

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139354.6 - phase: 0
(342 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
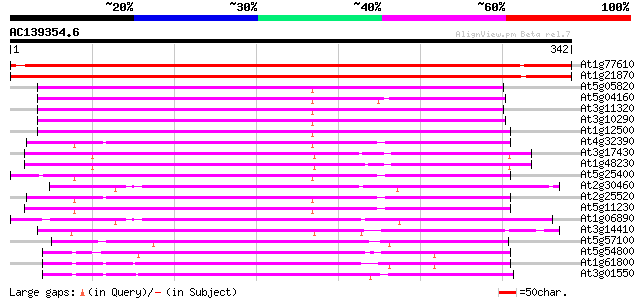
Score E
Sequences producing significant alignments: (bits) Value
At1g77610 unknown protein 618 e-177
At1g21870 glucose 6 phosphate/phosphate translocator, putative 611 e-175
At5g05820 phosphate/phosphoenolpyruvate translocator protein-like 184 6e-47
At5g04160 phosphate/phosphoenolpyruvate translocator - like protein 183 1e-46
At3g11320 unknown protein 182 3e-46
At3g10290 unknown protein 175 4e-44
At1g12500 phosphate/phosphoenolpyruvate translocator protein like 164 5e-41
At4g32390 putative protein 127 1e-29
At3g17430 unknown protein 127 1e-29
At1g48230 unknown protein 125 3e-29
At5g25400 putative protein 123 1e-28
At2g30460 integral membrane protein -like 123 2e-28
At2g25520 putative phosphate/phosphoenolpyruvate translocator pr... 123 2e-28
At5g11230 putative protein 122 2e-28
At1g06890 glucose-6-phosphate/phosphate-translocator like protein 121 6e-28
At3g14410 phosphate/phosphoenolpyruvate translocator protein like 120 1e-27
At5g57100 unknown protein 112 3e-25
At5g54800 glucose-6-phosphate/phosphate translocator 111 5e-25
At1g61800 Similar to glucose-6-phosphate/phosphate-translocator 106 2e-23
At3g01550 putative phosphate/phosphoenolpyruvate translocator 104 6e-23
>At1g77610 unknown protein
Length = 336
Score = 618 bits (1593), Expect = e-177
Identities = 304/343 (88%), Positives = 322/343 (93%), Gaps = 8/343 (2%)
Query: 1 MEEGLVCQWSVVRSLLAILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAY 60
MEEG S+ RSLLAILQWW FNVTVIIMNKWIFQKLDFKFPLSVSC+HFICS+IGAY
Sbjct: 1 MEEG-----SMFRSLLAILQWWGFNVTVIIMNKWIFQKLDFKFPLSVSCVHFICSSIGAY 55
Query: 61 VVIKVLKLKPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATT 120
+VIKVLKLKPLI VDP+DRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATT
Sbjct: 56 IVIKVLKLKPLIVVDPEDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATT 115
Query: 121 VVLQWLVWRKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAE 180
VVLQWLVWRKYFDWRIWASLVPIVGGILLTS+TELSFNMFGFCAALFGCLATSTKTILAE
Sbjct: 116 VVLQWLVWRKYFDWRIWASLVPIVGGILLTSVTELSFNMFGFCAALFGCLATSTKTILAE 175
Query: 181 ALLHGYKFDSINTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVL 240
+LLHGYKFDSINTVY+MAPFAT+I+ PALLLEG+GIL WF HP PW+A+III SSGVL
Sbjct: 176 SLLHGYKFDSINTVYYMAPFATMILGIPALLLEGSGILSWFEAHPAPWSALIIILSSGVL 235
Query: 241 AFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTF 300
AFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAV++SWLIFRNPISYMNAVGC ITLVGCTF
Sbjct: 236 AFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVMVSWLIFRNPISYMNAVGCGITLVGCTF 295
Query: 301 YGYVRNMISQQPAVPGTPRTPRTPRSKMELLPLV-NDKLDDKV 342
YGYVR+M+SQQ PGTPRTPRTPRSKMELLPLV NDKL+ KV
Sbjct: 296 YGYVRHMLSQQ--TPGTPRTPRTPRSKMELLPLVNNDKLEGKV 336
>At1g21870 glucose 6 phosphate/phosphate translocator, putative
Length = 341
Score = 611 bits (1575), Expect = e-175
Identities = 295/343 (86%), Positives = 321/343 (93%), Gaps = 3/343 (0%)
Query: 1 MEEG-LVCQWSVVRSLLAILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGA 59
MEEG L QW++ RSLL+ILQWW FNVTVIIMNKWIFQKLDFKFPLSVSC+HFICS+IGA
Sbjct: 1 MEEGSLWRQWTMFRSLLSILQWWGFNVTVIIMNKWIFQKLDFKFPLSVSCVHFICSSIGA 60
Query: 60 YVVIKVLKLKPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPAT 119
Y+VIKVLKLKPLI VDP+DRWRRIFPMSFVFCINIVLGN+SLRYIPVSFMQTIKS TPAT
Sbjct: 61 YIVIKVLKLKPLIVVDPEDRWRRIFPMSFVFCINIVLGNISLRYIPVSFMQTIKSLTPAT 120
Query: 120 TVVLQWLVWRKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILA 179
TVVLQWLVWRKYFDWRIWASLVPIVGGILLTSITELSFN+FGFCAALFGCLATSTKTILA
Sbjct: 121 TVVLQWLVWRKYFDWRIWASLVPIVGGILLTSITELSFNVFGFCAALFGCLATSTKTILA 180
Query: 180 EALLHGYKFDSINTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGV 239
E+LLHGYKFDSINTVY+MAPFAT+I+ PA LLE NGIL+WF HP PW+A+II+F+SGV
Sbjct: 181 ESLLHGYKFDSINTVYYMAPFATMILGLPAFLLERNGILDWFEAHPSPWSALIILFNSGV 240
Query: 240 LAFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCT 299
LAFCLNFSIFYVI STTAVTFNVAGNLKVAVAV +SW+IFRNPIS MNAVGC ITLVGCT
Sbjct: 241 LAFCLNFSIFYVIQSTTAVTFNVAGNLKVAVAVFVSWMIFRNPISPMNAVGCGITLVGCT 300
Query: 300 FYGYVRNMISQQPAVPGTPRTPRTPRSKMELLPLVNDKLDDKV 342
FYGYVR+M+SQQ PGTPRTPRTPR+KMEL+PLVNDKL+ K+
Sbjct: 301 FYGYVRHMLSQQQ--PGTPRTPRTPRNKMELIPLVNDKLESKI 341
>At5g05820 phosphate/phosphoenolpyruvate translocator protein-like
Length = 309
Score = 184 bits (467), Expect = 6e-47
Identities = 95/286 (33%), Positives = 172/286 (59%), Gaps = 2/286 (0%)
Query: 18 ILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQ 77
+ W++ N+ V+++NK++ FK+P+ ++ H ++ +YV I LK+ P+ ++ +
Sbjct: 15 VASWYSSNIGVLLLNKYLLSNYGFKYPIFLTMCHMTACSLLSYVAIAWLKMVPMQTIRSR 74
Query: 78 DRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIW 137
++ +I +S VFC+++V GN+SLR++PVSF Q I + TP T V +L+ RK W +
Sbjct: 75 VQFFKIAALSLVFCVSVVFGNISLRFLPVSFNQAIGATTPFFTAVFAYLMTRKKEAWLTY 134
Query: 138 ASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALL--HGYKFDSINTVY 195
+LVP+V G+++ S E SF++FGF + A + K++L LL G K +S+N +
Sbjct: 135 FTLVPVVTGVVIASGGEPSFHLFGFLMCIAATAARALKSVLQGILLSSEGEKLNSMNLLL 194
Query: 196 HMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLNFSIFYVIHST 255
+MAP A ++++ L++E N + ++ + + + + LA+ +N + F V + T
Sbjct: 195 YMAPIAVVLLLPATLIMEKNVVGITIALARDDFRIVWYLLFNSALAYLVNLTNFLVTNHT 254
Query: 256 TAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFY 301
+A+T V GN K AVAV++S LIF+NP+S +G ++T+ G Y
Sbjct: 255 SALTLQVLGNAKGAVAVVVSILIFKNPVSVTGMLGYSLTVCGVILY 300
>At5g04160 phosphate/phosphoenolpyruvate translocator - like protein
Length = 309
Score = 183 bits (465), Expect = 1e-46
Identities = 102/290 (35%), Positives = 169/290 (58%), Gaps = 8/290 (2%)
Query: 18 ILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQ 77
I+ W++ N+ V+++NK++ FKFP+ ++ H AI +Y+ I LKL PL + +
Sbjct: 16 IISWYSSNIGVLLLNKFLLSNYGFKFPIFLTMCHMSACAILSYISIVFLKLVPLQHLKSR 75
Query: 78 DRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIW 137
++ ++ +S VFC ++V GN+SLRY+PVSF Q + + TP T + +L+ K W +
Sbjct: 76 SQFLKVATLSIVFCASVVGGNISLRYLPVSFNQAVGATTPFFTALFAYLMTFKREAWVTY 135
Query: 138 ASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALL--HGYKFDSINTVY 195
+LVP+V G+++ S E F+ FGF + A + K++L LL G K +S+N +
Sbjct: 136 GALVPVVAGVVIASGGEPGFHWFGFIMCISATAARAFKSVLQGILLSSEGEKLNSMNLML 195
Query: 196 HMAPFATLIMVFPALLLEGNGILEWFSI---HPYPWAAMIIIFSSGVLAFCLNFSIFYVI 252
+M+P A + ++ L +E + I ++ H Y W I++ + V+A+ N F V
Sbjct: 196 YMSPIAVIALLPVTLFMEPDVISVTLTLAKQHQYMW---ILLLVNSVMAYSANLLNFLVT 252
Query: 253 HSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYG 302
T+A+T V GN K AVAV+IS LIF+NP++ M G +IT++G YG
Sbjct: 253 KHTSALTLQVLGNAKGAVAVVISILIFQNPVTVMGIGGYSITVLGVVAYG 302
>At3g11320 unknown protein
Length = 308
Score = 182 bits (461), Expect = 3e-46
Identities = 94/286 (32%), Positives = 170/286 (58%), Gaps = 2/286 (0%)
Query: 18 ILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQ 77
+ W++ N+ V+++NK++ FK+P+ ++ H ++ +YV I +K+ P+ ++ +
Sbjct: 15 VASWYSSNIGVLLLNKYLLSNYGFKYPIFLTMCHMTACSLLSYVAIAWMKMVPMQTIRSR 74
Query: 78 DRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIW 137
++ +I +S VFC+++V GN+SLR++PVSF Q I + TP T V +L+ K W +
Sbjct: 75 VQFLKIAALSLVFCVSVVFGNISLRFLPVSFNQAIGATTPFFTAVFAYLITFKREAWLTY 134
Query: 138 ASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALL--HGYKFDSINTVY 195
+LVP+V G+++ S +E SF++FGF + A + K++L LL G K +S+N +
Sbjct: 135 FTLVPVVTGVVIASGSEPSFHLFGFIMCIAATAARALKSVLQGILLSSEGEKLNSMNLLL 194
Query: 196 HMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLNFSIFYVIHST 255
+MAP A + ++ L++E N + ++ + + + + LA+ +N + F V T
Sbjct: 195 YMAPIAVVFLLPATLIMEKNVVGITIALARDDFRIVWYLLFNSALAYFVNLTNFLVTKHT 254
Query: 256 TAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFY 301
+A+T V GN K AVAV++S LIFRNP+S +G ++T+ G Y
Sbjct: 255 SALTLQVLGNAKGAVAVVVSILIFRNPVSVTGMLGYSLTVCGVILY 300
>At3g10290 unknown protein
Length = 355
Score = 175 bits (443), Expect = 4e-44
Identities = 96/287 (33%), Positives = 165/287 (57%), Gaps = 2/287 (0%)
Query: 18 ILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQ 77
I+ W+ N+ V+++NK++ FKFP+ ++ H AI +YV I LKL PL + +
Sbjct: 62 IILWYTSNIGVLLLNKFLLSNYGFKFPIFLTMCHMSACAILSYVSIVFLKLVPLQYLKSR 121
Query: 78 DRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIW 137
++ ++ +S VFC ++V GN+SLRY+PVSF Q + + TP T + +++ K W +
Sbjct: 122 SQFLKVATLSIVFCASVVGGNISLRYLPVSFNQAVGATTPFFTALFAYIMTFKREAWVTY 181
Query: 138 ASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALL--HGYKFDSINTVY 195
+LVP+V G+++ S E F+ FGF + A + K++L LL G + +S+N +
Sbjct: 182 GALVPVVTGVVIASGGEPGFHWFGFIMCISATAARAFKSVLQGILLSSEGERLNSMNLML 241
Query: 196 HMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLNFSIFYVIHST 255
+M+P A + ++ + +E + + ++ I++ + V+A+ N F V T
Sbjct: 242 YMSPIAVIALLPVTIFMEPDVMSVTLTLGRQHKYMYILLLVNSVMAYSANLLNFLVTKHT 301
Query: 256 TAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYG 302
+A+T V GN K AVAV+IS L+FRNP++ M G +IT++G YG
Sbjct: 302 SALTLQVLGNAKGAVAVVISILLFRNPVTVMGIGGYSITVLGVVAYG 348
>At1g12500 phosphate/phosphoenolpyruvate translocator protein like
Length = 361
Score = 164 bits (416), Expect = 5e-41
Identities = 91/290 (31%), Positives = 158/290 (54%), Gaps = 2/290 (0%)
Query: 18 ILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQ 77
I W+ N+ V+++NK++ F++P+ ++ H + A + VI + + P + +
Sbjct: 63 IAAWFGSNIGVLLLNKYLLFYYGFRYPIFLTMTHMLSCAAYSSAVINIAGIVPRQHILSR 122
Query: 78 DRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIW 137
++ +I +S +FC+++V GN SLRYIPVSF Q I + TP T V +L+ K ++
Sbjct: 123 RQFLKILSLSAIFCLSVVCGNTSLRYIPVSFNQAIGATTPFFTAVFSFLITCKTESTEVY 182
Query: 138 ASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALL--HGYKFDSINTVY 195
+L+P+V GI+L S +E SF++FGF + + K+++ +L K S+N +
Sbjct: 183 LALLPVVSGIVLASNSEPSFHLFGFLICVASTAGRALKSVVQGIILTSESEKLHSMNLLL 242
Query: 196 HMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLNFSIFYVIHST 255
+MAP A I++ L +EGN + + ++ + +A+ +N + F V T
Sbjct: 243 YMAPMAACILLPFTLYIEGNVLRVLIEKARTDPLIIFLLAGNATVAYLVNLTNFLVTKHT 302
Query: 256 TAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVR 305
+A+T V GN K AVA +S LIFRNP++ M G +T++G Y R
Sbjct: 303 SALTLQVLGNGKAAVAAGVSVLIFRNPVTVMGIAGFGVTIMGVVLYSEAR 352
>At4g32390 putative protein
Length = 350
Score = 127 bits (318), Expect = 1e-29
Identities = 76/300 (25%), Positives = 156/300 (51%), Gaps = 10/300 (3%)
Query: 11 VVRSLLAILQWWAFNVTVIIMNKWIFQK--LDFKFPLSVSCIHF-ICSAIGAYVVIKVLK 67
++ S + W + TVI+ NK+I K ++ FP++++ IH CS++ A ++IKV K
Sbjct: 15 ILLSYTYVAIWIFLSFTVIVYNKYILDKKMYNWPFPITLTMIHMAFCSSL-AVILIKVFK 73
Query: 68 LKPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLV 127
+ +S+ R + P+ ++ +++ L N + Y+ VSF+Q +K+ P + L+
Sbjct: 74 IVEPVSMSRDTYIRSVVPIGALYSLSLWLSNSAYIYLSVSFIQMLKALMPVAVYSIGVLL 133
Query: 128 WRKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALL--HG 185
++ F +++ I G+ + + E F+ +G L +T+ +L + LL G
Sbjct: 134 KKESFKSETMTNMLSISFGVAIAAYGEAKFDTWGVMLQLGAVAFEATRLVLIQILLTSKG 193
Query: 186 YKFDSINTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLN 245
+ I ++Y++AP + + FP + +E + E S H +I ++ V AF LN
Sbjct: 194 INLNPITSLYYVAPCCLVFLFFPWIFVELPILRETSSFH----FDFVIFGTNSVCAFALN 249
Query: 246 FSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVR 305
++F ++ T+A+T NVAG +K + + SW + ++ ++ +N G + +G +Y + +
Sbjct: 250 LAVFLLVGKTSALTMNVAGVVKDWLLIAFSWSVIKDTVTPLNLFGYGLAFLGVAYYNHCK 309
>At3g17430 unknown protein
Length = 375
Score = 127 bits (318), Expect = 1e-29
Identities = 89/316 (28%), Positives = 162/316 (51%), Gaps = 12/316 (3%)
Query: 10 SVVRSLLAILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSC--IHFICSAIGAYVVIKVLK 67
++V + + +L + + VI+ NKW+ F FPL ++ IH + A+++I+V K
Sbjct: 8 TLVLTYIYLLIYIILSSGVILYNKWVLSPKYFNFPLPITLTMIHMGFAGFVAFLLIRVFK 67
Query: 68 LKPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLV 127
+ + + + + P+S F ++ GN + +I V+F+Q +K+ P T ++ +
Sbjct: 68 VVAPVKMTFEIYATCVVPISAFFASSLWFGNTAYLHISVAFIQMLKALMPVATFIMAVVC 127
Query: 128 WRKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLH--G 185
++++++ + G++++S E+ FN+ G + G A + + +L + LL G
Sbjct: 128 GTDKPRCDVFSNMLLVSVGVVISSYGEIHFNIVGTVYQVTGIFAEALRLVLTQVLLQKKG 187
Query: 186 YKFDSINTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLN 245
+ I ++Y++AP + + + P +LE +E I W I FS+ + A LN
Sbjct: 188 LTLNPITSLYYIAPCSFVFLALPWYVLE-KPTMEVSQIQFNFW----IFFSNALCALALN 242
Query: 246 FSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIF-RNPISYMNAVGCAITLVGCTFYGY- 303
FSIF VI T AVT VAG LK + + +S +IF + I+ +N G AI L G Y Y
Sbjct: 243 FSIFLVIGRTGAVTIRVAGVLKDWILIALSTVIFPESTITGLNITGYAIALCGVVMYNYI 302
Query: 304 -VRNMISQQPAVPGTP 318
VR++ + QP P
Sbjct: 303 KVRDVKASQPTADSLP 318
>At1g48230 unknown protein
Length = 367
Score = 125 bits (314), Expect = 3e-29
Identities = 88/316 (27%), Positives = 162/316 (50%), Gaps = 12/316 (3%)
Query: 10 SVVRSLLAILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSC--IHFICSAIGAYVVIKVLK 67
++V + + +L + + VI+ NKW+ F FPL ++ IH S A+++I+V K
Sbjct: 8 TLVLTYIYLLIYIILSSGVILYNKWVLSPKYFNFPLPITLTMIHMGFSGFVAFLLIRVFK 67
Query: 68 LKPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLV 127
+ + + + + P+S F ++ GN + +I V+F+Q +K+ P T ++ +
Sbjct: 68 VVSPVKMTFEIYVTCVVPISAFFASSLWFGNTAYLHISVAFIQMLKALMPVATFLMAVVC 127
Query: 128 WRKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLH--G 185
++ ++V + G++++S E++FN+ G + G A + + +L + LL G
Sbjct: 128 GTDKARCDVFMNMVLVSVGVVVSSYGEINFNVIGTVYQVMGIFAEALRLVLTQVLLQKKG 187
Query: 186 YKFDSINTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLN 245
+ + ++Y++AP + + + P +LE I + I W I FS+ + A LN
Sbjct: 188 LTLNPVTSLYYIAPCSFVFLSLPWYVLEKPNI-DVSQIQFNFW----IFFSNALCALALN 242
Query: 246 FSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIF-RNPISYMNAVGCAITLVGCTFYGY- 303
FSIF VI T AVT VAG LK + + +S +IF + I+ +N G AI L G Y Y
Sbjct: 243 FSIFLVIGRTGAVTIRVAGVLKDWILIALSTVIFPESTITGLNITGYAIALCGVVMYNYI 302
Query: 304 -VRNMISQQPAVPGTP 318
++++ + QP P
Sbjct: 303 KIKDVKAIQPTTDSLP 318
>At5g25400 putative protein
Length = 349
Score = 123 bits (309), Expect = 1e-28
Identities = 76/309 (24%), Positives = 157/309 (50%), Gaps = 10/309 (3%)
Query: 1 MEEGLVCQWSVVRSLLAILQWWAFNVTVIIMNKWIFQK--LDFKFPLSVSCIHFICSAIG 58
+ EG++ + + +AI W + TVI+ NK+I K D+ FP+S++ IH +
Sbjct: 7 LSEGVIKNIIISYTYVAI--WIFLSFTVIVYNKYILDKKMYDWPFPISLTMIHMSFCSTL 64
Query: 59 AYVVIKVLKLKPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPA 118
A+++IKV K +S+ R + P+ ++ +++ L N + Y+ VSF+Q +K+ P
Sbjct: 65 AFLLIKVFKFVEPVSMSRDTYLRSVVPIGALYSLSLWLSNSAYIYLSVSFIQMLKALMPV 124
Query: 119 TTVVLQWLVWRKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTIL 178
+ L ++ F +++ I G+ + + E F+++G L +T+ ++
Sbjct: 125 AVYSIGVLFKKEGFKSETMMNMLSISFGVAIAAYGEARFDVWGVILQLGAVAFEATRLVM 184
Query: 179 AEALL--HGYKFDSINTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFS 236
+ LL G + I ++Y++AP + P +++E + + S H +I +
Sbjct: 185 IQILLTSKGITLNPITSLYYVAPCCLAFLFIPWIVVEFPILRDTSSFH----FDYLIFGT 240
Query: 237 SGVLAFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLV 296
+ AF LN ++F ++ T+A+T NVAG +K + + SW + ++ ++ +N G I +
Sbjct: 241 NSFCAFALNLAVFLLVGKTSALTMNVAGVVKDWLLIAFSWSVIKDTVTPINLFGYGIAFL 300
Query: 297 GCTFYGYVR 305
G +Y + +
Sbjct: 301 GVAYYNHAK 309
>At2g30460 integral membrane protein -like
Length = 353
Score = 123 bits (308), Expect = 2e-28
Identities = 82/315 (26%), Positives = 155/315 (49%), Gaps = 15/315 (4%)
Query: 25 NVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVI--KVLKLKPLISVDPQDRWRR 82
+V+++I NK + L F F +++ H + + +V + K + KP DP R
Sbjct: 22 SVSIVICNKALISTLGFTFATTLTSWHLLVTFCSLHVALWMKFFEHKPF---DP----RA 74
Query: 83 IFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIWASLVP 142
+ + I+I L N+SL + V F Q K TVVL+ + +RK F +I SLV
Sbjct: 75 VLGFGVLNGISIGLLNLSLGFNSVGFYQMTKLAIIPCTVVLETIFFRKMFSRKIQFSLVI 134
Query: 143 IVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDSINTVYHMAPFAT 202
++ G+ + ++T+L NM G +L + T I+ + YK S +Y P+
Sbjct: 135 LLLGVGIATVTDLQLNMLGSVLSLLAVITTCVAQIMTNTIQKKYKVSSTQLLYQSCPYQA 194
Query: 203 LIMVFPALLLEGNGILEWFSIHPYPWAAMIIIF--SSGVLAFCLNFSIFYVIHSTTAVTF 260
+ + L+ G+L ++ + + + ++ F S +++ +NFS F VI T+ VT+
Sbjct: 195 ITLFVTGPFLD--GLLTNQNVFAFKYTSQVVFFIVLSCLISVSVNFSTFLVIGKTSPVTY 252
Query: 261 NVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVRNMISQQPAVPGTPRT 320
V G+LK + + +L+ ++ S+ N +G + ++G Y Y + +QQ A + +
Sbjct: 253 QVLGHLKTCLVLAFGYLLLKDAFSWRNILGILVAVIGMVLYSYYCTLETQQKATETSTQL 312
Query: 321 PRTPRSKMELLPLVN 335
P+ ++ + PLV+
Sbjct: 313 PQMDENEKD--PLVS 325
>At2g25520 putative phosphate/phosphoenolpyruvate translocator
protein
Length = 347
Score = 123 bits (308), Expect = 2e-28
Identities = 74/300 (24%), Positives = 157/300 (51%), Gaps = 10/300 (3%)
Query: 11 VVRSLLAILQWWAFNVTVIIMNKWIFQK--LDFKFPLSVSCIHF-ICSAIGAYVVIKVLK 67
++ S + W + TVI+ NK+I K ++ FP++++ IH CS++ A ++IKV K
Sbjct: 15 IILSYTYVAIWIFLSFTVIVYNKYILDKKMYNWPFPITLTMIHMGFCSSL-AVILIKVFK 73
Query: 68 LKPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLV 127
+ +S+ + R + P+ ++ +++ L N + Y+ VSF+Q +K+ P + L+
Sbjct: 74 VVEPVSMSRETYLRSVVPIGALYSLSLWLSNSAYIYLSVSFIQMLKALMPVAVYSIGVLL 133
Query: 128 WRKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALL--HG 185
++ F + +++ I G+ + + E F+ +G L +T+ +L + LL G
Sbjct: 134 KKETFKSQTMTNMLSISFGVAIAAYGEAKFDGWGVFLQLGAVAFEATRLVLIQILLTSKG 193
Query: 186 YKFDSINTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLN 245
+ I ++Y++AP + + P + +E + + S H +I ++ V AF LN
Sbjct: 194 INLNPITSLYYVAPCCLVFLSVPWIFVEFPVLRDTSSFH----FDFVIFGTNSVCAFALN 249
Query: 246 FSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVR 305
++F ++ T+A+T NVAG +K + + SW + ++ ++ +N G + +G +Y + +
Sbjct: 250 LAVFLLVGKTSALTMNVAGVVKDWLLIAFSWSVIKDTVTPINLFGYGLAFLGVGYYNHCK 309
>At5g11230 putative protein
Length = 351
Score = 122 bits (307), Expect = 2e-28
Identities = 74/300 (24%), Positives = 151/300 (49%), Gaps = 8/300 (2%)
Query: 10 SVVRSLLAILQWWAFNVTVIIMNKWIFQK--LDFKFPLSVSCIHFICSAIGAYVVIKVLK 67
++V S + W + TVI+ NK+I K ++ FP+S++ IH + A+++IKV K
Sbjct: 14 NIVLSYSYVAIWIFLSFTVIVYNKYILDKKMYNWPFPISLTMIHMSFCSTLAFLIIKVFK 73
Query: 68 LKPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLV 127
+ + + R + P+ ++ +++ L N + Y+ VSF+Q +K+ P + L
Sbjct: 74 FVEPVKMTRETYLRSVVPIGALYALSLWLSNSAYIYLSVSFIQMLKALMPVAVYSIGVLF 133
Query: 128 WRKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALL--HG 185
++ F +++ I G+ + + E F+++G L +T+ +L + LL G
Sbjct: 134 KKEGFKSDTMMNMLSISFGVAIAAYGEARFDVWGVILQLGAVAFEATRLVLIQILLGDKG 193
Query: 186 YKFDSINTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLN 245
K + I ++Y++AP + P + +E + + S H I ++ AF LN
Sbjct: 194 IKLNPITSLYYVAPCCLAFLFIPWIYVEFPVLRDTSSFH----LDYAIFGANSFCAFALN 249
Query: 246 FSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVR 305
++F ++ T+A+T NVAG +K + + SW + ++ ++ +N G I +G +Y + +
Sbjct: 250 LAVFLLVGKTSALTMNVAGVVKDWLLIAFSWSVIKDTVTPINLFGYGIAFLGVAYYNHAK 309
>At1g06890 glucose-6-phosphate/phosphate-translocator like protein
Length = 357
Score = 121 bits (303), Expect = 6e-28
Identities = 85/336 (25%), Positives = 164/336 (48%), Gaps = 18/336 (5%)
Query: 1 MEEGLVCQWSVVRSL-LAILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGA 59
M EG Q + +L L+++ +V+++I NK + L F F +++ H + +
Sbjct: 1 MSEGQKFQLGTIGALSLSVVS----SVSIVICNKALISTLGFTFATTLTSWHLLVTFCSL 56
Query: 60 YVVI--KVLKLKPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTP 117
+V + K+ + KP DP R + + I+I L N+SL + V F Q K
Sbjct: 57 HVALWMKMFEHKPF---DP----RAVMGFGILNGISIGLLNLSLGFNSVGFYQMTKLAII 109
Query: 118 ATTVVLQWLVWRKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTI 177
TV+L+ L +RK F +I SL ++ G+ + ++T+L NM G +L + T I
Sbjct: 110 PCTVLLETLFFRKKFSRKIQFSLTILLLGVGIATVTDLQLNMLGSVLSLLAVVTTCVAQI 169
Query: 178 LAEALLHGYKFDSINTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFS- 236
+ + +K S +Y P+ + + L+G +L ++ + + + ++ F
Sbjct: 170 MTNTIQKKFKVSSTQLLYQSCPYQAITLFVTGPFLDG--LLTNQNVFAFKYTSQVVFFIV 227
Query: 237 -SGVLAFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITL 295
S +++ +NFS F VI T+ VT+ V G+LK + + +++ R+P + N +G + +
Sbjct: 228 LSCLISVSVNFSTFLVIGKTSPVTYQVLGHLKTCLVLAFGYVLLRDPFDWRNILGILVAV 287
Query: 296 VGCTFYGYVRNMISQQPAVPGTPRTPRTPRSKMELL 331
+G Y Y ++ +QQ A + + P+ S+ + L
Sbjct: 288 IGMVVYSYYCSIETQQKASETSTQLPQMKESEKDPL 323
>At3g14410 phosphate/phosphoenolpyruvate translocator protein like
Length = 340
Score = 120 bits (301), Expect = 1e-27
Identities = 90/330 (27%), Positives = 163/330 (49%), Gaps = 28/330 (8%)
Query: 18 ILQWWAFNVTVIIMNKWIF--QKLDFKFPLSVSCIHFICSAIGAYVVIKVLKL-KPLISV 74
IL + A + I NKW+ ++++F +PL ++ +H I S++ +++ KVLK+ K +
Sbjct: 19 ILLYIALSSGQIFFNKWVLSSKEINFPYPLGLTLLHMIFSSVLCFLLTKVLKIVKVEEGM 78
Query: 75 DPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDW 134
+ + P+ +F + + LGN + YI V+F Q +K+ P +L +
Sbjct: 79 TLEIYVTSVIPIGAMFAMTLWLGNTAYLYISVAFAQMLKAIMPVAVFILGVAAGLEMMSC 138
Query: 135 RIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLH--GYKFDSIN 192
R+ + I G+L+ S EL+ N G + G + + + I E L+ G K + I+
Sbjct: 139 RMLLIMSIISFGVLVASYGELNINWIGVVYQMGGVVGEALRLIFMELLVKRKGIKLNPIS 198
Query: 193 TVYHMAPFATLIMVFPALLLE-----GNGILEWFSIHPYPWAAMIIIFS-SGVLAFCLNF 246
+Y+++P + + + P + LE GNG PW ++ + + + F LN
Sbjct: 199 LMYYVSPCSAICLFVPWIFLEKSKIDGNG----------PWNFHFVVLTLNSLCTFALNL 248
Query: 247 SIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRN-PISYMNAVGCAITLVGCTFYGYVR 305
S+F VI T+A+T VAG +K V VL+S L+F + ++ +N G AI + G Y
Sbjct: 249 SVFLVISHTSALTIRVAGVVKDWVVVLVSALLFADTKLTIINLFGYAIAIAGVAAYN--N 306
Query: 306 NMISQQPAVPGTPRTPRTPRSKMELLPLVN 335
+ + ++ + T TP E +PLV+
Sbjct: 307 HKLKKEASKVVTTETP----GDAESIPLVS 332
>At5g57100 unknown protein
Length = 390
Score = 112 bits (280), Expect = 3e-25
Identities = 73/284 (25%), Positives = 138/284 (47%), Gaps = 15/284 (5%)
Query: 26 VTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQDRWRRIFP 85
+++I MNKW+ + + F+FP+ ++ IH+I AY+++ +LK L+ P + P
Sbjct: 76 ISIIFMNKWVLKNIGFEFPVFLTFIHYIV----AYLLMALLKSFSLLPASPPSTKSSLLP 131
Query: 86 M---SFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIWASLVP 142
+ V ++ L NVSL+Y V F Q K + V ++L +RK + SL
Sbjct: 132 LYTLGIVMSLSTGLANVSLKYNSVGFYQMAKIAVTPSIVFAEFLWYRKRVSFMKVVSLTV 191
Query: 143 IVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDSINTVYHMAPFAT 202
+ G+ + ++T+L F++FG C A + ++T IL + + ++ ++ P
Sbjct: 192 VSVGVAVATVTDLQFSLFGACVAFAWIIPSATNKILWSNMQQRENWTALALMWKTTPITL 251
Query: 203 LIMVFPALLLEGNGILEWFSIHPYPWAA--MIIIFSSGVLAFCLNFSIFYVIHSTTAVTF 260
L +V L+ G L + W+ I S +L F L +S + +T+A+T
Sbjct: 252 LFLVSMIPFLDPPGALS------FNWSLTNTSAILVSALLGFFLQWSGALALGATSAITH 305
Query: 261 NVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYV 304
V G K V +L ++ IF + +++ G + ++G + Y Y+
Sbjct: 306 VVLGQFKTCVLLLGNYYIFGSNSGFISVGGAFVAIMGTSLYTYL 349
>At5g54800 glucose-6-phosphate/phosphate translocator
Length = 388
Score = 111 bits (278), Expect = 5e-25
Identities = 78/297 (26%), Positives = 142/297 (47%), Gaps = 25/297 (8%)
Query: 21 WWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQ--- 77
WWA NV I NK + + +P S + ++ A ++ ++ I P+
Sbjct: 104 WWALNVVFNIYNKKVLNA--YPYPWLTSTL-----SLAAGSLMMLISWAVGIVETPKTDF 156
Query: 78 DRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIW 137
D W+ +FP++ I V VS+ + VSF IKS PA +V++ + + F ++
Sbjct: 157 DFWKTLFPVAVAHTIGHVAATVSMSKVAVSFTHIIKSGEPAFSVLVSRFILGETFPTSVY 216
Query: 138 ASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDSINTVYHM 197
SL+PI+GG L+++TEL+FNM GF A+ LA + I ++ + G +N +
Sbjct: 217 LSLIPIIGGCALSALTELNFNMIGFMGAMISNLAFVFRNIFSKKGMKGKSVSGMNYYACL 276
Query: 198 APFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLNFSIFYVIHSTTA 257
+ + LI+ A+ +EG + W W + + + + S+FY +++ +
Sbjct: 277 SMLSLLILTPFAIAVEGPQM--WVD----GWQTALATVGPQFVWWVVAQSVFYHLYNQVS 330
Query: 258 ---------VTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVR 305
+TF+V +K ++ S +IFR P+ +NA+G AI ++G Y +
Sbjct: 331 YMSLDQISPLTFSVGNTMKRISVIVSSIIIFRTPVQPVNALGAAIAILGTFLYSQAK 387
>At1g61800 Similar to glucose-6-phosphate/phosphate-translocator
Length = 388
Score = 106 bits (265), Expect = 2e-23
Identities = 77/298 (25%), Positives = 137/298 (45%), Gaps = 27/298 (9%)
Query: 21 WWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQDRW 80
WWA NV I NK + F +P S + C ++ +V ++ D + W
Sbjct: 104 WWALNVVFNIYNKKVLNA--FPYPWLTSTLSLACGSL-MMLVSWATRIADAPKTD-LEFW 159
Query: 81 RRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIWASL 140
+ +FP++ I V VS+ + VSF IKS PA +V++ + F ++ SL
Sbjct: 160 KTLFPVAVAHTIGHVAATVSMSKVAVSFTHIIKSGEPAFSVLVSRFFMGETFPLPVYLSL 219
Query: 141 VPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDSINTVYHMAPF 200
+PI+GG L +ITEL+FN+ GF A+ LA + I ++ + G +N ++
Sbjct: 220 LPIIGGCALAAITELNFNITGFMGAMISNLAFVFRNIFSKKGMKGKSVSGMNYYACLSMM 279
Query: 201 ATLIMVFPALLLEGNGILEWFSIHPYPWAA----MIIIFSSGVLAFCLNFSIFYVIHSTT 256
+ +I+ ++ +EG P WAA + + + + S+FY +++
Sbjct: 280 SLVILTPFSIAVEG----------PQMWAAGWQNAVSQVGPNFVWWVVAQSVFYHLYNQV 329
Query: 257 A---------VTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVR 305
+ +TF++ +K ++ S +IF PI +NA+G AI + G Y +
Sbjct: 330 SYMSLDQISPLTFSIGNTMKRISVIVASIIIFHTPIQPVNALGAAIAIFGTFLYSQAK 387
>At3g01550 putative phosphate/phosphoenolpyruvate translocator
Length = 383
Score = 104 bits (260), Expect = 6e-23
Identities = 77/297 (25%), Positives = 141/297 (46%), Gaps = 18/297 (6%)
Query: 21 WWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQDRW 80
W+ N+ I NK + + + +P +V+ C + ++ +LKL P P ++
Sbjct: 86 WYLLNIYYNIFNKQVLRV--YPYPATVTAFQLGCGTL-MIAIMWLLKLHPRPKFSPS-QF 141
Query: 81 RRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIWASL 140
I ++ + +L NVSL + VSF TIK+ P TV+L L+ ++ I SL
Sbjct: 142 TVIVQLAVAHTLGNLLTNVSLGRVNVSFTHTIKAMEPFFTVLLSVLLLGEWPSLWIVCSL 201
Query: 141 VPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGY-KFDSINTVYHMAP 199
+PIV G+ L S TE SFN GFC+A+ + ++ +L++ + G D+IN +
Sbjct: 202 LPIVAGVSLASFTEASFNWIGFCSAMASNVTNQSRNVLSKKFMVGKDALDNINLFSIITI 261
Query: 200 FATLIMVFPALLLEGNGIL---------EWFSIHPYPWAAMIIIFSSGVLAFCLNFSIFY 250
+ +++V A+L++G + + S+ + I+ +GV +
Sbjct: 262 ISFILLVPLAILIDGFKVTPSHLQVATSQGLSVKEF----CIMSLLAGVCLHSYQQVSYM 317
Query: 251 VIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVRNM 307
++ + VT +V +K V + S L F+ P+S +N++G A L G Y + +
Sbjct: 318 ILEMVSPVTHSVGNCVKRVVVITSSILFFKTPVSPLNSIGTATALAGVYLYSRAKRV 374
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.331 0.142 0.464
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,656,594
Number of Sequences: 26719
Number of extensions: 304092
Number of successful extensions: 941
Number of sequences better than 10.0: 60
Number of HSP's better than 10.0 without gapping: 42
Number of HSP's successfully gapped in prelim test: 18
Number of HSP's that attempted gapping in prelim test: 835
Number of HSP's gapped (non-prelim): 62
length of query: 342
length of database: 11,318,596
effective HSP length: 100
effective length of query: 242
effective length of database: 8,646,696
effective search space: 2092500432
effective search space used: 2092500432
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC139354.6


