

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139354.8 + phase: 0 /pseudo
(481 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
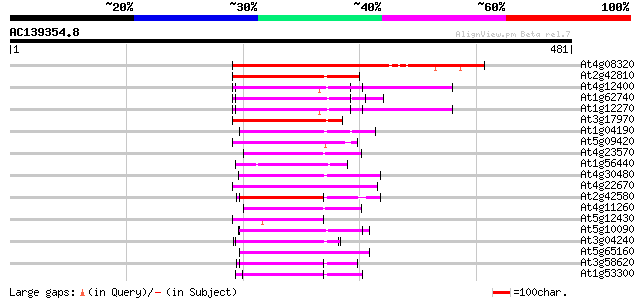
Score E
Sequences producing significant alignments: (bits) Value
At4g08320 unknown protein 228 7e-60
At2g42810 phosphoprotein phosphatase like protein 88 1e-17
At4g12400 stress-induced protein sti1 -like protein 80 2e-15
At1g62740 TPR-repeat protein 74 2e-13
At1g12270 putative protein 70 2e-12
At3g17970 phosphoprotein phosphatase, putative 65 6e-11
At1g04190 unknown protein 64 1e-10
At5g09420 putative subunit of TOC complex 63 4e-10
At4g23570 SGT1a 60 3e-09
At1g56440 hypothetical protein 60 3e-09
At4g30480 unknown protein 59 5e-09
At4g22670 HSP associated protein like 59 5e-09
At2g42580 unknown protein 59 8e-09
At4g11260 Sgt1 like protein 57 2e-08
At5g12430 putative protein 56 4e-08
At5g10090 putative protein 55 7e-08
At3g04240 O-linked GlcNAc transferase like protein 55 7e-08
At5g65160 unknown protein 54 1e-07
At3g58620 putative protein 53 4e-07
At1g53300 unknown protein 51 1e-06
>At4g08320 unknown protein
Length = 266
Score = 228 bits (580), Expect = 7e-60
Identities = 126/224 (56%), Positives = 162/224 (72%), Gaps = 12/224 (5%)
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
AMQS Y +A+ELY+ AIA+ +K+AV+YCNRAAAYTQIN +EAI+D L+SIEIDPNYSK
Sbjct: 25 AMQSNLYLEAVELYSFAIALTDKNAVFYCNRAAAYTQINMCSEAIKDCLKSIEIDPNYSK 84
Query: 252 AYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRADHNQNS 311
AYSRLGLAYYAQG Y +AI+KGFKKAL LDP+NESVKENIRVAE K+ EE+ R +QN+
Sbjct: 85 AYSRLGLAYYAQGKYAEAIEKGFKKALLLDPHNESVKENIRVAEQKIREEQQRQRRSQNT 144
Query: 312 RSQEFQNHYARGSRSHAAPASFGSMPFNPSNLASMFM-AAANGGQGSHSQEGQ----EDA 366
+ Q G + P+ F SMP NP +L SMFM A N G+HS+ + D
Sbjct: 145 STFYTQEPEMGGGQ--GIPSQF-SMPVNP-DLMSMFMNMAGNTFPGNHSRNNEGGAGGDG 200
Query: 367 NSSGANEPEIRFGGNVNVN---QDQIPQELRGAFQSVMHMFSGN 407
+GA+EPEI GGN+N++ +Q+P++L GA +S+M MF G+
Sbjct: 201 TRNGADEPEIHVGGNINIDLGGTEQMPEDLSGALRSMMQMFGGS 244
>At2g42810 phosphoprotein phosphatase like protein
Length = 484
Score = 87.8 bits (216), Expect = 1e-17
Identities = 43/109 (39%), Positives = 67/109 (61%), Gaps = 1/109 (0%)
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A + +Y AI+LY AI + +AVY+ NRA A+T++ Y AIQD+ ++IE+D YSK
Sbjct: 23 AFKGHKYSSAIDLYTKAIELNSNNAVYWANRAFAHTKLEEYGSAIQDASKAIEVDSRYSK 82
Query: 252 AYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLME 300
Y R G AY A G ++DA+ K F++ +L PN+ ++ E +M+
Sbjct: 83 GYYRRGAAYLAMGKFKDAL-KDFQQVKRLSPNDPDATRKLKECEKAVMK 130
>At4g12400 stress-induced protein sti1 -like protein
Length = 558
Score = 80.5 bits (197), Expect = 2e-15
Identities = 39/111 (35%), Positives = 66/111 (59%), Gaps = 1/111 (0%)
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A S Y AI + AI + + + Y NR+A+Y ++RY EA+ D+ ++IE+ P++SK
Sbjct: 12 AFSSGDYATAITHFTEAINLSPTNHILYSNRSASYASLHRYEEALSDAKKTIELKPDWSK 71
Query: 252 AYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEER 302
YSRLG A+ + +A+D +KK L++DP+NE +K + A + +
Sbjct: 72 GYSRLGAAFIGLSKFDEAVD-SYKKGLEIDPSNEMLKSGLADASRSRVSSK 121
Score = 60.8 bits (146), Expect = 2e-09
Identities = 30/99 (30%), Positives = 57/99 (57%), Gaps = 1/99 (1%)
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAY 253
+ ++Y +A++ Y+ AI Y NRAA YT++ E ++D+ + IE+DP+++K Y
Sbjct: 381 KEQKYPEAVKHYSEAIKRNPNDVRAYSNRAACYTKLGALPEGLKDAEKCIELDPSFTKGY 440
Query: 254 SRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIR 292
SR G + Y A++ +++ L+ DP N+ + +R
Sbjct: 441 SRKGAIQFFMKEYDKAMET-YQEGLKHDPKNQEFLDGVR 478
Score = 42.4 bits (98), Expect = 6e-04
Identities = 44/201 (21%), Positives = 86/201 (41%), Gaps = 14/201 (6%)
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A + K + A+E Y A+ + ++ Y NRAA Y ++ +Y E I+D +++E
Sbjct: 240 AYKKKDFGRAVEHYTKAMELDDEDISYLTNRAAVYLEMGKYEECIEDCDKAVERGRELRS 299
Query: 252 AYSRLGLAYYAQG-----------NYRDAIDKGFKKALQLDPNNESVKE-NIRVAEHKLM 299
+ + A +G ++ AI+ F+KAL N +++K+ N K +
Sbjct: 300 DFKMIARALTRKGSALVKMARCSKDFEPAIET-FQKALTEHRNPDTLKKLNDAEKVKKEL 358
Query: 300 EERHRADHNQNSRSQEFQNHYARGSRSHAAPASFG-SMPFNPSNLASMFMAAANGGQGSH 358
E++ D +E N + + + A + ++ NP+++ + AA +
Sbjct: 359 EQQEYFDPTIAEEEREKGNGFFKEQKYPEAVKHYSEAIKRNPNDVRAYSNRAACYTKLGA 418
Query: 359 SQEGQEDANSSGANEPEIRFG 379
EG +DA +P G
Sbjct: 419 LPEGLKDAEKCIELDPSFTKG 439
>At1g62740 TPR-repeat protein
Length = 571
Score = 73.9 bits (180), Expect = 2e-13
Identities = 37/114 (32%), Positives = 66/114 (57%), Gaps = 3/114 (2%)
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A S + A+ + AI + + V + NR+AA+ +N Y EA+ D+ +++E+ P++ K
Sbjct: 12 AFSSGDFNSAVNHFTDAINLTPTNHVLFSNRSAAHASLNHYDEALSDAKKTVELKPDWGK 71
Query: 252 AYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRA 305
YSRLG A+ + +A++ + K L++DP+NE +K + A+ K R RA
Sbjct: 72 GYSRLGAAHLGLNQFDEAVE-AYSKGLEIDPSNEGLKSGL--ADAKASASRSRA 122
Score = 66.6 bits (161), Expect = 3e-11
Identities = 33/99 (33%), Positives = 57/99 (57%), Gaps = 1/99 (1%)
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAY 253
+ ++Y DA+ Y AI K Y NRAA YT++ E ++D+ + IE+DP + K Y
Sbjct: 394 KEQKYPDAVRHYTEAIKRNPKDPRAYSNRAACYTKLGAMPEGLKDAEKCIELDPTFLKGY 453
Query: 254 SRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIR 292
SR G + Y +A++ ++K L+ DPNN+ + + ++
Sbjct: 454 SRKGAVQFFMKEYDNAMET-YQKGLEHDPNNQELLDGVK 491
Score = 43.1 bits (100), Expect = 3e-04
Identities = 30/129 (23%), Positives = 61/129 (47%), Gaps = 5/129 (3%)
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A + K + AI+ Y+ A+ I ++ Y NRAA + ++ +Y E I+D +++E
Sbjct: 253 AYKKKDFETAIQHYSTAMEIDDEDISYITNRAAVHLEMGKYDECIKDCDKAVERGRELRS 312
Query: 252 AYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRADHNQNS 311
Y + A +G A+ K K + +P ++ ++ + EH+ E R + + +
Sbjct: 313 DYKMVAKALTRKGT---ALGKMAKVSKDYEPVIQTYQK--ALTEHRNPETLKRLNEAERA 367
Query: 312 RSQEFQNHY 320
+ + Q Y
Sbjct: 368 KKELEQQEY 376
Score = 28.5 bits (62), Expect = 8.6
Identities = 17/61 (27%), Positives = 28/61 (45%)
Query: 219 YCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAYYAQGNYRDAIDKGFKKAL 278
Y AA+ +N++ EA++ + +EIDP+ S L A + R + F A
Sbjct: 73 YSRLGAAHLGLNQFDEAVEAYSKGLEIDPSNEGLKSGLADAKASASRSRASAPNPFGDAF 132
Query: 279 Q 279
Q
Sbjct: 133 Q 133
>At1g12270 putative protein
Length = 572
Score = 70.5 bits (171), Expect = 2e-12
Identities = 37/111 (33%), Positives = 62/111 (55%), Gaps = 1/111 (0%)
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A S + AI + AIA+ + V + NR+AA+ +++Y EA+ D+ +I++ P + K
Sbjct: 12 AFSSGDFTTAINHFTEAIALAPTNHVLFSNRSAAHASLHQYAEALSDAKETIKLKPYWPK 71
Query: 252 AYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEER 302
YSRLG A+ + A+ +KK L +DP NE++K + AE + R
Sbjct: 72 GYSRLGAAHLGLNQFELAV-TAYKKGLDVDPTNEALKSGLADAEASVARSR 121
Score = 59.3 bits (142), Expect = 5e-09
Identities = 30/99 (30%), Positives = 57/99 (57%), Gaps = 1/99 (1%)
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAY 253
+ ++Y +AI+ Y AI Y NRAA+YT++ E ++D+ + IE+DP +SK Y
Sbjct: 395 KEQKYPEAIKHYTEAIKRNPNDHKAYSNRAASYTKLGAMPEGLKDAEKCIELDPTFSKGY 454
Query: 254 SRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIR 292
SR + Y +A++ ++ L+ DP+N+ + + ++
Sbjct: 455 SRKAAVQFFLKEYDNAMET-YQAGLEHDPSNQELLDGVK 492
Score = 48.5 bits (114), Expect = 8e-06
Identities = 48/201 (23%), Positives = 85/201 (41%), Gaps = 14/201 (6%)
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A + K + AI+ Y+ AI I ++ Y NRAA Y ++ +Y E I+D +++E
Sbjct: 254 AYKKKDFETAIQHYSTAIEIDDEDISYLTNRAAVYLEMGKYNECIEDCNKAVERGRELRS 313
Query: 252 AYSRLGLAYYAQG-----------NYRDAIDKGFKKALQLDPNNESVKE-NIRVAEHKLM 299
Y + A +G +Y AI+ F+KAL N +++K N K
Sbjct: 314 DYKMVARALTRKGTALTKMAKCSKDYEPAIE-AFQKALTEHRNPDTLKRLNDAERAKKEW 372
Query: 300 EERHRADHNQNSRSQEFQNHYARGSRSHAAPASF-GSMPFNPSNLASMFMAAANGGQGSH 358
E++ D +E N + + + A + ++ NP++ + AA+ +
Sbjct: 373 EQKQYFDPKLGDEEREKGNDFFKEQKYPEAIKHYTEAIKRNPNDHKAYSNRAASYTKLGA 432
Query: 359 SQEGQEDANSSGANEPEIRFG 379
EG +DA +P G
Sbjct: 433 MPEGLKDAEKCIELDPTFSKG 453
>At3g17970 phosphoprotein phosphatase, putative
Length = 143
Score = 65.5 bits (158), Expect = 6e-11
Identities = 37/94 (39%), Positives = 57/94 (60%), Gaps = 1/94 (1%)
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A + K + AI LY+ AI + + +A YY NRAAAY ++ + +A +D ++I +D K
Sbjct: 38 AFKEKLWQKAIGLYSEAIKLSDNNATYYSNRAAAYLELGGFLQAEEDCTKAITLDKKNVK 97
Query: 252 AYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNE 285
AY R G A G+ + AI+ F+ AL L+PNN+
Sbjct: 98 AYLRRGTAREMLGDCKGAIE-DFRYALVLEPNNK 130
>At1g04190 unknown protein
Length = 328
Score = 64.3 bits (155), Expect = 1e-10
Identities = 38/116 (32%), Positives = 62/116 (52%), Gaps = 2/116 (1%)
Query: 198 YFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLG 257
+ A LY AI + +A Y NRAAA+ + + ++A+ D+ +I+++P + K Y R G
Sbjct: 31 FLKAAALYTQAIKLDPSNATLYSNRAAAFLSLVKLSKALADAETTIKLNPQWEKGYFRKG 90
Query: 258 LAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRADHNQNSRS 313
A Y DA+ F+ ALQ +P + V I+ +L +E+ RA +N RS
Sbjct: 91 CVLEAMEKYEDAL-AAFEMALQYNPQSTEVSRKIK-RLGQLQKEKQRAQELENLRS 144
>At5g09420 putative subunit of TOC complex
Length = 616
Score = 62.8 bits (151), Expect = 4e-10
Identities = 41/119 (34%), Positives = 62/119 (51%), Gaps = 16/119 (13%)
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A + KQ+ A+ Y AI + +A YYCNRAAA+ ++ + +A QD +++ ID K
Sbjct: 498 AYKGKQWNKAVNFYTEAIKLNGANATYYCNRAAAFLELCCFQQAEQDCTKAMLIDKKNVK 557
Query: 252 AYSRLGLAYYAQGNYRDA------------IDKGFKKALQLDPNNESVKENIRVAEHKL 298
AY R G A + Y++A I F+ AL L+P N++ K VAE +L
Sbjct: 558 AYLRRGTARESLVRYKEAAAGYWSVTLWLIISADFRHALVLEPQNKTAK----VAEKRL 612
>At4g23570 SGT1a
Length = 350
Score = 60.1 bits (144), Expect = 3e-09
Identities = 33/101 (32%), Positives = 55/101 (53%), Gaps = 1/101 (0%)
Query: 201 AIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAY 260
A++LY+ AI + A ++ +RA AY ++ +TEA+ D+ ++IE+DP+ +KAY R G A
Sbjct: 21 AVDLYSKAIDLDPNCAEFFADRAQAYIKLESFTEAVADANKAIELDPSLTKAYLRKGTAC 80
Query: 261 YAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEE 301
YR A +K + P+ K+ I + EE
Sbjct: 81 MKLEEYRTA-KTALEKGASITPSESKFKKLIDECNFLITEE 120
>At1g56440 hypothetical protein
Length = 476
Score = 60.1 bits (144), Expect = 3e-09
Identities = 35/96 (36%), Positives = 58/96 (59%), Gaps = 2/96 (2%)
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAY 253
+ K++ +AI+ Y+ +IA+ +AV Y NRA AY +I RY EA D ++ +D Y KAY
Sbjct: 96 KQKKFNEAIDCYSRSIAL-SPNAVTYANRAMAYLKIKRYREAEVDCTEALNLDDRYIKAY 154
Query: 254 SRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKE 289
SR A G ++A + + AL+L+P ++ +K+
Sbjct: 155 SRRATARKELGMIKEAKEDA-EFALRLEPESQELKK 189
>At4g30480 unknown protein
Length = 277
Score = 59.3 bits (142), Expect = 5e-09
Identities = 33/122 (27%), Positives = 63/122 (51%), Gaps = 1/122 (0%)
Query: 197 QYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRL 256
+Y A+EL E ++ Y NR + ++ + E I++ +++E++P Y+KA R
Sbjct: 127 KYAFALELVQELPESIELRSICYLNRGVCFLKLGKCEETIKECTKALELNPTYNKALVRR 186
Query: 257 GLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRADHNQNSRSQEF 316
A+ ++ DA+ KK L+LDP+N+ ++ IR E E+R + ++ +E
Sbjct: 187 AEAHEKLEHFEDAV-TDLKKILELDPSNDQARKGIRRLEPLAAEKREKMKEEAITKLKEM 245
Query: 317 QN 318
N
Sbjct: 246 GN 247
>At4g22670 HSP associated protein like
Length = 441
Score = 59.3 bits (142), Expect = 5e-09
Identities = 35/125 (28%), Positives = 61/125 (48%), Gaps = 1/125 (0%)
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A+ + +AIE AI + SA+ Y NRA+ Y ++ + AI+D+ ++EI+P+ +K
Sbjct: 133 ALSEGNFDEAIEHLTRAITLNPTSAIMYGNRASVYIKLKKPNAAIRDANAALEINPDSAK 192
Query: 252 AYSRLGLAYYAQGNYRDAI-DKGFKKALQLDPNNESVKENIRVAEHKLMEERHRADHNQN 310
Y G+A G + +A D + D +V + + HKL E R + D +
Sbjct: 193 GYKSRGMARAMLGEWAEAAKDLHLASTIDYDEEISAVLKKVEPNAHKLEEHRRKYDRLRK 252
Query: 311 SRSQE 315
R +
Sbjct: 253 EREDK 257
>At2g42580 unknown protein
Length = 691
Score = 58.5 bits (140), Expect = 8e-09
Identities = 29/72 (40%), Positives = 44/72 (60%)
Query: 198 YFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLG 257
+ +A+ LY+ AI I +A Y NRAAA T + R EA+++ L ++ IDP+YS+A+ RL
Sbjct: 236 FSEALSLYDRAILISPGNAAYRSNRAAALTALRRLGEAVKECLEAVRIDPSYSRAHQRLA 295
Query: 258 LAYYAQGNYRDA 269
Y G +A
Sbjct: 296 SLYLRLGEAENA 307
Score = 51.6 bits (122), Expect = 9e-07
Identities = 32/124 (25%), Positives = 64/124 (50%), Gaps = 7/124 (5%)
Query: 195 SKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYS 254
S ++ +A Y + + ++V YCNRAA + ++ + ++++D +++ P+Y KA
Sbjct: 471 SGRFSEACVAYGDGLKQDDSNSVLYCNRAACWYKLGLWEKSVEDCNHALKSQPSYIKALL 530
Query: 255 RLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRADHNQNSRSQ 314
R +Y G + DA+ K ++ + P + V E++ A+ LM + +Q S+S
Sbjct: 531 RRAASYGKLGRWEDAV-KDYEFLRRELPGDSEVAESLERAKTVLM------NRSQESKSL 583
Query: 315 EFQN 318
F N
Sbjct: 584 GFNN 587
>At4g11260 Sgt1 like protein
Length = 358
Score = 57.0 bits (136), Expect = 2e-08
Identities = 32/101 (31%), Positives = 55/101 (53%), Gaps = 1/101 (0%)
Query: 201 AIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAY 260
A++LY+ AI + A ++ +RA A +I+ +TEA+ D+ ++IE++P +KAY R G A
Sbjct: 21 AVDLYSKAIDLDPNCAAFFADRAQANIKIDNFTEAVVDANKAIELEPTLAKAYLRKGTAC 80
Query: 261 YAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEE 301
Y A +K + PN K+ I + ++ EE
Sbjct: 81 MKLEEYSTA-KAALEKGASVAPNEPKFKKMIDECDLRIAEE 120
>At5g12430 putative protein
Length = 1187
Score = 56.2 bits (134), Expect = 4e-08
Identities = 32/82 (39%), Positives = 48/82 (58%), Gaps = 4/82 (4%)
Query: 192 AMQSKQYFDAIELYNCAIAIYEKS----AVYYCNRAAAYTQINRYTEAIQDSLRSIEIDP 247
A QS ++ +A+E Y A+A +S AV +CNRAAAY + ++++AI D +I +D
Sbjct: 911 AFQSGRHTEAVEHYTAALACNVESRPFTAVCFCNRAAAYKALGQFSDAIADCSLAIALDQ 970
Query: 248 NYSKAYSRLGLAYYAQGNYRDA 269
NYSKA SR + +Y A
Sbjct: 971 NYSKAISRRATLFEMIRDYGQA 992
Score = 39.7 bits (91), Expect = 0.004
Identities = 24/61 (39%), Positives = 29/61 (47%), Gaps = 1/61 (1%)
Query: 219 YCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAYYAQGNYRDAIDKGFKKAL 278
Y NRAA + R EAI D + ID N+ K R Y + G DA + FKK L
Sbjct: 655 YSNRAATRMALGRMREAIADCTMASSIDSNFLKVQVRAANCYLSLGEIEDA-SRYFKKCL 713
Query: 279 Q 279
Q
Sbjct: 714 Q 714
>At5g10090 putative protein
Length = 594
Score = 55.5 bits (132), Expect = 7e-08
Identities = 33/111 (29%), Positives = 55/111 (48%)
Query: 198 YFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLG 257
+ +A+ LY AI+I K A Y N++AA T + R EA+ + +I IDP+Y +A+ RL
Sbjct: 253 FAEALALYEAAISIDPKKASYRSNKSAALTALGRILEAVFECREAIRIDPHYHRAHHRLA 312
Query: 258 LAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRADHN 308
Y G ++I + D + S + ++ +K E + D N
Sbjct: 313 NLYLRLGEVENSIYHFKHAGPEADQEDISKAKMVQTHLNKCTEAKRLRDWN 363
Score = 43.5 bits (101), Expect = 3e-04
Identities = 24/106 (22%), Positives = 55/106 (51%), Gaps = 1/106 (0%)
Query: 197 QYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRL 256
++ +A Y + +++V CNRAA +++ ++ A++D+ ++ + P Y+KA R
Sbjct: 486 RFQEACTAYGEGLDHDSRNSVLLCNRAACLSKMGQFDRAVEDTSAALAVRPGYTKARLRR 545
Query: 257 GLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEER 302
GN+ A+ ++ + P +E V + + A+ +L++ R
Sbjct: 546 ADCNAKLGNWESAVG-DYEILRKETPEDEEVIKGLSEAQKQLVKRR 590
Score = 28.9 bits (63), Expect = 6.6
Identities = 30/115 (26%), Positives = 46/115 (39%), Gaps = 11/115 (9%)
Query: 219 YCNRAAAYTQINRYTEAIQDSLRSIEIDPNYS-KAYSRLGLA---------YYAQGNYRD 268
Y +A A+ + R+ EA R D S K Y +G A + A G + +
Sbjct: 384 YALQAEAFLKTYRHQEADDALSRCPVFDGEMSTKYYGSIGYAGFLVVWAQVHMASGRFVE 443
Query: 269 AIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRADHNQNSRSQEFQNHYARG 323
A++ ++A +LD NN V +R A+ D + R QE Y G
Sbjct: 444 AVE-AIQRAGKLDGNNREVSMVLRRAQAVTAARSRGNDFFKAGRFQEACTAYGEG 497
>At3g04240 O-linked GlcNAc transferase like protein
Length = 977
Score = 55.5 bits (132), Expect = 7e-08
Identities = 34/90 (37%), Positives = 47/90 (51%), Gaps = 1/90 (1%)
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAY 253
Q Y DAI YN + I +A NR Y +I R TEAIQD + +I P ++A+
Sbjct: 405 QQGNYSDAISCYNEVLRIDPLAADALVNRGNTYKEIGRVTEAIQDYMHAINFRPTMAEAH 464
Query: 254 SRLGLAYYAQGNYRDAIDKGFKKALQLDPN 283
+ L AY G+ AI +K+AL L P+
Sbjct: 465 ANLASAYKDSGHVEAAI-TSYKQALLLRPD 493
Score = 42.7 bits (99), Expect = 4e-04
Identities = 27/90 (30%), Positives = 43/90 (47%), Gaps = 1/90 (1%)
Query: 193 MQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKA 252
M+S A++ Y A+ + Y N Y + R TEAI ++++ PN + A
Sbjct: 234 MESGDLNRALQYYKEAVKLKPAFPDAYLNLGNVYKALGRPTEAIMCYQHALQMRPNSAMA 293
Query: 253 YSRLGLAYYAQGNYRDAIDKGFKKALQLDP 282
+ + YY QG AI + +K+AL DP
Sbjct: 294 FGNIASIYYEQGQLDLAI-RHYKQALSRDP 322
Score = 40.0 bits (92), Expect = 0.003
Identities = 25/92 (27%), Positives = 47/92 (50%), Gaps = 1/92 (1%)
Query: 200 DAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLA 259
+AI Y A+ + SA+ + N A+ Y + + AI+ +++ DP + +AY+ LG A
Sbjct: 275 EAIMCYQHALQMRPNSAMAFGNIASIYYEQGQLDLAIRHYKQALSRDPRFLEAYNNLGNA 334
Query: 260 YYAQGNYRDAIDKGFKKALQLDPNNESVKENI 291
G +A+ + + + L L PN+ N+
Sbjct: 335 LKDIGRVDEAV-RCYNQCLALQPNHPQAMANL 365
Score = 37.7 bits (86), Expect = 0.014
Identities = 28/89 (31%), Positives = 46/89 (51%), Gaps = 1/89 (1%)
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAY 253
Q ++Y I A+ I + A Y N A A+ + AI+ L +IE+ PN++ A+
Sbjct: 99 QLQEYDMCIARNEEALRIQPQFAECYGNMANAWKEKGDTDRAIRYYLIAIELRPNFADAW 158
Query: 254 SRLGLAYYAQGNYRDAIDKGFKKALQLDP 282
S L AY +G +A + ++AL L+P
Sbjct: 159 SNLASAYMRKGRLSEA-TQCCQQALSLNP 186
Score = 35.8 bits (81), Expect = 0.054
Identities = 25/82 (30%), Positives = 42/82 (50%), Gaps = 1/82 (1%)
Query: 201 AIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAY 260
AI Y AI + A + N A+AY + R +EA Q +++ ++P A+S LG
Sbjct: 140 AIRYYLIAIELRPNFADAWSNLASAYMRKGRLSEATQCCQQALSLNPLLVDAHSNLGNLM 199
Query: 261 YAQGNYRDAIDKGFKKALQLDP 282
AQG +A + +A+++ P
Sbjct: 200 KAQGLIHEAY-SCYLEAVRIQP 220
Score = 35.4 bits (80), Expect = 0.070
Identities = 21/83 (25%), Positives = 36/83 (43%), Gaps = 1/83 (1%)
Query: 200 DAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLA 259
+A+ YN +A+ N Y + N A ++ + S ++ L +
Sbjct: 343 EAVRCYNQCLALQPNHPQAMANLGNIYMEWNMMGPASSLFKATLAVTTGLSAPFNNLAII 402
Query: 260 YYAQGNYRDAIDKGFKKALQLDP 282
Y QGNY DAI + + L++DP
Sbjct: 403 YKQQGNYSDAI-SCYNEVLRIDP 424
Score = 30.0 bits (66), Expect = 3.0
Identities = 23/94 (24%), Positives = 37/94 (38%), Gaps = 1/94 (1%)
Query: 201 AIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAY 260
A L+ +A+ + + N A Y Q Y++AI + IDP + A G Y
Sbjct: 378 ASSLFKATLAVTTGLSAPFNNLAIIYKQQGNYSDAISCYNEVLRIDPLAADALVNRGNTY 437
Query: 261 YAQGNYRDAIDKGFKKALQLDPNNESVKENIRVA 294
G +AI + + A+ P N+ A
Sbjct: 438 KEIGRVTEAI-QDYMHAINFRPTMAEAHANLASA 470
>At5g65160 unknown protein
Length = 593
Score = 54.3 bits (129), Expect = 1e-07
Identities = 31/111 (27%), Positives = 55/111 (48%)
Query: 198 YFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLG 257
+ +A+ LY+ AIAI A Y N++AA T + R +A+ + +I I+P+Y +A+ RLG
Sbjct: 252 FAEALALYDAAIAIDPNKAAYRSNKSAALTALGRILDAVFECREAIRIEPHYHRAHHRLG 311
Query: 258 LAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRADHN 308
Y G +I + D + + + ++ +K E + D N
Sbjct: 312 NLYLRLGEVEKSIYHFKHSGPEADREDIAKAKTVQTHLNKCTEAKRLRDWN 362
Score = 36.2 bits (82), Expect = 0.041
Identities = 23/107 (21%), Positives = 52/107 (48%), Gaps = 1/107 (0%)
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAY 253
+S ++ +A Y + +++V CNRAA +++ ++ ++I+D ++ + P Y KA
Sbjct: 482 KSGRFQEACAAYGEGLDHDPRNSVLLCNRAACRSKLGQFDKSIEDCTAALSVRPGYGKAR 541
Query: 254 SRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLME 300
R + A+ ++ + P +E V + A+ +LM+
Sbjct: 542 LRRADCNAKIEKWELAVG-DYEILKKESPEDEQVIRGLSEAQQQLMK 587
>At3g58620 putative protein
Length = 677
Score = 52.8 bits (125), Expect = 4e-07
Identities = 25/72 (34%), Positives = 42/72 (57%)
Query: 198 YFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLG 257
Y +A+ LY+ AI++ ++ Y NRAAA R EA+++ L ++ DP+Y++A+ RL
Sbjct: 227 YAEALALYDRAISLSPENPAYRSNRAAALAASGRLEEAVKECLEAVRCDPSYARAHQRLA 286
Query: 258 LAYYAQGNYRDA 269
Y G +A
Sbjct: 287 SLYLRLGEAENA 298
Score = 51.2 bits (121), Expect = 1e-06
Identities = 27/104 (25%), Positives = 57/104 (53%), Gaps = 1/104 (0%)
Query: 195 SKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYS 254
S +Y +A Y + + ++V YCNRAA + ++ + +++ D +++ I P+Y+KA
Sbjct: 462 SGRYSEASVAYGDGLKLDAFNSVLYCNRAACWFKLGMWEKSVDDCNQALRIQPSYTKALL 521
Query: 255 RLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKL 298
R +Y G + DA+ + ++ + P + V E+++ A + L
Sbjct: 522 RRAASYGKLGRWEDAV-RDYEVLRKELPGDSEVAESLQRARNAL 564
>At1g53300 unknown protein
Length = 699
Score = 51.2 bits (121), Expect = 1e-06
Identities = 26/70 (37%), Positives = 41/70 (58%)
Query: 200 DAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLA 259
+A++LY+ AIA+ +A Y NRAAA ++R EA+++ ++ DPNY +A+ RL L
Sbjct: 245 EALKLYDRAIALSPTNAAYRSNRAAALIGLSRIGEAVKECEDAVRSDPNYGRAHHRLALL 304
Query: 260 YYAQGNYRDA 269
G A
Sbjct: 305 LIRLGQVNSA 314
Score = 42.4 bits (98), Expect = 6e-04
Identities = 24/109 (22%), Positives = 57/109 (52%), Gaps = 1/109 (0%)
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAY 253
+S++Y +A Y + + +A+ YCNRAA + ++ + +I+D +++ P+Y+K
Sbjct: 477 KSERYTEASSAYAEGLRLDPCNAILYCNRAACWFKLGMWERSIEDCNQALRYQPSYTKPL 536
Query: 254 SRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEER 302
R + + A+ ++ ++ P+++ V E++ A+ L + R
Sbjct: 537 LRRAASNSKMERWGAAV-SDYEALIRELPHDKEVAESLFHAQVALKKSR 584
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.132 0.387
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,918,338
Number of Sequences: 26719
Number of extensions: 474987
Number of successful extensions: 1685
Number of sequences better than 10.0: 88
Number of HSP's better than 10.0 without gapping: 42
Number of HSP's successfully gapped in prelim test: 46
Number of HSP's that attempted gapping in prelim test: 1556
Number of HSP's gapped (non-prelim): 151
length of query: 481
length of database: 11,318,596
effective HSP length: 103
effective length of query: 378
effective length of database: 8,566,539
effective search space: 3238151742
effective search space used: 3238151742
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC139354.8


