

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
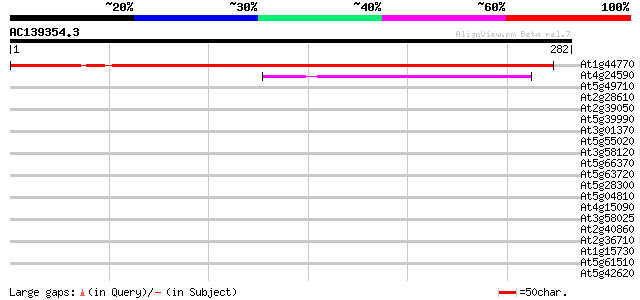
Query= AC139354.3 + phase: 0
(282 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At1g44770 unknown protein 305 1e-83
At4g24590 unknown protein 78 6e-15
At5g49710 putative protein 32 0.38
At2g28610 putative homeodomain transcription factor 32 0.38
At2g39050 similar to stress responsive lectin-like cDNAs from rice 31 0.65
At5g39990 unknown protein 31 0.86
At3g01370 unknown protein 31 0.86
At5g55020 putative transcription factor MYB120 (MYB120) 30 1.5
At3g58120 bZip transcription factor AtbZip61 30 1.9
At5g66370 putative protein 29 2.5
At5g63720 unknown protein 29 2.5
At5g28300 GTL1 - like protein 29 2.5
At5g04810 membrane-associated salt-inducible protein-like 29 2.5
At4g15090 unknown protein 29 2.5
At3g58025 putative protein 29 2.5
At2g40860 putative protein phosphatase 2C 29 2.5
At2g36710 putative pectinesterase 29 2.5
At1g15730 PRLI-interacting factor L 29 2.5
At5g61510 quinone oxidoreductase - like protein 29 3.2
At5g42620 major surface like glycoprotein 29 3.2
>At1g44770 unknown protein
Length = 271
Score = 305 bits (782), Expect = 1e-83
Identities = 150/274 (54%), Positives = 198/274 (71%), Gaps = 6/274 (2%)
Query: 1 MRTHNAVEMAKTVLEVADVAWTAVEYHHHHHRHHDPVVTHDDKYDDKDNNCPSDQDLEAL 60
M +H+A+E+ KTVLEVADVAWTAVE +HHHH H D H+ D P D++LEAL
Sbjct: 1 MTSHHAIEVTKTVLEVADVAWTAVETYHHHHHHQDE--NHESTNPISD---PRDRELEAL 55
Query: 61 KLENRRLRNLLDQNLKLLNNLSESNSFLSNCPPDLHVRLAATVRSDEYLTRLKVLQEETA 120
+ ENRRLR LL+ NLKL L+ES +F +CP DL+ RL V S ++L RL+ L++ +
Sbjct: 56 RQENRRLRTLLESNLKLFETLAESAAFSHDCPSDLYARLVTMVTSRDFLARLENLRQALS 115
Query: 121 SGG-NQFPFKEATEVDYQSADILINVDSKEPSWWVWVADETDPINFEECSGIDDESYLII 179
+G NQFPFKE TE D ++ ++LI +D +EPSWWV V D+ P N EE S ID+E Y+++
Sbjct: 116 NGTQNQFPFKEPTEDDVKTVEVLIEMDHQEPSWWVLVTDDMVPSNVEEQSAIDNEHYIVV 175
Query: 180 SEEHVVDGVANFMARCIMSNPKARNMSPEELQNNLSKAFAGTSKLEKVLDIWAAGKLFYA 239
+EEHV+D VA+F+A+CIMSNPKA+N+ PEELQ L + SK+ KV+DIW AGK+FY
Sbjct: 176 NEEHVIDAVAHFLAKCIMSNPKAKNLKPEELQKLLVQEVTALSKVGKVVDIWHAGKMFYT 235
Query: 240 LSTWGLALAGLYQTRSLLKVAAKGVHSGGKLALK 273
LSTWGLA GLYQ R +LK+AAKGVH+ K+ L+
Sbjct: 236 LSTWGLAFGGLYQARGVLKIAAKGVHATSKVVLR 269
>At4g24590 unknown protein
Length = 241
Score = 77.8 bits (190), Expect = 6e-15
Identities = 40/135 (29%), Positives = 70/135 (51%), Gaps = 5/135 (3%)
Query: 128 FKEATEVDYQSADILINVDSKEPSWWVWVADETDPINFEECSGIDDESYLIISEEHVVDG 187
F+E E+ S ++ DS W D + EEC G ++ Y+++ EE + DG
Sbjct: 99 FREKIELAQVSVPQVLAEDSSS-----WDVVSEDDLWDEECVGQTEDDYVVVREEDIADG 153
Query: 188 VANFMARCIMSNPKARNMSPEELQNNLSKAFAGTSKLEKVLDIWAAGKLFYALSTWGLAL 247
+A FMA + S + +++SP++LQ LS F+ + K+ W K+ Y +++W
Sbjct: 154 IACFMATYLSSLKQTKDISPDQLQKALSTMFSVKKRKGKLRKAWEGSKVIYNVASWSATA 213
Query: 248 AGLYQTRSLLKVAAK 262
G+YQ +L +A+K
Sbjct: 214 IGIYQNPMILSIASK 228
>At5g49710 putative protein
Length = 208
Score = 32.0 bits (71), Expect = 0.38
Identities = 14/52 (26%), Positives = 28/52 (52%)
Query: 155 WVADETDPINFEECSGIDDESYLIISEEHVVDGVANFMARCIMSNPKARNMS 206
W D + +E +E Y+++ EE + +G+A FMA + S + +++S
Sbjct: 115 WDVVSEDDLWDDETMAQREEDYVLVREEDIAEGIACFMATYLQSLKQTKDLS 166
>At2g28610 putative homeodomain transcription factor
Length = 244
Score = 32.0 bits (71), Expect = 0.38
Identities = 13/28 (46%), Positives = 14/28 (49%), Gaps = 3/28 (10%)
Query: 26 YHHHHHRHHDPVVTHDDKYDDKDNNCPS 53
YHHHHH HH H YD +C S
Sbjct: 114 YHHHHHNHHH---NHHRPYDHMSFDCCS 138
>At2g39050 similar to stress responsive lectin-like cDNAs from
rice
Length = 317
Score = 31.2 bits (69), Expect = 0.65
Identities = 12/26 (46%), Positives = 16/26 (61%), Gaps = 4/26 (15%)
Query: 24 VEYHHHHHRHHDPVVTHDDKYDDKDN 49
+E+HH HHRHH DD DD+ +
Sbjct: 1 MEHHHQHHRHHQ----RDDGEDDRQS 22
>At5g39990 unknown protein
Length = 447
Score = 30.8 bits (68), Expect = 0.86
Identities = 9/18 (50%), Positives = 13/18 (72%)
Query: 27 HHHHHRHHDPVVTHDDKY 44
HHHHH HH +V+ + K+
Sbjct: 17 HHHHHHHHSNIVSSERKW 34
>At3g01370 unknown protein
Length = 1020
Score = 30.8 bits (68), Expect = 0.86
Identities = 40/161 (24%), Positives = 62/161 (37%), Gaps = 13/161 (8%)
Query: 58 EALK--LENRRLRNLLDQNLKLLNNLSESNSFLSNCPPDLHVRLAATVRSDEYLTRLKVL 115
EALK +E +R ++L LKL NN+ E N L V +AT + ++
Sbjct: 695 EALKRSVEAQRRKSLKLHVLKLSNNIEELNRQL--------VEDSATNETWSDGESSNMM 746
Query: 116 QEETASGGNQFPFKEATEVDYQSADILINVDSKEPSWWVWVADETDPINFEECSGIDDES 175
EE + P K +++ + L S E +W E DP+ +DES
Sbjct: 747 VEEETENQHTEPEKAREKIELGYSSDLSVPSSGEENWEDDSEGEVDPLTTSSQEYQEDES 806
Query: 176 YLIISEEH---VVDGVANFMARCIMSNPKARNMSPEELQNN 213
S+ H +D AN + A + L +N
Sbjct: 807 ESASSQRHEGNSLDSTANLSVFAETGSANASSFHDRSLPHN 847
>At5g55020 putative transcription factor MYB120 (MYB120)
Length = 523
Score = 30.0 bits (66), Expect = 1.5
Identities = 18/74 (24%), Positives = 25/74 (33%), Gaps = 3/74 (4%)
Query: 25 EYHHHHHRHHDPVVTHDDKY---DDKDNNCPSDQDLEALKLENRRLRNLLDQNLKLLNNL 81
+ H+HHH HH H Y N PS L + N + + + N L
Sbjct: 157 QQHNHHHHHHQQQQQHQQMYFQPQSSQRNTPSSSPLPSPTPANAKSSSSFTFHTTTANLL 216
Query: 82 SESNSFLSNCPPDL 95
+ N P L
Sbjct: 217 HPLSPHTPNTPSQL 230
>At3g58120 bZip transcription factor AtbZip61
Length = 329
Score = 29.6 bits (65), Expect = 1.9
Identities = 20/75 (26%), Positives = 31/75 (40%), Gaps = 10/75 (13%)
Query: 27 HHHHHRHHD------PVVTHDDKYDDKDNNCPSDQD----LEALKLENRRLRNLLDQNLK 76
+H+HH HH P + + D+N SD D ++ N+ +QN
Sbjct: 104 NHNHHHHHSINGNVGPTRSSSNTSTPSDHNSLSDDDNNKEAPPSDHDHHMDNNVANQNNA 163
Query: 77 LLNNLSESNSFLSNC 91
NN +ES+ S C
Sbjct: 164 AGNNYNESDEVQSQC 178
>At5g66370 putative protein
Length = 130
Score = 29.3 bits (64), Expect = 2.5
Identities = 9/22 (40%), Positives = 14/22 (62%)
Query: 15 EVADVAWTAVEYHHHHHRHHDP 36
+ A++ + HHHHH+HH P
Sbjct: 70 KAAEMTFVKTGGHHHHHQHHYP 91
>At5g63720 unknown protein
Length = 492
Score = 29.3 bits (64), Expect = 2.5
Identities = 8/10 (80%), Positives = 9/10 (90%)
Query: 26 YHHHHHRHHD 35
+HHHHH HHD
Sbjct: 373 HHHHHHHHHD 382
Score = 27.7 bits (60), Expect = 7.2
Identities = 10/18 (55%), Positives = 11/18 (60%), Gaps = 5/18 (27%)
Query: 26 YHHHHHRHHDPVVTHDDK 43
+HHHHH HH H DK
Sbjct: 371 HHHHHHHHH-----HHDK 383
Score = 27.3 bits (59), Expect = 9.5
Identities = 7/9 (77%), Positives = 8/9 (88%)
Query: 26 YHHHHHRHH 34
+HHHHH HH
Sbjct: 368 FHHHHHHHH 376
>At5g28300 GTL1 - like protein
Length = 619
Score = 29.3 bits (64), Expect = 2.5
Identities = 8/10 (80%), Positives = 9/10 (90%)
Query: 26 YHHHHHRHHD 35
+HHHHH HHD
Sbjct: 62 HHHHHHHHHD 71
>At5g04810 membrane-associated salt-inducible protein-like
Length = 952
Score = 29.3 bits (64), Expect = 2.5
Identities = 15/49 (30%), Positives = 22/49 (44%), Gaps = 3/49 (6%)
Query: 8 EMAKTVLEVADVAWTAVEYHHHHHRHHDPVVTHDDKYDDKDNNCPSDQD 56
E+ +T+ + W++ HHHHH D V D DD D D +
Sbjct: 894 ELTETLQKTFPPDWSS---HHHHHGFLDQVSDVDSDEDDVDGEDGEDDE 939
>At4g15090 unknown protein
Length = 768
Score = 29.3 bits (64), Expect = 2.5
Identities = 23/85 (27%), Positives = 42/85 (49%), Gaps = 10/85 (11%)
Query: 35 DPVVTHDDKYDD--------KDNNCPSDQDLE-ALKLENRRLRNLLDQNLKLLNNLSESN 85
D + T +Y+D + C S+++ AL+ L+N +D N NN++ESN
Sbjct: 560 DQIQTRVQRYNDLCSRATELSEEGCVSEENYNIALRTLVETLKNCVDMN-NARNNITESN 618
Query: 86 SFLSNCPPDLHVRLAATVRSDEYLT 110
S L+N + ++ A V++ + T
Sbjct: 619 SQLNNGTHEEENQVMAGVKATKKKT 643
>At3g58025 putative protein
Length = 173
Score = 29.3 bits (64), Expect = 2.5
Identities = 8/15 (53%), Positives = 11/15 (73%)
Query: 26 YHHHHHRHHDPVVTH 40
+HHHHH HH+ T+
Sbjct: 124 HHHHHHHHHNSTATN 138
Score = 27.3 bits (59), Expect = 9.5
Identities = 7/11 (63%), Positives = 9/11 (81%)
Query: 24 VEYHHHHHRHH 34
+ +HHHHH HH
Sbjct: 120 LHHHHHHHHHH 130
>At2g40860 putative protein phosphatase 2C
Length = 404
Score = 29.3 bits (64), Expect = 2.5
Identities = 17/43 (39%), Positives = 24/43 (55%)
Query: 214 LSKAFAGTSKLEKVLDIWAAGKLFYALSTWGLALAGLYQTRSL 256
LSKA T E+ I G++ + + TW +A AGL TRS+
Sbjct: 271 LSKAHLATCIDERNRVIGEGGRIEWLVDTWRVAPAGLQVTRSI 313
>At2g36710 putative pectinesterase
Length = 407
Score = 29.3 bits (64), Expect = 2.5
Identities = 7/12 (58%), Positives = 11/12 (91%)
Query: 26 YHHHHHRHHDPV 37
+HHHH+ HH+P+
Sbjct: 60 HHHHHYHHHEPI 71
Score = 27.3 bits (59), Expect = 9.5
Identities = 8/16 (50%), Positives = 10/16 (62%)
Query: 26 YHHHHHRHHDPVVTHD 41
+HHHHH HH H+
Sbjct: 54 HHHHHHHHHHHYHHHE 69
Score = 27.3 bits (59), Expect = 9.5
Identities = 7/10 (70%), Positives = 9/10 (90%)
Query: 25 EYHHHHHRHH 34
++HHHHH HH
Sbjct: 51 KHHHHHHHHH 60
>At1g15730 PRLI-interacting factor L
Length = 448
Score = 29.3 bits (64), Expect = 2.5
Identities = 13/34 (38%), Positives = 14/34 (40%)
Query: 25 EYHHHHHRHHDPVVTHDDKYDDKDNNCPSDQDLE 58
E+ H H HH THD C D DLE
Sbjct: 334 EHEHEHEHHHSHDHTHDPGVGSVSIVCEGDLDLE 367
>At5g61510 quinone oxidoreductase - like protein
Length = 406
Score = 28.9 bits (63), Expect = 3.2
Identities = 8/9 (88%), Positives = 9/9 (99%)
Query: 26 YHHHHHRHH 34
+HHHHHRHH
Sbjct: 7 HHHHHHRHH 15
>At5g42620 major surface like glycoprotein
Length = 841
Score = 28.9 bits (63), Expect = 3.2
Identities = 12/45 (26%), Positives = 21/45 (46%)
Query: 29 HHHRHHDPVVTHDDKYDDKDNNCPSDQDLEALKLENRRLRNLLDQ 73
HHH HH + + ++C DQ +E K R++ ++ Q
Sbjct: 38 HHHHHHQRIAVEGVESGVASHSCIHDQIIEQRKRPGRKVYSVTPQ 82
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.315 0.131 0.386
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,345,515
Number of Sequences: 26719
Number of extensions: 272471
Number of successful extensions: 1670
Number of sequences better than 10.0: 52
Number of HSP's better than 10.0 without gapping: 35
Number of HSP's successfully gapped in prelim test: 17
Number of HSP's that attempted gapping in prelim test: 1427
Number of HSP's gapped (non-prelim): 123
length of query: 282
length of database: 11,318,596
effective HSP length: 98
effective length of query: 184
effective length of database: 8,700,134
effective search space: 1600824656
effective search space used: 1600824656
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC139354.3


