

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
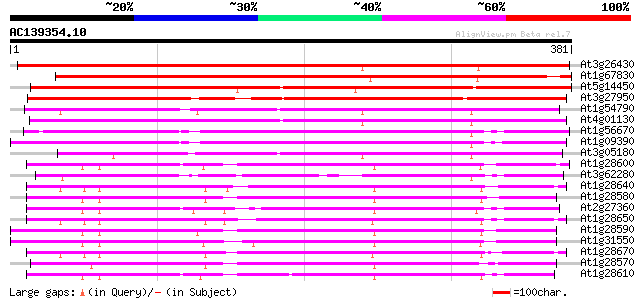
Query= AC139354.10 - phase: 0
(381 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At3g26430 nodulin, putative 392 e-109
At1g67830 ENOD8-like protein 368 e-102
At5g14450 early nodule-specific protein - like 362 e-100
At3g27950 early nodule-specific protein, putative 313 8e-86
At1g54790 285 3e-77
At4g01130 putative acetyltransferase 268 5e-72
At1g56670 lipase homolog (Lip-4) 264 7e-71
At1g09390 putative lipase 260 1e-69
At3g05180 putative nodulin 257 8e-69
At1g28600 lipase like protein 206 2e-53
At3g62280 putative protein 199 3e-51
At1g28640 lipase, putative 190 1e-48
At1g28580 lipase like protein 187 1e-47
At2g27360 putative lipase 186 2e-47
At1g28650 lipase, putative 186 2e-47
At1g28590 putative lipase (At1g28590) 185 4e-47
At1g31550 unknown protein 181 8e-46
At1g28670 lipase 181 8e-46
At1g28570 lipase like protein 180 1e-45
At1g28610 lipase, putative 178 4e-45
>At3g26430 nodulin, putative
Length = 380
Score = 392 bits (1006), Expect = e-109
Identities = 186/380 (48%), Positives = 260/380 (67%), Gaps = 5/380 (1%)
Query: 6 LSRHIPLVTLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPSPYGETY 65
+ ++ LV ++L C+ P + +C+FPAIF+FG SN DTGGL+A+F P P G+T+
Sbjct: 1 METNLLLVKCVLLASCLIHPRACSPSCNFPAIFNFGDSNSDTGGLSASFGQAPYPNGQTF 60
Query: 66 FHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSILPN 125
FH +GRFSDGR+I+DFIA LPYL+ +L+S+GSNF+HGANFA+ GST+ P + +
Sbjct: 61 FHSPSGRFSDGRLIIDFIAEELGLPYLNAFLDSIGSNFSHGANFATAGSTVRPPNATIAQ 120
Query: 126 GKLSPFSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGF 185
+SP SL +Q +QF +FI++++LIR++GGVF L+PK++YFS+ALY FDIGQNDLT G
Sbjct: 121 SGVSPISLDVQLVQFSDFITRSQLIRNRGGVFKKLLPKKEYFSQALYTFDIGQNDLTAGL 180
Query: 186 FGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANF--PSAIK 243
N T Q+ A +PD+ + I+ +Y+ G R FWIH T P GC P +L F P++
Sbjct: 181 KLNMTSDQIKAYIPDVHDQLSNVIRKVYSKGGRRFWIHNTAPLGCLPYVLDRFPVPASQI 240
Query: 244 DSYGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELP 303
D++GCA NE+++Y+N +LK + ELR LS AA TYVDIY+ K +L T +K GF P
Sbjct: 241 DNHGCAIPRNEIARYYNSELKRRVIELRKELSEAAFTYVDIYSIKLTLITQAKKLGFRYP 300
Query: 304 FVACCGYGGEYNIG--VGCGASININGTKIV-AGSCKNPSTRIIWDGVHYTEAANEIVFS 360
VACCG+GG+YN + CGA + I G +IV A SC + S R+ WDG+H+TE N +F
Sbjct: 301 LVACCGHGGKYNFNKLIKCGAKVMIKGKEIVLAKSCNDVSFRVSWDGIHFTETTNSWIFQ 360
Query: 361 QILTGVFNDPPISLDRACYR 380
QI G F+DPP+ + AC R
Sbjct: 361 QINDGAFSDPPLPVKSACTR 380
>At1g67830 ENOD8-like protein
Length = 372
Score = 368 bits (944), Expect = e-102
Identities = 174/355 (49%), Positives = 241/355 (67%), Gaps = 13/355 (3%)
Query: 32 CDFPAIFSFGASNVDTGGLAAAFRAPPSPYGETYFHRSTGRFSDGRIILDFIARSFRLPY 91
C FPAIF+FG SN DTGGL+AAF P+G ++F GR+ DGR+++DFIA S LPY
Sbjct: 26 CHFPAIFNFGDSNSDTGGLSAAFGQAGPPHGSSFFGSPAGRYCDGRLVIDFIAESLGLPY 85
Query: 92 LSPYLNSLGSNFTHGANFASGGSTINIPKSILPNGKLSPFSLQIQYIQFKEFISKTKLIR 151
LS +L+S+GSNF+HGANFA+ GS I S L SPFSL +Q++QF F ++++ +R
Sbjct: 86 LSAFLDSVGSNFSHGANFATAGSPIRALNSTLRQSGFSPFSLDVQFVQFYNFHNRSQTVR 145
Query: 152 DQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNKTIQQVNATVPDIVNNYIENIKN 211
+GGV+ T++P+ D FSKALY FDIGQNDLT G+F NKT++QV VP+I++ ++ IKN
Sbjct: 146 SRGGVYKTMLPESDSFSKALYTFDIGQNDLTAGYFANKTVEQVETEVPEIISQFMNAIKN 205
Query: 212 IYNLGARSFWIHGTGPKGCAPVILANFPSAIK--DSYGCAKQYNEVSQYFNFKLKEALAE 269
IY G R FWIH TGP GC ++ FP+ DS+GC N ++Q FN LK+A+ E
Sbjct: 206 IYGQGGRYFWIHNTGPIGCLAYVIERFPNKASDFDSHGCVSPLNHLAQQFNHALKQAVIE 265
Query: 270 LRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACCGYGGEYNI--GVGCGASININ 327
LRS+LS AAITYVD+Y+ K+ LF + + +GF+ V+CCG+GG+YN G+GCG +
Sbjct: 266 LRSSLSEAAITYVDVYSLKHELFVHAQGHGFKGSLVSCCGHGGKYNYNKGIGCGMKKIVK 325
Query: 328 GTKIVAGS-CKNPSTRIIWDGVHYTEAANEIVFSQILTGVFNDPPISLDRACYRK 381
G ++ G C P ++WDGVH+T+AAN+ +F +I G L +AC R+
Sbjct: 326 GKEVYIGKPCDEPDKAVVWDGVHFTQAANKFIFDKIAPG--------LSKACKRQ 372
>At5g14450 early nodule-specific protein - like
Length = 389
Score = 362 bits (929), Expect = e-100
Identities = 179/372 (48%), Positives = 238/372 (63%), Gaps = 7/372 (1%)
Query: 15 LIVLVLCITPPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPSPYGETYFHRSTGRFS 74
L++ + +T + C FPAI++FG SN DTGG++AAF PYG+ +FHR TGR S
Sbjct: 20 LLLCLFAVTTSVSVQPTCTFPAIYNFGDSNSDTGGISAAFEPIRDPYGQGFFHRPTGRDS 79
Query: 75 DGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSILPNGKLSPFSLQ 134
DGR+ +DFIA LPYLS YLNSLGSNF HGANFA+GGSTI + +SPFSL
Sbjct: 80 DGRLTIDFIAERLGLPYLSAYLNSLGSNFRHGANFATGGSTIRRQNETIFQYGISPFSLD 139
Query: 135 IQYIQFKEFISKTKLIRDQ--GGVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNKTIQ 192
+Q QF +F +++ L+ Q +P+++ F+KALY FDIGQNDL++G F ++
Sbjct: 140 MQIAQFDQFKARSALLFTQIKSRYDREKLPRQEEFAKALYTFDIGQNDLSVG-FRTMSVD 198
Query: 193 QVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPV---ILANFPSAIKDSYGCA 249
Q+ AT+PDIVN+ ++NIY G R+FW+H TGP GC PV + D GC
Sbjct: 199 QLKATIPDIVNHLASAVRNIYQQGGRTFWVHNTGPFGCLPVNMFYMGTPAPGYLDKSGCV 258
Query: 250 KQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACCG 309
K NE++ FN KLKE + LR L+ AAITYVD+YT KY + +NP+K GF P CCG
Sbjct: 259 KAQNEMAMEFNRKLKETVINLRKELTQAAITYVDVYTAKYEMMSNPKKLGFANPLKVCCG 318
Query: 310 YGGEYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFSQILTGVFND 369
Y +Y+ + CG +N T+I GSC NP + WDGVHYTEAAN+ V + L G+ D
Sbjct: 319 YHEKYD-HIWCGNKGKVNNTEIYGGSCPNPVMAVSWDGVHYTEAANKHVADRTLNGLLTD 377
Query: 370 PPISLDRACYRK 381
PP+ + RACYR+
Sbjct: 378 PPVPITRACYRQ 389
>At3g27950 early nodule-specific protein, putative
Length = 361
Score = 313 bits (803), Expect = 8e-86
Identities = 167/367 (45%), Positives = 228/367 (61%), Gaps = 19/367 (5%)
Query: 13 VTLIVLVLCITPPIFA-TKNCDFPAIFSFGASNVDTGGLAAAFRAPPSPYGETYFHRSTG 71
+ +IVL+L T + A + +C+FPA+F+FG SN DTG ++AA P P G +F RS G
Sbjct: 8 LAIIVLLLGFTEKLSALSSSCNFPAVFNFGDSNSDTGAISAAIGEVPPPNGVAFFGRSAG 67
Query: 72 RFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSILPNGKLSPF 131
R SDGR+I+DFI + LPYL+PYL+S+G+N+ HGANFA+GGS I + SPF
Sbjct: 68 RHSDGRLIIDFITENLTLPYLTPYLDSVGANYRHGANFATGGSCIRPTLACF-----SPF 122
Query: 132 SLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNKTI 191
L Q QF F ++T + +Q + FSKALY DIGQNDL IGF N T
Sbjct: 123 HLGTQVSQFIHFKTRTLSLYNQ----------TNDFSKALYTLDIGQNDLAIGF-QNMTE 171
Query: 192 QQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPSAIKDSYGCAKQ 251
+Q+ AT+P I+ N+ +K +Y GAR F IH TGP GC P +L FP+ +D YGC K
Sbjct: 172 EQLKATIPLIIENFTIALKLLYKEGARFFSIHNTGPTGCLPYLLKAFPAIPRDPYGCLKP 231
Query: 252 YNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACCGYG 311
N V+ FN +LK + +L+ L S+ TYVD+Y+ KY+L T + GF PF CC
Sbjct: 232 LNNVAIEFNKQLKNKITQLKKELPSSFFTYVDVYSAKYNLITKAKALGFIDPFDYCC--V 289
Query: 312 GEYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFSQILTGVFNDPP 371
G G+GCG +I +NGT++ + SC+N I WDG+HYTE AN +V ++IL G +DPP
Sbjct: 290 GAIGRGMGCGKTIFLNGTELYSSSCQNRKNFISWDGIHYTETANMLVANRILDGSISDPP 349
Query: 372 ISLDRAC 378
+ +AC
Sbjct: 350 LPTQKAC 356
>At1g54790
Length = 383
Score = 285 bits (729), Expect = 3e-77
Identities = 157/377 (41%), Positives = 223/377 (58%), Gaps = 21/377 (5%)
Query: 11 PLVTLIVLVLCITPPIFATKNCD----FPAIFSFGASNVDTGGL-AAAFRAPPSPYGETY 65
P+V + +VL I +F + +PAI +FG SN DTG L +A PYG+TY
Sbjct: 3 PIVNNLSIVLFILISLFLPSSFSIILKYPAIINFGDSNSDTGNLISAGIENVNPPYGQTY 62
Query: 66 FHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLG-SNFTHGANFASGGSTINIPKSILP 124
F+ +GR+ DGR+I+DF+ LP+L+PYL+SLG NF G NFA+ GSTI LP
Sbjct: 63 FNLPSGRYCDGRLIVDFLLDEMDLPFLNPYLDSLGLPNFKKGCNFAAAGSTI------LP 116
Query: 125 NG--KLSPFSLQIQYIQFKEFISKT-KLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDL 181
+SPFS +Q QF F S+ +L+ G + +P DY+SK LY+ DIGQND+
Sbjct: 117 ANPTSVSPFSFDLQISQFIRFKSRAIELLSKTGRKYEKYLPPIDYYSKGLYMIDIGQNDI 176
Query: 182 TIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANF--P 239
G F +KT+ QV A++P I+ + +K +Y G R+ WIH TGP GC +A F
Sbjct: 177 A-GAFYSKTLDQVLASIPSILETFEAGLKRLYEEGGRNIWIHNTGPLGCLAQNIAKFGTD 235
Query: 240 SAIKDSYGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYG 299
S D +GC +N+ ++ FN +L + ++ A +TYVDI++ K +L N ++G
Sbjct: 236 STKLDEFGCVSSHNQAAKLFNLQLHAMSNKFQAQYPDANVTYVDIFSIKSNLIANYSRFG 295
Query: 300 FELPFVACCGYGG---EYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANE 356
FE P +ACCG GG Y+ + CG + ++G + A +C + S I WDG+HYTEAANE
Sbjct: 296 FEKPLMACCGVGGAPLNYDSRITCGQTKVLDGISVTAKACNDSSEYINWDGIHYTEAANE 355
Query: 357 IVFSQILTGVFNDPPIS 373
V SQILTG ++DPP S
Sbjct: 356 FVSSQILTGKYSDPPFS 372
>At4g01130 putative acetyltransferase
Length = 382
Score = 268 bits (684), Expect = 5e-72
Identities = 144/370 (38%), Positives = 205/370 (54%), Gaps = 6/370 (1%)
Query: 14 TLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPSPYGETYFHRSTGRF 73
+L+VL++ + CDF AIF+FG SN DTGG AAF A P+G TYF + GR
Sbjct: 12 SLLVLIIVMLYGHKGDSKCDFEAIFNFGDSNSDTGGFWAAFPAQSGPWGMTYFKKPAGRA 71
Query: 74 SDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSILPNGKLSPFSL 133
SDGR+I+DF+A+S +P+LSPYL S+GS+F HGANFA+ ST+ +P + L +SPFSL
Sbjct: 72 SDGRLIIDFLAKSLGMPFLSPYLQSIGSDFRHGANFATLASTVLLPNTSLFVSGISPFSL 131
Query: 134 QIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNKTIQQ 193
IQ Q K+F ++P + F K+LY F IGQND T + +++
Sbjct: 132 AIQLNQMKQFKVNVDESHSLDRPGLKILPSKIVFGKSLYTFYIGQNDFTSN-LASIGVER 190
Query: 194 VNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANF--PSAIKDSYGCAKQ 251
V +P ++ IK IY +G R+F + P GC P IL + A D YGC
Sbjct: 191 VKLYLPQVIGQIAGTIKEIYGIGGRTFLVLNLAPVGCYPAILTGYTHTDADLDKYGCLIP 250
Query: 252 YNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACCGYG 311
N+ +Y+N L + L++ R+ L +A + Y+D + LF +P+ YG + ACCGYG
Sbjct: 251 VNKAVKYYNTLLNKTLSQTRTELKNATVIYLDTHKILLDLFQHPKSYGMKHGIKACCGYG 310
Query: 312 G---EYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFSQILTGVFN 368
G +N + CG + I A +C +P + WDG+H TEAAN + IL G +
Sbjct: 311 GRPYNFNQKLFCGNTKVIGNFSTTAKACHDPHNYVSWDGIHATEAANHHISMAILDGSIS 370
Query: 369 DPPISLDRAC 378
PP L+ C
Sbjct: 371 YPPFILNNLC 380
>At1g56670 lipase homolog (Lip-4)
Length = 373
Score = 264 bits (674), Expect = 7e-71
Identities = 153/376 (40%), Positives = 201/376 (52%), Gaps = 23/376 (6%)
Query: 10 IPLVTLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPS-PYGETYFHR 68
I LV+L +L+L P A P IF+FG SN DTGGL A P P G +F R
Sbjct: 16 ITLVSLALLIL--RQPSRAASCTARPVIFNFGDSNSDTGGLVAGLGYPIGFPNGRLFFRR 73
Query: 69 STGRFSDGRIILDFIARSFRLPYLSPYLNSLG-SNFTHGANFASGGSTINIPKSILPNGK 127
STGR SDGR+++DF+ +S L PYL+SLG + F +GANFA GS +PK++
Sbjct: 74 STGRLSDGRLLIDFLCQSLNTSLLRPYLDSLGRTRFQNGANFAIAGSP-TLPKNV----- 127
Query: 128 LSPFSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGFFG 187
PFSL IQ QF F S++ + + + F ALY+ DIGQND+ F
Sbjct: 128 --PFSLNIQVKQFSHFKSRSLELASSSNSLKGMFISNNGFKNALYMIDIGQNDIARSFAR 185
Query: 188 NKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPSAIKDSYG 247
+ Q +P I+ +IK +Y+ G R FWIH TGP GC P L+ S D +G
Sbjct: 186 GNSYSQTVKLIPQIITEIKSSIKRLYDEGGRRFWIHNTGPLGCLPQKLSMVKSKDLDQHG 245
Query: 248 CAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVAC 307
C YN + FN L ELR+ L A I Y+DIY KYSL N +YGF+ P +AC
Sbjct: 246 CLVSYNSAATLFNQGLDHMCEELRTELRDATIIYIDIYAIKYSLIANSNQYGFKSPLMAC 305
Query: 308 CGYGG---EYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFSQILT 364
CGYGG YN+ + CG G+ + C+ S I WDG+HYTE AN IV ++L+
Sbjct: 306 CGYGGTPYNYNVKITCGH----KGSNV----CEEGSRFISWDGIHYTETANAIVAMKVLS 357
Query: 365 GVFNDPPISLDRACYR 380
++ PP C R
Sbjct: 358 MHYSKPPTPFHFFCRR 373
>At1g09390 putative lipase
Length = 370
Score = 260 bits (664), Expect = 1e-69
Identities = 151/384 (39%), Positives = 205/384 (53%), Gaps = 22/384 (5%)
Query: 1 MAKIELSRHIPLVTLIVLVLCITPPIFATKNCDFP-AIFSFGASNVDTGGLAAAFRAPPS 59
MA + L H L+ L+ +L + + C P IF+FG SN DTGGL A
Sbjct: 1 MATLSLHSHSFLLVLLPFILILRQNLAVAGGCQVPPVIFNFGDSNSDTGGLVAGLGYSIG 60
Query: 60 -PYGETYFHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSL-GSNFTHGANFASGGSTIN 117
P G ++F RSTGR SDGR+++DF+ +S L+PYL+SL GS F +GANFA GS+
Sbjct: 61 LPNGRSFFQRSTGRLSDGRLVIDFLCQSLNTSLLNPYLDSLVGSKFQNGANFAIVGSS-T 119
Query: 118 IPKSILPNGKLSPFSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIG 177
+P+ + PF+L IQ +QF F S+ + ++ E F ALY+ DIG
Sbjct: 120 LPRYV-------PFALNIQLMQFLHFKSRALELASISDPLKEMMIGESGFRNALYMIDIG 172
Query: 178 QNDLTIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILAN 237
QND+ F + +V +P++++ IK +Y+ G R FW+H TGP GC P L+
Sbjct: 173 QNDIADSFSKGLSYSRVVKLIPNVISEIKSAIKILYDEGGRKFWVHNTGPLGCLPQKLSM 232
Query: 238 FPSAIKDSYGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEK 297
S D +GC YN ++ FN L +LR+ L A I YVDIY KY L N
Sbjct: 233 VHSKGFDKHGCLATYNAAAKLFNEGLDHMCRDLRTELKEANIVYVDIYAIKYDLIANSNN 292
Query: 298 YGFELPFVACCGYGG---EYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAA 354
YGFE P +ACCGYGG YN+ + CG G+K SC S I WDG+HYTE A
Sbjct: 293 YGFEKPLMACCGYGGPPYNYNVNITCGN----GGSK----SCDEGSRFISWDGIHYTETA 344
Query: 355 NEIVFSQILTGVFNDPPISLDRAC 378
N IV ++L+ + PP C
Sbjct: 345 NAIVAMKVLSMQHSTPPTPFHFFC 368
>At3g05180 putative nodulin
Length = 379
Score = 257 bits (656), Expect = 8e-69
Identities = 143/351 (40%), Positives = 198/351 (55%), Gaps = 13/351 (3%)
Query: 33 DFPAIFSFGASNVDTGGLAAAFRAPPSPYGETYFHRS--TGRFSDGRIILDFIARSFRLP 90
DFPA+F+FG SN DTG L++ P P E F RS +GRF +GR+I+DF+ + P
Sbjct: 33 DFPAVFNFGDSNSDTGELSSGLGFLPQPSYEITFFRSPTSGRFCNGRLIVDFLMEAIDRP 92
Query: 91 YLSPYLNSLG-SNFTHGANFASGGSTINIPKSILPNGKLSPFSLQIQYIQFKEFISKTKL 149
YL PYL+S+ + G NFA+ STI + SPF +Q QF F SK
Sbjct: 93 YLRPYLDSISRQTYRRGCNFAAAASTIQKANA----ASYSPFGFGVQVSQFITFKSKVLQ 148
Query: 150 IRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNKTIQQVNATVPDIVNNYIENI 209
+ Q +P E +FS LY+FDIGQND+ G F KT+ QV A VP I++ + + I
Sbjct: 149 LIQQDEELQRYLPSEYFFSNGLYMFDIGQNDIA-GAFYTKTVDQVLALVPIILDIFQDGI 207
Query: 210 KNIYNLGARSFWIHGTGPKGCAPVILANF--PSAIKDSYGCAKQYNEVSQYFNFKLKEAL 267
K +Y GAR++WIH TGP GC +++ F + D +GC +N+ ++ FN +L
Sbjct: 208 KRLYAEGARNYWIHNTGPLGCLAQVVSIFGEDKSKLDEFGCVSDHNQAAKLFNLQLHGLF 267
Query: 268 AELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACCGYGG---EYNIGVGCGASI 324
+L ++ TYVDI++ K L N KYGF+ + CCG GG Y+ VGCG +
Sbjct: 268 KKLPQQYPNSRFTYVDIFSIKSDLILNHSKYGFDHSIMVCCGTGGPPLNYDDQVGCGKTA 327
Query: 325 NINGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFSQILTGVFNDPPISLD 375
NGT I A C + S + WDG+HYTEAAN V ILTG +++ SL+
Sbjct: 328 RSNGTIITAKPCYDSSKYVNWDGIHYTEAANRFVALHILTGKYSETASSLN 378
>At1g28600 lipase like protein
Length = 393
Score = 206 bits (523), Expect = 2e-53
Identities = 139/386 (36%), Positives = 188/386 (48%), Gaps = 37/386 (9%)
Query: 12 LVTLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTG---GLAAAFRAPPS---PYGETY 65
LV + L +T T +F +I SFG S DTG GL+ + P + PYGET+
Sbjct: 7 LVIFLFSTLFVTIVSSETPCPNFKSIISFGDSIADTGNLVGLSDRNQLPVTAFPPYGETF 66
Query: 66 FHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSILPN 125
FH TGR DGRII+DFIA LPY+ PY S NF G NFA G+T + S L
Sbjct: 67 FHHPTGRSCDGRIIMDFIAEFVGLPYVPPYFGSKNRNFDKGVNFAVAGATA-LKSSFLKK 125
Query: 126 GKLSP---FSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIF-DIGQNDL 181
+ P SL +Q FK+ + + + D AL + +IG ND
Sbjct: 126 RGIQPHTNVSLGVQLKSFKKSLP---------NLCGSPSDCRDMIGNALILMGEIGGNDY 176
Query: 182 TIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPSA 241
FF K +++V VP ++ + I + +G ++F + G P GC+ V L + ++
Sbjct: 177 NFPFFNRKPVKEVEELVPFVIASISSTITELIGMGGKTFLVPGEFPIGCSVVYLTLYKTS 236
Query: 242 IKDSY----GCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEK 297
KD Y GC K N+ +Y + KLK L LR I Y D Y +F P K
Sbjct: 237 NKDEYDPSTGCLKWLNKFGEYHSEKLKVELNRLRKLYPHVNIIYADYYNSLLRIFKEPAK 296
Query: 298 YGF-ELPFVACCGYGGEYNIGV--GCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAA 354
+GF E PF ACCG GG YN CG+ + SCK+PS + WDGVH TEAA
Sbjct: 297 FGFMERPFPACCGIGGPYNFNFTRKCGS--------VGVKSCKDPSKYVGWDGVHMTEAA 348
Query: 355 NEIVFSQILTGVFNDPPISLDRACYR 380
+ + IL G + +PP DR+C R
Sbjct: 349 YKWIADGILNGPYANPP--FDRSCLR 372
>At3g62280 putative protein
Length = 343
Score = 199 bits (505), Expect = 3e-51
Identities = 133/369 (36%), Positives = 187/369 (50%), Gaps = 50/369 (13%)
Query: 18 LVLCITPPIFATKNCDF-----PAIFSFGASNVDTGGLAAAFRAPPS-PYGETYFHRSTG 71
LVLC++ + + + P + +FG SN DTGG+ A P P+G T+FHR TG
Sbjct: 13 LVLCLSLLVCSNSETSYKSNKKPILINFGDSNSDTGGVLAGVGLPIGLPHGITFFHRGTG 72
Query: 72 RFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSILPNGKLSPF 131
R DGR+I+DF ++ YLSPYL+SL NF G NFA G+T LP + F
Sbjct: 73 RLGDGRLIVDFYCEHLKMTYLSPYLDSLSPNFKRGVNFAVSGAT------ALP---IFSF 123
Query: 132 SLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGFF-GNKT 190
L IQ QF F ++++ + G + ++ F ALY+ DIGQNDL + + N T
Sbjct: 124 PLAIQIRQFVHFKNRSQELISSG---RRDLIDDNGFRNALYMIDIGQNDLLLALYDSNLT 180
Query: 191 IQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPSAIKDSYGCAK 250
V +P ++ + I+ G + +H + D GC +
Sbjct: 181 YAPVVEKIPSMLLEIKKAIQ-----GELAIHLHN---------------DSDLDPIGCFR 220
Query: 251 QYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACCGY 310
+NEV++ FN L ELRS A + YVDIY+ KY L + + YGF P +ACCGY
Sbjct: 221 VHNEVAKAFNKGLLSLCNELRSQFKDATLVYVDIYSIKYKLSADFKLYGFVDPLMACCGY 280
Query: 311 GG---EYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFSQILTGVF 367
GG Y+ CG G+ I C++ + I+WDGVHYTEAAN V +LT +
Sbjct: 281 GGRPNNYDRKATCGQP----GSTI----CRDVTKAIVWDGVHYTEAANRFVVDAVLTNRY 332
Query: 368 NDPPISLDR 376
+ P SLDR
Sbjct: 333 SYPKNSLDR 341
>At1g28640 lipase, putative
Length = 390
Score = 190 bits (482), Expect = 1e-48
Identities = 131/388 (33%), Positives = 187/388 (47%), Gaps = 40/388 (10%)
Query: 12 LVTLIVLVLCITPPIFATKNCD---FPAIFSFGASNVDTGG---LAAAFRAPPS---PYG 62
L++ +LVL T I A+ F +I SFG S DTG L+ P S PYG
Sbjct: 8 LISSFLLVLYSTTIIVASSESRCRRFKSIISFGDSIADTGNYLHLSDVNHLPQSAFLPYG 67
Query: 63 ETYFHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSI 122
E++FH +GR+SDGR+I+DFIA LPY+ Y S +F G NFA G+T +
Sbjct: 68 ESFFHPPSGRYSDGRLIIDFIAEFLGLPYVPSYFGSQNVSFDQGINFAVYGATALDRVFL 127
Query: 123 LPNGKLSPF---SLQIQYIQFKEFISK--TKLIRDQGGVFATLIPKEDYFSKALYIFDIG 177
+ G S F SL +Q FK+ + T RD +E + + +IG
Sbjct: 128 VGKGIESDFTNVSLSVQLNIFKQILPNLCTSSSRD---------CREMLGDSLILMGEIG 178
Query: 178 QNDLTIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILAN 237
ND FF K+I ++ VP ++ I ++ +LG ++F + G P GC P L
Sbjct: 179 VNDYNYPFFEGKSINEIKQLVPLVIKAISSAIVDLIDLGGKTFLVPGNFPLGCYPAYLTL 238
Query: 238 FPSAIKDSY----GCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFT 293
F +A ++ + GC + NE +Y N +LK L L+ I Y D Y + L+
Sbjct: 239 FQTAAEEDHDPFTGCIPRLNEFGEYHNEQLKTELKRLQELYDHVNIIYADYYNSLFRLYQ 298
Query: 294 NPEKYGFE-LPFVACCGYGGEYNIGVG--CGASININGTKIVAGSCKNPSTRIIWDGVHY 350
P KYGF+ P ACCG GG+YN +G CG C+NPS + WDG H
Sbjct: 299 EPVKYGFKNRPLAACCGVGGQYNFTIGKECGHR--------GVSCCQNPSEYVNWDGYHL 350
Query: 351 TEAANEIVFSQILTGVFNDPPISLDRAC 378
TEA ++ + IL G + P + D +C
Sbjct: 351 TEATHQKMAQVILNGTYASP--AFDWSC 376
>At1g28580 lipase like protein
Length = 390
Score = 187 bits (474), Expect = 1e-47
Identities = 126/377 (33%), Positives = 178/377 (46%), Gaps = 34/377 (9%)
Query: 12 LVTLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTG---GLAAAFRAPPS---PYGETY 65
LV + + +T TK +F +I SFG S DTG GL+ P PYGE +
Sbjct: 13 LVFIFLSTFVVTNVSSETKCREFKSIISFGDSIADTGNLLGLSDPKDLPHMAFPPYGENF 72
Query: 66 FHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSILPN 125
FH TGRFS+GR+I+DFIA LP + P+ S +NF G NFA GG+T +
Sbjct: 73 FHHPTGRFSNGRLIIDFIAEFLGLPLVPPFYGSHNANFEKGVNFAVGGATALERSFLEDR 132
Query: 126 GKLSPF---SLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIF-DIGQNDL 181
G P+ SL +Q FKE + + + D AL + +IG ND
Sbjct: 133 GIHFPYTNVSLGVQLNSFKESLP---------SICGSPSDCRDMIENALILMGEIGGNDY 183
Query: 182 TIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPSA 241
FF +K I+++ +P ++ I + +G R+F + G P GC+ + L + ++
Sbjct: 184 NYAFFVDKGIEEIKELMPLVITTISSAITELIGMGGRTFLVPGEFPVGCSVLYLTSHQTS 243
Query: 242 IKDSY----GCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEK 297
+ Y GC K N+ + +L+ L L+ I Y D Y + L+ P K
Sbjct: 244 NMEEYDPLTGCLKWLNKFGENHGEQLRAELNRLQKLYPHVNIIYADYYNALFHLYQEPAK 303
Query: 298 YGF-ELPFVACCGYGGEYNIGVG--CGASININGTKIVAGSCKNPSTRIIWDGVHYTEAA 354
+GF P ACCG GG YN VG CG I SC +PS + WDGVH TEAA
Sbjct: 304 FGFMNRPLSACCGAGGPYNYTVGRKCGTDI--------VESCDDPSKYVAWDGVHMTEAA 355
Query: 355 NEIVFSQILTGVFNDPP 371
++ IL G + PP
Sbjct: 356 YRLMAEGILNGPYAIPP 372
>At2g27360 putative lipase
Length = 394
Score = 186 bits (472), Expect = 2e-47
Identities = 131/384 (34%), Positives = 186/384 (48%), Gaps = 43/384 (11%)
Query: 12 LVTLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTG---GLAAAFRAPPS---PYGETY 65
L++ + IT T+ +F +I SFG S DTG GL++ P S PYGET+
Sbjct: 9 LLSFFISTFLITVVTSQTRCRNFKSIISFGDSITDTGNLLGLSSPNDLPESAFPPYGETF 68
Query: 66 FHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSIL-- 123
FH +GRFSDGR+I+DFIA +P++ P+ S NF G NFA GG+T + S+L
Sbjct: 69 FHHPSGRFSDGRLIIDFIAEFLGIPHVPPFYGSKNGNFEKGVNFAVGGATA-LECSVLEE 127
Query: 124 --PNGKLSPFSLQIQYIQFKEFI-----SKTKLIRDQGGVFATLIPKEDYFSKALYIFDI 176
+ S SL Q FKE + S + RD E+ F + I +I
Sbjct: 128 KGTHCSQSNISLGNQLKSFKESLPYLCGSSSPDCRDM---------IENAF---ILIGEI 175
Query: 177 GQNDLTIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILA 236
G ND F K I++V VP ++ I + ++GAR+F + G P GC+ L
Sbjct: 176 GGNDYNFPLFDRKNIEEVKELVPLVITTISSAISELVDMGARTFLVPGNFPLGCSVAYLT 235
Query: 237 NFPSAIKDSY----GCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLF 292
+ + K+ Y GC N+ S Y N +L+ L LR+ I Y D Y L
Sbjct: 236 LYETPNKEEYNPLTGCLTWLNDFSVYHNEQLQAELKRLRNLYPHVNIIYGDYYNTLLRLM 295
Query: 293 TNPEKYG-FELPFVACCGYGGEYN--IGVGCGASININGTKIVAGSCKNPSTRIIWDGVH 349
P K+G + P ACCG GG YN + CG+ G + C +PS + WDG+H
Sbjct: 296 QEPSKFGLMDRPLPACCGLGGPYNFTFSIKCGS----KGVEY----CSDPSKYVNWDGIH 347
Query: 350 YTEAANEIVFSQILTGVFNDPPIS 373
TEAA + + +LTG + PP +
Sbjct: 348 MTEAAYKWISEGVLTGPYAIPPFN 371
>At1g28650 lipase, putative
Length = 385
Score = 186 bits (471), Expect = 2e-47
Identities = 128/390 (32%), Positives = 187/390 (47%), Gaps = 45/390 (11%)
Query: 12 LVTLIVLVLCITPPIFATKNCD---FPAIFSFGASNVDTGG---LAAAFRAPPS---PYG 62
L+ +L+L T + A+ + +I SFG S DTG L+ P + PYG
Sbjct: 10 LIVSFLLILYYTTIVVASSESRCRRYKSIISFGDSIADTGNYVHLSNVNNLPQAAFLPYG 69
Query: 63 ETYFHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSI 122
E++FH +GR+SDGR+++DFIA LPY+ PY S +F G NFA G+T +
Sbjct: 70 ESFFHPPSGRYSDGRLVIDFIAEFLGLPYVPPYFGSQNVSFNQGINFAVYGATALDRAFL 129
Query: 123 LPNGKLSPF---SLQIQYIQFKEFI-----SKTKLIRDQGGVFATLIPKEDYFSKALYIF 174
+ G S F SL +Q FK+ + S T+ R+ G + +
Sbjct: 130 VKQGIKSDFTNISLSVQLNTFKQILPNLCASSTRDCREMLG------------DSLILMG 177
Query: 175 DIGQNDLTIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVI 234
+IG ND FF K+I ++ VP I+ I ++ +LG ++F + G P GC+
Sbjct: 178 EIGGNDYNYPFFEGKSINEIKELVPLIIKAISSAIVDLIDLGGKTFLVPGNFPIGCSTAY 237
Query: 235 LANFPSAIKDS---YGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSL 291
L F +A + GC N+ ++ N +LK L +L+ I Y D Y Y L
Sbjct: 238 LTLFQTATVEHDPFTGCIPWLNKFGEHHNEQLKIELKQLQKLYPHVNIIYADYYNSLYGL 297
Query: 292 FTNPEKYGFE-LPFVACCGYGGEYNIGVG--CGASININGTKIVAGSCKNPSTRIIWDGV 348
F P KYGF+ P ACCG GG+YN +G CG NG C+NPS + WDG
Sbjct: 298 FQEPAKYGFKNRPLAACCGVGGQYNFTIGKECGE----NGVSY----CQNPSEYVNWDGY 349
Query: 349 HYTEAANEIVFSQILTGVFNDPPISLDRAC 378
H TEA + + +L G + P + D +C
Sbjct: 350 HLTEATYQKMAQGLLNGRYTTP--AFDWSC 377
>At1g28590 putative lipase (At1g28590)
Length = 403
Score = 185 bits (469), Expect = 4e-47
Identities = 126/388 (32%), Positives = 182/388 (46%), Gaps = 34/388 (8%)
Query: 1 MAKIELSRHIPLVTLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTG---GLAAAFRAP 57
MA ++ + LV I+ L +T T+ +F +I SFG S DTG GL+ P
Sbjct: 1 MASLDSLPAMKLVRFILSTLLVTSVNSQTQCRNFKSIISFGDSIADTGNLLGLSDPNDLP 60
Query: 58 PS---PYGETYFHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGS 114
S PYGET+FH TGR+SDGR+I+DFIA P + P+ +NF G NFA G+
Sbjct: 61 ASAFPPYGETFFHHPTGRYSDGRLIIDFIAEFLGFPLVPPFYGCQNANFKKGVNFAVAGA 120
Query: 115 TINIPKSILPNG---KLSPFSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKAL 171
T P + G ++ SL +Q F E + + + D AL
Sbjct: 121 TALEPSFLEERGIHSTITNVSLSVQLRSFTESLP---------NLCGSPSDCRDMIENAL 171
Query: 172 YIF-DIGQNDLTIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGC 230
+ +IG ND F K +++V VP ++ I + +G R+F + G P G
Sbjct: 172 ILMGEIGGNDYNFALFQRKPVKEVEELVPFVIATISSAITELVCMGGRTFLVPGNFPIGY 231
Query: 231 APVILANFPSAIKDSY----GCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYT 286
+ L + ++ K+ Y GC K N+ S+Y+N +L+E L LR I Y D Y
Sbjct: 232 SASYLTLYKTSNKEEYDPLTGCLKWLNDFSEYYNKQLQEELNGLRKLYPHVNIIYADYYN 291
Query: 287 PKYSLFTNPEKYGF-ELPFVACCGYGGEYNIGVG--CGASININGTKIVAGSCKNPSTRI 343
LF P K+GF P ACCG GG YN CG+ + C +PS +
Sbjct: 292 ALLRLFQEPAKFGFMNRPLPACCGVGGSYNFNFSRRCGS--------VGVEYCDDPSQYV 343
Query: 344 IWDGVHYTEAANEIVFSQILTGVFNDPP 371
+DG+H TEAA ++ +L G + PP
Sbjct: 344 NYDGIHMTEAAYRLISEGLLKGPYAIPP 371
>At1g31550 unknown protein
Length = 394
Score = 181 bits (458), Expect = 8e-46
Identities = 126/389 (32%), Positives = 183/389 (46%), Gaps = 37/389 (9%)
Query: 1 MAKIELSRHIPLVTLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTG---GLAAAFRAP 57
MA ++ + L +L + L ++ ++ +F +I SFG S DTG GL+ P
Sbjct: 1 MASLDSHVLMKLGSLFLSTLFVSIVSSESQCRNFESIISFGDSIADTGNLLGLSDHNNLP 60
Query: 58 PS---PYGETYFHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGS 114
S PYGET+FH TGRFSDGR+I+DFIA LPY+ PY S NF G NFA +
Sbjct: 61 MSAFPPYGETFFHHPTGRFSDGRLIIDFIAEFLGLPYVPPYFGSTNGNFEKGVNFAVASA 120
Query: 115 TINIPKSILPNGKLSP--FSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKE--DYFSKA 170
T + G P FSL +Q FK+ + +P + D A
Sbjct: 121 TALESSFLEEKGYHCPHNFSLGVQLKIFKQSLPN-----------LCGLPSDCRDMIGNA 169
Query: 171 LYIF-DIGQNDLTIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKG 229
L + +IG ND FF + + +V VP +++ I + +G R+F + G P G
Sbjct: 170 LILMGEIGANDYNFPFFQLRPLDEVKELVPLVISTISSAITELIGMGGRTFLVPGGFPLG 229
Query: 230 CAPVILANFPSAIKDSY----GCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIY 285
C+ L ++ + Y GC K N+ +Y + +L+E L LR I Y D Y
Sbjct: 230 CSVAFLTLHQTSNMEEYDPLTGCLKWLNKFGEYHSEQLQEELNRLRKLNPHVNIIYADYY 289
Query: 286 TPKYSLFTNPEKYGF-ELPFVACCGYGGEYNIGV--GCGASININGTKIVAGSCKNPSTR 342
L P KYGF ACCG GG YN + CG+ + +C +PS
Sbjct: 290 NASLRLGREPSKYGFINRHLSACCGVGGPYNFNLSRSCGS--------VGVEACSDPSKY 341
Query: 343 IIWDGVHYTEAANEIVFSQILTGVFNDPP 371
+ WDG+H TEAA++ + ++ G + PP
Sbjct: 342 VAWDGLHMTEAAHKSMADGLVKGPYAIPP 370
>At1g28670 lipase
Length = 384
Score = 181 bits (458), Expect = 8e-46
Identities = 134/386 (34%), Positives = 185/386 (47%), Gaps = 36/386 (9%)
Query: 12 LVTLIVLVLCITPPIFATKNCD---FPAIFSFGASNVDTGG---LAAAFRAPPS---PYG 62
L++ +LVL T I A+ F +I SFG S DTG L+ P S PYG
Sbjct: 8 LISSFLLVLYSTTIIVASSESRCRRFKSIISFGDSIADTGNYLHLSDVNHLPQSAFLPYG 67
Query: 63 ETYFHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSI 122
E++FH +GR S+GR+I+DFIA LPY+ PY S +F G NFA G+T +
Sbjct: 68 ESFFHPPSGRASNGRLIIDFIAEFLGLPYVPPYFGSQNVSFEQGINFAVYGATALDRAFL 127
Query: 123 LPNGKLSPF---SLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQN 179
L G S F SL +Q FK+ + +T KE + + +IG N
Sbjct: 128 LGKGIESDFTNVSLSVQLDTFKQILPNL-------CASSTRDCKEMLGDSLILMGEIGGN 180
Query: 180 DLTIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFP 239
D FF K+I ++ VP IV I ++ +LG ++F + G P GC+ L F
Sbjct: 181 DYNYPFFEGKSINEIKELVPLIVKAISSAIVDLIDLGGKTFLVPGGFPTGCSAAYLTLFQ 240
Query: 240 S-AIKDS---YGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNP 295
+ A KD GC NE ++ N +LK L L+ I Y D + Y + P
Sbjct: 241 TVAEKDQDPLTGCYPLLNEFGEHHNEQLKTELKRLQKFYPHVNIIYADYHNSLYRFYQEP 300
Query: 296 EKYGFE-LPFVACCGYGGEYNIGVG--CGASININGTKIVAGSCKNPSTRIIWDGVHYTE 352
KYGF+ P ACCG GG+YN +G CG +N C+NPS + WDG H TE
Sbjct: 301 AKYGFKNKPLAACCGVGGKYNFTIGKECGYE-GVN-------YCQNPSEYVNWDGYHLTE 352
Query: 353 AANEIVFSQILTGVFNDPPISLDRAC 378
AA + + IL G + P + D +C
Sbjct: 353 AAYQKMTEGILNGPYATP--AFDWSC 376
>At1g28570 lipase like protein
Length = 389
Score = 180 bits (457), Expect = 1e-45
Identities = 121/372 (32%), Positives = 175/372 (46%), Gaps = 30/372 (8%)
Query: 15 LIVLVLCITPPIFATKNCDFPAIFSFGASNVDTGGLAAAF------RAPPSPYGETYFHR 68
LI+ C+T + +F +I SFG S DTG L A + PYGET+FH
Sbjct: 13 LILSTFCLTTVNSEPQCHNFKSIISFGDSIADTGNLLALSDPTNLPKVAFLPYGETFFHH 72
Query: 69 STGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSILPNGKL 128
TGRFS+GR+I+DFIA P + P+ S +NF G NFA GG+T + G
Sbjct: 73 PTGRFSNGRLIIDFIAEFLGFPLVPPFYGSQNANFEKGVNFAVGGATALERSFLEERGIH 132
Query: 129 SPF---SLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIF-DIGQNDLTIG 184
P+ SL +Q FKE + + + D +L + +IG ND
Sbjct: 133 FPYTNVSLAVQLSSFKESLP---------NLCVSPSDCRDMIENSLILMGEIGGNDYNYA 183
Query: 185 FFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPSAIKD 244
FF K I+++ VP ++ I + +G ++F + G P GC+ L+ + ++ +
Sbjct: 184 FFVGKNIEEIKELVPLVIETISSAITELIGMGGKTFLVPGEFPLGCSVAYLSLYQTSNIE 243
Query: 245 SY----GCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGF 300
Y GC K N+ S+Y + +L+ L L+ I Y D Y L P K+GF
Sbjct: 244 EYDPLTGCLKWLNKFSEYHDEQLQAELNRLQKLYPHVNIIYADYYNTLLRLAQEPAKFGF 303
Query: 301 -ELPFVACCGYGGEYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVF 359
P ACC GG +N +G GT+ V C +PS + WDGVH TEAA ++
Sbjct: 304 ISRPLPACCALGGPFNFTLG-----RKRGTQ-VPECCDDPSKYVSWDGVHMTEAAYRLMA 357
Query: 360 SQILTGVFNDPP 371
IL G + PP
Sbjct: 358 EGILKGPYAIPP 369
>At1g28610 lipase, putative
Length = 383
Score = 178 bits (452), Expect = 4e-45
Identities = 129/377 (34%), Positives = 179/377 (47%), Gaps = 37/377 (9%)
Query: 12 LVTLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTG---GLAAAFRAPPS---PYGETY 65
LV+ + L +T T+ + +I SFG S DTG GL+ P + PYGET+
Sbjct: 7 LVSFFLSTLFVTIVSSQTQCRNLESIISFGDSITDTGNLVGLSDRNHLPVTAFLPYGETF 66
Query: 66 FHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSILPN 125
FH TGR +GRII+DFIA LP++ P+ S NF G NFA G+T + SIL
Sbjct: 67 FHHPTGRSCNGRIIIDFIAEFLGLPHVPPFYGSKNGNFEKGVNFAVAGATA-LETSILEK 125
Query: 126 GKL----SPFSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIF-DIGQND 180
+ S SL IQ FKE + + + D A I +IG ND
Sbjct: 126 RGIYYPHSNISLGIQLKTFKESLP---------NLCGSPTDCRDMIGNAFIIMGEIGGND 176
Query: 181 LTIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPS 240
FF NKT +V VP ++ I + ++G R+F + G P GC+ L + +
Sbjct: 177 FNFAFFVNKT-SEVKELVPLVITKISSAIVELVDMGGRTFLVPGNFPLGCSATYLTLYQT 235
Query: 241 AIKDSY----GCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPE 296
+ K+ Y GC N+ S+Y+N KL+ L L I Y D + L+ P
Sbjct: 236 SNKEEYDPLTGCLTWLNDFSEYYNEKLQAELNRLSKLYPHVNIIYGDYFNALLRLYQEPS 295
Query: 297 KYGF-ELPFVACCGYGGEYNIGVG--CGASININGTKIVAGSCKNPSTRIIWDGVHYTEA 353
K+GF + P ACCG GG YN + CG+ G K C +PS + WDGVH TEA
Sbjct: 296 KFGFMDRPLPACCGLGGPYNFTLSKKCGSV----GVKY----CSDPSKYVNWDGVHMTEA 347
Query: 354 ANEIVFSQILTGVFNDP 370
A + + +L G + P
Sbjct: 348 AYKWIADGLLKGPYTIP 364
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.141 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,879,119
Number of Sequences: 26719
Number of extensions: 388915
Number of successful extensions: 1327
Number of sequences better than 10.0: 115
Number of HSP's better than 10.0 without gapping: 107
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 808
Number of HSP's gapped (non-prelim): 127
length of query: 381
length of database: 11,318,596
effective HSP length: 101
effective length of query: 280
effective length of database: 8,619,977
effective search space: 2413593560
effective search space used: 2413593560
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC139354.10


