

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138130.15 + phase: 0
(174 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
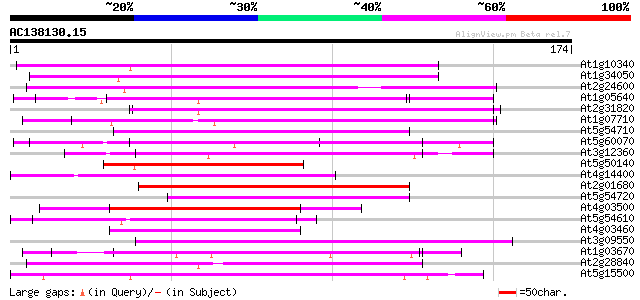
Score E
Sequences producing significant alignments: (bits) Value
At1g10340 putative protein 70 4e-13
At1g34050 putative protein 67 6e-12
At2g24600 unknown protein 65 2e-11
At1g05640 unknown protein 61 4e-10
At2g31820 ankyrin-like protein 59 1e-09
At1g07710 hypothetical protein 58 3e-09
At5g54710 putative protein 55 3e-08
At5g60070 ankyrin-like protein 54 4e-08
At3g12360 unknown protein 50 5e-07
At5g50140 ankyrin-like protein 50 6e-07
At4g14400 unknown protein 46 9e-06
At2g01680 unknown protein 46 1e-05
At5g54720 putative protein 45 3e-05
At4g03500 hypothetical protein 44 3e-05
At5g54610 ankyrin-repeat-containing protein-like 44 4e-05
At4g03460 hypothetical protein 44 6e-05
At3g09550 putative ankyrin 42 1e-04
At1g03670 hypothetical protein 42 1e-04
At2g28840 unknown protein 41 4e-04
At5g15500 unknown protein 40 5e-04
>At1g10340 putative protein
Length = 578
Score = 70.5 bits (171), Expect = 4e-13
Identities = 40/133 (30%), Positives = 68/133 (51%), Gaps = 2/133 (1%)
Query: 3 QEFFNAIKNNDISTFSSIVKVREGILNQRTDDTF--NTPLHLASKYGCIEMVSEIVRLCP 60
Q F+AI ND+ F +V+ E L +R ++ NT LH+A+K+G E+VS+I+ L P
Sbjct: 2 QPIFHAILKNDLPAFLELVEDSESSLEERNEEEHLNNTVLHMAAKFGHRELVSKIIELRP 61
Query: 61 DMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLV 120
+VS+ N TP+H A +V ++M +LE N + +AC ++
Sbjct: 62 SLVSSRNAYRNTPLHLAAILGDVNIVMQMLETGLEVCSARNINNHTPLHLACRSNSIEAA 121
Query: 121 NLLLNLSEIVEPG 133
L+ ++ + G
Sbjct: 122 RLIAEKTQSIGLG 134
Score = 36.2 bits (82), Expect = 0.009
Identities = 31/124 (25%), Positives = 56/124 (45%), Gaps = 14/124 (11%)
Query: 33 DDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEV 92
D + +T LH A G E+ + ++ L + A N N +P+H A + +V +L L+
Sbjct: 168 DGSQSTLLHHACDKGDFELTTILLGLDQGLEEALNPNGLSPLHLAVLRGSVVILEEFLDK 227
Query: 93 NPTAACKLNPTCKSAFFVACSHGHLDL---------VNLLLNLSEIVEPGLAGFDQACFH 143
P + + P+ ++ F +A + ++D +N + L + E G H
Sbjct: 228 VPLSFSSITPSKETVFHLAARNKNMDAFVFMAESLGINSQILLQQTDESG-----NTVLH 282
Query: 144 IAAS 147
IAAS
Sbjct: 283 IAAS 286
Score = 34.3 bits (77), Expect = 0.035
Identities = 27/125 (21%), Positives = 53/125 (41%), Gaps = 5/125 (4%)
Query: 27 ILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVS-----AENENMETPIHEACRQE 81
++ ++T L LA G +V I+ PD+ E+ + T +H AC +
Sbjct: 123 LIAEKTQSIGLGELILAISSGSTSIVGTILERFPDLAREEAWVVEDGSQSTLLHHACDKG 182
Query: 82 NVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQAC 141
+ ++ +LL ++ LNP S +A G + ++ L+ + + +
Sbjct: 183 DFELTTILLGLDQGLEEALNPNGLSPLHLAVLRGSVVILEEFLDKVPLSFSSITPSKETV 242
Query: 142 FHIAA 146
FH+AA
Sbjct: 243 FHLAA 247
Score = 26.2 bits (56), Expect = 9.6
Identities = 14/43 (32%), Positives = 25/43 (57%), Gaps = 3/43 (6%)
Query: 5 FFNAIKNNDISTFSSIVK---VREGILNQRTDDTFNTPLHLAS 44
F A +N ++ F + + + IL Q+TD++ NT LH+A+
Sbjct: 243 FHLAARNKNMDAFVFMAESLGINSQILLQQTDESGNTVLHIAA 285
>At1g34050 putative protein
Length = 573
Score = 66.6 bits (161), Expect = 6e-12
Identities = 39/129 (30%), Positives = 69/129 (53%), Gaps = 2/129 (1%)
Query: 7 NAIKNNDISTFSSIVKVREGILNQRT--DDTFNTPLHLASKYGCIEMVSEIVRLCPDMVS 64
+AI ND+ST ++ + +L +R D T LHLA++ G E+V I++LCP +V
Sbjct: 23 DAILANDVSTLLALAEGNLSVLRERYHWDSLGGTVLHLATELGHKEIVEAIIKLCPSLVG 82
Query: 65 AENENMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLL 124
N + +TP+H A R + ++ +L +N ++AF VAC + + D+ +L+L
Sbjct: 83 VTNLDGDTPLHFAARWGHATIVAQILASGYAEFTPVNGRGETAFVVACRYTNPDVASLIL 142
Query: 125 NLSEIVEPG 133
+ + G
Sbjct: 143 EETSSITIG 151
Score = 34.7 bits (78), Expect = 0.027
Identities = 19/85 (22%), Positives = 39/85 (45%)
Query: 33 DDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEV 92
D +TPLH A +E+ ++ + + N++ TP+H A + ++ +L +
Sbjct: 181 DGELSTPLHHACNANNLEITKMLLEIDESLAERVNKDGFTPLHLAAMKCSIPILKEFSDK 240
Query: 93 NPTAACKLNPTCKSAFFVACSHGHL 117
P L P ++ F +A H ++
Sbjct: 241 APRYFDILTPAKETVFHLAAEHKNI 265
Score = 32.7 bits (73), Expect = 0.10
Identities = 12/42 (28%), Positives = 25/42 (58%)
Query: 60 PDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAACKLN 101
P + + + TP+H AC N+++ +LLE++ + A ++N
Sbjct: 174 PKLAWNADGELSTPLHHACNANNLEITKMLLEIDESLAERVN 215
Score = 30.8 bits (68), Expect = 0.39
Identities = 20/85 (23%), Positives = 35/85 (40%), Gaps = 1/85 (1%)
Query: 7 NAIKNNDISTFSSIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAE 66
+A N++ ++++ E + + D F TPLHLA+ I ++ E P
Sbjct: 190 HACNANNLEITKMLLEIDESLAERVNKDGF-TPLHLAAMKCSIPILKEFSDKAPRYFDIL 248
Query: 67 NENMETPIHEACRQENVKVLMLLLE 91
ET H A +N+ + E
Sbjct: 249 TPAKETVFHLAAEHKNILAFYFMAE 273
>At2g24600 unknown protein
Length = 548
Score = 65.1 bits (157), Expect = 2e-11
Identities = 42/148 (28%), Positives = 75/148 (50%), Gaps = 9/148 (6%)
Query: 6 FNAIKNNDISTFSSIVKVREGILNQRTDD--TFNTPLHLASKYGCIEMVSEIVRLCPDMV 63
F+AI ND+ F +V+ RE L +R+++ T NT LH+A+K G E+V++I+ L P ++
Sbjct: 5 FDAILQNDLPAFLGLVEARESSLEERSEEQNTNNTVLHVAAKLGHRELVAKIIELRPSLL 64
Query: 64 SAENENMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLL 123
S+ N +TP+H A +V ++M +L+ N ++ HL V++
Sbjct: 65 SSRNAYGDTPLHLAALLGDVNIVMQMLDTGLELYSARNNKNQTPL-------HLAFVSIF 117
Query: 124 LNLSEIVEPGLAGFDQACFHIAASRGHT 151
+ ++ + D + A S G T
Sbjct: 118 MEAAKFIVEKTNSVDLDELNFALSSGST 145
Score = 38.9 bits (89), Expect = 0.001
Identities = 33/119 (27%), Positives = 57/119 (47%), Gaps = 4/119 (3%)
Query: 33 DDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEV 92
D + +T LH A G +E+ S ++ L + A N +P+H A ++ +V +L ++
Sbjct: 168 DGSRSTLLHYACDKGDLELTSILLGLNQGLEEALNSKGLSPLHLAVQRGSVIILEEFMDK 227
Query: 93 NPTAACKLNPTCKSAFFVACSHGHLD-LVNLLLNLSEIVEPGLAGFDQ---ACFHIAAS 147
+P + C P+ ++ F +A + + D V + NL L DQ HIAAS
Sbjct: 228 SPLSFCVRTPSKETVFHLAARNKNTDAFVFMAENLGTSSPILLKKKDQQGNTVLHIAAS 286
>At1g05640 unknown protein
Length = 627
Score = 60.8 bits (146), Expect = 4e-10
Identities = 42/155 (27%), Positives = 71/155 (45%), Gaps = 6/155 (3%)
Query: 2 DQEFFNAIKNNDISTFSSIVKVREGI-----LNQRTDDTFNTPLHLASKYGCIEMVSEIV 56
D A + ++ +++ GI L+ + + TPL+ A++ G +V E++
Sbjct: 114 DSPLHLAARTGNLGKVMELIRACNGIEELKELSSKQNLEGETPLYSAAENGHSLVVEEML 173
Query: 57 R-LCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHG 115
+ + D S + N P H A +Q +++ L LLE P A ++ +C +A A S G
Sbjct: 174 KHMDLDTASVKARNGFDPFHVAAKQGHIEALKKLLETFPNLAMTVDLSCTTALHTAASQG 233
Query: 116 HLDLVNLLLNLSEIVEPGLAGFDQACFHIAASRGH 150
H D+VNLLL + + H AA GH
Sbjct: 234 HTDVVNLLLKTDSHLAKIAKNNGKTALHSAARMGH 268
Score = 44.3 bits (103), Expect = 3e-05
Identities = 32/116 (27%), Positives = 54/116 (45%), Gaps = 12/116 (10%)
Query: 9 IKNNDISTFSSIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENE 68
+K+ D+ T S VK R G P H+A+K G IE + +++ P++ +
Sbjct: 173 LKHMDLDTAS--VKARNGF----------DPFHVAAKQGHIEALKKLLETFPNLAMTVDL 220
Query: 69 NMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLL 124
+ T +H A Q + V+ LLL+ + A K+A A GH ++V L+
Sbjct: 221 SCTTALHTAASQGHTDVVNLLLKTDSHLAKIAKNNGKTALHSAARMGHREVVKSLI 276
Score = 43.1 bits (100), Expect = 8e-05
Identities = 27/93 (29%), Positives = 48/93 (51%)
Query: 31 RTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLL 90
RTD T LH+A K +V E+V+ P ++S E+ TP+H A + +K++ L+
Sbjct: 285 RTDKKGQTALHMAVKGQNEGIVLELVKPDPAILSVEDSKGNTPLHTATNKGRIKIVRCLV 344
Query: 91 EVNPTAACKLNPTCKSAFFVACSHGHLDLVNLL 123
+ +N +A +A G+ +LV++L
Sbjct: 345 SFDGINLNAMNKAGDTALDIAEKIGNPELVSVL 377
Score = 30.8 bits (68), Expect = 0.39
Identities = 25/90 (27%), Positives = 43/90 (47%), Gaps = 1/90 (1%)
Query: 8 AIKNNDISTFSSIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAEN 67
A+K + +VK IL+ D NTPLH A+ G I++V +V ++A N
Sbjct: 297 AVKGQNEGIVLELVKPDPAILSVE-DSKGNTPLHTATNKGRIKIVRCLVSFDGINLNAMN 355
Query: 68 ENMETPIHEACRQENVKVLMLLLEVNPTAA 97
+ +T + A + N +++ +L E A
Sbjct: 356 KAGDTALDIAEKIGNPELVSVLKEAGAATA 385
>At2g31820 ankyrin-like protein
Length = 662
Score = 58.9 bits (141), Expect = 1e-09
Identities = 36/114 (31%), Positives = 58/114 (50%), Gaps = 1/114 (0%)
Query: 38 TPLHLASKYGCIEMVSEIVR-LCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTA 96
TPL+ A++ G +V E+++ + + S N P H A +Q +++VL +LLE P
Sbjct: 191 TPLYTAAENGHSIVVEEMLKHMDLETASIAARNGFDPFHVAAKQGHLEVLKILLETFPNL 250
Query: 97 ACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQACFHIAASRGH 150
A + +C +A A + GH+D+VNLLL + + H AA GH
Sbjct: 251 AMTTDLSCTTALHTAATQGHIDVVNLLLETDSNLAKIAKNNGKTALHSAARMGH 304
Score = 45.8 bits (107), Expect = 1e-05
Identities = 28/114 (24%), Positives = 52/114 (45%)
Query: 39 PLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAAC 98
P H+A+K G +E++ ++ P++ + + T +H A Q ++ V+ LLLE + A
Sbjct: 227 PFHVAAKQGHLEVLKILLETFPNLAMTTDLSCTTALHTAATQGHIDVVNLLLETDSNLAK 286
Query: 99 KLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQACFHIAASRGHTG 152
K+A A GH+++V L+ + Q H+A + G
Sbjct: 287 IAKNNGKTALHSAARMGHVEVVKSLIGKDPSIGFRTDKKGQTALHMAVKGQNDG 340
Score = 38.1 bits (87), Expect = 0.002
Identities = 31/128 (24%), Positives = 60/128 (46%), Gaps = 12/128 (9%)
Query: 28 LNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLM 87
L TD + T LH A+ G I++V+ ++ ++ N +T +H A R +V+V+
Sbjct: 250 LAMTTDLSCTTALHTAATQGHIDVVNLLLETDSNLAKIAKNNGKTALHSAARMGHVEVVK 309
Query: 88 LLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFD------QAC 141
L+ +P+ + + ++A +A G D + + E+V+P +A
Sbjct: 310 SLIGKDPSIGFRTDKKGQTALHMAVK-GQNDGI-----VVELVKPDVAVLSVEDNKGNTP 363
Query: 142 FHIAASRG 149
HIA ++G
Sbjct: 364 LHIATNKG 371
Score = 36.6 bits (83), Expect = 0.007
Identities = 25/93 (26%), Positives = 45/93 (47%)
Query: 31 RTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLL 90
RTD T LH+A K +V E+V+ ++S E+ TP+H A + +K++ L+
Sbjct: 321 RTDKKGQTALHMAVKGQNDGIVVELVKPDVAVLSVEDNKGNTPLHIATNKGRIKIVRCLV 380
Query: 91 EVNPTAACKLNPTCKSAFFVACSHGHLDLVNLL 123
+N + V+ G+ +LV++L
Sbjct: 381 SFEGINLNPINKAGDTPLDVSEKIGNAELVSVL 413
Score = 32.0 bits (71), Expect = 0.17
Identities = 23/90 (25%), Positives = 45/90 (49%), Gaps = 1/90 (1%)
Query: 8 AIKNNDISTFSSIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAEN 67
A+K + +VK +L+ D+ NTPLH+A+ G I++V +V ++ N
Sbjct: 333 AVKGQNDGIVVELVKPDVAVLSVE-DNKGNTPLHIATNKGRIKIVRCLVSFEGINLNPIN 391
Query: 68 ENMETPIHEACRQENVKVLMLLLEVNPTAA 97
+ +TP+ + + N +++ +L E A
Sbjct: 392 KAGDTPLDVSEKIGNAELVSVLKEAGAATA 421
>At1g07710 hypothetical protein
Length = 543
Score = 57.8 bits (138), Expect = 3e-09
Identities = 39/136 (28%), Positives = 68/136 (49%), Gaps = 6/136 (4%)
Query: 20 IVKVREGILNQ---RTDDTFNTPLHLASKYGCIEMVSEIVRLCPDM--VSAENENMETPI 74
+ K RE LNQ + + + T L++A++YG +E+V E++ C D+ V + N
Sbjct: 47 LTKTRESELNQLLGKQNQSGETALYVAAEYGDVEIVKEMIN-CYDLALVEIKARNGFDAF 105
Query: 75 HEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGL 134
H A +Q ++ VL +L E + A ++ + +A A + GH ++VN LL L +
Sbjct: 106 HIAAKQGDLDVLKVLAEAHSELAMTVDLSNTTALHTAATQGHTEVVNFLLELGSSLAGIA 165
Query: 135 AGFDQACFHIAASRGH 150
+ H A+ GH
Sbjct: 166 KSNGKTALHSASRNGH 181
Score = 47.8 bits (112), Expect = 3e-06
Identities = 35/147 (23%), Positives = 66/147 (44%), Gaps = 12/147 (8%)
Query: 5 FFNAIKNNDISTFSSIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVS 64
F A K D+ + + L D + T LH A+ G E+V+ ++ L +
Sbjct: 105 FHIAAKQGDLDVLKVLAEAHSE-LAMTVDLSNTTALHTAATQGHTEVVNFLLELGSSLAG 163
Query: 65 AENENMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLL 124
N +T +H A R +VKV+ LL P A +++ ++A +A ++++V L+
Sbjct: 164 IAKSNGKTALHSASRNGHVKVIKALLASEPAIAIRMDKKGQTALHMAVKGTNVEVVEELI 223
Query: 125 NLSEIVEPGLAGFDQACFHIAASRGHT 151
D++ +IA ++G+T
Sbjct: 224 KA-----------DRSSINIADTKGNT 239
>At5g54710 putative protein
Length = 652
Score = 54.7 bits (130), Expect = 3e-08
Identities = 27/92 (29%), Positives = 49/92 (52%)
Query: 33 DDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEV 92
D+ +T LH A E ++++ LCP +VS N + TP+H A N+ +L +LE
Sbjct: 65 DEEQSTLLHKAVTQRNEEYATKVIDLCPSLVSVTNVDGNTPLHLAAEIGNINILWKMLET 124
Query: 93 NPTAACKLNPTCKSAFFVACSHGHLDLVNLLL 124
K+N ++AF +AC + +++ +L+
Sbjct: 125 GEAECMKINKQGQTAFILACLNNNVNSARILV 156
Score = 31.2 bits (69), Expect = 0.30
Identities = 22/99 (22%), Positives = 46/99 (46%), Gaps = 3/99 (3%)
Query: 51 MVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLL--EVNPTAACKLNPTCKSAF 108
++ I+ P+++ +E T +H+AC+ N+++ LL +VN A K++ +
Sbjct: 178 IIDSILEKFPNLILDADEEQSTLLHKACKSGNLEMARTLLDVDVNQEIAEKVDKDGLTPL 237
Query: 109 FVACSHGHLDLV-NLLLNLSEIVEPGLAGFDQACFHIAA 146
A +G ++++ L G + FH+AA
Sbjct: 238 HRAVINGSVEILKEFLCKAPSSFNITTQGTIETVFHLAA 276
Score = 30.0 bits (66), Expect = 0.66
Identities = 17/55 (30%), Positives = 28/55 (50%)
Query: 37 NTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLE 91
NTPLHLA++ G I ++ +++ N+ +T AC NV +L+E
Sbjct: 103 NTPLHLAAEIGNINILWKMLETGEAECMKINKQGQTAFILACLNNNVNSARILVE 157
Score = 29.3 bits (64), Expect = 1.1
Identities = 19/98 (19%), Positives = 44/98 (44%), Gaps = 5/98 (5%)
Query: 51 MVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFV 110
++ +++ P +V +E T +H+A Q N + ++++ P+ N + +
Sbjct: 49 IIDKMLEKFPSLVLDVDEEQSTLLHKAVTQRNEEYATKVIDLCPSLVSVTNVDGNTPLHL 108
Query: 111 ACSHGHLDLVNLLLNLSE-----IVEPGLAGFDQACFH 143
A G+++++ +L E I + G F AC +
Sbjct: 109 AAEIGNINILWKMLETGEAECMKINKQGQTAFILACLN 146
>At5g60070 ankyrin-like protein
Length = 548
Score = 53.9 bits (128), Expect = 4e-08
Identities = 40/160 (25%), Positives = 81/160 (50%), Gaps = 14/160 (8%)
Query: 2 DQEFFNAIKNNDISTFSSIV--------KVREGILNQRTDDTFNTPLHLASKYGCIEMVS 53
D + +A++ D S I+ ++R+ L ++ + T L++A++YG ++V+
Sbjct: 33 DSQLLSAVRRGDFSAVKEILSNHMESEDELRD--LLRKQNQCGETALYVAAEYGDADVVA 90
Query: 54 EIVRLCPDMVSAENE--NMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVA 111
E+++ D+ AE + N P H A +Q + VL +L+E +P + ++ + +A A
Sbjct: 91 ELIKYY-DLEDAETKARNGFDPFHIAAKQGELDVLRVLMEEHPELSMTVDLSNTTALHTA 149
Query: 112 CSHGHLDLVNLLLNLSEIVEPGLAGFD-QACFHIAASRGH 150
+ GH+++V LL + +A + + H AA GH
Sbjct: 150 AAQGHVEVVEYLLEAAGSSLAAIAKSNGKTALHSAARNGH 189
Score = 47.0 bits (110), Expect = 5e-06
Identities = 26/91 (28%), Positives = 52/91 (56%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAA 97
T LH A++ G E+V IV + PD + ++ +TP+H A + +++ V++ L++ + ++
Sbjct: 179 TALHSAARNGHAEVVKAIVAVEPDTATRTDKKGQTPLHMAVKGQSIDVVVELMKGHRSSL 238
Query: 98 CKLNPTCKSAFFVACSHGHLDLVNLLLNLSE 128
+ +A VA G + +V LLL+ +E
Sbjct: 239 NMADSKGNTALHVATRKGRIKIVELLLDNNE 269
Score = 43.5 bits (101), Expect = 6e-05
Identities = 27/90 (30%), Positives = 46/90 (51%), Gaps = 1/90 (1%)
Query: 7 NAIKNNDISTFSSIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAE 66
+A +N +IV V RTD TPLH+A K I++V E+++ ++
Sbjct: 183 SAARNGHAEVVKAIVAVEPDTAT-RTDKKGQTPLHMAVKGQSIDVVVELMKGHRSSLNMA 241
Query: 67 NENMETPIHEACRQENVKVLMLLLEVNPTA 96
+ T +H A R+ +K++ LLL+ N T+
Sbjct: 242 DSKGNTALHVATRKGRIKIVELLLDNNETS 271
Score = 36.2 bits (82), Expect = 0.009
Identities = 22/94 (23%), Positives = 45/94 (47%), Gaps = 1/94 (1%)
Query: 28 LNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSA-ENENMETPIHEACRQENVKVL 86
L+ D + T LH A+ G +E+V ++ ++A N +T +H A R + +V+
Sbjct: 134 LSMTVDLSNTTALHTAAAQGHVEVVEYLLEAAGSSLAAIAKSNGKTALHSAARNGHAEVV 193
Query: 87 MLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLV 120
++ V P A + + ++ +A +D+V
Sbjct: 194 KAIVAVEPDTATRTDKKGQTPLHMAVKGQSIDVV 227
>At3g12360 unknown protein
Length = 590
Score = 50.4 bits (119), Expect = 5e-07
Identities = 24/89 (26%), Positives = 51/89 (56%)
Query: 40 LHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAACK 99
LHLA++ G +E++ ++ P + ++ +T +H A + ++ +V+ LLL+ +P +
Sbjct: 236 LHLAARQGHVEVIKALLSKDPQLARRIDKKGQTALHMAVKGQSSEVVKLLLDADPAIVMQ 295
Query: 100 LNPTCKSAFFVACSHGHLDLVNLLLNLSE 128
+ +C +A VA ++V LLL+L +
Sbjct: 296 PDKSCNTALHVATRKKRAEIVELLLSLPD 324
Score = 47.4 bits (111), Expect = 4e-06
Identities = 34/138 (24%), Positives = 70/138 (50%), Gaps = 10/138 (7%)
Query: 18 SSIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCP-DMVSAENENMETPIHE 76
+ + ++R I+N+ ++ T L A+ G +++V E+++ + ++ +N + P+H
Sbjct: 112 AEVAEIRASIVNE-VNELGETALFTAADKGHLDVVKELLKYSSRESIAKKNRSGYDPLHI 170
Query: 77 ACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLL----NLSEIVEP 132
A Q + ++ +LL+ + T + P+ + A GH ++VN LL NL EI
Sbjct: 171 AAIQGHHAIVEVLLDHDATLSQTFGPSNATPLVSAAMRGHTEVVNQLLSKAGNLLEISRS 230
Query: 133 GLAGFDQACFHIAASRGH 150
++ H+AA +GH
Sbjct: 231 N----NKNALHLAARQGH 244
>At5g50140 ankyrin-like protein
Length = 535
Score = 50.1 bits (118), Expect = 6e-07
Identities = 26/63 (41%), Positives = 40/63 (63%), Gaps = 1/63 (1%)
Query: 30 QRTDDTFN-TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLML 88
+ D++F T LHLA K G E+V +IV + P +VS+ N +TP+H A R + +L+L
Sbjct: 20 EEQDESFGGTFLHLAVKLGNEELVKKIVEIHPSLVSSTNTKSDTPLHLAARLGHTSILLL 79
Query: 89 LLE 91
+LE
Sbjct: 80 MLE 82
Score = 27.7 bits (60), Expect = 3.3
Identities = 29/120 (24%), Positives = 49/120 (40%), Gaps = 10/120 (8%)
Query: 15 STFSSIVKVREGILNQR------TDDTFNTPLHLASKYGCIEMVSEIVRLCP-DMVSAEN 67
ST SI + E + N D F TPLH A G +E ++ + P S
Sbjct: 83 STAESIESLEETVPNDLKLAEMVNKDGF-TPLHCAVMNGSVETLTAFINKAPLSFDSVTL 141
Query: 68 ENMETPIHEACRQENVKVLMLLLEVNPTAAC--KLNPTCKSAFFVACSHGHLDLVNLLLN 125
+ ET H A R + ++ + + + +L+ + A S G L LV+ +++
Sbjct: 142 QTSETVFHLAARHKKMEAFIFMAKNANLRRLLYELDGEGNTVLHAAASVGFLSLVSYIVH 201
>At4g14400 unknown protein
Length = 670
Score = 46.2 bits (108), Expect = 9e-06
Identities = 26/101 (25%), Positives = 52/101 (50%), Gaps = 1/101 (0%)
Query: 1 MDQEFFNAIKNNDISTFSSIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCP 60
M E F + N + + + + + +T ++ LH+A+K+G +E+V EI+ CP
Sbjct: 67 MTPEIFGGMSNGEKECLEKL-RSNGTPMERVKSNTGDSILHIAAKWGHLELVKEIIFECP 125
Query: 61 DMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAACKLN 101
++ +N + +TP+H A + KV+ L+ +A L+
Sbjct: 126 CLLFEQNSSRQTPLHVATHGGHTKVVEALVASVTSALASLS 166
Score = 28.1 bits (61), Expect = 2.5
Identities = 19/75 (25%), Positives = 32/75 (42%), Gaps = 6/75 (8%)
Query: 27 ILNQRT------DDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQ 80
ILN+ T D + P+H A+K E++ E ++ CP N + +H A +
Sbjct: 315 ILNRSTKGVYVCDQDGSFPIHSAAKNEHYEIIKEFIKRCPASKYLLNRLGQNILHVAAKN 374
Query: 81 ENVKVLMLLLEVNPT 95
E +L+ T
Sbjct: 375 EASLTAYMLMHDKDT 389
>At2g01680 unknown protein
Length = 532
Score = 45.8 bits (107), Expect = 1e-05
Identities = 22/84 (26%), Positives = 51/84 (60%)
Query: 41 HLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAACKL 100
H+A+K G + +V E++RL P++ + + +P++ A Q++++++ +L+V+P+ A +
Sbjct: 99 HVAAKRGHLGIVKELLRLWPELCRICDASNTSPLYAAAVQDHLEIVNAMLDVDPSCAMIV 158
Query: 101 NPTCKSAFFVACSHGHLDLVNLLL 124
K++ A +G L +V L+
Sbjct: 159 RKNGKTSLHTAGRYGLLRIVKALI 182
Score = 37.4 bits (85), Expect = 0.004
Identities = 20/92 (21%), Positives = 46/92 (49%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAA 97
T LH A +YG + +V ++ +V +++ +T +H A + +++V+ +L+ + T
Sbjct: 164 TSLHTAGRYGLLRIVKALIEKDAAIVGVKDKKGQTALHMAVKGRSLEVVEEILQADYTIL 223
Query: 98 CKLNPTCKSAFFVACSHGHLDLVNLLLNLSEI 129
+ + +A +A + +LLL + I
Sbjct: 224 NERDRKGNTALHIATRKARPQITSLLLTFTAI 255
Score = 30.8 bits (68), Expect = 0.39
Identities = 28/164 (17%), Positives = 70/164 (42%), Gaps = 26/164 (15%)
Query: 3 QEFFNAIKNNDISTFSSIVKVREG--ILNQRT------------DDTFNTPLHLASKYGC 48
Q FF+++++ D+S +V G ++++ + +D T +++++
Sbjct: 12 QAFFSSVRSGDLSQLQQLVDNLTGDELIDESSPCSAVAELMSVQNDAGETAVYISAAENL 71
Query: 49 IEMVSEIVRLCP-DMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSA 107
++ ++R + V +++ H A ++ ++ ++ LL + P + + S
Sbjct: 72 EDIFRYLIRFSSLETVKIRSKSDMNAFHVAAKRGHLGIVKELLRLWPELCRICDASNTSP 131
Query: 108 FFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQACFHIAASRGHT 151
+ A HL++VN +L++ D +C I G T
Sbjct: 132 LYAAAVQDHLEIVNAMLDV-----------DPSCAMIVRKNGKT 164
Score = 29.3 bits (64), Expect = 1.1
Identities = 22/95 (23%), Positives = 46/95 (48%), Gaps = 1/95 (1%)
Query: 31 RTDDTFNT-PLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLL 89
R D NT PL+ A+ +E+V+ ++ + P +N +T +H A R ++++ L
Sbjct: 122 RICDASNTSPLYAAAVQDHLEIVNAMLDVDPSCAMIVRKNGKTSLHTAGRYGLLRIVKAL 181
Query: 90 LEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLL 124
+E + + ++A +A L++V +L
Sbjct: 182 IEKDAAIVGVKDKKGQTALHMAVKGRSLEVVEEIL 216
Score = 29.3 bits (64), Expect = 1.1
Identities = 16/57 (28%), Positives = 27/57 (47%), Gaps = 1/57 (1%)
Query: 101 NPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAG-FDQACFHIAASRGHTGENKE 156
N ++A +++ + D+ L+ S + + D FH+AA RGH G KE
Sbjct: 56 NDAGETAVYISAAENLEDIFRYLIRFSSLETVKIRSKSDMNAFHVAAKRGHLGIVKE 112
>At5g54720 putative protein
Length = 185
Score = 44.7 bits (104), Expect = 3e-05
Identities = 23/75 (30%), Positives = 38/75 (50%)
Query: 50 EMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFF 109
E ++I+ LCP +V N + TP+H A N +L +L K+N ++AF
Sbjct: 82 EYATKIIDLCPSLVRVANVDGNTPLHLAAEIGNEFILWKMLRCGEADCRKINKQGQTAFI 141
Query: 110 VACSHGHLDLVNLLL 124
+AC + H+ + LL
Sbjct: 142 LACLNNHVAVALTLL 156
Score = 27.7 bits (60), Expect = 3.3
Identities = 17/55 (30%), Positives = 29/55 (51%)
Query: 37 NTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLE 91
NTPLHLA++ G ++ +++R N+ +T AC +V V + LL+
Sbjct: 103 NTPLHLAAEIGNEFILWKMLRCGEADCRKINKQGQTAFILACLNNHVAVALTLLQ 157
>At4g03500 hypothetical protein
Length = 652
Score = 44.3 bits (103), Expect = 3e-05
Identities = 27/100 (27%), Positives = 48/100 (48%), Gaps = 4/100 (4%)
Query: 10 KNNDISTFSSIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENEN 69
K N + + +S + V ++N R NT LHLA+ G + +V I++ CP ++ N
Sbjct: 80 KENYLRSNNSYISVAPTLVNDRG----NTILHLAASSGHVSLVRYIIQKCPGLLLKSNMM 135
Query: 70 METPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFF 109
E +H A ++ V+ L++ +C P K +F
Sbjct: 136 GEVALHLAAEAGHLDVVWNLIDFINDISCTNLPVAKRIYF 175
Score = 42.0 bits (97), Expect = 2e-04
Identities = 17/59 (28%), Positives = 36/59 (60%)
Query: 32 TDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLL 90
+DD P H+A+KYG ++++ EI++ CP+ + + + + +H A + +KV+ +L
Sbjct: 310 SDDDGLFPTHMAAKYGHVQILEEILKHCPEAIELLDRDGQNILHLAAKYGKLKVIKFIL 368
Score = 33.9 bits (76), Expect = 0.046
Identities = 17/54 (31%), Positives = 32/54 (58%), Gaps = 5/54 (9%)
Query: 40 LHLASKYGCIEMVSEIVRLCPD-----MVSAENENMETPIHEACRQENVKVLML 88
LHLA+KYG ++++ I+ C D +++ ++ N TP+H A + KV+ +
Sbjct: 352 LHLAAKYGKLKVIKFILSCCKDKNKKKLINEQDVNGNTPLHLATINWHPKVVSM 405
>At5g54610 ankyrin-repeat-containing protein-like
Length = 426
Score = 43.9 bits (102), Expect = 4e-05
Identities = 26/97 (26%), Positives = 50/97 (50%), Gaps = 3/97 (3%)
Query: 1 MDQEFFNAIKNNDISTFSSIVKVREGILNQRTD--DTFNTPLHLASKYGCIEMVSEIVRL 58
MD + ++ + S+++ IL Q+ D +TPLH AS G +++ E++ L
Sbjct: 1 MDSKLLLVTQSGSVDDLYSLIQAAPDIL-QKVDVLPIIHTPLHEASSAGKLDLAMELMIL 59
Query: 59 CPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPT 95
P NE +P+H A + V++ + L++V+P+
Sbjct: 60 KPSFAKKLNEYGLSPLHLAVENDQVELALELVKVDPS 96
Score = 42.4 bits (98), Expect = 1e-04
Identities = 22/82 (26%), Positives = 43/82 (51%), Gaps = 1/82 (1%)
Query: 8 AIKNNDISTFSSIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAEN 67
A++N+ + +VKV ++ R TPLHL +K G ++++++ + CP+ + N
Sbjct: 78 AVENDQVELALELVKVDPSLVRIRGRGGM-TPLHLVAKKGDVDLLTDFLLACPESIKDVN 136
Query: 68 ENMETPIHEACRQENVKVLMLL 89
N ET +H + + L +L
Sbjct: 137 VNGETILHITIMNDKYEQLKVL 158
Score = 37.7 bits (86), Expect = 0.003
Identities = 33/131 (25%), Positives = 60/131 (45%), Gaps = 10/131 (7%)
Query: 37 NTPLHLASKYGCIEMVSEIVRLCPDMVSAEN--ENMETPIHEACRQENVKVLMLLLEVNP 94
++ L L ++ G ++ + +++ PD++ + + TP+HEA + + M L+ + P
Sbjct: 2 DSKLLLVTQSGSVDDLYSLIQAAPDILQKVDVLPIIHTPLHEASSAGKLDLAMELMILKP 61
Query: 95 TAACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGF----DQACFHIAASRGH 150
+ A KLN S +A + D V L L L + V+P L H+ A +G
Sbjct: 62 SFAKKLNEYGLSPLHLAVEN---DQVELALELVK-VDPSLVRIRGRGGMTPLHLVAKKGD 117
Query: 151 TGENKEFSLLC 161
+F L C
Sbjct: 118 VDLLTDFLLAC 128
Score = 34.3 bits (77), Expect = 0.035
Identities = 18/64 (28%), Positives = 33/64 (51%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAA 97
+PLHLA + +E+ E+V++ P +V TP+H ++ +V +L L P +
Sbjct: 73 SPLHLAVENDQVELALELVKVDPSLVRIRGRGGMTPLHLVAKKGDVDLLTDFLLACPESI 132
Query: 98 CKLN 101
+N
Sbjct: 133 KDVN 136
>At4g03460 hypothetical protein
Length = 637
Score = 43.5 bits (101), Expect = 6e-05
Identities = 17/59 (28%), Positives = 35/59 (58%)
Query: 32 TDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLL 90
+DD + P+H+A KYG ++++ I++ CPD + + + +H A + ++VL +L
Sbjct: 306 SDDDGSFPIHMAVKYGYVKILKAILKRCPDALELLDRENQNVLHVAAKNGKIEVLKFIL 364
Score = 38.9 bits (89), Expect = 0.001
Identities = 31/109 (28%), Positives = 52/109 (47%), Gaps = 6/109 (5%)
Query: 8 AIKNNDISTFSSIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPD-----M 62
A+K + +I+K L + N LH+A+K G IE++ I+R C D +
Sbjct: 317 AVKYGYVKILKAILKRCPDALELLDRENQNV-LHVAAKNGKIEVLKFILRCCKDKNKEKL 375
Query: 63 VSAENENMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVA 111
++ E+ N TP+H A + + KV+ +L N LN +A +A
Sbjct: 376 INEEDANGNTPLHLATKNWHPKVVSMLTWDNRVDLKTLNHDGVTALDIA 424
Score = 30.8 bits (68), Expect = 0.39
Identities = 24/100 (24%), Positives = 48/100 (48%), Gaps = 5/100 (5%)
Query: 30 QRTDDTFNTPLHLASKYGCIEMVSEIVRL-----CPDMVSAENENMETPIHEACRQENVK 84
Q +++ ++ LASK G +V ++ D V +++ PIH A + VK
Sbjct: 265 QHSNNGSSSTSTLASKIGGRSIVHGAMKARRKDKALDSVYVSDDDGSFPIHMAVKYGYVK 324
Query: 85 VLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLL 124
+L +L+ P A L+ ++ VA +G ++++ +L
Sbjct: 325 ILKAILKRCPDALELLDRENQNVLHVAAKNGKIEVLKFIL 364
Score = 28.5 bits (62), Expect = 1.9
Identities = 26/102 (25%), Positives = 39/102 (37%), Gaps = 20/102 (19%)
Query: 53 SEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVAC 112
SE V + P ++A ET + + N+ A +N + +A
Sbjct: 92 SETVPMGPKTIAAVRAGDETYLRDMKFDVNI------------ALSSVNDHGNTMLHLAA 139
Query: 113 SHGHLDLVNLLLNLSEIVEPGLA----GFDQACFHIAASRGH 150
+ GH DLV +LN PGL + H+AA GH
Sbjct: 140 AAGHTDLVCYILN----AYPGLLMKSNSMGEVALHVAAGAGH 177
>At3g09550 putative ankyrin
Length = 436
Score = 42.4 bits (98), Expect = 1e-04
Identities = 27/117 (23%), Positives = 53/117 (45%)
Query: 40 LHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAACK 99
LHLA++ G +++V ++ P + ++ +T +H A + + +V+ LLL +P
Sbjct: 79 LHLAARQGHVDIVRTLLDKDPQLARRTDKKGQTSLHMAVKGVSSQVVRLLLRADPAIVML 138
Query: 100 LNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQACFHIAASRGHTGENKE 156
+ + +A ++VN LL L + L + + IA H+ E E
Sbjct: 139 PDKFGNTVLHIATRKKRAEIVNELLQLPDTNVNALTRDHKTAYDIAEGLTHSEETAE 195
Score = 38.9 bits (89), Expect = 0.001
Identities = 27/111 (24%), Positives = 50/111 (44%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAA 97
TPL A+ G E+V+E++ ++ N + +H A RQ +V ++ LL+ +P A
Sbjct: 43 TPLVSAATRGHSEVVNELLAKDSSLLEISRSNGKNALHLAARQGHVDIVRTLLDKDPQLA 102
Query: 98 CKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQACFHIAASR 148
+ + +++ +A +V LLL + F HIA +
Sbjct: 103 RRTDKKGQTSLHMAVKGVSSQVVRLLLRADPAIVMLPDKFGNTVLHIATRK 153
Score = 36.2 bits (82), Expect = 0.009
Identities = 24/84 (28%), Positives = 44/84 (51%)
Query: 28 LNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLM 87
L +RTD T LH+A K ++V ++R P +V ++ T +H A R++ +++
Sbjct: 101 LARRTDKKGQTSLHMAVKGVSSQVVRLLLRADPAIVMLPDKFGNTVLHIATRKKRAEIVN 160
Query: 88 LLLEVNPTAACKLNPTCKSAFFVA 111
LL++ T L K+A+ +A
Sbjct: 161 ELLQLPDTNVNALTRDHKTAYDIA 184
Score = 35.8 bits (81), Expect = 0.012
Identities = 24/78 (30%), Positives = 40/78 (50%), Gaps = 2/78 (2%)
Query: 74 IHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVN-LLLNLSEIVEP 132
+H AC Q + ++ LLLE P + + + + A + GH ++VN LL S ++E
Sbjct: 11 LHIACSQGHRSIVQLLLEHEPQLSKTVAQSNATPLVSAATRGHSEVVNELLAKDSSLLEI 70
Query: 133 GLAGFDQACFHIAASRGH 150
+ A H+AA +GH
Sbjct: 71 SRSNGKNA-LHLAARQGH 87
Score = 30.0 bits (66), Expect = 0.66
Identities = 16/45 (35%), Positives = 23/45 (50%)
Query: 107 AFFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQACFHIAASRGHT 151
A +ACS GH +V LLL + +A + AA+RGH+
Sbjct: 10 ALHIACSQGHRSIVQLLLEHEPQLSKTVAQSNATPLVSAATRGHS 54
>At1g03670 hypothetical protein
Length = 643
Score = 42.4 bits (98), Expect = 1e-04
Identities = 32/128 (25%), Positives = 60/128 (46%), Gaps = 7/128 (5%)
Query: 5 FFNAIKNNDISTFSSIVKVREGILNQRTDDTFNTPLHLASKYGCIE----MVSEIVRLCP 60
+F I DI +++ G++ R ++ T L + GC E +++E +
Sbjct: 248 YFLVIALTDI--LGIVLRQDPGLIELRNEEG-RTCLSYGASMGCYEGIRYILAEFDKAAS 304
Query: 61 DMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLV 120
+ +++ TPIH A ++ +V+++ L+ P + LN C++ F VA G +V
Sbjct: 305 SLCYVADDDGFTPIHMAAKEGHVRIIKEFLKHCPDSRELLNNQCQNIFHVAAIAGKSKVV 364
Query: 121 NLLLNLSE 128
LL L E
Sbjct: 365 KYLLKLDE 372
Score = 40.4 bits (93), Expect = 5e-04
Identities = 30/120 (25%), Positives = 59/120 (49%), Gaps = 14/120 (11%)
Query: 14 ISTFSSIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPD------MVSAEN 67
ISTF +++ Q + T LH+A++ G + +V +VR + ++A++
Sbjct: 92 ISTFPNLL--------QNVNLMGETTLHVAARAGSLNIVEILVRFITESSSYDAFIAAKS 143
Query: 68 ENMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLLNLS 127
+N +T +H A + ++V+V L+ V + N S ++A G+ +LV +L S
Sbjct: 144 KNGDTALHAALKGKHVEVAFCLVSVKHDVSFDKNNDEASPLYMAVEAGYHELVLKMLESS 203
Score = 40.4 bits (93), Expect = 5e-04
Identities = 29/115 (25%), Positives = 51/115 (44%), Gaps = 7/115 (6%)
Query: 33 DDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEV 92
DD TP+H+A+K G + ++ E ++ CPD N + H A KV+ LL++
Sbjct: 311 DDDGFTPIHMAAKEGHVRIIKEFLKHCPDSRELLNNQCQNIFHVAAIAGKSKVVKYLLKL 370
Query: 93 NPTAAC--KLNPTCKSAFFVACSHGHLDLVNLL-----LNLSEIVEPGLAGFDQA 140
+ + + + +A H + +VN+L +NL + G D A
Sbjct: 371 DEGKRMMNEQDINGNTPLHLATKHRYPIVVNMLTWNDGINLRALNNEGFTALDIA 425
>At2g28840 unknown protein
Length = 456
Score = 40.8 bits (94), Expect = 4e-04
Identities = 29/125 (23%), Positives = 59/125 (47%), Gaps = 5/125 (4%)
Query: 6 FNAIKNNDISTFSSIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVR--LCPDMV 63
F +++ DI T ++ +LNQ T ++ LH+A+ G IE++S ++ PD++
Sbjct: 16 FASVQCGDIITIRRVMATEPSLLNQTTPYDRHSVLHVAAANGQIEILSLLLERFTNPDLL 75
Query: 64 SAENENMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLL 123
N + +TP+ A + + L EV + ++ A +GH + V +
Sbjct: 76 ---NRHKQTPLMLAAMYGRISCVKKLAEVGANILMFDSVNRRTCLHYAAYYGHANCVQAI 132
Query: 124 LNLSE 128
L+ ++
Sbjct: 133 LSAAQ 137
Score = 27.7 bits (60), Expect = 3.3
Identities = 18/61 (29%), Positives = 28/61 (45%), Gaps = 3/61 (4%)
Query: 33 DDTFNTPLHLASKY---GCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLL 89
DD TPLHLA++ C+ ++ + L S TP+H A R ++ + L
Sbjct: 155 DDKGATPLHLAARQRRPECVNVLLDSGSLVCASTSVYGSPGSTPLHLAARSGSIDCVRKL 214
Query: 90 L 90
L
Sbjct: 215 L 215
>At5g15500 unknown protein
Length = 457
Score = 40.4 bits (93), Expect = 5e-04
Identities = 39/151 (25%), Positives = 69/151 (44%), Gaps = 6/151 (3%)
Query: 1 MDQEFFNAI-KNNDISTFSSIVKVREGILNQRTDDTF-NTPLHLASKYGCIEMVSEIVRL 58
MDQ A K+ +I ++ +L++ F NTPLH+A+ G E E++ L
Sbjct: 1 MDQRSLEAAAKSGNIDLLYELIHEDPYVLDKTDHVPFVNTPLHVAAVNGKTEFAMEMMNL 60
Query: 59 CPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLD 118
P N + TP+H A + +++ +++V+P+ + VA S +D
Sbjct: 61 KPSFARKLNADGLTPLHLAVEHGHFWLVLEVVKVDPSLVRIKGRHGMTPLLVAVSRKKID 120
Query: 119 LVN-LLLNLSE-IVEPGLAGFDQACFHIAAS 147
L++ L E IV+ + G + HIA +
Sbjct: 121 LMSEFFLGCPESIVDANVNG--ENALHIAVN 149
Score = 35.0 bits (79), Expect = 0.021
Identities = 19/77 (24%), Positives = 40/77 (51%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAA 97
TPLHLA ++G +V E+V++ P +V + + TP+ A ++ + ++ P +
Sbjct: 74 TPLHLAVEHGHFWLVLEVVKVDPSLVRIKGRHGMTPLLVAVSRKKIDLMSEFFLGCPESI 133
Query: 98 CKLNPTCKSAFFVACSH 114
N ++A +A ++
Sbjct: 134 VDANVNGENALHIAVNN 150
Score = 28.9 bits (63), Expect = 1.5
Identities = 18/57 (31%), Positives = 30/57 (52%), Gaps = 4/57 (7%)
Query: 38 TPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEAC----RQENVKVLMLLL 90
TPL +A I+++SE CP+ + N N E +H A ++E + VL +L+
Sbjct: 108 TPLLVAVSRKKIDLMSEFFLGCPESIVDANVNGENALHIAVNNYDQREGLSVLKVLM 164
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.323 0.137 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,850,995
Number of Sequences: 26719
Number of extensions: 143955
Number of successful extensions: 551
Number of sequences better than 10.0: 66
Number of HSP's better than 10.0 without gapping: 43
Number of HSP's successfully gapped in prelim test: 23
Number of HSP's that attempted gapping in prelim test: 359
Number of HSP's gapped (non-prelim): 183
length of query: 174
length of database: 11,318,596
effective HSP length: 92
effective length of query: 82
effective length of database: 8,860,448
effective search space: 726556736
effective search space used: 726556736
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 56 (26.2 bits)
Medicago: description of AC138130.15


