

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138130.19 - phase: 0
(881 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
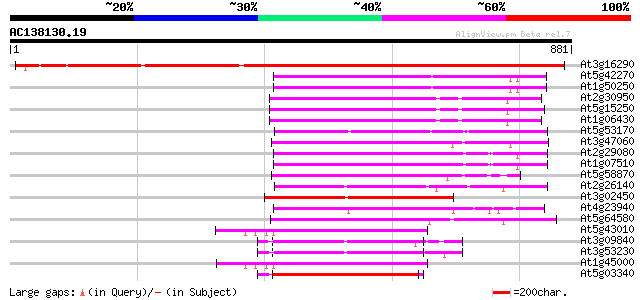
Score E
Sequences producing significant alignments: (bits) Value
At3g16290 putative FtsH-like metalloprotease 1083 0.0
At5g42270 cell division protein FtsH 313 2e-85
At1g50250 putative chloroplast FtsH protease 309 4e-84
At2g30950 zinc dependent protease (VAR2) 303 3e-82
At5g15250 FtsH-like protein Pftf precursor - like 296 4e-80
At1g06430 cell division protease FtsH, putative 293 2e-79
At5g53170 cell division protein FtsH protease-like 283 2e-76
At3g47060 FtsH metalloprotease - like protein 266 4e-71
At2g29080 AAA-type like ATPase 265 6e-71
At1g07510 unknown protein 261 9e-70
At5g58870 cell division protein - like 260 3e-69
At2g26140 FtsH like protease 258 1e-68
At3g02450 cell division protein FtsH-like protein 245 9e-65
At4g23940 cell division protein - like 243 3e-64
At5g64580 unknown protein 232 8e-61
At5g43010 26S proteasome AAA-ATPase subunit RPT4a (gb|AAF22524.1) 194 2e-49
At3g09840 putative transitional endoplasmic reticulum ATPase 193 4e-49
At3g53230 CDC48 - like protein 192 7e-49
At1g45000 26S proteasome AAA-ATPase subunit RPT4a like protein 192 9e-49
At5g03340 transitional endoplasmic reticulum ATPase 191 1e-48
>At3g16290 putative FtsH-like metalloprotease
Length = 876
Score = 1083 bits (2802), Expect = 0.0
Identities = 561/874 (64%), Positives = 691/874 (78%), Gaps = 26/874 (2%)
Query: 9 FPPSSSSSKLKKFPK-----PSKFRSISSQIQTPES---ENDEKNQKNLNFNNLNFLKFT 60
FP SSS P+ P + SIS Q + + E+ + N K N +N L
Sbjct: 5 FPLHSSSPSQFLSPENRQRLPRNYPSISCQNNSATNVVHEDGDDNDK-AKTNQVNLLAIP 63
Query: 61 VTLTVISASLPQAATAVAAAGKKRAPRKASTKKVEALSIEEVKTWIEGLPIVSERIPYTE 120
+TLT+ISASL + + A A +++ +K K EAL++E++K W + LP+VS RIPYT+
Sbjct: 64 ITLTIISASLAKPSFAAAKVTERKRTQK---KPQEALTLEQLKAWSKDLPVVSNRIPYTD 120
Query: 121 IAELKNLGMLKHIVKPSAVELRERAVAVLVVLEDSRVLRTVLPNVESDRKFWGLWDELKI 180
I LK G LKH++KP + LR++A VLVVLEDSRVLRTVLP++E +++FW WDEL I
Sbjct: 121 ILSLKAEGKLKHVIKPPNLSLRQKAEPVLVVLEDSRVLRTVLPSLEGNKRFWEQWDELGI 180
Query: 181 ENLCVNAYSPPVKVPEIPLSVLARIWLSLPFHKPLVEFVNRFQPKKKSKKELALREARMQ 240
+ CVNAY+PPVK P +P L +W K + +PKK+SK+ L+ R
Sbjct: 181 DVQCVNAYTPPVKRPPVPSPYLGFLW------KVPAYMLTWVKPKKESKRAAELKRMRED 234
Query: 241 LQRQKKEEVVKTMKEREMIERNERNKKREAENEKRMR-RRKEYKEKMVEVKANEFFNTTI 299
+RQ+KEE+ +ER M+E+ + +K++ E +KR R+K+Y+E + E + N +
Sbjct: 235 FKRQRKEEIETMKEERVMMEKTMKAQKKQQERKKRKAVRKKKYEESLREARKNYRDMADM 294
Query: 300 WTRMAKDKMAINGIGVLFFVIFYRTVVVSYKKQKKDYEDRIKIQKADAEERRKMREMEAE 359
W R+A+D +G++FF IFYR VV++Y+KQKKDYEDR+KI+KA+A+ER+KMRE+E E
Sbjct: 295 WARLAQDPNVATALGLVFFYIFYRVVVLNYRKQKKDYEDRLKIEKAEADERKKMRELERE 354
Query: 360 MGWSEAGGDEDESELVKEG--EENPYLKMTKEFMKSGARVRRAQNRRLPQYLERGVDVKF 417
M G E+E E V+EG E+NPYL+M +FMKSGARVRRA N+RLP+YLERGVDVKF
Sbjct: 355 ME-----GIEEEDEEVEEGTGEKNPYLQMAMQFMKSGARVRRASNKRLPEYLERGVDVKF 409
Query: 418 TDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEAGV 477
TDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEAGV
Sbjct: 410 TDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEAGV 469
Query: 478 NFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGGQER 537
NFFSISASQFVEIYVGVGASRVR+LYQEA+ENAPSVVFIDELDAVGR+RGLIKGSGGQER
Sbjct: 470 NFFSISASQFVEIYVGVGASRVRALYQEARENAPSVVFIDELDAVGRERGLIKGSGGQER 529
Query: 538 DATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRIEIL 597
DATLNQLLV LDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPG IGR+EIL
Sbjct: 530 DATLNQLLVSLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGLIGRMEIL 589
Query: 598 KVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQAAQM 657
+VHARKKP+AED+DY VASMTDGMVGAELANIVE+AAINMMRD RTE++TDDLLQAAQ+
Sbjct: 590 QVHARKKPMAEDLDYMAVASMTDGMVGAELANIVEIAAINMMRDGRTELTTDDLLQAAQI 649
Query: 658 EERGMLDRKERSKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGRELGYVRTMLE 717
EERGMLDRK+RS E W QVAINEAAMAV A+N P+ NIE++TI PRAGRELGYVR ++
Sbjct: 650 EERGMLDRKDRSLETWRQVAINEAAMAVVAVNFPDMKNIEFLTINPRAGRELGYVRVKMD 709
Query: 718 SINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWAETADNARVAARMYMIGGL 777
I F +GML+RQS+ DHITVQLAPRAADE+W+G+DQLSTIWAET+DNAR AAR ++GGL
Sbjct: 710 HIKFKEGMLSRQSILDHITVQLAPRAADELWYGEDQLSTIWAETSDNARSAARSLVLGGL 769
Query: 778 SDKYRGVSNFWVTDRINEIDLEAMKILNLCYERAKEILQQNKTLMDTLVNELVVKKTLTK 837
SDK+ G++NFWV DRIN+ID+EA++ILN+CYERAKEIL +N+TLMD +V +LV KK+LTK
Sbjct: 770 SDKHHGLNNFWVADRINDIDVEALRILNMCYERAKEILGRNRTLMDEVVEKLVQKKSLTK 829
Query: 838 EDIVRLVQLHGHAKPIPISVLDIRDAKHKELQEI 871
++ LV+L+G +KP+P S+L++R K EL+E+
Sbjct: 830 QEFFTLVELYGSSKPMPPSILELRKIKRLELEEM 863
>At5g42270 cell division protein FtsH
Length = 704
Score = 313 bits (803), Expect = 2e-85
Identities = 188/449 (41%), Positives = 266/449 (58%), Gaps = 22/449 (4%)
Query: 415 VKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGE 474
V F DVAG + +LEL+E+V F + + Y G KIP G LL GPPG GKTLLA+AVAGE
Sbjct: 247 VTFGDVAGADQAKLELQEVVDFLKNPDKYTALGAKIPKGCLLVGPPGTGKTLLARAVAGE 306
Query: 475 AGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGG 534
AGV FFS +AS+FVE++VGVGASRVR L+++AK AP +VFIDE+DAVGR+RG G G
Sbjct: 307 AGVPFFSCAASEFVELFVGVGASRVRDLFEKAKSKAPCIVFIDEIDAVGRQRGAGMGGGN 366
Query: 535 QERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRI 594
ER+ T+NQLL +DGF G VI +A+TNRPD+LD AL+RPGRFDR++ + +P GR+
Sbjct: 367 DEREQTINQLLTEMDGFSGNSGVIVLAATNRPDVLDSALLRPGRFDRQVTVDRPDVAGRV 426
Query: 595 EILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQA 654
+ILKVH+R K I +DVDYE VA T G GA+L N++ AAI R E+S D++ A
Sbjct: 427 QILKVHSRGKAIGKDVDYEKVARRTPGFTGADLQNLMNEAAILAARRELKEISKDEISDA 486
Query: 655 AQMEERGMLDRKER--SKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGRELGYV 712
+ G ++K S+EK VA +EA A+ +P +D + I+I PR G+ G
Sbjct: 487 LERIIAGP-EKKNAVVSEEKKRLVAYHEAGHALVGALMPEYDPVAKISIIPR-GQAGGLT 544
Query: 713 RTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLST-IWAETADNARVAARM 771
G+ +R L + + V L R A+E+ FG + ++T + +RVA +M
Sbjct: 545 FFAPSEERLESGLYSRSYLENQMAVALGGRVAEEVIFGDENVTTGASNDFMQVSRVARQM 604
Query: 772 YMIGGLSDKYRGVS------NFWVTDRINE-----------IDLEAMKILNLCYERAKEI 814
G S K V+ N ++ ++ +D E +++ Y RAKEI
Sbjct: 605 VERFGFSKKIGQVAVGGAGGNPFLGQSMSSQKDYSMATADVVDAEVRELVEKAYVRAKEI 664
Query: 815 LQQNKTLMDTLVNELVVKKTLTKEDIVRL 843
+ ++ L L+ K+T+ E+ + L
Sbjct: 665 ITTQIDILHKLAQLLIEKETVDGEEFMSL 693
>At1g50250 putative chloroplast FtsH protease
Length = 716
Score = 309 bits (792), Expect = 4e-84
Identities = 183/449 (40%), Positives = 268/449 (58%), Gaps = 22/449 (4%)
Query: 415 VKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGE 474
V F DVAG + +LEL+E+V F + + Y G KIP G LL GPPG GKTLLA+AVAGE
Sbjct: 259 VSFADVAGADQAKLELQEVVDFLKNPDKYTALGAKIPKGCLLVGPPGTGKTLLARAVAGE 318
Query: 475 AGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGG 534
AGV FFS +AS+FVE++VGVGASRVR L+++AK AP +VFIDE+DAVGR+RG G G
Sbjct: 319 AGVPFFSCAASEFVELFVGVGASRVRDLFEKAKSKAPCIVFIDEIDAVGRQRGAGMGGGN 378
Query: 535 QERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRI 594
ER+ T+NQLL +DGF G VI +A+TNRPD+LD AL+RPGRFDR++ + +P GR+
Sbjct: 379 DEREQTINQLLTEMDGFSGNSGVIVLAATNRPDVLDSALLRPGRFDRQVTVDRPDVAGRV 438
Query: 595 EILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQA 654
+IL+VH+R K + +DVD++ VA T G GA+L N++ AAI R E+S D++ A
Sbjct: 439 KILQVHSRGKALGKDVDFDKVARRTPGFTGADLQNLMNEAAILAARRELKEISKDEISDA 498
Query: 655 AQMEERGMLDRKER--SKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGRELGYV 712
+ G ++K S+EK VA +EA A+ +P +D + I+I PR G+ G
Sbjct: 499 LERIIAGP-EKKNAVVSEEKKRLVAYHEAGHALVGALMPEYDPVAKISIIPR-GQAGGLT 556
Query: 713 RTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLST-IWAETADNARVAARM 771
G+ +R L + + V L R A+E+ FG + ++T + +RVA +M
Sbjct: 557 FFAPSEERLESGLYSRSYLENQMAVALGGRVAEEVIFGDENVTTGASNDFMQVSRVARQM 616
Query: 772 YMIGGLSDKYRGVS------NFWVTDRINE-----------IDLEAMKILNLCYERAKEI 814
G S K V+ N ++ +++ +D E +++ Y+RA EI
Sbjct: 617 IERFGFSKKIGQVAVGGPGGNPFMGQQMSSQKDYSMATADIVDAEVRELVEKAYKRATEI 676
Query: 815 LQQNKTLMDTLVNELVVKKTLTKEDIVRL 843
+ + ++ L L+ K+T+ E+ + L
Sbjct: 677 ITTHIDILHKLAQLLIEKETVDGEEFMSL 705
>At2g30950 zinc dependent protease (VAR2)
Length = 695
Score = 303 bits (776), Expect = 3e-82
Identities = 191/450 (42%), Positives = 264/450 (58%), Gaps = 33/450 (7%)
Query: 409 LERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLA 468
+E V F DVAG+ + + + E+V+F E + G KIP G+LL GPPG GKTLLA
Sbjct: 218 MEPNTGVTFDDVAGVDEAKQDFMEVVEFLKKPERFTAVGAKIPKGVLLIGPPGTGKTLLA 277
Query: 469 KAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGL 528
KA+AGEAGV FFSIS S+FVE++VGVGASRVR L+++AKENAP +VF+DE+DAVGR+RG
Sbjct: 278 KAIAGEAGVPFFSISGSEFVEMFVGVGASRVRDLFKKAKENAPCIVFVDEIDAVGRQRGT 337
Query: 529 IKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKP 588
G G ER+ TLNQLL +DGFEG VI +A+TNR DILD AL+RPGRFDR++ + P
Sbjct: 338 GIGGGNDEREQTLNQLLTEMDGFEGNTGVIVVAATNRADILDSALLRPGRFDRQVSVDVP 397
Query: 589 GFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVST 648
GR +ILKVHA K DV EI+A T G GA+LAN++ AAI R +RT +S+
Sbjct: 398 DVKGRTDILKVHAGNKKFDNDVSLEIIAMRTPGFSGADLANLLNEAAILAGRRARTSISS 457
Query: 649 ---DDLLQ--AAQMEERGMLDRKERSKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAP 703
DD + A ME M D K +S VA +E AV P D ++ +T+ P
Sbjct: 458 KEIDDSIDRIVAGMEGTVMTDGKSKS-----LVAYHEVGHAVCGTLTPGHDAVQKVTLIP 512
Query: 704 RAGRELGYVRTMLESINFND-GMLTRQSLFDHITVQLAPRAADEMWFGKDQLST-IWAET 761
R G R + I +D ++++Q LF I L RAA+E+ FG +++T +
Sbjct: 513 R-----GQARGLTWFIPSDDPTLISKQQLFARIVGGLGGRAAEEIIFGDSEVTTGAVGDL 567
Query: 762 ADNARVAARMYMIGGLSD---------------KYRGVSNFWVTDRINE-IDLEAMKILN 805
+A +M G+SD R ++ +++++ E ID K+ +
Sbjct: 568 QQITGLARQMVTTFGMSDIGPWSLMDSSAQSDVIMRMMARNSMSEKLAEDIDSAVKKLSD 627
Query: 806 LCYERAKEILQQNKTLMDTLVNELVVKKTL 835
YE A ++ N+ MD LV L+ K+T+
Sbjct: 628 SAYEIALSHIKNNREAMDKLVEVLLEKETI 657
>At5g15250 FtsH-like protein Pftf precursor - like
Length = 687
Score = 296 bits (757), Expect = 4e-80
Identities = 183/455 (40%), Positives = 268/455 (58%), Gaps = 34/455 (7%)
Query: 409 LERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLA 468
+E + F DVAG+ + + + EEIV+F E + G KIP G+LL GPPG GKTLLA
Sbjct: 214 MEPNTGITFEDVAGVDEAKQDFEEIVEFLKTPEKFSALGAKIPKGVLLTGPPGTGKTLLA 273
Query: 469 KAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGL 528
KA+AGEAGV FFS+S S+F+E++VGVGASR R L+ +AK N+P +VFIDE+DAVGR RG
Sbjct: 274 KAIAGEAGVPFFSLSGSEFIEMFVGVGASRARDLFNKAKANSPCIVFIDEIDAVGRMRGT 333
Query: 529 IKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKP 588
G G ER+ TLNQ+L +DGF G VI IA+TNRP+ILD AL+RPGRFDR++ + P
Sbjct: 334 GIGGGNDEREQTLNQILTEMDGFAGNTGVIVIAATNRPEILDSALLRPGRFDRQVSVGLP 393
Query: 589 GFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAI---NMMRDSRTE 645
GR EILKVH+R K + +DV ++A T G GA+LAN++ AAI +D T
Sbjct: 394 DIRGREEILKVHSRSKKLDKDVSLSVIAMRTPGFSGADLANLMNEAAILAGRRGKDKITL 453
Query: 646 VSTDDLLQ--AAQMEERGMLDRKERSKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAP 703
DD + A ME M+D K ++ VA +E A+ A D ++ +T+ P
Sbjct: 454 TEIDDSIDRIVAGMEGTKMIDGKSKA-----IVAYHEVGHAICATLTEGHDPVQKVTLVP 508
Query: 704 RAGRELGYVRTMLESINFND-GMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWA-ET 761
R G R + + D ++++Q LF I L RAA+++ FG+ +++T A +
Sbjct: 509 R-----GQARGLTWFLPGEDPTLVSKQQLFARIVGGLGGRAAEDVIFGEPEITTGAAGDL 563
Query: 762 ADNARVAARMYMIGGLSD----------------KYRGVSNFWVTDRINE-IDLEAMKIL 804
+A +M + G+S+ R ++ +++++ E ID KI+
Sbjct: 564 QQVTEIARQMVTMFGMSEIGPWALTDPAVKQNDVVLRMLARNSMSEKLAEDIDSCVKKII 623
Query: 805 NLCYERAKEILQQNKTLMDTLVNELVVKKTLTKED 839
YE AK+ ++ N+ +D LV+ L+ K+TLT ++
Sbjct: 624 GDAYEVAKKHVRNNREAIDKLVDVLLEKETLTGDE 658
>At1g06430 cell division protease FtsH, putative
Length = 649
Score = 293 bits (751), Expect = 2e-79
Identities = 182/449 (40%), Positives = 263/449 (58%), Gaps = 33/449 (7%)
Query: 409 LERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLA 468
+E V F DVAG+ + + + E+V+F E + G +IP G+LL GPPG GKTLLA
Sbjct: 211 MEPNTGVTFDDVAGVDEAKQDFMEVVEFLKKPERFTAVGARIPKGVLLVGPPGTGKTLLA 270
Query: 469 KAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGL 528
KA+AGEAGV FFSIS S+FVE++VGVGASRVR L+++AKENAP +VF+DE+DAVGR+RG
Sbjct: 271 KAIAGEAGVPFFSISGSEFVEMFVGVGASRVRDLFKKAKENAPCIVFVDEIDAVGRQRGT 330
Query: 529 IKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKP 588
G G ER+ TLNQLL +DGFEG VI +A+TNR DILD AL+RPGRFDR++ + P
Sbjct: 331 GIGGGNDEREQTLNQLLTEMDGFEGNTGVIVVAATNRADILDSALLRPGRFDRQVSVDVP 390
Query: 589 GFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVST 648
GR +ILKVH+ K V E++A T G GA+LAN++ AAI R +T +S+
Sbjct: 391 DVKGRTDILKVHSGNKKFESGVSLEVIAMRTPGFSGADLANLLNEAAILAGRRGKTAISS 450
Query: 649 ---DDLLQ--AAQMEERGMLDRKERSKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAP 703
DD + A ME M D K +S VA +E A+ P D ++ +T+ P
Sbjct: 451 KEIDDSIDRIVAGMEGTVMTDGKSKS-----LVAYHEVGHAICGTLTPGHDAVQKVTLIP 505
Query: 704 RAGRELGYVRTMLESINFND-GMLTRQSLFDHITVQLAPRAADEMWFGKDQLST-IWAET 761
R G R + I +D ++++Q LF I L RAA+E+ FG+ +++T ++
Sbjct: 506 R-----GQARGLTWFIPSDDPTLISKQQLFARIVGGLGGRAAEEVIFGESEVTTGAVSDL 560
Query: 762 ADNARVAARMYMIGGLSD---------------KYRGVSNFWVTDRI-NEIDLEAMKILN 805
+A +M G+S+ R ++ +++++ N+ID + +
Sbjct: 561 QQITGLAKQMVTTFGMSEIGPWSLMDSSEQSDVIMRMMARNSMSEKLANDIDTAVKTLSD 620
Query: 806 LCYERAKEILQQNKTLMDTLVNELVVKKT 834
YE A ++ N+ MD +V L+ K+T
Sbjct: 621 KAYEIALSQIRNNREAMDKIVEILLEKET 649
>At5g53170 cell division protein FtsH protease-like
Length = 806
Score = 283 bits (725), Expect = 2e-76
Identities = 181/434 (41%), Positives = 260/434 (59%), Gaps = 16/434 (3%)
Query: 417 FTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEAG 476
F DV G + ELEE+V++ + + R G K+P GILL G PG GKTLLAKA+AGEAG
Sbjct: 361 FKDVKGCDDAKQELEEVVEYLKNPSKFTRLGGKLPKGILLTGAPGTGKTLLAKAIAGEAG 420
Query: 477 VNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGGQE 536
V FF + S+F E++VGVGA RVRSL+Q AK+ AP ++FIDE+DAVG R +G
Sbjct: 421 VPFFYRAGSEFEEMFVGVGARRVRSLFQAAKKKAPCIIFIDEIDAVGSTRKQWEG----H 476
Query: 537 RDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRIEI 596
TL+QLLV +DGFE +I +A+TN PDILDPAL RPGRFDR I +P P GR EI
Sbjct: 477 TKKTLHQLLVEMDGFEQNEGIIVMAATNLPDILDPALTRPGRFDRHIVVPSPDVRGREEI 536
Query: 597 LKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQAAQ 656
L+++ + KP++EDVD + +A T G GA+LAN+V +AAI + ++S++ L A
Sbjct: 537 LELYLQGKPMSEDVDVKAIARGTPGFNGADLANLVNIAAIKAAVEGAEKLSSEQLEFAKD 596
Query: 657 MEERGMLDRKER--SKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGRELGYVRT 714
G +RK S++ + A +E+ A+ A+N I TI PR G LG V T
Sbjct: 597 RIVMG-TERKTMFVSEDSKKLTAYHESGHAIVALNTKGAHPIHKATIMPR-GSALGMV-T 653
Query: 715 MLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWAETADNARVAARMYMI 774
L S ++ ++++ L + V + R A+E+ FG D ++T + A A+ YM+
Sbjct: 654 QLPS--NDETSVSKRQLLARLDVCMGGRVAEELIFGLDHITTGASSDLSQATELAQ-YMV 710
Query: 775 G--GLSDKYRGV--SNFWVTDRINEIDLEAMKILNLCYERAKEILQQNKTLMDTLVNELV 830
G+S+ V +D + ID E +K+L YER K +L++++ + TL N L+
Sbjct: 711 SSCGMSEAIGPVHIKERPSSDMQSRIDAEVVKLLREAYERVKSLLKRHEKQLHTLANALL 770
Query: 831 VKKTLTKEDIVRLV 844
+TLT EDI R++
Sbjct: 771 EYETLTAEDIKRIL 784
>At3g47060 FtsH metalloprotease - like protein
Length = 802
Score = 266 bits (680), Expect = 4e-71
Identities = 174/460 (37%), Positives = 254/460 (54%), Gaps = 27/460 (5%)
Query: 412 GVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAV 471
G + F DVAG+ + + ELEEIV+F + E Y R G + P G+LL G PG GKTLLAKAV
Sbjct: 319 GETITFADVAGVDEAKEELEEIVEFLRNPEKYVRLGARPPRGVLLVGLPGTGKTLLAKAV 378
Query: 472 AGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKR-GLIK 530
AGEA V F S SAS+FVE+YVG+GASRVR L+ AK+ APS++FIDE+DAV + R G +
Sbjct: 379 AGEAEVPFISCSASEFVELYVGMGASRVRDLFARAKKEAPSIIFIDEIDAVAKSRDGKFR 438
Query: 531 GSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGF 590
ER+ TLNQLL +DGF+ VI + +TNR D+LDPAL RPGRFDR + + P
Sbjct: 439 MGSNDEREQTLNQLLTEMDGFDSNSAVIVLGATNRADVLDPALRRPGRFDRVVTVETPDK 498
Query: 591 IGRIEILKVHARKK--PIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVST 648
IGR IL+VH KK P+ +DV+ +ASMT G GA+LAN+V AA+ R ++T V
Sbjct: 499 IGRESILRVHVSKKELPLGDDVNLGSIASMTTGFTGADLANLVNEAALLAGRKNKTNVEK 558
Query: 649 DDLLQAAQMEERGMLDRKERSK-EKWEQVAINEAAMAVAAMNLPNF----DNIEYITIAP 703
D +QA + G+ + R K + VA +EA AV + N +E ++I P
Sbjct: 559 IDFIQAVERSIAGIEKKSARLKGNEKAVVARHEAGHAVVGTAVANLLTGQPRVEKLSILP 618
Query: 704 RAGRELGYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWAETAD 763
R G LG+ T + + + +L L + L RAA+E+ + + + +
Sbjct: 619 RTGGALGF--TYIPPTSEDRYLLFIDELLGRLVTLLGGRAAEEVVYSGRISTGAFDDIRR 676
Query: 764 NARVAARMYMIGGLSDKYRGVS--------------NFWVTDRINEIDL---EAMKILNL 806
+A + GL+ K VS + W D+ +DL E +L
Sbjct: 677 ATDMAYKAVAEYGLNQKIGPVSVATLSGGGIDDSGGSPWGRDQGKLVDLVQKEVTILLQS 736
Query: 807 CYERAKEILQQNKTLMDTLVNELVVKKTLTKEDIVRLVQL 846
+ A +++ N +++ L +L K+ + E++ + + +
Sbjct: 737 ALDVALSVVRANPDVLEGLGAQLEEKEKVEGEELQKWLSM 776
>At2g29080 AAA-type like ATPase
Length = 809
Score = 265 bits (678), Expect = 6e-71
Identities = 173/449 (38%), Positives = 257/449 (56%), Gaps = 29/449 (6%)
Query: 415 VKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGE 474
+ F DVAG + + E+ E V F + + Y G KIP G LL GPPG GKTLLAKA AGE
Sbjct: 319 IYFKDVAGCDEAKQEIMEFVHFLKNPKKYEDLGAKIPKGALLVGPPGTGKTLLAKATAGE 378
Query: 475 AGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGG 534
+GV F SIS S F+E++VGVG SRVR L+QEA++ APS++FIDE+DA+GR RG G
Sbjct: 379 SGVPFLSISGSDFMEMFVGVGPSRVRHLFQEARQAAPSIIFIDEIDAIGRARGRGGLGGN 438
Query: 535 QERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRI 594
ER++TLNQLLV +DGF V+ +A TNRPDILD AL+RPGRFDR+I I KP GR
Sbjct: 439 DERESTLNQLLVEMDGFGTTAGVVVLAGTNRPDILDKALLRPGRFDRQITIDKPDIKGRD 498
Query: 595 EILKVHARKKPIAEDVDY--EIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLL 652
+I K++ +K + + Y + +A++T G GA++AN+ AA+ R V T
Sbjct: 499 QIFKIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAARHEGATV-TMAHF 557
Query: 653 QAAQMEERGMLDRKERSKEKWEQ--VAINEAAMAVAAMNLPNFDNIEYITIAPRAGRELG 710
++A G L++K R K E+ VA +E+ AV L + + + +TI PR LG
Sbjct: 558 ESAIDRVIGGLEKKNRVISKLERRTVAYHESGHAVVGWFLEHAEPLLKVTIVPRGTAALG 617
Query: 711 YVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWAETADNARVAAR 770
+ + + + ++T++ LFD + L RAA+++ GK +ST D +V
Sbjct: 618 FA----QYVPNENLLMTKEQLFDMTCMTLGGRAAEQVLIGK--IST--GAQNDLEKVTKM 669
Query: 771 MY---MIGGLSDKYRGVSNFWVTDRINE------------IDLEAMKILNLCYERAKEIL 815
Y + G SDK G+ +F D + ID E + YER E++
Sbjct: 670 TYAQVAVYGFSDKV-GLLSFPPRDDGYDFSKPYSNKTGAIIDEEVRDWVAKAYERTVELV 728
Query: 816 QQNKTLMDTLVNELVVKKTLTKEDIVRLV 844
+++K + + L+ K+ L ++D+++++
Sbjct: 729 EEHKVKVAEIAELLLEKEVLHQDDLLKIL 757
>At1g07510 unknown protein
Length = 813
Score = 261 bits (668), Expect = 9e-70
Identities = 171/447 (38%), Positives = 258/447 (57%), Gaps = 26/447 (5%)
Query: 415 VKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGE 474
+ F DVAG + + E+ E V F + + Y G KIP G LL GPPG GKTLLAKA AGE
Sbjct: 324 IYFKDVAGCEEAKQEIMEFVHFLQNPKKYEDLGAKIPKGALLVGPPGTGKTLLAKATAGE 383
Query: 475 AGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGG 534
+ V F SIS S F+E++VGVG SRVR+L+QEA++ APS++FIDE+DA+GR RG SGG
Sbjct: 384 SAVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIIFIDEIDAIGRARGRGGFSGG 443
Query: 535 -QERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGR 593
ER++TLNQLLV +DGF V+ +A TNRPDILD AL+RPGRFDR+I I KP GR
Sbjct: 444 NDERESTLNQLLVEMDGFGTTAGVVVLAGTNRPDILDKALLRPGRFDRQITIDKPDIKGR 503
Query: 594 IEILKVHARKKPIAEDVDY--EIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDL 651
+I +++ +K + + Y + +A++T G GA++AN+ AA+ R V T
Sbjct: 504 DQIFQIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAARHEGATV-TMAH 562
Query: 652 LQAAQMEERGMLDRKERSKEKWEQ--VAINEAAMAVAAMNLPNFDNIEYITIAPRAGREL 709
+A G L++K R K E+ VA +E+ AVA L + + + +TI PR L
Sbjct: 563 FDSAIDRVIGGLEKKNRVISKLERRTVAYHESGHAVAGWFLEHAEPLLKVTIVPRGTAAL 622
Query: 710 GYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWAETADNARVAA 769
G+ + + + ++T++ LFD + L RAA+++ G+ +ST D +V
Sbjct: 623 GFA----QYVPNENLLMTKEQLFDMTCMTLGGRAAEQVLIGR--IST--GAQNDLEKVTK 674
Query: 770 RMY---MIGGLSDKYRGVSNFWVTDRINE---------IDLEAMKILNLCYERAKEILQQ 817
Y + G SDK +S D ++ ID E + + Y+R E++++
Sbjct: 675 MTYAQVAVYGFSDKIGLLSFPQREDEFSKPYSNRTGAMIDEEVREWVGKAYKRTVELIEE 734
Query: 818 NKTLMDTLVNELVVKKTLTKEDIVRLV 844
+K + + L+ K+ L ++D+ +++
Sbjct: 735 HKEQVAQIAELLLEKEVLHQDDLTKVL 761
>At5g58870 cell division protein - like
Length = 806
Score = 260 bits (664), Expect = 3e-69
Identities = 162/398 (40%), Positives = 228/398 (56%), Gaps = 22/398 (5%)
Query: 412 GVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAV 471
G + F DVAG+ + + ELEEIV+F + + Y R G + P G+LL G PG GKTLLAKAV
Sbjct: 323 GETITFADVAGVDEAKEELEEIVEFLKNPDRYVRLGARPPRGVLLVGLPGTGKTLLAKAV 382
Query: 472 AGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKR-GLIK 530
AGE+ V F S SAS+FVE+YVG+GASRVR L+ AK+ APS++FIDE+DAV + R G +
Sbjct: 383 AGESDVPFISCSASEFVELYVGMGASRVRDLFARAKKEAPSIIFIDEIDAVAKSRDGKFR 442
Query: 531 GSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGF 590
ER+ TLNQLL +DGF+ VI + +TNR D+LDPAL RPGRFDR + + P
Sbjct: 443 MVSNDEREQTLNQLLTEMDGFDSSSAVIVLGATNRADVLDPALRRPGRFDRVVTVESPDK 502
Query: 591 IGRIEILKVHARKK--PIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVST 648
+GR ILKVH KK P+ +DV+ +ASMT G GA+LAN+V AA+ R S+ V
Sbjct: 503 VGRESILKVHVSKKELPLGDDVNLASIASMTTGFTGADLANLVNEAALLAGRKSKMTVDK 562
Query: 649 DDLLQAAQMEERGMLDRKERSK-EKWEQVAINEAAMAV----AAMNLPNFDNIEYITIAP 703
D + A + G+ + R K + VA +EA AV A L +E ++I P
Sbjct: 563 IDFIHAVERSIAGIEKKTARLKGSEKAVVARHEAGHAVVGTAVASLLSGQSRVEKLSILP 622
Query: 704 RAGRELGYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWAETAD 763
R+G LG+ T + + + +L L + L RAA+E+ + I D
Sbjct: 623 RSGGALGF--TYIPPTHEDRYLLFIDELHGRLVTLLGGRAAEEVVYS----GRISTGALD 676
Query: 764 NARVAARMYMIGGLSDKYRGVSNFWVTDRINEIDLEAM 801
+ R A M Y+ V+ + + ++I + + +
Sbjct: 677 DIRRATDM--------AYKAVAEYGLNEKIGPVSVATL 706
>At2g26140 FtsH like protease
Length = 717
Score = 258 bits (659), Expect = 1e-68
Identities = 165/441 (37%), Positives = 246/441 (55%), Gaps = 22/441 (4%)
Query: 416 KFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEA 475
KF+DV G+ + + ELEEIV + + + R G K+P G+LL GPPG GKT+LA+A+AGEA
Sbjct: 225 KFSDVKGVDEAKAELEEIVHYLRDPKRFTRLGGKLPKGVLLVGPPGTGKTMLARAIAGEA 284
Query: 476 GVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGGQ 535
GV FFS S S+F E++VGVGA RVR L+ AK+ +P ++FIDE+DA+G R Q
Sbjct: 285 GVPFFSCSGSEFEEMFVGVGARRVRDLFSAAKKCSPCIIFIDEIDAIGGSR---NPKDQQ 341
Query: 536 ERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRIE 595
TLNQ+LV LDGF+ +I +A+TN P+ LD ALVRPGRFDR I +P P GR +
Sbjct: 342 YMKMTLNQMLVELDGFKQNEGIIVVAATNFPESLDKALVRPGRFDRHIVVPNPDVEGRRQ 401
Query: 596 ILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQAA 655
IL+ H K AEDVD I+A T G GA+LAN+V VAA+ D +V+ DL A
Sbjct: 402 ILESHMSKVLKAEDVDLMIIARGTPGFSGADLANLVNVAALKAAMDGSKDVTMSDLEFA- 460
Query: 656 QMEERGMLDRKER----SKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGRELGY 711
++R M+ + + S E + A +E A+ A++ + TI PR G LG
Sbjct: 461 --KDRIMMGSERKSAVISDESRKLTAFHEGGHALVAIHTEGALPVHKATIVPR-GMALGM 517
Query: 712 VRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWAETADNARVAARM 771
V + + ++ ++R+ + + V + R A+E+ FG+ ++++ + + A AR
Sbjct: 518 VSQLPDK---DETSISRKQMLARLDVCMGGRVAEELIFGESEVTSGASSDLEQATKLARA 574
Query: 772 YM--------IGGLSDKYRGVSNFWVTDRINEIDLEAMKILNLCYERAKEILQQNKTLMD 823
+ +G ++ Y T+ I+ E ++L Y AK IL +
Sbjct: 575 MVTKFGMSKEVGLVAHNYDDNGKSMSTETRLLIESEVKQLLEKAYNNAKTILTVYNKELH 634
Query: 824 TLVNELVVKKTLTKEDIVRLV 844
L N L+ +TL+ + I L+
Sbjct: 635 ALANALLQHETLSGKQIKELL 655
>At3g02450 cell division protein FtsH-like protein
Length = 622
Score = 245 bits (625), Expect = 9e-65
Identities = 138/301 (45%), Positives = 195/301 (63%), Gaps = 6/301 (1%)
Query: 400 AQNRRLPQYLERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGP 459
A N + + V F DV G+ + EL EIV Y++ G ++P G+LL GP
Sbjct: 316 ASNSPAKKRRSKNPTVGFDDVEGVDSAKDELVEIVSCLQGSINYKKLGARLPRGVLLVGP 375
Query: 460 PGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDEL 519
PG GKTLLA+AVAGEAGV FFS+SAS+FVE++VG GA+R+R L+ A++N+PS++FIDEL
Sbjct: 376 PGTGKTLLARAVAGEAGVPFFSVSASEFVELFVGRGAARIRDLFNAARKNSPSIIFIDEL 435
Query: 520 DAVGRKRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRF 579
DAVG KRG S ERD TLNQLL +DGFE +VI IA+TNRP+ LD AL RPGRF
Sbjct: 436 DAVGGKRG---RSFNDERDQTLNQLLTEMDGFESDTKVIVIAATNRPEALDSALCRPGRF 492
Query: 580 DRKIFIPKPGFIGRIEILKVHARKKPIAED--VDYEIVASMTDGMVGAELANIVEVAAIN 637
RK+ + +P GR +IL +H R P+ ED + ++VAS+T G VGA+LANIV AA+
Sbjct: 493 SRKVLVAEPDQEGRRKILAIHLRDVPLEEDAFLICDLVASLTPGFVGADLANIVNEAALL 552
Query: 638 MMRDSRTEVSTDDLLQAAQMEERGMLDRKERSKEKWEQVAINEAAM-AVAAMNLPNFDNI 696
R V+ +D+++A + + G+ D++ R + +++ M ++A N P+ D +
Sbjct: 553 AARRGGEAVAREDIMEAIERAKFGINDKEARPRTLGNELSKMFPWMPSLARRNGPDQDGL 612
Query: 697 E 697
+
Sbjct: 613 Q 613
>At4g23940 cell division protein - like
Length = 946
Score = 243 bits (621), Expect = 3e-64
Identities = 167/475 (35%), Positives = 258/475 (54%), Gaps = 56/475 (11%)
Query: 415 VKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGE 474
VKF DVAG+ + EL+E+VK+ + +++ + G+K P G+LL GPPG GKTL+AKA+AGE
Sbjct: 427 VKFADVAGIDEAVDELQELVKYLKNPDLFDKMGIKPPHGVLLEGPPGCGKTLVAKAIAGE 486
Query: 475 AGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVG-RKRGLIK--- 530
AGV F+ ++ S+FVE+ VGVG++R+R L++ AK N PSV+FIDE+DA+ R++G+ K
Sbjct: 487 AGVPFYQMAGSEFVEVLVGVGSARIRDLFKRAKVNKPSVIFIDEIDALATRRQGIFKENS 546
Query: 531 ----GSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIP 586
+ QER+ TLNQLL+ LDGF+ VI + +TNR D+LDPAL+RPGRFDRKI +
Sbjct: 547 DQLYNAATQERETTLNQLLIELDGFDTGKGVIFLGATNRRDLLDPALLRPGRFDRKIRVR 606
Query: 587 KPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEV 646
P GR++ILK+HA K +++ VD AS G GA+LA +V+ AA+ +R + +
Sbjct: 607 PPNAKGRLDILKIHASKVKMSDSVDLSSYASNLPGWSGAKLAQLVQEAALVAVRKTHNSI 666
Query: 647 STDDLLQAAQMEERGMLD-RKERSKEKWEQVAINEAAMAVAAMNLPNFDN-----IEYIT 700
D+ A G E + + A E +A+ + L ++N + ++
Sbjct: 667 LQSDMDDAVDRLTVGPTRIGLELGHQGQCRRATTEVGVAITSHLLLRYENAKIERCDRVS 726
Query: 701 IAPRAGRELGYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKD-------- 752
I PR V L+ ++ G L + L + V L RAA+E+ +G D
Sbjct: 727 IIPRGQTLSQVVFHRLDDESYMFGRLPQ--LLHRLQVLLGGRAAEEVIYGSDTSKASVDY 784
Query: 753 ---------QLSTIWAETADNAR---------------VAARMYMIGGLSDKYRGVS--- 785
++ TIW +N V R+ G L D Y V
Sbjct: 785 LSDASWLARKILTIW--NLENPMVIHGEPPPWRKRPQFVGPRLDFEGSLYDDYDLVEPPV 842
Query: 786 NFWVTDRINEIDLEAMKILNLCYERAKEILQQNKTLMDTLVNELVVKKTLTKEDI 840
NF + D E+ + ++++ Y + +L+QN+T + V L+ +K ++ E I
Sbjct: 843 NFNMDD---EVAHRSEELISQMYNKTVSLLRQNQTALLKTVKVLLNQKEISGEAI 894
>At5g64580 unknown protein
Length = 855
Score = 232 bits (591), Expect = 8e-61
Identities = 159/464 (34%), Positives = 243/464 (52%), Gaps = 18/464 (3%)
Query: 410 ERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAK 469
E V F D AG I+ EL+EIV+ + E ++ +G+ P G+LL GPPG GKTLLAK
Sbjct: 308 EEKTGVTFDDFAGQEYIKRELQEIVRILKNDEEFQNKGIYCPKGVLLHGPPGTGKTLLAK 367
Query: 470 AVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLI 529
A+AGEAG+ FF+ + + FVE++VGV ASRV+ L+ ++ APS++FIDE+DA+G KRG
Sbjct: 368 AIAGEAGLPFFAANGTDFVEMFVGVAASRVKDLFASSRSYAPSIIFIDEIDAIGSKRGGP 427
Query: 530 K-GSGGQERDATLNQLLVCLDGFE-GRGEVITIASTNRPDILDPALVRPGRFDRKIFIPK 587
G GG ER+ L Q+L +DGF+ +V+ I +TNR DILDPAL+R GRFD+ I +
Sbjct: 428 DIGGGGAEREQGLLQILTEMDGFKVTTSQVLVIGATNRLDILDPALLRKGRFDKIIRVGL 487
Query: 588 PGFIGRIEILKVHARKKPI-AEDVDYEI---VASMTDGMVGAELANIVEVAAINMMRDSR 643
P GR+ ILKVHAR K +ED E+ VA T+ GAEL N++ A I R
Sbjct: 488 PSKDGRLAILKVHARNKFFRSEDEKEELLQEVAENTEDFTGAELQNVLNEAGILTARKDL 547
Query: 644 TEVSTDDLLQAAQME----ERGMLDRKERSKEKWEQVAINEAAMAVAAMNLPNFDNIEYI 699
+ ++LL+A + + E G D E +E ++A EAA+AV A LP+ +Y
Sbjct: 548 DYIGREELLEALKRQKGTFETGQEDSTEVPEELKLRLAYREAAVAVLACYLPD----QYR 603
Query: 700 TIAPRAGRELGYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWA 759
I+ + M S + + I APR +E FG + L I A
Sbjct: 604 PISETDINSIRSQPNMRYSETSGRVFARKSDYVNSIIRACAPRVVEEEMFGIENLCWISA 663
Query: 760 ETADNARVAARMYMI----GGLSDKYRGVSNFWVTDRINEIDLEAMKILNLCYERAKEIL 815
++ A A ++ Y V + + +++ + + E+ IL
Sbjct: 664 KSTLEASQRAEFLILQTGMTAFGKAYYRNQRDLVPNLVPKLEALRDEYMRFAVEKCSSIL 723
Query: 816 QQNKTLMDTLVNELVVKKTLTKEDIVRLVQLHGHAKPIPISVLD 859
Q+ ++ ++ + + L+ K + ++I + P+ +D
Sbjct: 724 QEYQSALEEITDVLLEKGEIKADEIWNIYNTAPRIPQKPVRPVD 767
>At5g43010 26S proteasome AAA-ATPase subunit RPT4a (gb|AAF22524.1)
Length = 399
Score = 194 bits (493), Expect = 2e-49
Identities = 127/368 (34%), Positives = 199/368 (53%), Gaps = 34/368 (9%)
Query: 323 RTVVVS-YKK---QKKDYEDRIKIQKADAEERRK-MREMEAEMGWSEAGGD--------- 368
RT VS Y+K Q K+ E R++ + + +K + E ++ ++ G
Sbjct: 11 RTAAVSEYRKKLLQHKELESRVRTARENLRGAKKEFNKTEDDLKSLQSVGQIIGEVLRPL 70
Query: 369 EDESELVKEGEENPYL-----KMTKEFMKSGARVRRAQN-----RRLPQYLERGV----- 413
++E +VK Y+ K+ KE + SG RV R LP+ ++ V
Sbjct: 71 DNERLIVKASSGPRYVVGCRSKVDKEKLTSGTRVVLDMTTLTIMRALPREVDPVVYNMLH 130
Query: 414 ----DVKFTDVAGLGKIRLELEEIVKF-FTHGEMYRRRGVKIPGGILLCGPPGVGKTLLA 468
++ ++ V GLG EL E ++ + E++ R G+K P G+LL GPPG GKTLLA
Sbjct: 131 EDPGNISYSAVGGLGDQIRELRESIELPLMNPELFLRVGIKPPKGVLLYGPPGTGKTLLA 190
Query: 469 KAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGL 528
+A+A NF + +S ++ Y+G A +R ++ A+E+ P ++F+DE+DA+G +R
Sbjct: 191 RAIASNIDANFLKVVSSAIIDKYIGESARLIREMFNYAREHQPCIIFMDEIDAIGGRRFS 250
Query: 529 IKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKP 588
S +E TL +LL LDGF+ G+V I +TNRPD+LDPAL+RPGR DRKI IP P
Sbjct: 251 EGTSADREIQRTLMELLNQLDGFDNLGKVKMIMATNRPDVLDPALLRPGRLDRKIEIPLP 310
Query: 589 GFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVST 648
R++ILK+HA ++DYE + + +G GA+L NI A + +R R V
Sbjct: 311 NEQSRMDILKIHAAGIAKHGEIDYEAIVKLAEGFNGADLRNICTEAGMFAIRAERDYVIH 370
Query: 649 DDLLQAAQ 656
+D ++A +
Sbjct: 371 EDFMKAVR 378
>At3g09840 putative transitional endoplasmic reticulum ATPase
Length = 809
Score = 193 bits (490), Expect = 4e-49
Identities = 105/263 (39%), Positives = 158/263 (59%), Gaps = 11/263 (4%)
Query: 389 EFMKSGARVRRAQNRRLPQYLERGVDVKFTDVAGLGKIRLELEEIVKF-FTHGEMYRRRG 447
E G V+R RL DV + DV G+ K ++ E+V+ H ++++ G
Sbjct: 185 EIFCEGEPVKREDEERLD-------DVGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIG 237
Query: 448 VKIPGGILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAK 507
VK P GILL GPPG GKTL+A+AVA E G FF I+ + + G S +R ++EA+
Sbjct: 238 VKPPKGILLYGPPGSGKTLIARAVANETGAFFFCINGPEIMSKLAGESESNLRKAFEEAE 297
Query: 508 ENAPSVVFIDELDAVGRKRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPD 567
+NAPS++FIDE+D++ KR + + G+ ++QLL +DG + R VI + +TNRP+
Sbjct: 298 KNAPSIIFIDEIDSIAPKR---EKTNGEVERRIVSQLLTLMDGLKSRAHVIVMGATNRPN 354
Query: 568 ILDPALVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAEL 627
+DPAL R GRFDR+I I P IGR+E+L++H + +AEDVD E ++ T G VGA+L
Sbjct: 355 SIDPALRRFGRFDREIDIGVPDEIGRLEVLRIHTKNMKLAEDVDLERISKDTHGYVGADL 414
Query: 628 ANIVEVAAINMMRDSRTEVSTDD 650
A + AA+ +R+ + +D
Sbjct: 415 AALCTEAALQCIREKMDVIDLED 437
Score = 181 bits (458), Expect = 2e-45
Identities = 109/305 (35%), Positives = 166/305 (53%), Gaps = 15/305 (4%)
Query: 414 DVKFTDVAGLGKIRLELEEIVKF-FTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVA 472
+V + D+ GL ++ EL+E V++ H E + + G+ G+L GPPG GKTLLAKA+A
Sbjct: 476 NVSWNDIGGLENVKRELQETVQYPVEHPEKFEKFGMSPSKGVLFYGPPGCGKTLLAKAIA 535
Query: 473 GEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGS 532
E NF S+ + + ++ G + VR ++ +A+++AP V+F DELD++ +RG G
Sbjct: 536 NECQANFISVKGPELLTMWFGESEANVREIFDKARQSAPCVLFFDELDSIATQRGGGSGG 595
Query: 533 -GGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFI 591
GG D LNQLL +DG + V I +TNRPDI+D AL+RPGR D+ I+IP P
Sbjct: 596 DGGGAADRVLNQLLTEMDGMNAKKTVFIIGATNRPDIIDSALLRPGRLDQLIYIPLPDED 655
Query: 592 GRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAA-----INMMRDSRTEV 646
R+ I K RK PIA+DVD +A T G GA++ I + A N+ +D E
Sbjct: 656 SRLNIFKAALRKSPIAKDVDIGALAKYTQGFSGADITEICQRACKYAIRENIEKDIEKEK 715
Query: 647 STDDLLQAAQMEERGMLDRKERSKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAG 706
+ +A MEE G+ + E +E+ +M A ++ + D +Y A
Sbjct: 716 RRSENPEA--MEEDGVDEVSEIKAAHFEE------SMKYARRSVSDADIRKYQAFAQTLQ 767
Query: 707 RELGY 711
+ G+
Sbjct: 768 QSRGF 772
>At3g53230 CDC48 - like protein
Length = 815
Score = 192 bits (488), Expect = 7e-49
Identities = 105/263 (39%), Positives = 158/263 (59%), Gaps = 11/263 (4%)
Query: 389 EFMKSGARVRRAQNRRLPQYLERGVDVKFTDVAGLGKIRLELEEIVKF-FTHGEMYRRRG 447
E G ++R RL + V + DV G+ K ++ E+V+ H ++++ G
Sbjct: 186 EIFCEGEPIKREDEERLDE-------VGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIG 238
Query: 448 VKIPGGILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAK 507
VK P GILL GPPG GKTL+A+AVA E G FF I+ + + G S +R ++EA+
Sbjct: 239 VKPPKGILLYGPPGSGKTLIARAVANETGAFFFCINGPEIMSKLAGESESNLRKAFEEAE 298
Query: 508 ENAPSVVFIDELDAVGRKRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPD 567
+NAPS++FIDE+D++ KR + + G+ ++QLL +DG + R VI + +TNRP+
Sbjct: 299 KNAPSIIFIDEIDSIAPKR---EKTHGEVERRIVSQLLTLMDGLKSRAHVIVMGATNRPN 355
Query: 568 ILDPALVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAEL 627
+DPAL R GRFDR+I I P IGR+E+L++H + +AEDVD E V+ T G VGA+L
Sbjct: 356 SIDPALRRFGRFDREIDIGVPDEIGRLEVLRIHTKNMKLAEDVDLERVSKDTHGYVGADL 415
Query: 628 ANIVEVAAINMMRDSRTEVSTDD 650
A + AA+ +R+ + DD
Sbjct: 416 AALCTEAALQCIREKMDVIDLDD 438
Score = 185 bits (469), Expect = 1e-46
Identities = 106/304 (34%), Positives = 167/304 (54%), Gaps = 15/304 (4%)
Query: 414 DVKFTDVAGLGKIRLELEEIVKF-FTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVA 472
+V + D+ GL ++ EL+E V++ H E + + G+ G+L GPPG GKTLLAKA+A
Sbjct: 477 NVSWEDIGGLENVKRELQETVQYPVEHPEKFEKFGMSPSKGVLFYGPPGCGKTLLAKAIA 536
Query: 473 GEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGS 532
E NF SI + + ++ G + VR ++ +A+++AP V+F DELD++ +RG G
Sbjct: 537 NECQANFISIKGPELLTMWFGESEANVREIFDKARQSAPCVLFFDELDSIATQRGNSVGD 596
Query: 533 GGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIG 592
G D LNQLL +DG + V I +TNRPDI+DPAL+RPGR D+ I+IP P
Sbjct: 597 AGGAADRVLNQLLTEMDGMNAKKTVFIIGATNRPDIIDPALLRPGRLDQLIYIPLPDEES 656
Query: 593 RIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLL 652
R +I K RK P+A+DVD +A T G GA++ I + + +R++ +
Sbjct: 657 RYQIFKSCLRKSPVAKDVDLRALAKYTQGFSGADITEICQRSCKYAIREN---------I 707
Query: 653 QAAQMEERGMLDRKERSKEKWEQVAINEA-----AMAVAAMNLPNFDNIEYITIAPRAGR 707
+ +ER + E +E E++A +A +M A ++ + D +Y A +
Sbjct: 708 EKDIEKERKRAESPEAMEEDEEEIAEIKAGHFEESMKYARRSVSDADIRKYQAFAQTLQQ 767
Query: 708 ELGY 711
G+
Sbjct: 768 SRGF 771
>At1g45000 26S proteasome AAA-ATPase subunit RPT4a like protein
Length = 399
Score = 192 bits (487), Expect = 9e-49
Identities = 124/364 (34%), Positives = 196/364 (53%), Gaps = 33/364 (9%)
Query: 326 VVSYKKQ---KKDYEDRIKIQKADAEERRK-MREMEAEMGWSEAGGD---------EDES 372
V Y+K+ K+ E R++ + + +K + E ++ ++ G ++E
Sbjct: 15 VTDYRKKLLHHKELESRVRTARENLRAAKKEFNKTEDDLKSLQSVGQIIGEVLRPLDNER 74
Query: 373 ELVKEGEENPYL-----KMTKEFMKSGARVRRAQN-----RRLPQYLERGV--------- 413
+VK Y+ K+ KE + SG RV R LP+ ++ V
Sbjct: 75 LIVKASSGPRYVVGCRSKVDKEKLTSGTRVVLDMTTLTIMRALPREVDPVVYNMLHEDPG 134
Query: 414 DVKFTDVAGLGKIRLELEEIVKF-FTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVA 472
++ ++ V GLG EL E ++ + E++ R G+K P G+LL GPPG GKTLLA+A+A
Sbjct: 135 NISYSAVGGLGDQIRELRESIELPLMNPELFLRVGIKPPKGVLLYGPPGTGKTLLARAIA 194
Query: 473 GEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGS 532
NF + +S ++ Y+G A +R ++ A+E+ P ++F+DE+DA+G +R S
Sbjct: 195 SNIDANFLKVVSSAIIDKYIGESARLIREMFNYAREHQPCIIFMDEIDAIGGRRFSEGTS 254
Query: 533 GGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIG 592
+E TL +LL LDGF+ G+V I +TNRPD+LDPAL+RPGR DRKI IP P
Sbjct: 255 ADREIQRTLMELLNQLDGFDQLGKVKMIMATNRPDVLDPALLRPGRLDRKIEIPLPNEQS 314
Query: 593 RIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLL 652
R+EILK+HA ++DYE + + +G GA+L NI A + +R R V +D +
Sbjct: 315 RMEILKIHASGIAKHGEIDYEAIVKLGEGFNGADLRNICTEAGMFAIRAERDYVIHEDFM 374
Query: 653 QAAQ 656
+A +
Sbjct: 375 KAVR 378
>At5g03340 transitional endoplasmic reticulum ATPase
Length = 810
Score = 191 bits (486), Expect = 1e-48
Identities = 104/263 (39%), Positives = 158/263 (59%), Gaps = 11/263 (4%)
Query: 389 EFMKSGARVRRAQNRRLPQYLERGVDVKFTDVAGLGKIRLELEEIVKF-FTHGEMYRRRG 447
E G V+R RL + V + DV G+ K ++ E+V+ H ++++ G
Sbjct: 185 EIFCEGEPVKREDEERLDE-------VGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIG 237
Query: 448 VKIPGGILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAK 507
VK P GILL GPPG GKTL+A+AVA E G FF I+ + + G S +R ++EA+
Sbjct: 238 VKPPKGILLYGPPGSGKTLIARAVANETGAFFFCINGPEIMSKLAGESESNLRKAFEEAE 297
Query: 508 ENAPSVVFIDELDAVGRKRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPD 567
+NAPS++FIDE+D++ KR + + G+ ++QLL +DG + R VI + +TNRP+
Sbjct: 298 KNAPSIIFIDEIDSIAPKR---EKTNGEVERRIVSQLLTLMDGLKSRAHVIVMGATNRPN 354
Query: 568 ILDPALVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAEL 627
+DPAL R GRFDR+I I P IGR+E+L++H + +AEDVD E ++ T G VGA+L
Sbjct: 355 SIDPALRRFGRFDREIDIGVPDEIGRLEVLRIHTKNMKLAEDVDLERISKDTHGYVGADL 414
Query: 628 ANIVEVAAINMMRDSRTEVSTDD 650
A + AA+ +R+ + +D
Sbjct: 415 AALCTEAALQCIREKMDVIDLED 437
Score = 182 bits (463), Expect = 5e-46
Identities = 92/230 (40%), Positives = 138/230 (60%), Gaps = 1/230 (0%)
Query: 414 DVKFTDVAGLGKIRLELEEIVKF-FTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVA 472
+V + D+ GL ++ EL+E V++ H E + + G+ G+L GPPG GKTLLAKA+A
Sbjct: 476 NVSWEDIGGLENVKRELQETVQYPVEHPEKFEKFGMSPSKGVLFYGPPGCGKTLLAKAIA 535
Query: 473 GEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGS 532
E NF S+ + + ++ G + VR ++ +A+++AP V+F DELD++ +RG G
Sbjct: 536 NECQANFISVKGPELLTMWFGESEANVREIFDKARQSAPCVLFFDELDSIATQRGNSAGD 595
Query: 533 GGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIG 592
G D LNQLL +DG + V I +TNRPDI+D AL+RPGR D+ I+IP P
Sbjct: 596 AGGAADRVLNQLLTEMDGMNAKKTVFIIGATNRPDIIDSALLRPGRLDQLIYIPLPDEDS 655
Query: 593 RIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDS 642
R+ I K RK P+A+DVD +A T G GA++ I + A +R++
Sbjct: 656 RLNIFKACLRKSPVAKDVDVTALAKYTQGFSGADITEICQRACKYAIREN 705
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.134 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,748,616
Number of Sequences: 26719
Number of extensions: 808957
Number of successful extensions: 5843
Number of sequences better than 10.0: 280
Number of HSP's better than 10.0 without gapping: 157
Number of HSP's successfully gapped in prelim test: 128
Number of HSP's that attempted gapping in prelim test: 4530
Number of HSP's gapped (non-prelim): 879
length of query: 881
length of database: 11,318,596
effective HSP length: 108
effective length of query: 773
effective length of database: 8,432,944
effective search space: 6518665712
effective search space used: 6518665712
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 65 (29.6 bits)
Medicago: description of AC138130.19


