

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138453.15 + phase: 0 /pseudo/partial
(195 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
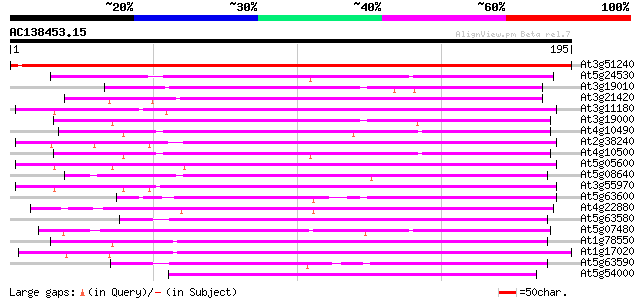
Score E
Sequences producing significant alignments: (bits) Value
At3g51240 flavanone 3-hydroxylase (FH3) 309 5e-85
At5g24530 flavanone 3-hydroxylase-like protein 107 5e-24
At3g19010 oxidase like protein 97 5e-21
At3g21420 unknown protein 95 2e-20
At3g11180 putative leucoanthocyanidin dioxygenase 95 2e-20
At3g19000 unknown protein 95 3e-20
At4g10490 putative flavanone 3-beta-hydroxylase 94 6e-20
At2g38240 putative anthocyanidin synthase 91 5e-19
At4g10500 putative Fe(II)/ascorbate oxidase 90 7e-19
At5g05600 leucoanthocyanidin dioxygenase-like protein 90 9e-19
At5g08640 flavonol synthase (FLS) (sp|Q96330) 89 1e-18
At3g55970 leucoanthocyanidin dioxygenase -like protein 89 1e-18
At5g63600 1-aminocyclopropane-1-carboxylic acid oxidase-like pro... 89 2e-18
At4g22880 putative leucoanthocyanidin dioxygenase (LDOX) 86 2e-17
At5g63580 flavonol synthase 84 4e-17
At5g07480 putative protein 80 5e-16
At1g78550 unknown protein 80 5e-16
At1g17020 SRG1-like protein 80 5e-16
At5g63590 flavonol synthase 80 9e-16
At5g54000 ethylene-forming-enzyme-like dioxygenase 79 2e-15
>At3g51240 flavanone 3-hydroxylase (FH3)
Length = 358
Score = 309 bits (792), Expect = 5e-85
Identities = 148/195 (75%), Positives = 170/195 (86%), Gaps = 1/195 (0%)
Query: 1 MAPVKTLKYLAQEKTLESSFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCNK 60
MAP TL LA E L S F+RD ERPKVAYN FS+EIP+ISLAGIDDVDG R E C +
Sbjct: 1 MAP-GTLTELAGESKLNSKFVRDEDERPKVAYNVFSDEIPVISLAGIDDVDGKRGEICRQ 59
Query: 61 IVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHL 120
IVEACENWGIFQ+VDHGVD L+++MTRLA+ FF LPPE+KLRFD+SGGKKGGFIVSSHL
Sbjct: 60 IVEACENWGIFQVVDHGVDTNLVADMTRLARDFFALPPEDKLRFDMSGGKKGGFIVSSHL 119
Query: 121 QGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAM 180
QGE V+DWRE++ YFSYP++ RDYSRWP+KPEGW +VTE+YSE+LMSL+CKLLEVLSEAM
Sbjct: 120 QGEAVQDWREIVTYFSYPVRNRDYSRWPDKPEGWVKVTEEYSERLMSLACKLLEVLSEAM 179
Query: 181 GLEKEALTKACVDMD 195
GLEKE+LT ACVDMD
Sbjct: 180 GLEKESLTNACVDMD 194
>At5g24530 flavanone 3-hydroxylase-like protein
Length = 341
Score = 107 bits (266), Expect = 5e-24
Identities = 54/176 (30%), Positives = 104/176 (58%), Gaps = 7/176 (3%)
Query: 15 TLESSFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIV 74
TL +++R +S+RP+++ + + P+I L+ D R +I +AC +G FQ++
Sbjct: 14 TLPENYVRPISDRPRLSEVSQLEDFPLIDLSSTD-----RSFLIQQIHQACARFGFFQVI 68
Query: 75 DHGVDQKLISEMTRLAKGFFDLPPEEKLR-FDLSGGKKGGFIVSSHLQGEPVRDWREMMI 133
+HGV++++I EM +A+ FF + EEK++ + K S +++ E V +WR+ +
Sbjct: 69 NHGVNKQIIDEMVSVAREFFSMSMEEKMKLYSDDPTKTTRLSTSFNVKKEEVNNWRDYLR 128
Query: 134 YFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEALTK 189
YPI + + WP+ P +KE+ +YS ++ + K+ E++SE++GLEK+ + K
Sbjct: 129 LHCYPI-HKYVNEWPSNPPSFKEIVSKYSREVREVGFKIEELISESLGLEKDYMKK 183
>At3g19010 oxidase like protein
Length = 349
Score = 97.1 bits (240), Expect = 5e-21
Identities = 55/165 (33%), Positives = 88/165 (53%), Gaps = 16/165 (9%)
Query: 34 NFSNEIPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGF 93
N EIP+I L+ +DD + ++ ++I +ACE WG FQ+++HGV + + K F
Sbjct: 23 NQPEEIPVIDLSRLDDPEDVQ-NVISEIGDACEKWGFFQVLNHGVPSDARQRVEKTVKMF 81
Query: 94 FDLPPEEKLRFDLSGGKKGGFIVSSHLQGEPVRDWREMM-IYFSYPI------------K 140
FDLP EEK++ G+ H + V+DW+E+ IYF P+
Sbjct: 82 FDLPMEEKIKVKRDDVNPVGYHDGEHTKN--VKDWKEVFDIYFKDPMVIPSTTDPEDEGL 139
Query: 141 QRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKE 185
+ Y++WP P ++E E Y+ L+ KLLE++S ++GL KE
Sbjct: 140 RLVYNKWPQSPSDFREACEVYARHAEKLAFKLLELISLSLGLPKE 184
>At3g21420 unknown protein
Length = 364
Score = 95.1 bits (235), Expect = 2e-20
Identities = 52/172 (30%), Positives = 97/172 (56%), Gaps = 7/172 (4%)
Query: 20 FIRDVSERPKVAYN----NFSNEIPIISLAGID--DVDGLRIETCNKIVEACENWGIFQI 73
FIR+ ER V + + ++IP+I L+ + D D E K+ +ACE+WG FQ+
Sbjct: 32 FIREEYERGVVVSSLKTHHLHHQIPVIDLSKLSKPDNDDFFFEIL-KLSQACEDWGFFQV 90
Query: 74 VDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQGEPVRDWREMMI 133
++HG++ +++ ++ +A FFD+P EEK ++ + G G+ + + DW M
Sbjct: 91 INHGIEVEVVEDIEEVASEFFDMPLEEKKKYPMEPGTVQGYGQAFIFSEDQKLDWCNMFA 150
Query: 134 YFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKE 185
+P + R+ WP+KP + E E YS+++ L +LL+ ++ ++GL++E
Sbjct: 151 LGVHPPQIRNPKLWPSKPARFSESLEGYSKEIRELCKRLLKYIAISLGLKEE 202
>At3g11180 putative leucoanthocyanidin dioxygenase
Length = 400
Score = 95.1 bits (235), Expect = 2e-20
Identities = 54/193 (27%), Positives = 100/193 (50%), Gaps = 6/193 (3%)
Query: 3 PVKTLKYLAQEK--TLESSFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGL---RIET 57
P+ ++ LA+ +L +I+ S+RP+ + E+ I++ I D+D L +
Sbjct: 52 PIVRVQSLAESNLTSLPDRYIKPPSQRPQTTIIDHQPEVADINIP-IIDLDSLFSGNEDD 110
Query: 58 CNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVS 117
+I EAC WG FQ+++HGV +L+ K FF+LP E K + S G+
Sbjct: 111 KKRISEACREWGFFQVINHGVKPELMDAARETWKSFFNLPVEAKEVYSNSPRTYEGYGSR 170
Query: 118 SHLQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLS 177
++ + DW + P+ +D+++WP+ P +E+ ++Y ++L+ L +L+ +LS
Sbjct: 171 LGVEKGAILDWNDYYYLHFLPLALKDFNKWPSLPSNIREMNDEYGKELVKLGGRLMTILS 230
Query: 178 EAMGLEKEALTKA 190
+GL E L +A
Sbjct: 231 SNLGLRAEQLQEA 243
>At3g19000 unknown protein
Length = 352
Score = 94.7 bits (234), Expect = 3e-20
Identities = 55/191 (28%), Positives = 96/191 (49%), Gaps = 20/191 (10%)
Query: 16 LESSFIRDVSERPKVAYNN-----FSNEIPIISLAGIDDVDGLRIETCNKIVEACENWGI 70
L+ +FI+ RP N FS+EIP I L+ ++D + +I EAC+ WG
Sbjct: 4 LDEAFIQAPEHRPNTHLTNSGDFIFSDEIPTIDLSSLEDTHHDKTAIAKEIAEACKRWGF 63
Query: 71 FQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQGEPVRDWRE 130
FQ+++HG+ L + + A FF+L EEK + G+ H + VRDW+E
Sbjct: 64 FQVINHGLPSALRHRVEKTAAEFFNLTTEEKRKVKRDEVNPMGYHDEEHTKN--VRDWKE 121
Query: 131 MMIYFSYPIK-------------QRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLS 177
+ +F ++ ++WP P ++EV ++Y+ ++ L+ +LLE++S
Sbjct: 122 IFDFFLQDSTIVPASPEPEDTELRKLTNQWPQNPSHFREVCQEYAREVEKLAFRLLELVS 181
Query: 178 EAMGLEKEALT 188
++GL + LT
Sbjct: 182 ISLGLPGDRLT 192
>At4g10490 putative flavanone 3-beta-hydroxylase
Length = 348
Score = 93.6 bits (231), Expect = 6e-20
Identities = 55/173 (31%), Positives = 99/173 (56%), Gaps = 5/173 (2%)
Query: 18 SSFIRDVSERPKVAYNNFSNE-IPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIVDH 76
S+++R VS+RPK++ S + IP+I L + + R + N+ AC + G FQI +H
Sbjct: 20 SNYVRPVSDRPKMSEVQTSGDSIPLIDLHDLHGPN--RADIINQFAHACSSCGFFQIKNH 77
Query: 77 GVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSS-HLQGEPVRDWREMMIYF 135
GV ++ I +M A+ FF E+++ + KK + +S ++ E V +WR+ +
Sbjct: 78 GVPEETIKKMMNAAREFFRQSESERVKHYSADTKKTTRLSTSFNVSKEKVSNWRDFLRLH 137
Query: 136 SYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEALT 188
YPI+ + WP+ P ++EVT +Y+ + +L LLE +SE++GL K+ ++
Sbjct: 138 CYPIEDF-INEWPSTPISFREVTAEYATSVRALVLTLLEAISESLGLAKDRVS 189
>At2g38240 putative anthocyanidin synthase
Length = 353
Score = 90.5 bits (223), Expect = 5e-19
Identities = 55/193 (28%), Positives = 101/193 (51%), Gaps = 10/193 (5%)
Query: 3 PVKTLKYLAQE--KTLESSFIRDVSERP--KVAYNNFSNEIPIISLAGI-DDVDGLRIET 57
P+ +++ L+Q T+ + +++ +RP ++ EIP++ + + +GLR+
Sbjct: 8 PIVSVQSLSQTGVPTVPNRYVKPAHQRPVFNTTQSDAGIEIPVLDMNDVWGKPEGLRL-- 65
Query: 58 CNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVS 117
+ ACE WG FQ+V+HGV L+ + + FF+LP EEK ++ S G+
Sbjct: 66 ---VRSACEEWGFFQMVNHGVTHSLMERVRGAWREFFELPLEEKRKYANSPDTYEGYGSR 122
Query: 118 SHLQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLS 177
+ + DW + P R+ S+WP++P +E+ E+Y E++ L +L E LS
Sbjct: 123 LGVVKDAKLDWSDYFFLNYLPSSIRNPSKWPSQPPKIRELIEKYGEEVRKLCERLTETLS 182
Query: 178 EAMGLEKEALTKA 190
E++GL+ L +A
Sbjct: 183 ESLGLKPNKLMQA 195
>At4g10500 putative Fe(II)/ascorbate oxidase
Length = 349
Score = 90.1 bits (222), Expect = 7e-19
Identities = 49/175 (28%), Positives = 96/175 (54%), Gaps = 5/175 (2%)
Query: 16 LESSFIRDVSERPKVAYNNFSNE-IPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIV 74
+ S+++R +S+RP ++ S + IP+I L + + R ++ AC +G FQI
Sbjct: 20 IPSNYVRPISDRPNLSEVESSGDSIPLIDLRDLHGPN--RAVIVQQLASACSTYGFFQIK 77
Query: 75 DHGVDQKLISEMTRLAKGFFDLPPEEKLR-FDLSGGKKGGFIVSSHLQGEPVRDWREMMI 133
+HGV +++M +A+ FF P E+++ + K S ++ + V +WR+ +
Sbjct: 78 NHGVPDTTVNKMQTVAREFFHQPESERVKHYSADPTKTTRLSTSFNVGADKVLNWRDFLR 137
Query: 134 YFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEALT 188
+PI+ WP+ P ++EVT +Y+ + +L +LLE +SE++GLE + ++
Sbjct: 138 LHCFPIEDF-IEEWPSSPISFREVTAEYATSVRALVLRLLEAISESLGLESDHIS 191
>At5g05600 leucoanthocyanidin dioxygenase-like protein
Length = 371
Score = 89.7 bits (221), Expect = 9e-19
Identities = 55/192 (28%), Positives = 97/192 (49%), Gaps = 4/192 (2%)
Query: 3 PVKTLKYLAQEK--TLESSFIRDVSERPKVAYNN-FSNEIPIISLAGIDDVDGLRIETCN 59
P+ ++ LA+ +L +I+ S RP + + IPII L G+ +GL +
Sbjct: 23 PIVRVQSLAESNLSSLPDRYIKPASLRPTTTEDAPTATNIPIIDLEGLFSEEGLSDDVIM 82
Query: 60 -KIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSS 118
+I EAC WG FQ+V+HGV +L+ + FF +P K + S G+
Sbjct: 83 ARISEACRGWGFFQVVNHGVKPELMDAARENWREFFHMPVNAKETYSNSPRTYEGYGSRL 142
Query: 119 HLQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSE 178
++ DW + P +D+++WP+ P +EV ++Y E+L+ LS +++ VLS
Sbjct: 143 GVEKGASLDWSDYYFLHLLPHHLKDFNKWPSFPPTIREVIDEYGEELVKLSGRIMRVLST 202
Query: 179 AMGLEKEALTKA 190
+GL+++ +A
Sbjct: 203 NLGLKEDKFQEA 214
>At5g08640 flavonol synthase (FLS) (sp|Q96330)
Length = 336
Score = 89.0 bits (219), Expect = 1e-18
Identities = 53/170 (31%), Positives = 87/170 (51%), Gaps = 6/170 (3%)
Query: 20 FIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIVDHGVD 79
FIR E+P A F P I + + D D + +V+A E WG+FQ+V+HG+
Sbjct: 23 FIRSEKEQP--AITTFRGPTPAIPVVDLSDPDEESVRRA--VVKASEEWGLFQVVNHGIP 78
Query: 80 QKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQGEP--VRDWREMMIYFSY 137
+LI + + + FF+LP EK K + LQ +P + W + + + +
Sbjct: 79 TELIRRLQDVGRKFFELPSSEKESVAKPEDSKDIEGYGTKLQKDPEGKKAWVDHLFHRIW 138
Query: 138 PIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEAL 187
P +Y WP P ++EV E+Y+ + LS LL +LS+ +GL+++AL
Sbjct: 139 PPSCVNYRFWPKNPPEYREVNEEYAVHVKKLSETLLGILSDGLGLKRDAL 188
>At3g55970 leucoanthocyanidin dioxygenase -like protein
Length = 363
Score = 89.0 bits (219), Expect = 1e-18
Identities = 53/196 (27%), Positives = 105/196 (53%), Gaps = 9/196 (4%)
Query: 3 PVKTLKYLAQEK--TLESSFIRDVSERPKVAYNNFSNE----IPIISLAGI--DDVDGLR 54
P+ ++ L++ + + +++ +S+RP + +N N IPII L + DD+ L+
Sbjct: 10 PIVRVQSLSESNLGAIPNRYVKPLSQRPNITPHNKHNPQTTTIPIIDLGRLYTDDLT-LQ 68
Query: 55 IETCNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGF 114
+T ++I +AC G FQ+V+HG+ +L+ + + FF+LP E K S G+
Sbjct: 69 AKTLDEISKACRELGFFQVVNHGMSPQLMDQAKATWREFFNLPMELKNMHANSPKTYEGY 128
Query: 115 IVSSHLQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLE 174
++ + DW + P +DY++WP+ P +E+ E Y ++++ L L++
Sbjct: 129 GSRLGVEKGAILDWSDYYYLHYQPSSLKDYTKWPSLPLHCREILEDYCKEMVKLCENLMK 188
Query: 175 VLSEAMGLEKEALTKA 190
+LS+ +GL+++ L A
Sbjct: 189 ILSKNLGLQEDRLQNA 204
>At5g63600 1-aminocyclopropane-1-carboxylic acid oxidase-like
protein
Length = 325
Score = 88.6 bits (218), Expect = 2e-18
Identities = 50/157 (31%), Positives = 89/157 (55%), Gaps = 17/157 (10%)
Query: 38 EIPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLP 97
++P++ L+ + D D L ++V+A E WG+FQ+V+HG+ +L+ ++ + FF+LP
Sbjct: 32 DVPVVDLS-VSDEDFL----VREVVKASEEWGVFQVVNHGIPTELMRQLQMVGTQFFELP 86
Query: 98 PEEKLRF----DLSGGKKGGFIVSSHLQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEG 153
EK D G KK ++L G + +W E + + P +Y WP P
Sbjct: 87 DAEKETVAKEEDFEGYKK------NYLGG--INNWDEHLFHRLSPPSIINYKYWPKNPPQ 138
Query: 154 WKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEALTKA 190
++EVTE+Y++ + L+ K+L LSE +GL++E T++
Sbjct: 139 YREVTEEYTKHMKRLTEKILGWLSEGLGLQRETFTQS 175
>At4g22880 putative leucoanthocyanidin dioxygenase (LDOX)
Length = 356
Score = 85.5 bits (210), Expect = 2e-17
Identities = 53/185 (28%), Positives = 92/185 (49%), Gaps = 8/185 (4%)
Query: 8 KYLAQEKTLESSFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETC-NKIVEACE 66
+Y+ ++ LES I DV K ++P I L I+ D E C ++ +A
Sbjct: 21 EYIRPKEELES--INDVFLEEK---KEDGPQVPTIDLKNIESDDEKIRENCIEELKKASL 75
Query: 67 NWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRF--DLSGGKKGGFIVSSHLQGEP 124
+WG+ +++HG+ L+ + + + FF L EEK ++ D + GK G+
Sbjct: 76 DWGVMHLINHGIPADLMERVKKAGEEFFSLSVEEKEKYANDQATGKIQGYGSKLANNASG 135
Query: 125 VRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEK 184
+W + + +YP ++RD S WP P + E T +Y++ L L+ K+ + LS +GLE
Sbjct: 136 QLEWEDYFFHLAYPEEKRDLSIWPKTPSDYIEATSEYAKCLRLLATKVFKALSVGLGLEP 195
Query: 185 EALTK 189
+ L K
Sbjct: 196 DRLEK 200
>At5g63580 flavonol synthase
Length = 310
Score = 84.3 bits (207), Expect = 4e-17
Identities = 47/149 (31%), Positives = 77/149 (51%), Gaps = 5/149 (3%)
Query: 39 IPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPP 98
IPII L+ +D+ + +V+ E WGIF +V+HG+ LI + + FF+LP
Sbjct: 19 IPIIDLSNLDEE-----LVAHAVVKGSEEWGIFHVVNHGIPMDLIQRLKDVGTQFFELPE 73
Query: 99 EEKLRFDLSGGKKGGFIVSSHLQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVT 158
EK G K +++L+ W E + + +P ++ WP P ++EV
Sbjct: 74 TEKKAVAKQDGSKDFEGYTTNLKYVKGEVWTENLFHRIWPPTCINFDYWPKNPPQYREVI 133
Query: 159 EQYSEKLMSLSCKLLEVLSEAMGLEKEAL 187
E+Y+++ LS ++L LSE +GL EAL
Sbjct: 134 EEYTKETKKLSERILGYLSEGLGLPSEAL 162
>At5g07480 putative protein
Length = 356
Score = 80.5 bits (197), Expect = 5e-16
Identities = 49/183 (26%), Positives = 101/183 (54%), Gaps = 10/183 (5%)
Query: 11 AQEKTLE---SSFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCNKIVEACEN 67
AQE++L ++ S +P ++ S +P I ++ + D R ++ AC++
Sbjct: 16 AQERSLPYVPDCYVVPPSSKP---CDSNSGIVPTIDVSRLKGGDDERRGVIQELSLACQH 72
Query: 68 WGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQG--EPV 125
G FQIV+HG++Q ++ + +A FF+LP +EK +F +S S+ L+ + +
Sbjct: 73 LGFFQIVNHGINQNILDDALEVANSFFELPAKEKKQF-MSNDVYAPVRYSTSLKDGLDTI 131
Query: 126 RDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKE 185
+ WR + ++++P+ R WP P G++E ++ E++ LS +L+ ++E++GL ++
Sbjct: 132 QFWRIFLKHYAHPL-HRWIHLWPENPPGYREKMGKFCEEVRKLSIELMGAITESLGLGRD 190
Query: 186 ALT 188
L+
Sbjct: 191 YLS 193
>At1g78550 unknown protein
Length = 356
Score = 80.5 bits (197), Expect = 5e-16
Identities = 45/174 (25%), Positives = 90/174 (50%), Gaps = 2/174 (1%)
Query: 15 TLESSFIRDVSERPKVAYNN-FSNEIPIISLAGIDDVDGLRIETCNKIVEACENWGIFQI 73
T+ ++R E+ ++ ++ S+EIP+I + + V + E K+ AC++WG FQ+
Sbjct: 28 TIPPRYVRVDQEKTEILNDSSLSSEIPVIDMTRLCSVSAMDSEL-KKLDFACQDWGFFQL 86
Query: 74 VDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQGEPVRDWREMMI 133
V+HG+D + ++ + FF+LP +EK + G+ GF + + DW +M I
Sbjct: 87 VNHGIDSSFLEKLETEVQEFFNLPMKEKQKLWQRSGEFEGFGQVNIVSENQKLDWGDMFI 146
Query: 134 YFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEAL 187
+ PI+ R + P ++E E YS ++ S++ L ++ + ++ E +
Sbjct: 147 LTTEPIRSRKSHLFSKLPPPFRETLETYSSEVKSIAKILFAKMASVLEIKHEEM 200
>At1g17020 SRG1-like protein
Length = 358
Score = 80.5 bits (197), Expect = 5e-16
Identities = 50/196 (25%), Positives = 102/196 (51%), Gaps = 5/196 (2%)
Query: 4 VKTLKYLAQEKTLES---SFIRDVSERPKVAYN-NFSNEIPIISLAGIDDVDGLRIETCN 59
V +++ + +EKT+ + ++R ++ +V + + EIPII + + + E
Sbjct: 14 VPSVQEMVKEKTITTVPPRYVRSDQDKTEVDDDFDVKIEIPIIDMKRLCSSTTMDSEV-E 72
Query: 60 KIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSH 119
K+ AC+ WG FQ+V+HG+D + ++ + FF+LP EEK +F + GF +
Sbjct: 73 KLDFACKEWGFFQLVNHGIDSSFLDKVKSEIQDFFNLPMEEKKKFWQRPDEIEGFGQAFV 132
Query: 120 LQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEA 179
+ + DW ++ + P++ R +P P +++ E YS ++ S++ L+ ++ A
Sbjct: 133 VSEDQKLDWADLFFHTVQPVELRKPHLFPKLPLPFRDTLEMYSSEVQSVAKILIAKMARA 192
Query: 180 MGLEKEALTKACVDMD 195
+ ++ E L K D+D
Sbjct: 193 LEIKPEELEKLFDDVD 208
>At5g63590 flavonol synthase
Length = 308
Score = 79.7 bits (195), Expect = 9e-16
Identities = 49/159 (30%), Positives = 82/159 (50%), Gaps = 17/159 (10%)
Query: 36 SNEIPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFD 95
S +IP+I L+ D+ + +V+A + WGIFQ+V+HG+ +LI + ++ FF+
Sbjct: 11 SLDIPVIDLSNPDEE-----LVASAVVKASQEWGIFQVVNHGIPTELILRLLQVGMEFFE 65
Query: 96 LPPEEKL-------RFDLSGGKKGGFIVSSHLQGEPVRDWREMMIYFSYPIKQRDYSRWP 148
LP EK D+ G + L+G W + + + +P + ++ WP
Sbjct: 66 LPETEKEAVAKPEDSLDIEGYRTK---YQKDLEGR--NAWVDHLFHRIWPPSRVNHKFWP 120
Query: 149 NKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEAL 187
P + EV E+Y+ + LS K++E LSE +GL EAL
Sbjct: 121 KNPPEYIEVNEEYASHIKKLSEKIMEWLSEGLGLRHEAL 159
>At5g54000 ethylene-forming-enzyme-like dioxygenase
Length = 349
Score = 79.0 bits (193), Expect = 2e-15
Identities = 38/128 (29%), Positives = 65/128 (50%)
Query: 56 ETCNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFI 115
E +K+ A WG+ Q+++HG+ + L+ ++ L K FF LP +EK ++ GF
Sbjct: 63 EELSKLHSAISTWGVVQVMNHGISEALLDKIHELTKQFFVLPTKEKQKYAREISSFQGFG 122
Query: 116 VSSHLQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEV 175
L + V DW + + +YP QR WP P G++E +Y+ K + K +
Sbjct: 123 NDMILSDDQVLDWVDRLYLITYPEDQRQLKFWPENPSGFRETLHEYTMKQQLVVEKFFKA 182
Query: 176 LSEAMGLE 183
L+ ++ LE
Sbjct: 183 LARSLELE 190
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.136 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,408,529
Number of Sequences: 26719
Number of extensions: 187359
Number of successful extensions: 634
Number of sequences better than 10.0: 103
Number of HSP's better than 10.0 without gapping: 91
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 476
Number of HSP's gapped (non-prelim): 107
length of query: 195
length of database: 11,318,596
effective HSP length: 94
effective length of query: 101
effective length of database: 8,807,010
effective search space: 889508010
effective search space used: 889508010
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 57 (26.6 bits)
Medicago: description of AC138453.15


