

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139600.8 + phase: 0
(318 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
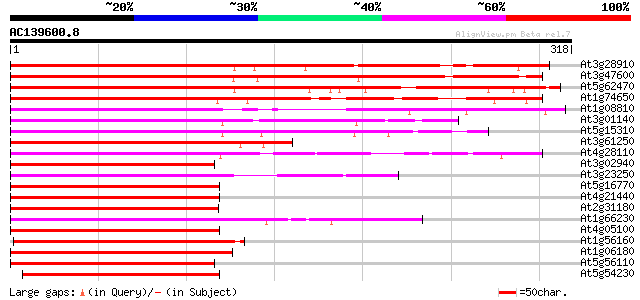
Score E
Sequences producing significant alignments: (bits) Value
At3g28910 MYB family transcription factor (hsr1), putative 337 7e-93
At3g47600 putative transcription factor (MYB94) 313 6e-86
At5g62470 MYB96 transcription factor-like protein 310 7e-85
At1g74650 putative MYB family transcription factor 295 2e-80
At1g08810 transcription factor like protein 252 2e-67
At3g01140 putative Myb-related transcription factor 200 1e-51
At5g15310 myb-related protein - like 194 5e-50
At3g61250 putative transcription factor (MYB17) 185 2e-47
At4g28110 putative transcription factor (MYB41) 182 2e-46
At3g02940 MYB family transcription factor like protein 181 4e-46
At3g23250 myb-related transcription factor like protein 181 5e-46
At5g16770 putative transcription factor (MYB9) 181 6e-46
At4g21440 myb-related protein M4 180 1e-45
At2g31180 putative transcription factor (MYB14) 180 1e-45
At1g66230 myb-related transcription factor, putative 177 5e-45
At4g05100 MYB - like protein 175 3e-44
At1g56160 putative transcription factor (MYB72) 175 3e-44
At1g06180 MYB-related protein 172 2e-43
At5g56110 Atmyb103 (gb|AAD40692.1) 172 3e-43
At5g54230 Myb-related transcription factor-like protein 171 5e-43
>At3g28910 MYB family transcription factor (hsr1), putative
Length = 323
Score = 337 bits (863), Expect = 7e-93
Identities = 182/322 (56%), Positives = 217/322 (66%), Gaps = 28/322 (8%)
Query: 1 MVRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNY 60
MVRPPCCDK GVKKGPWTPEEDIILV+YIQEHGPGNW+AVPTNTGLLRCSKSCRLRWTNY
Sbjct: 1 MVRPPCCDKGGVKKGPWTPEEDIILVTYIQEHGPGNWRAVPTNTGLLRCSKSCRLRWTNY 60
Query: 61 LRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKLKKLE 120
LRPGIKRGNFT+ EEKMI+HLQALLGNRWAAIA+YLPQRTDNDIKNYWNT+LKKKL K+
Sbjct: 61 LRPGIKRGNFTEHEEKMIVHLQALLGNRWAAIASYLPQRTDNDIKNYWNTHLKKKLNKVN 120
Query: 121 TSTSSE---SCLGHDEFSVS----QPIARGQWERRLQTDIHMAKKALSEALQP------E 167
+ E S L S S I+RGQWERRLQTDIH+AKKALSEAL P
Sbjct: 121 QDSHQELDRSSLSSSPSSSSANSNSNISRGQWERRLQTDIHLAKKALSEALSPAVAPIIT 180
Query: 168 KSTSSSNLMLPLESNFSSESSFCSTKPTTQSLSYASSADNIARLLKGWMKNKPKEGSNGN 227
+ ++++ + SS S F T+ T S +YASS +NIA+LLKGW+KN PK ++ +
Sbjct: 181 STVTTTSSSAESRRSTSSASGFLRTQET--STTYASSTENIAKLLKGWVKNSPKTQNSAD 238
Query: 228 NTNVTQNYETSASSEGMEKGSTSVELSETFESLFGYESFDSSNSDSTTLFQDESKPENIG 287
T+ E S +G E + F+S + FD S + E+KP+ G
Sbjct: 239 QIASTEVKEVIKSDDGK-------ECAGAFQS---FSEFDHSYQQAGVSPDHETKPDITG 288
Query: 288 ---EIMPFSMLEKWLLDDGACQ 306
+S+ EKWL +D Q
Sbjct: 289 CCSNQSQWSLFEKWLFEDSGGQ 310
>At3g47600 putative transcription factor (MYB94)
Length = 333
Score = 313 bits (803), Expect = 6e-86
Identities = 175/321 (54%), Positives = 212/321 (65%), Gaps = 26/321 (8%)
Query: 1 MVRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNY 60
M RPPCCDK GVKKGPWTPEEDIILVSYIQEHGPGNW++VPT+TGL RCSKSCRLRWTNY
Sbjct: 1 MGRPPCCDKIGVKKGPWTPEEDIILVSYIQEHGPGNWRSVPTHTGLRRCSKSCRLRWTNY 60
Query: 61 LRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKLKKLE 120
LRPGIKRGNFT+ EEKMI+HLQALLGNRWAAIA+YLP+RTDNDIKNYWNT+LKKKLKK+
Sbjct: 61 LRPGIKRGNFTEHEEKMILHLQALLGNRWAAIASYLPERTDNDIKNYWNTHLKKKLKKMN 120
Query: 121 TSTSS--ESCLGHDEFSVSQP---------IARGQWERRLQTDIHMAKKALSEALQPEKS 169
S S + L + +FS+S ++GQWERRLQTDI+MAK+AL +AL +K
Sbjct: 121 DSCDSTINNGLDNKDFSISNKNTTSHQSSNSSKGQWERRLQTDINMAKQALCDALSIDKP 180
Query: 170 TSSSNLMLP-LESNFSSESSFCSTKPTT-------QSLSYASSADNIARLLKGWMKNKPK 221
+ +N +P L SS SS +T TT S YASSA+NIARLL+ +MK+ PK
Sbjct: 181 QNPTNFSIPDLGYGPSSSSSSTTTTTTTTRNTNPYPSGVYASSAENIARLLQNFMKDTPK 240
Query: 222 EGSNGNNTNVTQNYETSASSEGMEKGSTSVELSETFESLFGYESFDSSNSDSTTLFQDES 281
T+ASS +G + SLF + S D + + D +
Sbjct: 241 TSVPLPVAATEMAITTAASSPSTTEG----DGEGIDHSLFSFNSIDEAEEKPKLIDHDIN 296
Query: 282 KPENIGEIMPFSMLEKWLLDD 302
G + S+ EKWL D+
Sbjct: 297 GLITQGSL---SLFEKWLFDE 314
>At5g62470 MYB96 transcription factor-like protein
Length = 352
Score = 310 bits (794), Expect = 7e-85
Identities = 188/353 (53%), Positives = 224/353 (63%), Gaps = 50/353 (14%)
Query: 1 MVRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNY 60
M RPPCC+K GVKKGPWTPEEDIILVSYIQEHGPGNW++VPT+TGL RCSKSCRLRWTNY
Sbjct: 1 MGRPPCCEKIGVKKGPWTPEEDIILVSYIQEHGPGNWRSVPTHTGLRRCSKSCRLRWTNY 60
Query: 61 LRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKLKKLE 120
LRPGIKRGNFT+ EEK I+HLQALLGNRWAAIA+YLP+RTDNDIKNYWNT+LKKKLKK+
Sbjct: 61 LRPGIKRGNFTEHEEKTIVHLQALLGNRWAAIASYLPERTDNDIKNYWNTHLKKKLKKIN 120
Query: 121 TSTSSE----SCLGHDEFSVSQPIARGQWERRLQTDIHMAKKALSEALQPEK--STSSSN 174
S + S Q +GQWERRLQTDI+MAK+AL EAL +K ST SS+
Sbjct: 121 ESGEEDNDGVSSSNTSSQKNHQSTNKGQWERRLQTDINMAKQALCEALSLDKPSSTLSSS 180
Query: 175 LMLPLE-------SNFSS---------ESSFCSTKPTTQSLS--------YASSADNIAR 210
LP NFSS SS ST TT S + YASSA+NIAR
Sbjct: 181 SSLPTPVITQQNIRNFSSALLDRCYDPSSSSSSTTTTTTSNTTNPYPSGVYASSAENIAR 240
Query: 211 LLKGWMKNKPKEGSNGNNTNVTQNYETSASSEGMEKGSTSVELSETFE-SLFGYESFDSS 269
LL+ +MK+ PK +T + + S G + S E E FE S F + S D +
Sbjct: 241 LLQDFMKDTPKA--------LTLSSSSPVSETGPLTAAVSEEGGEGFEQSFFSFNSMDET 292
Query: 270 N--SDSTTLFQDE-SKPE-----NIGEIM--PFSMLEKWLLDDGACQEKVGLS 312
+ T+ F D+ KPE + G I S+ EKWL D+ + E VG++
Sbjct: 293 QNLTQETSFFHDQVIKPEITMDQDHGLISQGSLSLFEKWLFDEQS-HEMVGMA 344
>At1g74650 putative MYB family transcription factor
Length = 330
Score = 295 bits (755), Expect = 2e-80
Identities = 170/339 (50%), Positives = 210/339 (61%), Gaps = 68/339 (20%)
Query: 1 MVRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNY 60
M RPPCC+K VKKGPWTPEEDIILVSYIQ+HGPGNW++VP NTGLLRCSKSCRLRWTNY
Sbjct: 1 MGRPPCCEKIEVKKGPWTPEEDIILVSYIQQHGPGNWRSVPANTGLLRCSKSCRLRWTNY 60
Query: 61 LRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKL---- 116
LRPGIKRGNFT EEKMIIHLQALLGNRWAAIA+YLPQRTDNDIKNYWNT+LKKKL
Sbjct: 61 LRPGIKRGNFTQPEEKMIIHLQALLGNRWAAIASYLPQRTDNDIKNYWNTHLKKKLVMMK 120
Query: 117 --------KKLETSTSSESCLGHDE--FSVSQPIARGQWERRLQTDIHMAKKALSEALQP 166
K +T SC ++ + +GQWE++LQTDI+MAK+AL +AL
Sbjct: 121 FQNGIINENKTNLATDISSCNNNNNGCNHNKRTTNKGQWEKKLQTDINMAKQALFQALSL 180
Query: 167 EKSTSSSNLMLPLESNFSSESSFCSTKPTTQS-LSYASSADNIARLLKGWMKNKPKEGSN 225
++ +S ++P + + S KP S +YASS DNI++LL+ W + S+
Sbjct: 181 DQPSS----LIPPDPD--------SPKPHHHSTTTYASSTDNISKLLQNWTSS----SSS 224
Query: 226 GNNTNVTQNYETSASSEGMEKGSTSVELSETFESLFGYESFDSSNSDS------------ 273
NT+ N +S+ EG LF + S SSNS+S
Sbjct: 225 KPNTSSVSNNRSSSPGEG---------------GLFDHHSLFSSNSESGSVDEKLNLMSE 269
Query: 274 TTLFQDESKPENIGEIMP----------FSMLEKWLLDD 302
T++F+ ESKP+ E P S++EKWL DD
Sbjct: 270 TSMFKGESKPDIDMEATPTTTTTDDQGSLSLIEKWLFDD 308
>At1g08810 transcription factor like protein
Length = 280
Score = 252 bits (643), Expect = 2e-67
Identities = 152/332 (45%), Positives = 189/332 (56%), Gaps = 72/332 (21%)
Query: 1 MVRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNY 60
M RPPCCDK G+KKGPWTPEEDIILVSYIQEHGPGNW++VPTNTGLLRCSKSCRLRWTNY
Sbjct: 1 MGRPPCCDKIGIKKGPWTPEEDIILVSYIQEHGPGNWRSVPTNTGLLRCSKSCRLRWTNY 60
Query: 61 LRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKLKKLE 120
LRPGIKRGNFT EE MIIHLQALLGN+WA+IA+YLPQRTDNDIKNYWNT+LKKKL K +
Sbjct: 61 LRPGIKRGNFTPHEEGMIIHLQALLGNKWASIASYLPQRTDNDIKNYWNTHLKKKLNKSD 120
Query: 121 TSTSSESCLGHDEFSVSQPIARGQWERRLQTDIHMAKKALSEALQPEKSTSSSNLMLPLE 180
+ DE S S+ IA LQT
Sbjct: 121 S----------DERSRSENIA-------LQTS---------------------------- 135
Query: 181 SNFSSESSFCSTKPTTQSLS-YASSADNIARLLKGWMKNKPKEGSN--------GNNTNV 231
ST+ T S YASS +NI+RLL+GWM+ PK ++ N TN
Sbjct: 136 ----------STRNTINHRSTYASSTENISRLLEGWMRASPKSSTSTTFLEHKMQNRTNN 185
Query: 232 TQNYETSASSEGMEKGSTSVELSETF--ESLFGYESFDSSNSDSTTLFQ-DESKPENIGE 288
++ + +GS S+ + G ++ +++N D D ++
Sbjct: 186 FIDHHSDQFPYEQLQGSWEEGHSKGINGDDDQGIKNSENNNGDDVHHEDGDHEDDDDHNA 245
Query: 289 IMPFSMLEKWLLDD-----GACQEKVGLSEIN 315
P + +EKWLL++ G +E L E++
Sbjct: 246 TPPLTFIEKWLLEETSTTGGQMEEMSHLMELS 277
>At3g01140 putative Myb-related transcription factor
Length = 345
Score = 200 bits (508), Expect = 1e-51
Identities = 119/270 (44%), Positives = 160/270 (59%), Gaps = 23/270 (8%)
Query: 1 MVRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNY 60
M R PCCDK G+KKGPWTPEED L++YI+EHG G+W+++P GL RC KSCRLRWTNY
Sbjct: 1 MGRSPCCDKAGLKKGPWTPEEDQKLLAYIEEHGHGSWRSLPEKAGLQRCGKSCRLRWTNY 60
Query: 61 LRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKLKKL- 119
LRP IKRG FT QEE+ II L ALLGNRW+AIA +LP+RTDN+IKNYWNT+LKK+L K+
Sbjct: 61 LRPDIKRGKFTVQEEQTIIQLHALLGNRWSAIATHLPKRTDNEIKNYWNTHLKKRLIKMG 120
Query: 120 ---------ETSTSSESCLGHDEFSVSQPIARGQWE-RRLQTDIHMAKKALSEALQPEKS 169
+ SS + + ++S QWE RL+ + +A+++ LQ ++
Sbjct: 121 IDPVTHKHKNETLSSSTGQSKNAATLSH---MAQWESARLEAEARLARESKLLHLQHYQN 177
Query: 170 TSSSN-LMLPLESNFSSESSFCSTKPT----TQSLSYASSADNIARLLKGWMKNKPKEGS 224
++ N P + F+ ++S TKP Q L +S + L P + S
Sbjct: 178 NNNLNKSAAPQQHCFTQKTSTNWTKPNQGNGDQQLESPTSTVTFSENLL-MPLGIPTDSS 236
Query: 225 NGNNTNVTQNYETSASSEGMEKGSTSVELS 254
N N N E+SA E STS ++S
Sbjct: 237 RNRNNN---NNESSAMIELAVSSSTSSDVS 263
>At5g15310 myb-related protein - like
Length = 326
Score = 194 bits (493), Expect = 5e-50
Identities = 122/294 (41%), Positives = 166/294 (55%), Gaps = 33/294 (11%)
Query: 1 MVRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNY 60
M R PCCDK G+KKGPWTPEED L++YI+EHG G+W+++P GL RC KSCRLRWTNY
Sbjct: 1 MGRSPCCDKLGLKKGPWTPEEDQKLLAYIEEHGHGSWRSLPEKAGLHRCGKSCRLRWTNY 60
Query: 61 LRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKLKKL- 119
LRP IKRG F QEE+ II L ALLGNRW+AIA +LP+RTDN+IKNYWNT+LKK+L K+
Sbjct: 61 LRPDIKRGKFNLQEEQTIIQLHALLGNRWSAIATHLPKRTDNEIKNYWNTHLKKRLVKMG 120
Query: 120 ----ETSTSSESCLGHDEFSVSQPIA--RGQWE-RRLQTDIHMAKKALSEALQPEKSTSS 172
+E+ L S + I QWE RL+ + +A+++ LQ ++ +S
Sbjct: 121 IDPVTHKPKNETPLSSLGLSKNAAILSHTAQWESARLEAEARLARESKLLHLQHYQTKTS 180
Query: 173 SNLMLPLESNFSSESSFCSTKP---------TTQSLSYASSADNIARLLK------GWMK 217
S S +TKP T ++S++ ++I ++ G
Sbjct: 181 SQPHHHHGFTHKSLLPNWTTKPHEDQQQLESPTSTVSFSEMKESIPAKIEFVGSSTGVTL 240
Query: 218 NKPKEGSNGNNTNVTQNYETSASSEGMEKGSTSVELSETFESLFGYESFDSSNS 271
K E N+T +ET+ EG+E+G T + L G +S D S S
Sbjct: 241 MKEPEHDWINST--MHEFETTQMGEGIEEGFTGL--------LLGGDSIDRSFS 284
>At3g61250 putative transcription factor (MYB17)
Length = 299
Score = 185 bits (470), Expect = 2e-47
Identities = 90/166 (54%), Positives = 119/166 (71%), Gaps = 6/166 (3%)
Query: 1 MVRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNY 60
M R PCCDK G+KKGPWTPEED +LV++I+++G G+W+ +P GLLRC KSCRLRWTNY
Sbjct: 1 MGRTPCCDKIGLKKGPWTPEEDEVLVAHIKKNGHGSWRTLPKLAGLLRCGKSCRLRWTNY 60
Query: 61 LRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKLKKLE 120
LRP IKRG FT EEK++I L A+LGNRWAAIAA LP RTDN+IKN WNT+LKK+L +
Sbjct: 61 LRPDIKRGPFTADEEKLVIQLHAILGNRWAAIAAQLPGRTDNEIKNLWNTHLKKRLLSMG 120
Query: 121 TSTSSESCL---GHDEFSVSQPIAR--GQWE-RRLQTDIHMAKKAL 160
+ L G + + S P R QWE R++ + ++++++
Sbjct: 121 LDPRTHEPLPSYGLAKQAPSSPTTRHMAQWESARVEAEARLSRESM 166
>At4g28110 putative transcription factor (MYB41)
Length = 282
Score = 182 bits (463), Expect = 2e-46
Identities = 121/308 (39%), Positives = 161/308 (51%), Gaps = 35/308 (11%)
Query: 1 MVRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNY 60
M R PCCDK GVKKGPWT EED L+ YI+ HGPGNW+ +P N GL RC KSCRLRWTNY
Sbjct: 1 MGRSPCCDKNGVKKGPWTAEEDQKLIDYIRFHGPGNWRTLPKNAGLHRCGKSCRLRWTNY 60
Query: 61 LRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKLKK-- 118
LRP IKRG F+ +EE+ II L +++GN+W+AIAA LP RTDN+IKN+WNT+++K+L +
Sbjct: 61 LRPDIKRGRFSFEEEETIIQLHSVMGNKWSAIAARLPGRTDNEIKNHWNTHIRKRLVRSG 120
Query: 119 LETSTSSESCLGHDEFSVSQPIARGQWERRLQTDIHMAKKALSEALQPEKSTSSSNLMLP 178
++ T S D S+ + Q + S L P+ +S L+LP
Sbjct: 121 IDPVTHSPRLDLLDLSSLLSALFN-------QPNFSAVATHASSLLNPDVLRLAS-LLLP 172
Query: 179 LESNFSSESSFCSTKPTTQSLSYASSADNIARLLKGWMKNKPKEGSNGNNTNVTQNYETS 238
L++ S T + S SS + V NYETS
Sbjct: 173 LQNPNPVYPSNLDQNLQTPNTSSESSQ------------------PQAETSTVPTNYETS 214
Query: 239 ASSEGMEKGSTSVELSETFESLFGYESFDSSNSDSTTLF---QDESKPE-NIGEIMPFSM 294
S E M V L++ L ESFD + ST + Q+ + E N F +
Sbjct: 215 -SLEPMNARLDDVGLADVLPPL--SESFDLDSLMSTPMSSPRQNSIEAETNSSTFFDFGI 271
Query: 295 LEKWLLDD 302
E ++LDD
Sbjct: 272 PEDFILDD 279
>At3g02940 MYB family transcription factor like protein
Length = 321
Score = 181 bits (460), Expect = 4e-46
Identities = 77/116 (66%), Positives = 99/116 (84%)
Query: 1 MVRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNY 60
M R PCCD+ G+KKGPWTPEED L+++I++HG G+W+A+P GL RC KSCRLRWTNY
Sbjct: 1 MGRSPCCDESGLKKGPWTPEEDQKLINHIRKHGHGSWRALPKQAGLNRCGKSCRLRWTNY 60
Query: 61 LRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKL 116
LRP IKRGNFT +EE+ II+L +LLGN+W++IA +LP RTDN+IKNYWNT+++KKL
Sbjct: 61 LRPDIKRGNFTAEEEQTIINLHSLLGNKWSSIAGHLPGRTDNEIKNYWNTHIRKKL 116
>At3g23250 myb-related transcription factor like protein
Length = 285
Score = 181 bits (459), Expect = 5e-46
Identities = 100/220 (45%), Positives = 128/220 (57%), Gaps = 25/220 (11%)
Query: 1 MVRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNY 60
M R PCC+K G+K+GPWTPEED ILVS+I HG NW+A+P GLLRC KSCRLRW NY
Sbjct: 1 MGRAPCCEKMGLKRGPWTPEEDQILVSFILNHGHSNWRALPKQAGLLRCGKSCRLRWMNY 60
Query: 61 LRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKLKKLE 120
L+P IKRGNFT +EE II L +LGNRW+AIAA LP RTDN+IKN W+T+LKK+L+ +
Sbjct: 61 LKPDIKRGNFTKEEEDAIISLHQILGNRWSAIAAKLPGRTDNEIKNVWHTHLKKRLEDYQ 120
Query: 121 TSTSSESCLGHDEFSVSQPIARGQWERRLQTDIHMAKKALSEALQPEKSTSSSNLMLPLE 180
+ S + K SE++ +++ S L
Sbjct: 121 PAKPKTS------------------------NKKKGTKPKSESVITSSNSTRSESELADS 156
Query: 181 SNFSSESSFCSTKPTTQSLSYASSADNIARLLKGWMKNKP 220
SN S ES F ST P+T +S + + + M NKP
Sbjct: 157 SNPSGESLF-STSPSTSEVSSMTLISHDGYSNEINMDNKP 195
>At5g16770 putative transcription factor (MYB9)
Length = 336
Score = 181 bits (458), Expect = 6e-46
Identities = 78/119 (65%), Positives = 99/119 (82%)
Query: 1 MVRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNY 60
M R PCCD+ G+KKGPWT EED L+ +IQ+HG G+W+A+P GL RC KSCRLRWTNY
Sbjct: 1 MGRSPCCDENGLKKGPWTQEEDDKLIDHIQKHGHGSWRALPKQAGLNRCGKSCRLRWTNY 60
Query: 61 LRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKLKKL 119
LRP IKRGNFT++EE+ II+L +LLGN+W++IA LP RTDN+IKNYWNT+L+KKL ++
Sbjct: 61 LRPDIKRGNFTEEEEQTIINLHSLLGNKWSSIAGNLPGRTDNEIKNYWNTHLRKKLLQM 119
>At4g21440 myb-related protein M4
Length = 350
Score = 180 bits (456), Expect = 1e-45
Identities = 78/119 (65%), Positives = 96/119 (80%)
Query: 1 MVRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNY 60
M R PCC+K G+KKGPWT EED LV YIQ+HG GNW+ +P N GL RC KSCRLRWTNY
Sbjct: 1 MARSPCCEKNGLKKGPWTSEEDQKLVDYIQKHGYGNWRTLPKNAGLQRCGKSCRLRWTNY 60
Query: 61 LRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKLKKL 119
LRP IKRG F+ +EE+ II L + LGN+W+AIAA LP RTDN+IKN+WNT+++KKL ++
Sbjct: 61 LRPDIKRGRFSFEEEETIIQLHSFLGNKWSAIAARLPGRTDNEIKNFWNTHIRKKLLRM 119
>At2g31180 putative transcription factor (MYB14)
Length = 249
Score = 180 bits (456), Expect = 1e-45
Identities = 81/118 (68%), Positives = 96/118 (80%)
Query: 1 MVRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNY 60
M R PCC+K GVK+GPWTPEED IL++YI +G NW+A+P + GLLRC KSCRLRW NY
Sbjct: 1 MGRAPCCEKMGVKRGPWTPEEDQILINYIHLYGHSNWRALPKHAGLLRCGKSCRLRWINY 60
Query: 61 LRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKLKK 118
LRP IKRGNFT QEE+ II+L LGNRW+AIAA LP RTDN+IKN W+T+LKK+L K
Sbjct: 61 LRPDIKRGNFTPQEEQTIINLHESLGNRWSAIAAKLPGRTDNEIKNVWHTHLKKRLSK 118
>At1g66230 myb-related transcription factor, putative
Length = 282
Score = 177 bits (450), Expect = 5e-45
Identities = 103/247 (41%), Positives = 141/247 (56%), Gaps = 15/247 (6%)
Query: 1 MVRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNY 60
M R PCCDK G+KKGPWT EED L+++I +G W+AVP +GLLRC KSCRLRWTNY
Sbjct: 1 MGRQPCCDKVGLKKGPWTAEEDRKLINFILTNGQCCWRAVPKLSGLLRCGKSCRLRWTNY 60
Query: 61 LRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKLKKLE 120
LRP +KRG +D EEKM+I L + LGNRW+ IA++LP RTDN+IKN+WNT++KKKL+K+
Sbjct: 61 LRPDLKRGLLSDYEEKMVIDLHSQLGNRWSKIASHLPGRTDNEIKNHWNTHIKKKLRKMG 120
Query: 121 TSTSSESCLGHDEFSVSQPIARGQ---------WERRLQTDIHMAKKALSEALQPEKSTS 171
+ L E +P+ + Q ER L+ +I + L E+L ++
Sbjct: 121 IDPLTHKPLSIVEKEDEEPLKKLQNNTVPFQETMERPLENNIKNISR-LEESLGDDQ-FM 178
Query: 172 SSNLMLPLES----NFSSESSFCSTKPTTQSLSYASSADNIARLLKGWMKNKPKEGSNGN 227
NL +E S CS + S S +S + N + LK + + GN
Sbjct: 179 EINLEYGVEDVPLIETESLDLICSNSTMSSSTSTSSHSSNDSSFLKDLQFPEFEWSDYGN 238
Query: 228 NTNVTQN 234
+ N N
Sbjct: 239 SNNDNNN 245
>At4g05100 MYB - like protein
Length = 324
Score = 175 bits (444), Expect = 3e-44
Identities = 77/120 (64%), Positives = 97/120 (80%), Gaps = 1/120 (0%)
Query: 1 MVRPPCCDKE-GVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTN 59
M R PCC+K+ G+KKGPWTPEED L+ YI HG GNW+ +P N GL RC KSCRLRWTN
Sbjct: 1 MGRSPCCEKKNGLKKGPWTPEEDQKLIDYINIHGYGNWRTLPKNAGLQRCGKSCRLRWTN 60
Query: 60 YLRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKLKKL 119
YLRP IKRG F+ +EE+ II L +++GN+W+AIAA LP RTDN+IKNYWNT+++K+L K+
Sbjct: 61 YLRPDIKRGRFSFEEEETIIQLHSIMGNKWSAIAARLPGRTDNEIKNYWNTHIRKRLLKM 120
>At1g56160 putative transcription factor (MYB72)
Length = 296
Score = 175 bits (443), Expect = 3e-44
Identities = 76/131 (58%), Positives = 100/131 (76%), Gaps = 2/131 (1%)
Query: 3 RPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNYLR 62
R PCCDK VK+GPW+P+ED+ L+++IQ+HG NW+++P GLLRC KSCRLRW NYLR
Sbjct: 5 RAPCCDKNKVKRGPWSPQEDLTLITFIQKHGHQNWRSLPKLAGLLRCGKSCRLRWINYLR 64
Query: 63 PGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKLKKLETS 122
P +KRGNF+ +EE IIH LGN+W+ IA++LP RTDN+IKN WNT+LKK+L +S
Sbjct: 65 PDVKRGNFSKKEEDAIIHYHQTLGNKWSKIASFLPGRTDNEIKNVWNTHLKKRLTPSSSS 124
Query: 123 TSSESCLGHDE 133
+S S HD+
Sbjct: 125 SSLSST--HDQ 133
>At1g06180 MYB-related protein
Length = 246
Score = 172 bits (437), Expect = 2e-43
Identities = 80/126 (63%), Positives = 94/126 (74%)
Query: 1 MVRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNY 60
M R PCC+K G+KKGPW+ EED IL++YI HG NW+A+P GLLRC KSCRLRW NY
Sbjct: 1 MGRRPCCEKIGLKKGPWSAEEDRILINYISLHGHPNWRALPKLAGLLRCGKSCRLRWINY 60
Query: 61 LRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKLKKLE 120
LRP IKRGNFT EE II L LLGNRW+AIAA LP RTDN+IKN W+T+LKK+L +
Sbjct: 61 LRPDIKRGNFTPHEEDTIISLHQLLGNRWSAIAAKLPGRTDNEIKNVWHTHLKKRLHHSQ 120
Query: 121 TSTSSE 126
+ E
Sbjct: 121 DQNNKE 126
>At5g56110 Atmyb103 (gb|AAD40692.1)
Length = 320
Score = 172 bits (435), Expect = 3e-43
Identities = 75/116 (64%), Positives = 89/116 (76%)
Query: 1 MVRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNY 60
M R PCC+KE VK+G WTPEED L SYI +HG NW+ +P N GL RC KSCRLRWTNY
Sbjct: 1 MGRIPCCEKENVKRGQWTPEEDNKLASYIAQHGTRNWRLIPKNAGLQRCGKSCRLRWTNY 60
Query: 61 LRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKL 116
LRP +K G F++ EE +I+ ++LGNRW+ IAA LP RTDND+KNYWNT LKKKL
Sbjct: 61 LRPDLKHGQFSEAEEHIIVKFHSVLGNRWSLIAAQLPGRTDNDVKNYWNTKLKKKL 116
>At5g54230 Myb-related transcription factor-like protein
Length = 319
Score = 171 bits (433), Expect = 5e-43
Identities = 76/112 (67%), Positives = 90/112 (79%)
Query: 8 DKEGVKKGPWTPEEDIILVSYIQEHGPGNWKAVPTNTGLLRCSKSCRLRWTNYLRPGIKR 67
++ VKKGPWTPEED LV YIQ HGPG W+ +P N GL RC KSCRLRWTNYLRP IKR
Sbjct: 8 EESEVKKGPWTPEEDEKLVGYIQTHGPGKWRTLPKNAGLKRCGKSCRLRWTNYLRPDIKR 67
Query: 68 GNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKLKKL 119
G F+ QEE+ II L LLGN+W+AIA +LP RTDN+IKNYWNT++KKKL ++
Sbjct: 68 GEFSLQEEETIIQLHRLLGNKWSAIAIHLPGRTDNEIKNYWNTHIKKKLLRM 119
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.311 0.128 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,605,691
Number of Sequences: 26719
Number of extensions: 322857
Number of successful extensions: 1248
Number of sequences better than 10.0: 204
Number of HSP's better than 10.0 without gapping: 171
Number of HSP's successfully gapped in prelim test: 34
Number of HSP's that attempted gapping in prelim test: 931
Number of HSP's gapped (non-prelim): 254
length of query: 318
length of database: 11,318,596
effective HSP length: 99
effective length of query: 219
effective length of database: 8,673,415
effective search space: 1899477885
effective search space used: 1899477885
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC139600.8


