

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139600.10 + phase: 0
(81 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
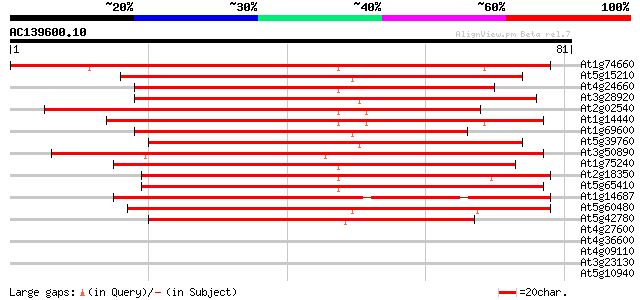
Score E
Sequences producing significant alignments: (bits) Value
At1g74660 unknown protein 94 9e-21
At5g15210 unknown protein 82 6e-17
At4g24660 unknown protein 82 6e-17
At3g28920 unknown protein 79 5e-16
At2g02540 unknown protein 78 7e-16
At1g14440 unknown protein 78 7e-16
At1g69600 hypothetical protein 77 1e-15
At5g39760 unknown protein 76 3e-15
At3g50890 unknown protein 75 4e-15
At1g75240 unknown protein 75 8e-15
At2g18350 unknown protein 74 1e-14
At5g65410 unknown protein 74 1e-14
At1g14687 unknown protein 71 8e-14
At5g60480 putative protein 70 2e-13
At5g42780 unknown protein 70 2e-13
At4g27600 carbohydrate kinase like protein 28 0.81
At4g36600 putative protein 26 3.1
At4g09110 putative protein 26 3.1
At3g23130 superman protein 26 4.0
At5g10940 unknown protein 25 5.3
>At1g74660 unknown protein
Length = 102
Score = 94.4 bits (233), Expect = 9e-21
Identities = 51/101 (50%), Positives = 65/101 (63%), Gaps = 23/101 (22%)
Query: 1 MRKRQVVVRR-----------------SEDPQRSVVKYGECQKNHAANVGGYAVDGCREF 43
M+KRQ+V+++ S + S V+Y ECQKNHAAN+GGYAVDGCREF
Sbjct: 2 MKKRQMVIKQRSRNSNTSSSWTTTSSSSSSSEISNVRYVECQKNHAANIGGYAVDGCREF 61
Query: 44 MPS----TNGSLTCAACGCHRNFHKREV--EVVSESSSPHS 78
M + T +L CAACGCHRNFH++EV EVV E S P++
Sbjct: 62 MAAGVEGTVDALRCAACGCHRNFHRKEVDTEVVCEYSPPNA 102
>At5g15210 unknown protein
Length = 271
Score = 81.6 bits (200), Expect = 6e-17
Identities = 38/64 (59%), Positives = 46/64 (71%), Gaps = 6/64 (9%)
Query: 17 SVVKYGECQKNHAANVGGYAVDGCREFMPSTN------GSLTCAACGCHRNFHKREVEVV 70
+V Y EC KNHAA +GG+A+DGC EFMPS + SLTCAACGCHRNFH+RE +
Sbjct: 52 AVATYKECLKNHAAGIGGHALDGCGEFMPSPSFNSNDPASLTCAACGCHRNFHRREEDPS 111
Query: 71 SESS 74
S S+
Sbjct: 112 SLSA 115
>At4g24660 unknown protein
Length = 220
Score = 81.6 bits (200), Expect = 6e-17
Identities = 35/56 (62%), Positives = 43/56 (76%), Gaps = 4/56 (7%)
Query: 19 VKYGECQKNHAANVGGYAVDGCREFMPS----TNGSLTCAACGCHRNFHKREVEVV 70
++Y EC KNHA N+GG+AVDGC EFMPS T +L CAACGCHRNFH++E E +
Sbjct: 47 IRYRECLKNHAVNIGGHAVDGCCEFMPSGEDGTLDALKCAACGCHRNFHRKETESI 102
>At3g28920 unknown protein
Length = 312
Score = 78.6 bits (192), Expect = 5e-16
Identities = 37/64 (57%), Positives = 43/64 (66%), Gaps = 6/64 (9%)
Query: 19 VKYGECQKNHAANVGGYAVDGCREFMPSTNG------SLTCAACGCHRNFHKREVEVVSE 72
V Y EC KNHAA +GG+A+DGC EFMPS + SL CAACGCHRNFH+RE + S
Sbjct: 50 VTYKECLKNHAAAIGGHALDGCGEFMPSPSSTPSDPTSLKCAACGCHRNFHRRETDDSSA 109
Query: 73 SSSP 76
P
Sbjct: 110 VPPP 113
>At2g02540 unknown protein
Length = 310
Score = 78.2 bits (191), Expect = 7e-16
Identities = 37/67 (55%), Positives = 46/67 (68%), Gaps = 4/67 (5%)
Query: 6 VVVRRSEDPQRSVVKYGECQKNHAANVGGYAVDGCREFMPS-TNGS---LTCAACGCHRN 61
++V + + V+KY EC KNHAA +GG A+DGC EFMPS GS LTC+ C CHRN
Sbjct: 72 IMVTNIKKEKPVVIKYKECLKNHAATMGGNAIDGCGEFMPSGEEGSIEALTCSVCNCHRN 131
Query: 62 FHKREVE 68
FH+RE E
Sbjct: 132 FHRRETE 138
>At1g14440 unknown protein
Length = 312
Score = 78.2 bits (191), Expect = 7e-16
Identities = 38/69 (55%), Positives = 51/69 (73%), Gaps = 6/69 (8%)
Query: 15 QRSVVKYGECQKNHAANVGGYAVDGCREFMPS-TNGS---LTCAACGCHRNFHKREV--E 68
++ ++KY EC KNHAA +GG A DGC EFMPS +GS LTC+AC CHRNFH++EV E
Sbjct: 84 KKPMIKYKECLKNHAAAMGGNATDGCGEFMPSGEDGSIEALTCSACNCHRNFHRKEVEGE 143
Query: 69 VVSESSSPH 77
+ + + SP+
Sbjct: 144 LAATAMSPY 152
>At1g69600 hypothetical protein
Length = 242
Score = 77.4 bits (189), Expect = 1e-15
Identities = 35/54 (64%), Positives = 40/54 (73%), Gaps = 6/54 (11%)
Query: 19 VKYGECQKNHAANVGGYAVDGCREFMPSTN------GSLTCAACGCHRNFHKRE 66
V Y EC KNHAAN+GG+A+DGC EFMPS SL CAACGCHRNFH+R+
Sbjct: 29 VCYKECLKNHAANLGGHALDGCGEFMPSPTATSTDPSSLRCAACGCHRNFHRRD 82
>At5g39760 unknown protein
Length = 334
Score = 76.3 bits (186), Expect = 3e-15
Identities = 35/60 (58%), Positives = 44/60 (73%), Gaps = 6/60 (10%)
Query: 21 YGECQKNHAANVGGYAVDGCREFMPSTNG------SLTCAACGCHRNFHKREVEVVSESS 74
Y EC KNHAA +GG+A+DGC EFMPS + SL CAACGCHRNFH+R+ + ++SS
Sbjct: 56 YKECLKNHAAALGGHALDGCGEFMPSPSSISSDPTSLKCAACGCHRNFHRRDPDNNNDSS 115
>At3g50890 unknown protein
Length = 249
Score = 75.5 bits (184), Expect = 4e-15
Identities = 37/78 (47%), Positives = 48/78 (61%), Gaps = 7/78 (8%)
Query: 7 VVRRSEDPQRSV---VKYGECQKNHAANVGGYAVDGCREFM----PSTNGSLTCAACGCH 59
+ + ++ P+ V KY ECQKNHAA+ GG+ VDGC EFM T G+L CAAC CH
Sbjct: 43 IKKENQKPKTRVDQGAKYRECQKNHAASTGGHVVDGCCEFMAGGEEGTLGALKCAACNCH 102
Query: 60 RNFHKREVEVVSESSSPH 77
R+FH++EV S H
Sbjct: 103 RSFHRKEVYGHRNSKQDH 120
>At1g75240 unknown protein
Length = 309
Score = 74.7 bits (182), Expect = 8e-15
Identities = 35/62 (56%), Positives = 43/62 (68%), Gaps = 4/62 (6%)
Query: 16 RSVVKYGECQKNHAANVGGYAVDGCREFMPS----TNGSLTCAACGCHRNFHKREVEVVS 71
+ V+Y EC KNHAA+VGG DGC EFMPS T +L CAAC CHRNFH++E++ V
Sbjct: 71 KPTVRYRECLKNHAASVGGSVHDGCGEFMPSGEEGTIEALRCAACDCHRNFHRKEMDGVG 130
Query: 72 ES 73
S
Sbjct: 131 SS 132
>At2g18350 unknown protein
Length = 262
Score = 74.3 bits (181), Expect = 1e-14
Identities = 37/66 (56%), Positives = 47/66 (71%), Gaps = 7/66 (10%)
Query: 20 KYGECQKNHAANVGGYAVDGCREFMPS----TNGSLTCAACGCHRNFHKREVE---VVSE 72
+Y ECQKNHAA+ GG+ VDGC EFM S T SL CAAC CHR+FH++E++ VV+
Sbjct: 81 RYRECQKNHAASSGGHVVDGCGEFMSSGEEGTVESLLCAACDCHRSFHRKEIDGLFVVNF 140
Query: 73 SSSPHS 78
+S HS
Sbjct: 141 NSFGHS 146
>At5g65410 unknown protein
Length = 279
Score = 73.9 bits (180), Expect = 1e-14
Identities = 32/62 (51%), Positives = 42/62 (67%), Gaps = 4/62 (6%)
Query: 20 KYGECQKNHAANVGGYAVDGCREFMPS----TNGSLTCAACGCHRNFHKREVEVVSESSS 75
++ EC KN A N+GG+AVDGC EFMP+ T +L CAACGCHRNFH++E+ +
Sbjct: 74 RFRECLKNQAVNIGGHAVDGCGEFMPAGIEGTIDALKCAACGCHRNFHRKELPYFHHAPP 133
Query: 76 PH 77
H
Sbjct: 134 QH 135
>At1g14687 unknown protein
Length = 168
Score = 71.2 bits (173), Expect = 8e-14
Identities = 30/63 (47%), Positives = 43/63 (67%), Gaps = 2/63 (3%)
Query: 16 RSVVKYGECQKNHAANVGGYAVDGCREFMPSTNGSLTCAACGCHRNFHKREVEVVSESSS 75
+S Y EC +NHAA +G YA+DGCRE+ + G L C ACGCHR++H+R ++V+S
Sbjct: 2 QSTCVYRECMRNHAAKLGSYAIDGCREYSQPSTGDL-CVACGCHRSYHRR-IDVISSPQI 59
Query: 76 PHS 78
H+
Sbjct: 60 NHT 62
>At5g60480 putative protein
Length = 191
Score = 70.1 bits (170), Expect = 2e-13
Identities = 35/68 (51%), Positives = 43/68 (62%), Gaps = 7/68 (10%)
Query: 18 VVKYGECQKNHAANVGGYAVDGCREFMPSTN------GSLTCAACGCHRNFHKRE-VEVV 70
VV Y EC KNHA ++GG+A+DGC EF P + SL C ACGCHRNFH+R +
Sbjct: 2 VVLYNECLKNHAVSLGGHALDGCGEFTPKSTTILTDPPSLRCDACGCHRNFHRRSPSDGF 61
Query: 71 SESSSPHS 78
S+ SP S
Sbjct: 62 SQHRSPPS 69
>At5g42780 unknown protein
Length = 242
Score = 70.1 bits (170), Expect = 2e-13
Identities = 31/49 (63%), Positives = 37/49 (75%), Gaps = 2/49 (4%)
Query: 21 YGECQKNHAANVGGYAVDGCREFMPST--NGSLTCAACGCHRNFHKREV 67
Y EC+KNHAA++G A DGC EF+ ST SL CAACGCHRNFH+ E+
Sbjct: 64 YYECRKNHAADIGTTAYDGCGEFVSSTGEEDSLNCAACGCHRNFHREEL 112
>At4g27600 carbohydrate kinase like protein
Length = 471
Score = 28.1 bits (61), Expect = 0.81
Identities = 20/65 (30%), Positives = 28/65 (42%), Gaps = 9/65 (13%)
Query: 6 VVVRRSEDPQR---------SVVKYGECQKNHAANVGGYAVDGCREFMPSTNGSLTCAAC 56
V+V + D QR SVV Y C + A + V+G +P T ++T A
Sbjct: 241 VIVLTTPDAQRTMLAYQGTSSVVNYDSCLASLIAKTNVFVVEGYLFELPDTIRTITKACE 300
Query: 57 GCHRN 61
HRN
Sbjct: 301 EAHRN 305
>At4g36600 putative protein
Length = 347
Score = 26.2 bits (56), Expect = 3.1
Identities = 19/68 (27%), Positives = 28/68 (40%), Gaps = 3/68 (4%)
Query: 11 SEDPQRSVVKYGECQKNHAANVGGYAVDGCREFMPSTNGSLTCAACGCHRNFHKREVEVV 70
+E + V E KN A A D + T + A G +NFHK E E
Sbjct: 22 TETAEDMVRNEAEHAKNAAETAKKMASDAAHDTKDKT---ASWAGWGFRQNFHKAEAEEA 78
Query: 71 SESSSPHS 78
+ES+ ++
Sbjct: 79 AESAKNYA 86
>At4g09110 putative protein
Length = 302
Score = 26.2 bits (56), Expect = 3.1
Identities = 10/23 (43%), Positives = 15/23 (64%)
Query: 50 SLTCAACGCHRNFHKREVEVVSE 72
SL+ AC H+ F++ EVE S+
Sbjct: 62 SLSMVACFLHKTFYRAEVEAASQ 84
>At3g23130 superman protein
Length = 204
Score = 25.8 bits (55), Expect = 4.0
Identities = 20/68 (29%), Positives = 28/68 (40%), Gaps = 14/68 (20%)
Query: 21 YGECQKNHAANVG------GYAVDGC-REFMPSTNGSLTCAACGCHRNFHKREVEVVSES 73
Y CQ++H +G Y C REF + A G H N H+R+ +
Sbjct: 27 YDNCQQDHDYLLGFSWPPRSYTCSFCKREFR-------SAQALGGHMNVHRRDRARLRLQ 79
Query: 74 SSPHSNGT 81
SP S+ T
Sbjct: 80 QSPSSSST 87
>At5g10940 unknown protein
Length = 757
Score = 25.4 bits (54), Expect = 5.3
Identities = 12/52 (23%), Positives = 26/52 (49%)
Query: 6 VVVRRSEDPQRSVVKYGECQKNHAANVGGYAVDGCREFMPSTNGSLTCAACG 57
+++ S+D + ++ Y + H+ + G A C +F+P T+ L + G
Sbjct: 64 LLISGSDDLRINIWNYSSRKLLHSIDTGHTANIFCTKFVPETSDELVVSGAG 115
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.129 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,914,294
Number of Sequences: 26719
Number of extensions: 65462
Number of successful extensions: 211
Number of sequences better than 10.0: 27
Number of HSP's better than 10.0 without gapping: 22
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 174
Number of HSP's gapped (non-prelim): 27
length of query: 81
length of database: 11,318,596
effective HSP length: 57
effective length of query: 24
effective length of database: 9,795,613
effective search space: 235094712
effective search space used: 235094712
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 52 (24.6 bits)
Medicago: description of AC139600.10


