 bits (62), Expect = 7.3
Identities = 15/41 (36%), Positives = 21/41 (50%), Gaps = 1/41 (2%)
Query: 29 RPSRVDFKVFASEGQVSEPDLSVKVNGLHMPNPFVIGSGPP 69
RPS D + + G + PD+S G+ P +IGS PP
Sbjct: 600 RPSMEDIVQYLN-GSLELPDISPNSPGIGSFTPLIIGSNPP 639
>At4g20850 putative protein
Length = 1396
Score = 28.1 bits (61), Expect = 9.6
Identities = 20/67 (29%), Positives = 29/67 (42%), Gaps = 9/67 (13%)
Query: 217 AKATVPVWA---KMTPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVE 273
A A VP W +M N T ++ P S C +A + + M GI ++ +E
Sbjct: 526 AVAPVPTWTLQRRMLMNGTSMASP------SACGAIALLLSAMKAEGIPVSPYSVRRALE 579
Query: 274 GYSTPGG 280
STP G
Sbjct: 580 NTSTPVG 586
>At2g42500 putative serine/threonine protein phosphatase catalytic
subunit, PP2A-3
Length = 313
Score = 28.1 bits (61), Expect = 9.6
Identities = 27/96 (28%), Positives = 40/96 (41%), Gaps = 3/96 (3%)
Query: 62 FVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYARLRANANGSAKGEI 121
F IG P TNY M D G+ V +TV+L A + R LR N ++
Sbjct: 73 FRIGGMCPDTNYLFMGDYVDRGYYSV--ETVTLLVALKMRYPQRITILRGNHESRQITQV 130
Query: 122 IGWQNIELISDRPLETMLKEFKQLKEEYPDRILIAS 157
G+ + E + + K F L + +P L+ S
Sbjct: 131 YGFYD-ECLRKYGNANVWKIFTDLFDYFPLTALVES 165
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.133 0.386
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,135,070
Number of Sequences: 26719
Number of extensions: 381231
Number of successful extensions: 1074
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 1071
Number of HSP's gapped (non-prelim): 13
length of query: 424
length of database: 11,318,596
effective HSP length: 102
effective length of query: 322
effective length of database: 8,593,258
effective search space: 2767029076
effective search space used: 2767029076
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC139706.5


