

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139356.2 + phase: 0
(180 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
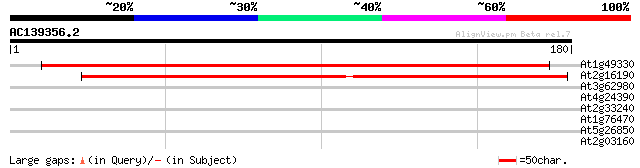
Sequences producing significant alignments: (bits) Value
At1g49330 hypothetical protein 172 9e-44
At2g16190 hypothetical protein 163 4e-41
At3g62980 transport inhibitor response 1 (TIR1) 29 1.2
At4g24390 transport inhibitor response-like protein 28 2.7
At2g33240 putative myosin heavy chain 28 2.7
At1g76470 putative cinnamoyl-CoA reductase 27 5.9
At5g26850 unknown protein 27 7.7
At2g03160 putative kinetechore (Skp1p-like) protein 27 7.7
>At1g49330 hypothetical protein
Length = 331
Score = 172 bits (436), Expect = 9e-44
Identities = 83/163 (50%), Positives = 110/163 (66%)
Query: 11 RRRSAPRSGPIPIPFIWATNHRAKIHNLNHLLQNKIFSITGDVQCKSCQTKFQMPFDLRT 70
R + +S I PF WATN R +I +L +L N+I +ITG+VQC+ C+ +Q+ ++LR
Sbjct: 157 RSTVSKKSDTISPPFPWATNRRGEIQSLEYLESNQITTITGEVQCRHCEKVYQVSYNLRE 216
Query: 71 KFDVISQYLVTNMNTMHDRAPKILMNPRLPRCAHCNQENSVKPVIAEKKKNINWLFLLLG 130
+F + ++ +T M DRA K P RC C +E +VKPVIAE+K INWLFLLLG
Sbjct: 217 RFAEVVKFYLTEKRKMRDRAHKDWAYPEQRRCELCGREKAVKPVIAERKSQINWLFLLLG 276
Query: 131 QMLGCCKLEQLKYFCKYNNHHRTGAKNRVLYLTYFELCQQLDP 173
Q LG C LEQLK FCK++ +HRTGAK+RVLYLTY LC+ L P
Sbjct: 277 QTLGFCTLEQLKNFCKHSKNHRTGAKDRVLYLTYMGLCKMLQP 319
>At2g16190 hypothetical protein
Length = 303
Score = 163 bits (413), Expect = 4e-41
Identities = 76/156 (48%), Positives = 101/156 (64%), Gaps = 2/156 (1%)
Query: 24 PFIWATNHRAKIHNLNHLLQNKIFSITGDVQCKSCQTKFQMPFDLRTKFDVISQYLVTNM 83
P+ WAT KI + L N I I+G V CK+C + ++L KF + Y+ N
Sbjct: 150 PYPWATKKPGKIQSFRDLSSNNINVISGQVHCKTCDRTDTVEYNLEEKFSELYGYIKVNK 209
Query: 84 NTMHDRAPKILMNPRLPRCAHCNQENSVKPVIAEKKKNINWLFLLLGQMLGCCKLEQLKY 143
M RAP P+L C C E +KPV++E+K+ INWLFLLLGQMLGCC L+QL+Y
Sbjct: 210 EEMRHRAPGSWSTPKLIPCRTCKSE--MKPVMSERKEEINWLFLLLGQMLGCCTLDQLRY 267
Query: 144 FCKYNNHHRTGAKNRVLYLTYFELCQQLDPSGPFNV 179
FC+ N+ HRTG+K+RV+Y+TY LC+QLDP GPFN+
Sbjct: 268 FCQLNSKHRTGSKDRVVYITYLSLCKQLDPEGPFNL 303
>At3g62980 transport inhibitor response 1 (TIR1)
Length = 594
Score = 29.3 bits (64), Expect = 1.2
Identities = 20/51 (39%), Positives = 28/51 (54%), Gaps = 4/51 (7%)
Query: 128 LLGQMLGCCKLEQLKYFCKY--NNHHRTGAKNRVLYLTYFELCQQLDPSGP 176
L+ +GC KLE + YFC+ N T A+NR +T F LC ++P P
Sbjct: 364 LVSVSMGCPKLESVLYFCRQMTNAALITIARNRP-NMTRFRLC-IIEPKAP 412
>At4g24390 transport inhibitor response-like protein
Length = 623
Score = 28.1 bits (61), Expect = 2.7
Identities = 15/36 (41%), Positives = 17/36 (46%), Gaps = 1/36 (2%)
Query: 134 GCCKLEQLKYFCK-YNNHHRTGAKNRVLYLTYFELC 168
GC KLE + YFC+ N T LT F LC
Sbjct: 416 GCRKLESILYFCQNMTNGAVTAMSENCPQLTVFRLC 451
>At2g33240 putative myosin heavy chain
Length = 1611
Score = 28.1 bits (61), Expect = 2.7
Identities = 13/51 (25%), Positives = 25/51 (48%)
Query: 42 LQNKIFSITGDVQCKSCQTKFQMPFDLRTKFDVISQYLVTNMNTMHDRAPK 92
L ++ G +CQTK PF + + +V+ +V +N +H + P+
Sbjct: 22 LDGEVVEANGQEIKVNCQTKTVSPFSPKQRDNVLVLKVVAKVNAVHPKDPE 72
>At1g76470 putative cinnamoyl-CoA reductase
Length = 317
Score = 26.9 bits (58), Expect = 5.9
Identities = 10/20 (50%), Positives = 14/20 (70%)
Query: 107 QENSVKPVIAEKKKNINWLF 126
+E V+P+ AEK KN+ W F
Sbjct: 275 KEKEVRPLSAEKLKNLGWKF 294
>At5g26850 unknown protein
Length = 970
Score = 26.6 bits (57), Expect = 7.7
Identities = 18/50 (36%), Positives = 28/50 (56%), Gaps = 2/50 (4%)
Query: 96 NP-RLPRCAHCNQENSVKPVIAEKKKNINWLFLLLGQMLGCCKLEQLKYF 144
NP R+P+ A +E K + +E+ K IN + +ML CK +Q+ YF
Sbjct: 64 NPIRIPKIAKFLEERCYKDLRSEQMKFINIVTEAYNKMLCHCK-DQMAYF 112
>At2g03160 putative kinetechore (Skp1p-like) protein
Length = 200
Score = 26.6 bits (57), Expect = 7.7
Identities = 23/73 (31%), Positives = 30/73 (40%), Gaps = 8/73 (10%)
Query: 61 KFQMPFDLRTKFDVISQYLVTNMNTMHDRAPKIL------MNPRLPRCAHCNQENSVKPV 114
KF FD++T FD+I N+ + D K + M P R N EN P
Sbjct: 118 KFMKDFDIKTIFDIILAANYLNVQGLFDLCSKTIADYIKDMTPEEVR-ELFNIENDFTPE 176
Query: 115 IAEKKKNIN-WLF 126
E +N N W F
Sbjct: 177 EEEAIRNENAWTF 189
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.325 0.138 0.449
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,272,159
Number of Sequences: 26719
Number of extensions: 167277
Number of successful extensions: 458
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 453
Number of HSP's gapped (non-prelim): 8
length of query: 180
length of database: 11,318,596
effective HSP length: 93
effective length of query: 87
effective length of database: 8,833,729
effective search space: 768534423
effective search space used: 768534423
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC139356.2


