

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138133.3 - phase: 0 /pseudo
(430 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
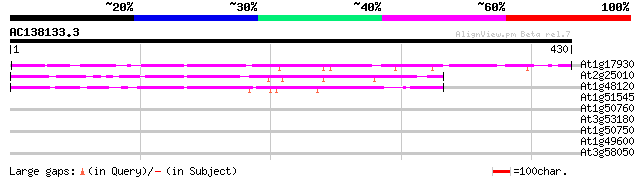
Score E
Sequences producing significant alignments: (bits) Value
At1g17930 unknown protein 96 4e-20
At2g25010 unknown protein 86 4e-17
At1g48120 serine/threonine phosphatase PP7, putative 70 3e-12
At1g51545 unknown protein 38 0.012
At1g50760 hypothetical protein 36 0.036
At3g53180 nodulin / glutamate-ammonia ligase - like protein 31 1.2
At1g50750 hypothetical protein 31 1.5
At1g49600 DNA binding protein ACBF, putative 30 2.0
At3g58050 putative protein 29 4.4
>At1g17930 unknown protein
Length = 478
Score = 95.9 bits (237), Expect = 4e-20
Identities = 111/448 (24%), Positives = 187/448 (40%), Gaps = 75/448 (16%)
Query: 2 PMGEMTVTLDDVACLTHLPIEGRMLAHGKKMPRHEGAALLMTYLGVSQNEAEKICNQEYG 61
P GEMT+TLD+V+ + L ++G+ + G K + + + + LG K+ E
Sbjct: 84 PCGEMTITLDEVSLILGLAVDGKPVV-GVKEKDEDPSQVCLRLLG-------KLPKGELS 135
Query: 62 GY-ISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRCYLLYLVGCLLFGDRS 120
G ++ L++ + A + +E+E Y R YL+Y+VG +F
Sbjct: 136 GNRVTAKWLKESFAECPKGATM-------KEIE-------YHTRAYLIYIVGSTIFATTD 181
Query: 121 NKRIELIYLTTMADGYAGMRNYF*GAMTLAYLYGELADACRPGHRALGGSVTLLTAWFLA 180
+I + YL D + Y GA LA+LY ++ +A + +GG +TLL W
Sbjct: 182 PSKISVDYLILFED-FEKAGEYAWGAAALAFLYRQIGNASQRSQSIIGGCLTLLQCWSYF 240
Query: 181 HFPGFFSVDLNTDYLENYPVAARWK---HQKGHGEGITYRSLLDRIQLDDVCWRPYEKHR 237
H ++D +P+A WK + + YR LD + +V W P+E
Sbjct: 241 H----LNIDRPKRTTRQFPLALLWKGRQQSRSKNDLFKYRKALDDLDPSNVSWCPFEGDL 296
Query: 238 EI--QDFEE--VFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIPRHPTDVRDLPPPSIV 293
+I Q F++ + S + G + V H P+R ++Q+G Q IP ++PP
Sbjct: 297 DIVPQSFKDNLLLGRSRTKLIGPKVVEWHFPDRCMKQFGLCQVIP------GEVPPRKNE 350
Query: 294 Q-----MFVDFGTHTLKADARGEQAGEDTWRVAD--GYVLWYARVSHPQILPPIPRDLPR 346
+ + D T + R E E+ D Y+ W+ ++ +P + RD
Sbjct: 351 KNHDEDLLEDMNTADEEWMRRRENIVENGGGNGDESEYMQWFNSIT----VPKLHRDTSL 406
Query: 347 PANEEQIIAEQWQRYEARSSPDTYDMVSGVVAYADAQLGQEEVMSMTPQ----QWFEAMT 402
A+ + A Q E S+ D+ + + +E VMSM+ W+EA T
Sbjct: 407 EADIMNVQAAILQFDEVASTLSLEDLHP-----EEREAIEEAVMSMSNALRVGDWYEAST 461
Query: 403 HMREQIAPILTRRRAQRPRRRHHQQDQD 430
T +R +RR QQ D
Sbjct: 462 ----------TNKR----KRREEQQQTD 475
>At2g25010 unknown protein
Length = 509
Score = 85.9 bits (211), Expect = 4e-17
Identities = 90/351 (25%), Positives = 146/351 (40%), Gaps = 52/351 (14%)
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRMLAHGKKMPRHEGAALLMTYLGVSQNEAEKICNQEY 60
+P+GEMT+TLD+VA + L I+G + K G + M G + N+E
Sbjct: 92 LPLGEMTITLDEVALVLGLEIDGDPIVGSK-----VGDEVAMDMCGRLLGKLPSAANKE- 145
Query: 61 GGYISYPRLRDFYTSYLGRANVLAGTEDPEELE-ELARVRTYCVRCYLLYLVGCLLFGDR 119
++ R++ ++L R +E PE+ ++ + T R YLLYL+G +F
Sbjct: 146 ---VNCSRVK---LNWLKR----TFSECPEDASFDVVKCHT---RAYLLYLIGSTIFATT 192
Query: 120 SNKRIELIYLTTMADGYAGMRNYF*GAMTLAYLYGELADACRPGHRALGGSVTLLTAWFL 179
++ + YL D + Y GA LA LY L +A + G +TLL W
Sbjct: 193 DGDKVSVKYLPLFED-FDQAGRYAWGAAALACLYRALGNASLKSQSNICGCLTLLQCW-- 249
Query: 180 AHFPGFFSVDLNTDYLEN--YPVAARWKHQ--KGHGEGITYRSLLDRIQLDDVCWRPYEK 235
+F +D+ +P+A WK + + + YR LD + + W PYE+
Sbjct: 250 ----SYFHLDIGRPEKSEACFPLALLWKGKGSRSKTDLSEYRRELDDLDPSKITWCPYER 305
Query: 236 HREI----QDFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIP---------RHPT 282
+ + + S + ++ H P+R LRQ+G Q IP
Sbjct: 306 FENLIPPHIKAKLILGRSKTTLVCFEKIELHFPDRCLRQFGKRQPIPLKVKRRDRKNRRL 365
Query: 283 DVRDLPPPSIVQMFVDFGTHTLKADARGEQAGEDTWRVADG-YVLWYARVS 332
D D + + + G H + + G V DG Y+ WYAR+S
Sbjct: 366 DDLDTSMSLACEEWAERGDHIVDSPGGGNV-------VDDGAYMEWYARIS 409
>At1g48120 serine/threonine phosphatase PP7, putative
Length = 1338
Score = 69.7 bits (169), Expect = 3e-12
Identities = 93/351 (26%), Positives = 143/351 (40%), Gaps = 60/351 (17%)
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRMLAHGKKMPRHEGAALLMTYLGVSQNEAEKICNQEY 60
+P GE+TVTL DV L L ++G + K + A L LG + +
Sbjct: 101 LPAGEITVTLQDVNILLGLRVDGPAVTGSTK---YNWADLCEDLLGHRPGPKDL-----H 152
Query: 61 GGYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYC-VRCYLLYLVGCLLFGDR 119
G ++S LR+ + + DP+E V C R ++L L+ L+GD+
Sbjct: 153 GSHVSLAWLRENFRNL---------PADPDE------VTLKCHTRAFVLALMSGFLYGDK 197
Query: 120 SNKRIELIYLTTMADGYAGMRNYF*GAMTLAYLYGELADACRPGHRALGGSVTLLTAWFL 179
S + L +L + D + + G+ TLA LY EL A + + G + LL W
Sbjct: 198 SKHDVALTFLPLLRD-FDEVAKLSWGSATLALLYRELCRASKRTVSTICGPLVLLQLWAW 256
Query: 180 AHF----PGFFSVDLNTDYLENY------PVAAR--WKHQKGHGEGIT-YRSLLDRIQLD 226
PG D+ Y++ P+ R H++ G+ YR D+ + +
Sbjct: 257 ERLHVGRPGRLK-DVGASYMDGIDGPLPDPLGCRASLSHKENPRGGLDFYRDQFDQQKDE 315
Query: 227 DVCWRPYE-----KHREIQDFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIPRHP 281
V W+PY K I E W + + V H P+RVLRQ+G QTIP
Sbjct: 316 QVIWQPYTPDLLAKIPLICVSGENIWRTVAPLICFDVVEWHRPDRVLRQFGLHQTIPAPC 375
Query: 282 TDVRDLPPPSIVQMFVDFGTHTLKADARGEQAGEDTWRVADGYVLWYARVS 332
+ + L H + D RG+ + + R + LW ARVS
Sbjct: 376 DNEKAL--------------HAI--DKRGKSEYDWSARHSRHIGLWEARVS 410
>At1g51545 unknown protein
Length = 629
Score = 37.7 bits (86), Expect = 0.012
Identities = 73/308 (23%), Positives = 111/308 (35%), Gaps = 65/308 (21%)
Query: 2 PMGEMTVTLDDVACLTHLPIEGRMLAHGKKMPRHEGAALLMTYLGVSQNEAEKICNQEYG 61
P GE T+TL+DV L ++G + A L + + S + EK E
Sbjct: 49 PWGEATITLEDVLVLLGFSVQGSPVF----------APLESSEMRDSVEKLEK-ARLENR 97
Query: 62 GYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRCYLLYLVGCLLFGDRSN 121
G R + +S+LGR +++E A +L + + +F D
Sbjct: 98 GQDGLVRQNLWVSSFLGRG---------DQMEHEA---------FLAFWLSQFVFPDMCR 139
Query: 122 KRIELIYLTTMADGYAGMRNYF*GAMTLAYLYGELADACRPGHRALGGSVTLLTAWFLAH 181
+ I L G R F A+ LA LY +L +VTL + + L
Sbjct: 140 RSISTKVLPMAVRLARGERIAFAPAV-LARLYRDLGQIQASAREKSTPNVTLKSLFKLVQ 198
Query: 182 ---FPGFFSVDLNTDYL-ENYPVAARWKHQKGHGEGITYRSLLDRIQLDDVCWRPYEKHR 237
+ F S + + P +RW Q R+ L D WRPY K
Sbjct: 199 LWAWERFKSTSPKARVIPKGEPRISRWHSQTSKNV---------RLNLVDFDWRPYTKPL 249
Query: 238 EIQD-----FEEVFWYS-------GWI----------MCGVRRVYRHLPERVLRQYGYVQ 275
+I + EE W + G++ + GV V + P RV Q+G Q
Sbjct: 250 QIWNPPRFYPEEAMWMTVDDNLDDGFVSFARCMRVSQLVGVGIVEDYYPNRVAMQFGLAQ 309
Query: 276 TIPRHPTD 283
+P TD
Sbjct: 310 DLPGLVTD 317
>At1g50760 hypothetical protein
Length = 649
Score = 36.2 bits (82), Expect = 0.036
Identities = 28/117 (23%), Positives = 44/117 (36%), Gaps = 28/117 (23%)
Query: 195 LENYPVAARWKHQKGHGEGITYRSLLDRIQLDDVCWRPYEKHREIQDFEEVFWYSGWIMC 254
L P A W K + R +LD ++D WRPY R + +++ +Y MC
Sbjct: 276 LAGKPRVAIWNKLKQQTSNV--RQILDNSKIDSFEWRPYT--RSVTNWKFPQFYPEKAMC 331
Query: 255 ------------------------GVRRVYRHLPERVLRQYGYVQTIPRHPTDVRDL 287
G+ +V + P RV Q+G +Q +P D +L
Sbjct: 332 VHVSPSLDEELISFARCIKDSELVGIEKVEHYFPNRVASQFGMLQDVPSPHVDRNNL 388
>At3g53180 nodulin / glutamate-ammonia ligase - like protein
Length = 845
Score = 31.2 bits (69), Expect = 1.2
Identities = 18/61 (29%), Positives = 31/61 (50%), Gaps = 1/61 (1%)
Query: 134 DGYAGMRNYF*GAMTLA-YLYGELADACRPGHRALGGSVTLLTAWFLAHFPGFFSVDLNT 192
DGYA Y+ GA ++ L+DAC G +L ++ F + GF+ ++++T
Sbjct: 328 DGYASPETYYLGAKKAREVIFLVLSDACASGDLSLMEAIDAAKDIFSRNSIGFYKLNIDT 387
Query: 193 D 193
D
Sbjct: 388 D 388
>At1g50750 hypothetical protein
Length = 816
Score = 30.8 bits (68), Expect = 1.5
Identities = 16/62 (25%), Positives = 30/62 (47%), Gaps = 1/62 (1%)
Query: 217 RSLLDRIQLDDVCWRPYEKHREIQDFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQT 276
R +LD +++D V W P + +++ + G+ V + P RV Q+G +Q
Sbjct: 258 RQILDSLKIDTV-WIPASPNLDVEFVSFARCIKVSQLVGIDNVEHYFPNRVASQFGMLQD 316
Query: 277 IP 278
+P
Sbjct: 317 VP 318
>At1g49600 DNA binding protein ACBF, putative
Length = 445
Score = 30.4 bits (67), Expect = 2.0
Identities = 21/94 (22%), Positives = 39/94 (41%), Gaps = 13/94 (13%)
Query: 338 PPIPRDLPRPANEEQIIAEQWQRYEARSSPDTYDMVSGVVAYADAQLG--QEEVMSMTPQ 395
PP+ + P P ++Q +QWQ+ + ++ + Y A + Q++ M M P
Sbjct: 25 PPLQQSTPPPQQQQQ---QQWQQQQ--------QWMAAMQQYPAAAMAMMQQQQMMMYPH 73
Query: 396 QWFEAMTHMREQIAPILTRRRAQRPRRRHHQQDQ 429
+ Q P Q+ +++HHQ Q
Sbjct: 74 PQYAPYNQAAYQQHPQFQYAAYQQQQQQHHQSQQ 107
>At3g58050 putative protein
Length = 1209
Score = 29.3 bits (64), Expect = 4.4
Identities = 18/51 (35%), Positives = 25/51 (48%)
Query: 64 ISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRCYLLYLVGCL 114
+SY +L+ F++ +A D + L E AR YC RC L L G L
Sbjct: 24 VSYNQLQKFWSELSPKARQELLKIDKQTLFEQARKNMYCSRCNGLLLEGFL 74
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.140 0.448
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,252,449
Number of Sequences: 26719
Number of extensions: 449593
Number of successful extensions: 1158
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 1138
Number of HSP's gapped (non-prelim): 12
length of query: 430
length of database: 11,318,596
effective HSP length: 102
effective length of query: 328
effective length of database: 8,593,258
effective search space: 2818588624
effective search space used: 2818588624
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC138133.3


