

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138133.4 + phase: 0
(476 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
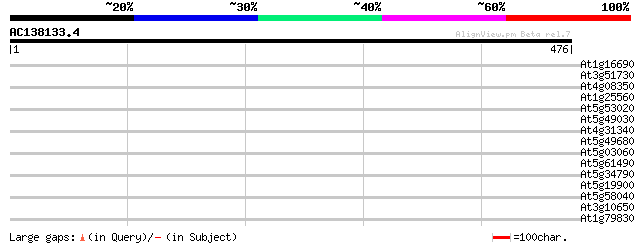
Score E
Sequences producing significant alignments: (bits) Value
At1g16690 Unknown protein 33 0.26
At3g51730 unknown protein 32 0.59
At4g08350 putative protein 32 1.0
At1g25560 unknown protein 31 1.3
At5g53020 putative protein 31 1.7
At5g49030 isoleucyl-tRNA synthetase 30 2.2
At4g31340 unknown protein 30 2.9
At5g49680 putative protein 30 3.8
At5g03060 putative protein 30 3.8
At5g61490 putative protein 29 5.0
At5g34790 putative protein 29 5.0
At5g19900 PRLI-interacting factor A 29 5.0
At5g58040 putative protein 28 8.5
At3g10650 hypothetical protein 28 8.5
At1g79830 unknown protein (At1g79830) 28 8.5
>At1g16690 Unknown protein
Length = 439
Score = 33.5 bits (75), Expect = 0.26
Identities = 40/163 (24%), Positives = 74/163 (44%), Gaps = 21/163 (12%)
Query: 327 KSVKHLGVRSGIAHEAYTQWVIDRAEEIGMPYPAMRYVSSSTP---SMPLPLLPTTQDMY 383
+++K+L ++ G+ H Y+ W R EI P +R + P + P + + +
Sbjct: 173 EALKYLSIKYGVFHAIYSYWKNKR--EIWQK-PILRRLQPPPPVNDTNPYNVFRPREKAH 229
Query: 384 QEHL-AMESRE---------KQVWKARYNQAENLIMTLDGRDEQKTHDNLMLKKELAKVR 433
+ H M+ RE +QV + +QA+ ++ L R+E+K D + + L +++
Sbjct: 230 RLHARRMQQRENNAQSFEKLRQV-RRNLDQAKTILEALIKREEKK-RDFMASEVSLQRIQ 287
Query: 434 RELEEKDELLMRDSKRARGRRDFFARYGDSDSE---SDDAPTT 473
+ + + ELL A R+G S+ E SDD TT
Sbjct: 288 LKYKNETELLEDSLALAGFPLSTAYRFGSSEDEFMDSDDPTTT 330
>At3g51730 unknown protein
Length = 213
Score = 32.3 bits (72), Expect = 0.59
Identities = 25/76 (32%), Positives = 33/76 (42%), Gaps = 17/76 (22%)
Query: 153 LLIYGLILFPNLDNFVDMNAIEIFHSKNPVPTLL-----------------ADTYHAIHD 195
LL+ GLIL + +FVD E +K V TL A+ +HD
Sbjct: 10 LLLLGLILVSDARSFVDSTISEKVSNKEDVCTLCEEYVTDALSYLEKNVTQAEIIEDLHD 69
Query: 196 RTLKGRGYILCCISLL 211
R + RGY CISL+
Sbjct: 70 RCSQLRGYSQQCISLV 85
>At4g08350 putative protein
Length = 1054
Score = 31.6 bits (70), Expect = 1.0
Identities = 21/65 (32%), Positives = 30/65 (45%), Gaps = 3/65 (4%)
Query: 414 RDEQKTHDNLMLKKELAKVRRELEEKDELLMRD---SKRARGRRDFFARYGDSDSESDDA 470
RDE D +L + E EE++E R S+R RGR +F Y + DS+ +D
Sbjct: 5 RDEDDELDGDYEALDLEEEEEEDEEEEEERGRGGGGSRRKRGRSNFIDDYAEEDSQEEDD 64
Query: 471 PTTSY 475
Y
Sbjct: 65 DDEDY 69
>At1g25560 unknown protein
Length = 361
Score = 31.2 bits (69), Expect = 1.3
Identities = 29/124 (23%), Positives = 49/124 (39%), Gaps = 33/124 (26%)
Query: 313 QRKAFMDAWSKVRRKSVKHLGVRSGIAHEAYTQWVIDRAEEIGMPYPAMRYVSSSTPS-- 370
++ + D + + RRK V G RSG+ Y + RA E+ + + TPS
Sbjct: 152 RKHTYADEFEQSRRKFVNGDGKRSGLETATYGNDAVLRAREV-------LFEKTVTPSDV 204
Query: 371 ----------------MPLPLLPTTQDMYQEH-----LAMESREKQVWKARY---NQAEN 406
PLP + T M + +E R +VW+ RY N +++
Sbjct: 205 GKLNRLVIPKQHAEKHFPLPAMTTAMGMNPSPTKGVLINLEDRTGKVWRFRYSYWNSSQS 264
Query: 407 LIMT 410
++T
Sbjct: 265 YVLT 268
>At5g53020 putative protein
Length = 720
Score = 30.8 bits (68), Expect = 1.7
Identities = 22/80 (27%), Positives = 41/80 (50%), Gaps = 3/80 (3%)
Query: 375 LLPTTQDMYQEHLAMESREKQVW---KARYNQAENLIMTLDGRDEQKTHDNLMLKKELAK 431
L T + QEH + R+K+ + + Q E + + + E K H+ L E+ K
Sbjct: 146 LRETQERDVQEHSSELWRQKKTFLELASSQRQLEAELSRANKQIEAKGHELEDLSLEINK 205
Query: 432 VRRELEEKDELLMRDSKRAR 451
+R++LE+KD +L K+++
Sbjct: 206 MRKDLEQKDRILAVMMKKSK 225
>At5g49030 isoleucyl-tRNA synthetase
Length = 1093
Score = 30.4 bits (67), Expect = 2.2
Identities = 22/61 (36%), Positives = 28/61 (45%), Gaps = 3/61 (4%)
Query: 312 GQRKAFMDAWSKVRRKSVKHLGV-RSGIAHEAYTQWVIDRAEEIGMPYPAMRYVSSSTPS 370
G R A MDA + V K V H V R + + W I R G+P PA +V + P
Sbjct: 548 GFRTATMDAINNV--KWVPHQAVNRISAMTSSRSDWCISRQRTWGVPIPAFYHVKTKEPL 605
Query: 371 M 371
M
Sbjct: 606 M 606
>At4g31340 unknown protein
Length = 437
Score = 30.0 bits (66), Expect = 2.9
Identities = 16/70 (22%), Positives = 36/70 (50%)
Query: 381 DMYQEHLAMESREKQVWKARYNQAENLIMTLDGRDEQKTHDNLMLKKELAKVRRELEEKD 440
++ + L +++EK +AR N+AE + L+ ++ N K ++ K+ R ++ +
Sbjct: 122 EVLKNFLEQKNKEKDSTEARTNEAEKKLRELNSSLDKLQKTNEEQKNKIGKLERAIKIAE 181
Query: 441 ELLMRDSKRA 450
E ++R A
Sbjct: 182 EEMLRTKLEA 191
>At5g49680 putative protein
Length = 1378
Score = 29.6 bits (65), Expect = 3.8
Identities = 28/117 (23%), Positives = 53/117 (44%), Gaps = 16/117 (13%)
Query: 368 TPSMPL-PLLPTTQDMYQEHLAMESREKQVWK--------ARYNQAENLIMTLDGRDEQK 418
TP + + PL T + +M SR+ QV AR +A N + L G ++ +
Sbjct: 725 TPDLKVKPLKELTFNSRNITASMTSRQFQVMTDVLSNLLFARLPKAHNDSLKLSGEEDDE 784
Query: 419 THDNL-------MLKKELAKVRRELEEKDELLMRDSKRARGRRDFFARYGDSDSESD 468
+ + + + ELAK+ E +E+D +++ D R + + + + + ESD
Sbjct: 785 VEEEIDEVVPDGIEEVELAKIELEAKERDRMMLLDDIRKLTQNESNSGNINLEKESD 841
>At5g03060 putative protein
Length = 292
Score = 29.6 bits (65), Expect = 3.8
Identities = 17/83 (20%), Positives = 42/83 (50%), Gaps = 1/83 (1%)
Query: 377 PTTQDMYQEHLAMESREKQVWKARYNQAENLIMTLDGRDEQKTHDNLMLKKELAKVRREL 436
PT +++ + + ++ ++ K Y E +++ +Q N ++K + K +EL
Sbjct: 23 PTRRNLVVQEEILATKH-EILKENYENLEKDYKSIEESFKQMNEMNEIMKFQYQKQIKEL 81
Query: 437 EEKDELLMRDSKRARGRRDFFAR 459
EEK L++D ++ R ++ + +
Sbjct: 82 EEKILSLLKDLEKERSEKEEYMK 104
>At5g61490 putative protein
Length = 260
Score = 29.3 bits (64), Expect = 5.0
Identities = 14/42 (33%), Positives = 23/42 (54%)
Query: 205 LCCISLLYRWFISHLPSSFHDNSENWSYSQRIMALTPNEVVW 246
LC SLL + + L +FH + S + +M+LT ++ VW
Sbjct: 118 LCNTSLLQPFMLDRLYDAFHVFQTDPSVQRMVMSLTSDKAVW 159
>At5g34790 putative protein
Length = 400
Score = 29.3 bits (64), Expect = 5.0
Identities = 33/116 (28%), Positives = 54/116 (46%), Gaps = 24/116 (20%)
Query: 364 VSSSTPSMPLPLLPTTQDMYQEHLAMESREKQVWKARYNQAEN--LIMTLDGR------- 414
+S S M LL +D Y + +M ++E+ +W ++ Q N +TL+ R
Sbjct: 38 ISKSVLKMMQGLL---KDPYPTYSSMPAKEQDLWLKQFAQEFNWDRNLTLEVRKAFDKDI 94
Query: 415 --------DEQKTHDNLMLKKELAK---VRRELEEKDELLMRDSKRARGRRDFFAR 459
+E K + M KK+L+K + + K+EL R+S RA RRD A+
Sbjct: 95 SSRYRGTINEWKQRISAMTKKKLSKSKTISPPVVMKEELKRRNS-RADARRDTLAK 149
>At5g19900 PRLI-interacting factor A
Length = 494
Score = 29.3 bits (64), Expect = 5.0
Identities = 21/71 (29%), Positives = 31/71 (43%)
Query: 402 NQAENLIMTLDGRDEQKTHDNLMLKKELAKVRRELEEKDELLMRDSKRARGRRDFFARYG 461
N EN + D Q +NL LK+ L + REL + L +R +D
Sbjct: 306 NVLENRVDDQDSHIAQLEEENLTLKERLFLMERELGDMRRRLQYLERRDMVAQDANEEVV 365
Query: 462 DSDSESDDAPT 472
+++SESD T
Sbjct: 366 ENESESDGDDT 376
>At5g58040 putative protein
Length = 1192
Score = 28.5 bits (62), Expect = 8.5
Identities = 18/77 (23%), Positives = 36/77 (46%), Gaps = 2/77 (2%)
Query: 383 YQEHLAMESREKQVWKARYNQAENLIMTLDGRDEQKTHDNLMLKKELAKVRRELEEKDEL 442
+++ ++ RE+QV + +E+ + +DE K D + K+ + ++ + +D
Sbjct: 868 HEDGISARGRERQVAVRGHRGSEDRSSRM--KDEYKASDKEHVTKDTLRHAKQTKRRDYP 925
Query: 443 LMRDSKRARGRRDFFAR 459
S RG DF AR
Sbjct: 926 GEESSSHHRGHEDFSAR 942
>At3g10650 hypothetical protein
Length = 1309
Score = 28.5 bits (62), Expect = 8.5
Identities = 24/92 (26%), Positives = 37/92 (40%), Gaps = 6/92 (6%)
Query: 216 ISHLPSSFHDNSENWSYSQRIMALTPNEVV-----WLTPAAQVKEIIMGCGDFLNVP-LL 269
+ H PS D + + S + TP + + AQ+ + MG P +L
Sbjct: 191 VRHPPSHERDRTHPDNGSMNTLVSTPPGSLRTLDECIASPAQLAKAYMGSRPSEVTPSML 250
Query: 270 GTRGGINYNPELAMRQFGFPMKSKPISLATSP 301
G RG + + + FP KS +SL T P
Sbjct: 251 GLRGQAGREDSVFLNRTPFPQKSPTMSLVTKP 282
>At1g79830 unknown protein (At1g79830)
Length = 918
Score = 28.5 bits (62), Expect = 8.5
Identities = 23/90 (25%), Positives = 41/90 (45%), Gaps = 2/90 (2%)
Query: 367 STPSMPLPL--LPTTQDMYQEHLAMESREKQVWKARYNQAENLIMTLDGRDEQKTHDNLM 424
S PS P+ L L Q + L E + + Y A+ TL+GR Q +
Sbjct: 670 SLPSSPIQLSCLRAEQGQLSKSLEKERQRAAENRQEYLAAKEEADTLEGRANQLEVEIRE 729
Query: 425 LKKELAKVRRELEEKDELLMRDSKRARGRR 454
L+++ + +E+ +EL+ +D +R + R
Sbjct: 730 LRRKHKQELQEVLLHNELIQKDLEREKASR 759
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.136 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,079,945
Number of Sequences: 26719
Number of extensions: 499603
Number of successful extensions: 1382
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 1372
Number of HSP's gapped (non-prelim): 19
length of query: 476
length of database: 11,318,596
effective HSP length: 103
effective length of query: 373
effective length of database: 8,566,539
effective search space: 3195319047
effective search space used: 3195319047
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC138133.4


