

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138087.1 + phase: 0
(1399 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
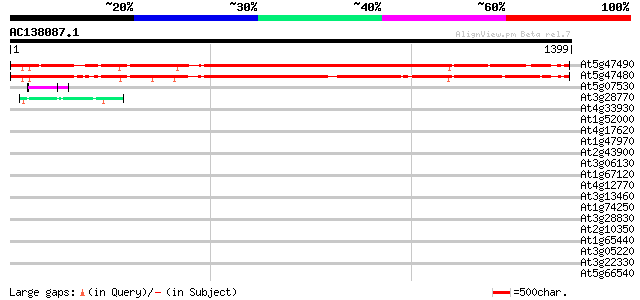
Score E
Sequences producing significant alignments: (bits) Value
At5g47490 putative protein 1173 0.0
At5g47480 putative protein 1138 0.0
At5g07530 glycine-rich protein atGRP-7 48 3e-05
At3g28770 hypothetical protein 44 5e-04
At4g33930 putative protein 42 0.002
At1g52000 42 0.002
At4g17620 hypothetical protein 41 0.004
At1g47970 Unknown protein (T2J15.12) 40 0.009
At2g43900 putative inositol polyphosphate 5'-phosphatase 39 0.027
At3g06130 unknown protein 38 0.036
At1g67120 hypothetical protein 38 0.036
At4g12770 auxilin-like protein 38 0.047
At3g13460 unknown protein 38 0.047
At1g74250 putative heat shock protein 38 0.047
At3g28830 hypothetical protein 37 0.080
At2g10350 hypothetical protein 37 0.080
At1g65440 37 0.080
At3g05220 unknown protein 37 0.10
At3g22330 DEAD-Box RNA helicase like protein 36 0.14
At5g66540 unknown protein 36 0.18
>At5g47490 putative protein
Length = 1361
Score = 1173 bits (3035), Expect = 0.0
Identities = 691/1464 (47%), Positives = 887/1464 (60%), Gaps = 172/1464 (11%)
Query: 1 MASNPPFHVEDQDDEDFFDKLVEDDVGNVN------------DEANDSDDVKAFSNLSIG 48
MAS F +EDQ DEDFFDKLV+D D+ +DSDD++AFSNLSIG
Sbjct: 1 MASASQFLLEDQTDEDFFDKLVDDAYSPTEAQASSSVTELKFDDESDSDDIRAFSNLSIG 60
Query: 49 -----GDDADVNASAFEN--SSGGGSGGEGKERKEEGDVKLDGGNVQEGSSSGCDGMMDR 101
G D +N + N ++ G SG G++ E + + + R
Sbjct: 61 KDPLGGGDGTLNEAILGNDVANEGASGSVGED--EPSSIAPEAVQFPHSDARELRDDEMR 118
Query: 102 SDHGMESRNSSGSSADKSNRRSSLDVKEKDWNAFNVDS--NGGAGSESYSDFFSEFGDQN 159
S+ + + N VKE DW +F+ D N G G SYSDFF+E
Sbjct: 119 SEVADMPLSETAKECTIVNEPGIPGVKELDWGSFDADLSVNDGRGFGSYSDFFTELDATA 178
Query: 160 GKGYDHDLNTEVKHANEIPGDQYAQTYNRDSNTEVKL---GNEIPSDGMNASVDYVQYQE 216
G N +V + GN + +D N SV + Q+Q
Sbjct: 179 G--------------------------NLQGKADVAVATGGNLVANDTNNTSVGFEQHQG 212
Query: 217 GQSYDASARNSTSGEDVNSSQYWESLYPGWKYDYNTGQWYQVDEHNATAATQGSSE---- 272
+D S SG+ V++SQ WE+LYPGWKYD +TGQW+QVD H+A+ +Q S E
Sbjct: 213 QLHHD-----SASGQYVDNSQSWENLYPGWKYDASTGQWFQVDGHDASMNSQESYENSTS 267
Query: 273 ------VNTAEVSYMQQTAQSAVAGTLAESAATETVPSWNQVSQGNNGYPEHMIFDPQYP 326
N ++V+Y +Q+ SAVAGT+ E V +WNQVSQ +NGYPEHM+FD QYP
Sbjct: 268 NWENVAANNSDVAYQRQSTASAVAGTV------ENVSTWNQVSQVSNGYPEHMVFDSQYP 321
Query: 327 GWYYDTIAQEWRSLETYHSSIQYAVQGHG----NGHASSGTFSHNDNSLYRDYGQVGYYE 382
GWYYDTIAQEWRSL++Y+ + Q Q + NG++ + +++++ Y +
Sbjct: 322 GWYYDTIAQEWRSLDSYNQAFQTTGQANDQQVQNGNSFTAVDHSRESNVHDVYDKNQILR 381
Query: 383 SQGVGSQAANNNWSGSYGINHQQDLDRHTTDTA-------TKSGGSAYGGNQQFDHSFGS 435
+Q Q+ + +W SY +QQ + + A T + S GGNQQ ++ + +
Sbjct: 382 TQKFDIQSQHGSWDQSYYDKNQQATNMWQPENAGAAEAAVTPASLSNSGGNQQVNNLYST 441
Query: 436 SNSVNKNQQNASSSFGSVPLYNKVNHGHGLVNGTVEVQRFAPSGNFGQHYNYSNTQFDEQ 495
+ + S VQ F P QH N +N +
Sbjct: 442 GPVAEQFKPYESG-----------------------VQSFIP-----QHMNVANVTQNGP 473
Query: 496 KNISNDYAESHQPFGYSNQSYQSGHQQSYAPNVGRSSAGRPPHALVTFGFGGKLIILKDS 555
+ SN + + + QS+QS Q ++P+ GRSS GRPPHALV FGFGGKLI++KD
Sbjct: 474 MSFSNGFYSRQESVDDAPQSFQSS--QLFSPSAGRSSDGRPPHALVNFGFGGKLILMKDD 531
Query: 556 S--LSSSTYGSQ-GAAQGSVSVLNLMEAVSGSIGSSSIGNGAGDYFRALGQQSIPGPLVG 612
S L +S++GSQ G S+SVLNL E +SGS SS+G + YF L QQS+PGPLVG
Sbjct: 532 SGSLQNSSFGSQKGTGGSSISVLNLAEVISGSASYSSLGENSLSYFSCLDQQSLPGPLVG 591
Query: 613 GSVGSKELNKWIDERIAHCGSPDMDYKKSERMRLLLSLLKIACQYYGKLRSPFGTDTILK 672
G+VGSK+L+KW+DERI +C S MD+ + + +++LLSLL+I+CQYYGKLRSPFG+D + K
Sbjct: 592 GNVGSKDLHKWLDERILNCESSYMDFSRGKLLKMLLSLLRISCQYYGKLRSPFGSDALQK 651
Query: 673 DNDTPGSAVAKLFASAKMSGKE--YGVLSHCLQNLPSEAQMRATASEVQNLLVSGKKKEA 730
+ D+ +AVAKLFA AK G + Y +S CLQ+LP E+QM+ TASEVQNLL SG+K EA
Sbjct: 652 ETDSAEAAVAKLFAIAKEDGVQNGYAPISQCLQHLPPESQMQVTASEVQNLLASGRKMEA 711
Query: 731 LQYAQEGQLWGPALVLASQLGEKFYVDTVKQMALRQLVAGSPLRTLCLLIAGQPAEVFSS 790
LQ AQEG LWGPALV+A+QLG++FYVDTVKQMALRQLV GSPLRTLCLL+AGQPAEVFS+
Sbjct: 712 LQCAQEGHLWGPALVIAAQLGQQFYVDTVKQMALRQLVPGSPLRTLCLLVAGQPAEVFST 771
Query: 791 DS-SNSGDPSAFNMPQNPAQFGSSGMLDDWEENLAVITSNRTKDDELVIIHLGDCLWKER 849
S S+ P + N+P QFG S MLD WEENL +IT+NRT DDELVI HLGDC+WKER
Sbjct: 772 GSTSDISFPGSVNLPPQQPQFGCSSMLDSWEENLGIITANRTTDDELVITHLGDCMWKER 831
Query: 850 SEITAAHICYLIAEANFESYSDSARLCLIGADHWKFPRTYASPEAIQRTELYEYSKVLGN 909
EI AAHICYLIA+ NF++YSD+ARLCL+GADHWK+PRTYASPEAIQRTELYEYSK LGN
Sbjct: 832 GEIIAAHICYLIADKNFDTYSDTARLCLVGADHWKYPRTYASPEAIQRTELYEYSKTLGN 891
Query: 910 SQFILLPFQPYKLIYAYMLAEVGKVSDSLKYCQAVLKSLKTGRAPEVETWKQRLSSLEER 969
SQ+ LL FQPYK++YA+MLAEVGK+S + KYCQAVLK LKTGR+PEVE WKQ +SSLEER
Sbjct: 892 SQYTLLTFQPYKVMYAHMLAEVGKLSTAQKYCQAVLKCLKTGRSPEVEMWKQFVSSLEER 951
Query: 970 IRTHQQGGYAANLAPGKLVGKLLNFFDSTAHRVVGGLPPPAP-SSQGNVHGNE-QNYQSG 1027
IR HQQGGY ANL P KLVG LLNFF S HR VGG+PPPAP S++GN+ GNE Q+ Q
Sbjct: 952 IRIHQQGGYTANLHPEKLVGVLLNFFGSKTHRPVGGMPPPAPHSTKGNLQGNEYQHQQQE 1011
Query: 1028 AHRVSNSQSTMAMSSLVPSGSMEPNGEWTADNNRMTKSNRSVSEPDFGRSPRQETSHDAQ 1087
A +++ SQS MSSL+P S+EP E RM RSVSEPDFGR+P QE + ++
Sbjct: 1012 ATKLAYSQSVNTMSSLMPPASVEPTHESGGSGRRMAVHTRSVSEPDFGRTPIQEMADSSK 1071
Query: 1088 GKASEGT-----------SRFSRFSFGSQLLQKTMGLVLKPRPGKQAKLGEKNKFYYDEN 1136
KA +G SRFSRF FG + + T+G VL R K+AKLG +N+FYYD+
Sbjct: 1072 EKAVDGVTKLKSSGSVAGSRFSRFGFG--IFKDTVGRVL-ARSSKEAKLGAENQFYYDDK 1128
Query: 1137 LKRWVEEGAAPPAEETALPPPPTTAAFQNGLTEYNLQSALKTEGPPSKEGSDLKTSNPEL 1196
LKRWVE G PPAEE ALPPPPT AFQN Y +S + G + N
Sbjct: 1129 LKRWVERGVEPPAEEAALPPPPTIGAFQNNSLGYENKSDMIPSNGNWSSGGPTPSEN--- 1185
Query: 1197 TPGIPPIPPGTNHFSARGRVGIRSRYVDTFN-QGGGNSANLFQSPSVPSAKPVVAANAKF 1255
+ GIPPI G+N FSARGR G+R+RYVDT+N G GNS + SPSV +AKP + A AKF
Sbjct: 1186 SSGIPPISHGSNQFSARGRTGVRARYVDTYNPPGRGNSHTMIPSPSVQTAKPPIPAKAKF 1245
Query: 1256 FIP-TPAPSSNEQTME-AIEENNQEDDLAYENPSTSYRNDWSFQSPKHASASTWQRCPSM 1313
F+P PA SN+Q ME A E QE+ A E ++S QR PSM
Sbjct: 1246 FVPAAPASFSNDQAMEPAAAETRQEEISADEVVASS----------GAPPPMMMQRYPSM 1295
Query: 1314 GNFANHEAVVS--GSNSRSPHSRRTVSWGGSTDVTYSPTKMREIMPLGEALGMPPSTYMS 1371
N + +S G N + P SRRT SW G+ + +++P PST+
Sbjct: 1296 DNIQRNGLGISVNGDNHQPPTSRRTASWSGNFNTSFTPP-------------TSPSTFKP 1342
Query: 1372 DDISSMRTSMKSGNFGEDLHEVDL 1395
++S S + GE+L EV+L
Sbjct: 1343 VLLNS-----SSSSLGEELQEVEL 1361
>At5g47480 putative protein
Length = 1350
Score = 1138 bits (2943), Expect = 0.0
Identities = 678/1464 (46%), Positives = 886/1464 (60%), Gaps = 183/1464 (12%)
Query: 1 MASNPPFHVEDQDDEDFFDKLVEDDVGNVN---------DEANDSDDVKAFSNLSI---- 47
MAS F ++DQ DEDFFDKLV+D D+ +DSDD KAF+NLS+
Sbjct: 1 MASTADFLLDDQTDEDFFDKLVDDSYTPTASSSAKELKFDDGSDSDDAKAFANLSVVDDV 60
Query: 48 -GGDDADVNASAFEN--SSGGGSGGEGKERKEEGDVKLDGGNVQEGSSSGCDGMMDRSDH 104
G D +N + N ++ G SG GKE V ++ D + RS+
Sbjct: 61 LGDGDVALNEAGLGNDVANEGTSGSVGKEEPSSSIAPEAVQFVNSDANRLRDVDVVRSEV 120
Query: 105 GMESRNSSGSSADKSNRRSSLDVKEKDWNAFNVDS--NGGAGSESYSDFFSEFGDQNGKG 162
+ +G ++ + S VKE DW +F DS N G G SYSDFF+E G
Sbjct: 121 DDMALTETGKESNIVDGSGSPGVKEVDWGSFYADSSVNDGGGFGSYSDFFTELDATAG-- 178
Query: 163 YDHDLNTEVKHANEIPGDQYAQTYNRDSNTEVKLGNEIPSDGMNASVDYVQYQEGQSYDA 222
++ + + A G+ A N NT V L N S + Q+Q +D
Sbjct: 179 ---NVQGQAEVAVATGGNLVA---NDTINTSVGLDN---------SAGFEQHQGQVQHD- 222
Query: 223 SARNSTSGEDVNSSQYWESLYPGWKYDYNTGQWYQVDEHNATAATQGSSEVNT------- 275
S SG+ V++SQ WE+LYPGWKYD +TGQWYQVD +AT +Q S +T
Sbjct: 223 ----SGSGQYVDNSQSWENLYPGWKYDASTGQWYQVDGQDATVNSQESYINSTGNWESVA 278
Query: 276 ---AEVSYMQQTAQSAVAGTLAESAATETVPSWNQVSQGNNGYPEHMIFDPQYPGWYYDT 332
++V+Y++Q+ SA+AGT E+V +WNQVSQ NGYPEHM+FD QYPGWYYDT
Sbjct: 279 ADNSDVAYLKQSTTSAMAGT------AESVSTWNQVSQVGNGYPEHMVFDAQYPGWYYDT 332
Query: 333 IAQEWRSLETYHSSIQYAVQGHG------NGHASSGTFSHNDNSLYRDYGQVGY-YESQG 385
IAQEWRSL++Y+ + Q V G NGHA + T+ +N S D +++Q
Sbjct: 333 IAQEWRSLDSYNQASQTTVTGQAHDQQVQNGHARTTTYHNNSQSSVYDVNNKNQTFKAQD 392
Query: 386 VGSQAANNNWSGSYGINHQQDLDR-------HTTDTATKSGGSAYGGNQQFDHSFGSSNS 438
Q + +W SY N+QQ + T S +GGNQQ ++ + + +
Sbjct: 393 FAIQGQHGSWDESYYANNQQAGNTWQPVNVGKAEPAVTSDSLSRFGGNQQVNNLYSTESV 452
Query: 439 VNKNQQNASSSFGSVPLYNKVNHGHGLVNGTVEVQRFAPSGNFGQHYNYSNTQFDEQKNI 498
+ + N T+ Q F P QH N ++ + +
Sbjct: 453 AEQFKPN-----------------------TIGAQSFIP-----QHMNVASATQNGPLSF 484
Query: 499 SNDYAESHQPFGYSNQSYQSGHQQSYAPNVGRSSAGRPPHALVTFGFGGKLIILKDS--S 556
SND Q ++ +S+Q+ Q ++P+VGRSS RPPHALV+FGFGGKLI++KD+ S
Sbjct: 485 SNDLYNRQQSVDHAQKSFQNN--QLFSPSVGRSSDRRPPHALVSFGFGGKLIVMKDNNGS 542
Query: 557 LSSSTYGSQGAAQGSVSVLNLMEAVSGSIGSSSIGNGAGDYFRALGQQSIPGPLVGGSVG 616
L ++++GSQG S++VLNL E +SGS SS G + YFR L QQS+PGPLVGG+VG
Sbjct: 543 LQNTSFGSQGIGGSSITVLNLAEVISGSASYSSPGEDSLSYFRCLHQQSLPGPLVGGNVG 602
Query: 617 SKELNKWIDERIAHCGSPDMDYKKSERMRLLLSLLKIACQYYGKLRSPFGTDTILKDNDT 676
SKEL+KWIDER+ HC S +MD+ + + +++LLSLL+I+CQYYGKLRSPFG+D K+ DT
Sbjct: 603 SKELHKWIDERLLHCESSNMDFSRGKLLKMLLSLLRISCQYYGKLRSPFGSDASQKETDT 662
Query: 677 PGSAVAKLFASAKMSGKE--YGVLSHCLQNLPSEAQMRATASEVQNLLVSGKKKEALQYA 734
P +AVAKLFA AK G + Y +S CLQ+LP E+QM+ TASEVQNLL SG+K EALQ A
Sbjct: 663 PEAAVAKLFAFAKKDGIQNGYAPISQCLQHLPPESQMQVTASEVQNLLASGRKMEALQCA 722
Query: 735 QEGQLWGPALVLASQLGEKFYVDTVKQMALRQLVAGSPLRTLCLLIAGQPAEVFSSDSSN 794
QEG LWGPALV+A+QLG++FYVDTVKQMALRQL+ GSPLRTLCLL+AGQPAEV + SS+
Sbjct: 723 QEGHLWGPALVIAAQLGDQFYVDTVKQMALRQLIPGSPLRTLCLLVAGQPAEVCPTGSSS 782
Query: 795 SGDPSAFNMPQNPAQFGSSGMLDDWEENLAVITSNRTKDDELVIIHLGDCLWKERSEITA 854
S MLD+WEENL +IT+NRT DD+LVIIHLGD +WKER EI A
Sbjct: 783 S-------------------MLDNWEENLGIITANRTTDDDLVIIHLGDSMWKERGEIIA 823
Query: 855 AHICYLIAEANFESYSDSARLCLIGADHWKFPRTYASPEAIQRTELYEYSKVLGNSQFIL 914
AHICYLIA+ NF+ YS+SARLCL+GADHWK PRTYASP+AIQRTELYEYSK LGNSQ+IL
Sbjct: 824 AHICYLIADKNFDPYSESARLCLVGADHWKCPRTYASPDAIQRTELYEYSKTLGNSQYIL 883
Query: 915 LPFQPYKLIYAYMLAEVGKVSDSLKYCQAVLKSLKTGRAPEVETWKQRLSSLEERIRTHQ 974
LPFQPYK+IYA+MLAEVGK+S + KYCQAV++ LKT R+ EVE WKQ SSLEERIR+HQ
Sbjct: 884 LPFQPYKIIYAHMLAEVGKLSTAQKYCQAVIRCLKTSRSSEVEMWKQFASSLEERIRSHQ 943
Query: 975 QGGYAANLAPGKLVGKLLNFFDSTAHRVVGGLPPPAP-SSQGNVHGNE-QNYQSGAHRVS 1032
+GG NLAP KLVGKLLN + G+PPPAP S+ GN NE Q+ Q A ++S
Sbjct: 944 EGG---NLAPAKLVGKLLN--------SLWGMPPPAPHSTTGNPQVNEYQHQQQEAAKLS 992
Query: 1033 NSQSTMAMSSLVPSGSMEPNGEWTADNNRMTKSNRSVSEPDFGRSPRQETSHDAQGKASE 1092
SQS MSSL+P S+EP EW + M +RSVSEPDF R+P Q+ + ++ KA +
Sbjct: 993 YSQSANTMSSLMPPASIEPVHEWGGNGRTMAAHSRSVSEPDFSRTPIQDQTDSSKDKAPD 1052
Query: 1093 G-----------TSRFSRFSFGSQLLQKTMGLVLKPRPGKQAKLGEKNKFYYDENLKRWV 1141
G +SRFSRF G +L+ T+G V R +AKLG +N+FYYD+NLKRWV
Sbjct: 1053 GVTQVKSTRKVPSSRFSRFGIG--ILKNTVGKVFPSRSSNEAKLGNENQFYYDDNLKRWV 1110
Query: 1142 EEGAAPPAEETALPPPPTTAAFQNGLTEYNLQSALKTEGPPSKEGSDLKTSNP-ELTPGI 1200
E G PPAEE ALPPPPT+ F++ + +S +K E PS + P E +PGI
Sbjct: 1111 ERGVEPPAEEAALPPPPTSVPFRSNSLGHENKSEIKNEMSPSSGSWSSGSPTPSENSPGI 1170
Query: 1201 PPIPPGTNHFSARGRVGIRSRYVDTFNQGGGNSANLFQSPSVPSAKPVVAANAKFFIP-T 1259
PP+ G+N FSARGR+G+R+RYVDT+NQG S++++QSP V S+KP + A AKFF+P
Sbjct: 1171 PPVSQGSNQFSARGRMGVRARYVDTYNQG---SSSMYQSPPVQSSKPPIPAKAKFFVPAA 1227
Query: 1260 PAPSSNEQTMEAI----EENNQEDDLAYENPSTSYRNDWSFQSPKHASASTWQRCPSMGN 1315
PA +N+Q ME++ + N D+ + + SFQSP S QR PS+ N
Sbjct: 1228 PASFANDQVMESVSAETRQENSGDEAVVGSAGAPGPSQASFQSPT-PSPIAMQRFPSVDN 1286
Query: 1316 FANHEAVVSGSNSRSPH--SRRTVSWGGSTDVT--YSPTKMREIMPLGEALGMPPSTYMS 1371
+ S N P SRRT SW GS + + SPT ST+
Sbjct: 1287 IRRSGSGTS-LNGDLPQSVSRRTASWSGSVNSSSFMSPTS--------------ASTFRP 1331
Query: 1372 DDISSMRTSMKSGNFGEDLHEVDL 1395
++S S + GE+L EV+L
Sbjct: 1332 SPLNS-----SSSSLGEELQEVEL 1350
>At5g07530 glycine-rich protein atGRP-7
Length = 543
Score = 48.1 bits (113), Expect = 3e-05
Identities = 29/99 (29%), Positives = 43/99 (43%)
Query: 48 GGDDADVNASAFENSSGGGSGGEGKERKEEGDVKLDGGNVQEGSSSGCDGMMDRSDHGME 107
GG +++ S E GG G + K +K + KL G SG + M S+ GM
Sbjct: 262 GGSESEEGMSGSEGGMSGGGGSKSKSKKSKLKAKLGKKKGMSGGMSGSEEGMSGSEGGMS 321
Query: 108 SRNSSGSSADKSNRRSSLDVKEKDWNAFNVDSNGGAGSE 146
S S S + KS ++ L K+ + G +GSE
Sbjct: 322 SGGGSKSKSKKSKLKAKLGKKKSMSGGMSGSEEGMSGSE 360
Score = 43.5 bits (101), Expect = 9e-04
Identities = 24/75 (32%), Positives = 33/75 (44%)
Query: 44 NLSIGGDDADVNASAFENSSGGGSGGEGKERKEEGDVKLDGGNVQEGSSSGCDGMMDRSD 103
++S G ++ S E GG GG+ K RK + L G SG +G M RS+
Sbjct: 344 SMSGGMSGSEEGMSGSEGGMSGGGGGKSKSRKSKLKANLGKKKCMSGGMSGSEGGMSRSE 403
Query: 104 HGMESRNSSGSSADK 118
G+ SG S K
Sbjct: 404 GGISGGGMSGGSGSK 418
Score = 40.4 bits (93), Expect = 0.007
Identities = 29/99 (29%), Positives = 40/99 (40%), Gaps = 1/99 (1%)
Query: 48 GGDDADVNASAFENSSGGGSGGEGKERKEEGDVKLDGGNVQEGSSSGCDGMMDRSDHGME 107
G D S E GG GG+ K K + KL+ G S +G M S+ GM
Sbjct: 220 GMSSGDEGMSGSEGGMSGGEGGKSKSGKGKLKAKLEKKKGMSGGSESEEG-MSGSEGGMS 278
Query: 108 SRNSSGSSADKSNRRSSLDVKEKDWNAFNVDSNGGAGSE 146
S S + KS ++ L K+ + G +GSE
Sbjct: 279 GGGGSKSKSKKSKLKAKLGKKKGMSGGMSGSEEGMSGSE 317
Score = 36.6 bits (83), Expect = 0.10
Identities = 33/122 (27%), Positives = 51/122 (41%), Gaps = 7/122 (5%)
Query: 47 IGGDDADVNASAFENSSGGGSGGEGKERKEEGDVKLDGGNVQEGSSSGCDGMMDRSDHGM 106
+ G + ++ S SSGGGS + K +K + KL G SG + M S+ GM
Sbjct: 306 MSGSEEGMSGSEGGMSSGGGS--KSKSKKSKLKAKLGKKKSMSGGMSGSEEGMSGSEGGM 363
Query: 107 ESRNSSGSSADKSNRRSSLDVKEKDWNAFNVDSNGGAGSESYSD-FFSEFGDQNGKGYDH 165
S + KS +++L K+ + +G G S S+ S G G G H
Sbjct: 364 SGGGGGKSKSRKSKLKANLGKKK----CMSGGMSGSEGGMSRSEGGISGGGMSGGSGSKH 419
Query: 166 DL 167
+
Sbjct: 420 KI 421
Score = 36.6 bits (83), Expect = 0.10
Identities = 25/74 (33%), Positives = 33/74 (43%), Gaps = 2/74 (2%)
Query: 47 IGGDDADVNASAFENSSGGGSGGEGKERKEEGDVK--LDGGNVQEGSSSGCDGMMDRSDH 104
+ G + V+ S S GG SGG G + K G L G ++ SG +G M S+
Sbjct: 446 MSGSEGGVSGSEGSMSGGGMSGGSGSKHKIGGGKHGGLRGKFGKKRGMSGSEGGMSGSEG 505
Query: 105 GMESRNSSGSSADK 118
GM SGS K
Sbjct: 506 GMSESGMSGSGGGK 519
Score = 34.7 bits (78), Expect = 0.40
Identities = 44/171 (25%), Positives = 71/171 (40%), Gaps = 25/171 (14%)
Query: 62 SSGGGSGGEGKERKEEGDVKLDGGNVQEGSSSGCDGMMDRSDHGMESRNSSGSSADKSNR 121
S GG S GK K EG +EG SSG +G M S+ GM S + K
Sbjct: 201 SGGGKSKFGGKGGKSEG---------EEGMSSGDEG-MSGSEGGMSGGEGGKSKSGKGKL 250
Query: 122 RSSLDVK---------EKDWNAFNVDSNGGAGSES---YSDFFSEFGDQNGKGYDHDLNT 169
++ L+ K E+ + +GG GS+S S ++ G + G +
Sbjct: 251 KAKLEKKKGMSGGSESEEGMSGSEGGMSGGGGSKSKSKKSKLKAKLGKKKGMSGGMSGSE 310
Query: 170 EVKHANE--IPGDQYAQTYNRDSNTEVKLG-NEIPSDGMNASVDYVQYQEG 217
E +E + +++ ++ S + KLG + S GM+ S + + EG
Sbjct: 311 EGMSGSEGGMSSGGGSKSKSKKSKLKAKLGKKKSMSGGMSGSEEGMSGSEG 361
Score = 30.4 bits (67), Expect = 7.5
Identities = 32/128 (25%), Positives = 44/128 (34%), Gaps = 13/128 (10%)
Query: 47 IGGDDADVNASAFENSSGGGSGGEGKERKEEGDV---------KLDGGNVQEGSSSGCDG 97
+ G + ++ S S GG SGG G + K G K G + G SG +G
Sbjct: 392 MSGSEGGMSRSEGGISGGGMSGGSGSKHKIGGGKHGGLGGKFGKKRGMSGSGGGMSGSEG 451
Query: 98 MMDRSDHGMESRNSSGSSADK----SNRRSSLDVKEKDWNAFNVDSNGGAGSESYSDFFS 153
+ S+ M SG S K + L K + G +GSE
Sbjct: 452 GVSGSEGSMSGGGMSGGSGSKHKIGGGKHGGLRGKFGKKRGMSGSEGGMSGSEGGMSESG 511
Query: 154 EFGDQNGK 161
G GK
Sbjct: 512 MSGSGGGK 519
>At3g28770 hypothetical protein
Length = 2081
Score = 44.3 bits (103), Expect = 5e-04
Identities = 66/283 (23%), Positives = 104/283 (36%), Gaps = 35/283 (12%)
Query: 24 DDVGNVNDEA-------NDSDDVKAFSNLSIGGDDADVNASAFENS---SGGGSGGEGKE 73
D+VG N + +S D K N GG + + +++ GG KE
Sbjct: 1596 DEVGKENSKTIEVKGRHEESKDGKTNEN---GGKEVSTEEGSKDSNIVERNGGKEDSIKE 1652
Query: 74 RKEEGD-VKLDGGNVQEGSSSGCDGMMDRSDHGMESRNSSGSSADKSNRRSSLDVKEKDW 132
E+G V+++GG DG ++ G E+ GS DK ++ KE
Sbjct: 1653 GSEDGKTVEINGGEELSTEEGSKDGKIEEGKEGKENSTKEGSKDDKI--EEGMEGKE--- 1707
Query: 133 NAFNVDSNGGAGSESYSD--FFSEFGDQNGKGYDHDLNTEVKHANEIPGDQYAQTYNRDS 190
N+ S G +E + D E G ++G +++ + EI G + +
Sbjct: 1708 NSTKESSKDGKINEIHGDKEATMEEGSKDGGTNSTGKDSKDSKSVEINGVKDDSLKDDSK 1767
Query: 191 NTEVKLGNEIPSDGMNASVDYVQYQEGQSYDASARNSTSGE------DVNSSQYWESLYP 244
N ++ N D + +V E Q D S NSTS E D N +
Sbjct: 1768 NGDINEINNGKEDSVKDNV-----TEIQGNDNSLTNSTSSEPNGDKLDTNKDSMKNNTME 1822
Query: 245 ---GWKYDYNTGQWYQVDEHNATAATQGSSEVNTAEVSYMQQT 284
G D G+ + E N + Q +V + E S QT
Sbjct: 1823 AQGGSNGDSTNGETEETKESNVSMNNQNMQDVGSNENSMNNQT 1865
Score = 40.0 bits (92), Expect = 0.009
Identities = 47/220 (21%), Positives = 79/220 (35%), Gaps = 27/220 (12%)
Query: 19 DKLVEDDVGNVNDEANDSDDVKAFSNLSIGGDDADVNASAFENSSGGGSGGEGKERKEEG 78
DK+ E G N S D K I D A+ E S GG+ GK+ K+
Sbjct: 1697 DKIEEGMEGKENSTKESSKDGK------INEIHGDKEATMEEGSKDGGTNSTGKDSKDSK 1750
Query: 79 DVKLDG------------GNVQEGSSSGCDGMMDRSDHGMESRNSSGSSADKSNRRSSLD 126
V+++G G++ E ++ D + D + NS +S LD
Sbjct: 1751 SVEINGVKDDSLKDDSKNGDINEINNGKEDSVKDNVTEIQGNDNSLTNSTSSEPNGDKLD 1810
Query: 127 VKEKDWNAFNVDSNGGAGSESYSDFFSEFGDQNGKGYDHDLNTEVKHANEI------PGD 180
+ +++ GG+ +S + E + N + ++ + N + GD
Sbjct: 1811 TNKDSMKNNTMEAQGGSNGDSTNGETEETKESNVSMNNQNMQDVGSNENSMNNQTTGTGD 1870
Query: 181 QYAQTYNRDSNTEVKLGNEIPSDGMNASVDYVQYQEGQSY 220
T ++TE E+ S N QE QS+
Sbjct: 1871 DIIST---TTDTESNTSKEVTSFISNLEEKSPGTQEFQSF 1907
Score = 32.7 bits (73), Expect = 1.5
Identities = 51/234 (21%), Positives = 90/234 (37%), Gaps = 31/234 (13%)
Query: 21 LVEDDVGNVNDEANDSDDVKAFSNLSIGGDDADVNASAFENSSGGGSGGEGK-------- 72
+VE + ++ +++ +N++ G + + N E++ G G EG
Sbjct: 180 VVEVKSKSSSEASSEESSSTEHNNVTTGSNMVETNGENSESTQEKGDGVEGSNGGDVSME 239
Query: 73 -------ERKEEGDVKLDGGNVQEGSSSGCDGMMDRSDHGMESRNSSGSSADKSNRRSSL 125
E +EG+ ++ G +E + + ++ G S G SA + N S
Sbjct: 240 NLQGNKVEDLKEGNNVVENGETKENNGENVESNNEKEVEG--QGESIGDSAIEKNLESKE 297
Query: 126 DVKEKDWNAFNVDSNGGAGSESYSDFFSEFGDQNGKGYDHDLNTEVKHANEIPGDQYAQT 185
DVK + V++ GS S ++ E NG D EV+ E D +
Sbjct: 298 DVKSE------VEAAKNDGS-SMTENLGEAQGNNGVS-TIDNEKEVEGQGESIEDSDIEK 349
Query: 186 YNRDSNTEVKLGNEIPSD---GMNASVDYVQYQEGQSYD--ASARNSTSGEDVN 234
N +S +VK E + M ++ Q G S + ++ N SGE N
Sbjct: 350 -NLESKEDVKSEVEAAKNAGSSMTGKLEEAQRNNGVSTNETMNSENKGSGESTN 402
>At4g33930 putative protein
Length = 343
Score = 42.0 bits (97), Expect = 0.002
Identities = 34/128 (26%), Positives = 51/128 (39%), Gaps = 6/128 (4%)
Query: 327 GWYYDTIAQEWRSLETYHSSIQYAVQGHGNGHASSGTFSHNDNSLYRDYGQVGYYESQGV 386
GW D W + S+ NGH+S S S + + G S G
Sbjct: 73 GWSSDGTDTNWGWGSSSGSNHSSGTGSTHNGHSSGSNHSSATGSTHNGHTSTGSNHSSGN 132
Query: 387 GSQ----AANNNWSGSYGINHQQDLDRHTTDTATKSGGSAYGGNQQFDHSFGSSNS--VN 440
GS+ ++ +N S S G NH + ++ S S+ G+ +HS GS++S V
Sbjct: 133 GSRHNGYSSGSNHSSSTGSNHSSSTGSTHNNHSSGSNHSSILGSTHKNHSSGSNHSSIVG 192
Query: 441 KNQQNASS 448
N SS
Sbjct: 193 STHNNHSS 200
Score = 31.2 bits (69), Expect = 4.4
Identities = 48/234 (20%), Positives = 76/234 (31%), Gaps = 36/234 (15%)
Query: 85 GNVQEGSSSGCDGMMDRSDHGMESRNSSGSSADKSNRRSSLDVKEK---DWNAFNVDSNG 141
G+ Q GS+ G G S NS GSS S S W++ D+N
Sbjct: 33 GSGQSGSNGGW---------GWRSGNSGGSSGSGSGGSDSNSGGSSWGWGWSSDGTDTNW 83
Query: 142 GAGSESYSDFFSEFGDQNGKGYDHDLNTEVKHANEIPGDQYAQTYNRDSNTEVKLGNEIP 201
G GS S G +H T H G ++ N G+
Sbjct: 84 GWGSSS--------------GSNHSSGTGSTHNGHSSGSNHSSATGSTHNGHTSTGSNHS 129
Query: 202 SDGMNASVDYVQYQEGQSYDASARNSTSGEDVNSSQYWESLYPGWKYDYNTGQWYQVDEH 261
S + Y S S +S++G N+ G + G ++
Sbjct: 130 SGNGSRHNGYSSGSNHSSSTGSNHSSSTGSTHNNHS------SGSNHSSILGSTHK---- 179
Query: 262 NATAATQGSSEVNTAEVSYMQQTAQSAVAGTLAESAATETVPSWNQVSQGNNGY 315
N ++ + SS V + ++ + S++ G+ A V+ NGY
Sbjct: 180 NHSSGSNHSSIVGSTHNNHSSGSNHSSITGSTHNHTAPIPAGRKIAVTVWKNGY 233
>At1g52000
Length = 730
Score = 42.0 bits (97), Expect = 0.002
Identities = 70/328 (21%), Positives = 123/328 (37%), Gaps = 40/328 (12%)
Query: 24 DDVGNVNDEANDSDDVKAFSNLSIGGDDADVNA-SAFENSSGGGSGGEGKERKEEGDVKL 82
+ V N+ D +++ K+ + G + NA S N GG+GG G + G K
Sbjct: 275 ETVSNIGDTESNAGGSKSNDGANNGASGIESNAGSTGTNFGAGGTGGIGDTESDAGGSKT 334
Query: 83 DGGNVQEGSSSGCDGMMDRSDHGMESRNSSGSSADKSN---RRSSLDVKEKDWNAFNV-D 138
+ GN G++ G G+ S+ G N SN +++ KE + A + +
Sbjct: 335 NSGN--GGTNDGASGI--GSNDGSTGTNPGAGGGTDSNIEGTENNVGGKETNPGASGIGN 390
Query: 139 SNGGAGSESYSDFFSEFGDQNGKGYDHDLNTEVKHANEIPGDQYAQTYN--------RDS 190
S+G G+ + G + G + N P AQ N D
Sbjct: 391 SDGSTGTSPEGTESNADGTKTNTGGKESNTGSESNTNSSPQKLEAQGGNGGNQWDDGTDH 450
Query: 191 NTEVKLGNEIPSDGM-NASVDYV---QYQEGQSYDASARNSTSGEDVNSSQYW----ESL 242
+ +K+ + G+ DYV Q +EG + R TS +++ + E L
Sbjct: 451 DGVMKIHVAVGGLGIEQIRFDYVKNGQLKEGPFHGVKGRGGTSTIEISHPDEYLVSVEGL 510
Query: 243 YPG--------WKYDYNTGQWYQVDEHNATAATQGSSEVNTAEVSYMQQTAQSAVAGTLA 294
Y ++ + +T Q++ + + TQ S +VN ++ A S + A
Sbjct: 511 YDSSNIIQGIQFQSNKHTSQYFGYEYYG--DGTQFSLQVNEKKIIGFHGFADSHLNSLGA 568
Query: 295 -----ESAATETVPSWNQVSQGNNGYPE 317
S+++ P N+V Y E
Sbjct: 569 YFVPISSSSSSLTPPPNKVKAQGGSYGE 596
Score = 31.2 bits (69), Expect = 4.4
Identities = 43/199 (21%), Positives = 75/199 (37%), Gaps = 25/199 (12%)
Query: 48 GGDDADVNASAFENSSGGG-------SGGEGKERKEEGDVKLDGGNVQ--EGSSSGCDGM 98
G + NA ++SSG +GG G+ GD + + G + +G+++G G+
Sbjct: 244 GNGGTEKNAGGSKSSSGSARTNPGASAGGNGETVSNIGDTESNAGGSKSNDGANNGASGI 303
Query: 99 MDRSDHGMESRN-SSGSSADKSNRRSSLDVKEKDWNAFNVDSNGGAGSESYSDFFSEFGD 157
S+ G N +G + + S D N+ N +N GA +D G
Sbjct: 304 --ESNAGSTGTNFGAGGTGGIGDTES--DAGGSKTNSGNGGTNDGASGIGSND--GSTGT 357
Query: 158 QNGKGYDHDLNTEVKHANEIPGDQYAQTYNRDSNTEVKLGNEIPSDGMNASVDYVQYQEG 217
G G D N E N + G + + N++ G NA +G
Sbjct: 358 NPGAGGGTDSNIEGTE-NNVGGKETNPGASGIGNSDGSTGTSPEGTESNA--------DG 408
Query: 218 QSYDASARNSTSGEDVNSS 236
+ + S +G + N++
Sbjct: 409 TKTNTGGKESNTGSESNTN 427
Score = 30.4 bits (67), Expect = 7.5
Identities = 29/114 (25%), Positives = 49/114 (42%), Gaps = 8/114 (7%)
Query: 355 GNGHASSGTFSHNDNSLYRDYGQ----VGYYESQGVGSQAANNNWSGSYGINHQQDLDRH 410
G +SSG+ N + G+ +G ES GS++ + +G+ GI + +
Sbjct: 253 GGSKSSSGSARTNPGASAGGNGETVSNIGDTESNAGGSKSNDGANNGASGI----ESNAG 308
Query: 411 TTDTATKSGGSAYGGNQQFDHSFGSSNSVNKNQQNASSSFGSVPLYNKVNHGHG 464
+T T +GG+ G+ + D +NS N + +S GS N G G
Sbjct: 309 STGTNFGAGGTGGIGDTESDAGGSKTNSGNGGTNDGASGIGSNDGSTGTNPGAG 362
>At4g17620 hypothetical protein
Length = 1225
Score = 41.2 bits (95), Expect = 0.004
Identities = 30/146 (20%), Positives = 60/146 (40%), Gaps = 6/146 (4%)
Query: 10 EDQDDEDFFDKLVEDDVGNVNDEANDSDDVKAFSNLSIGGDDADVNASAFENSSGGGSGG 69
ED D++ + GN + ++ S + S+ + G + D + ++S GSG
Sbjct: 706 EDSDNDSRSSRSKSSSSGNASSSSSSSSSSSSSSSAAGGDGEGDGGGADSGSASDSGSGS 765
Query: 70 EGKERKEEGDVKLDGGNVQEGSSSGCDGMMDRSDHGMESRNSSGSSADKSNRRS-----S 124
G ++E GD K++ + SG + + + ++ GS ++ + S
Sbjct: 766 SG-GKEEHGDDKVESYRSNDNGESGVYPYEEEEEEEEDEKDLFGSDNEEYTKTPAISTYS 824
Query: 125 LDVKEKDWNAFNVDSNGGAGSESYSD 150
+ V W+ N GG G +S+
Sbjct: 825 IPVLPAGWSNDNHGGRGGMGRGRWSN 850
Score = 30.4 bits (67), Expect = 7.5
Identities = 26/95 (27%), Positives = 37/95 (38%), Gaps = 2/95 (2%)
Query: 100 DRSDHGMESRNSSGSSADKSNRRSSLDVKEKDWNAFNVDSNG-GAGSESYSDFFSEFGDQ 158
D + SR+ S SS + S+ SS +A D G G G++S S S G
Sbjct: 707 DSDNDSRSSRSKSSSSGNASSSSSSSSSSSSSSSAAGGDGEGDGGGADSGSASDSGSGSS 766
Query: 159 NGKGYDHDLNTEVKHANEIPGDQYAQTYNRDSNTE 193
GK D E +N+ G+ Y + E
Sbjct: 767 GGKEEHGDDKVESYRSND-NGESGVYPYEEEEEEE 800
>At1g47970 Unknown protein (T2J15.12)
Length = 198
Score = 40.0 bits (92), Expect = 0.009
Identities = 35/141 (24%), Positives = 62/141 (43%), Gaps = 15/141 (10%)
Query: 9 VEDQDDEDFFDKLVEDDVGNVNDEANDSDDVKAFSNLSIGGDDADVNASAFENSSGGGSG 68
V D+DD+D + +DD D+ +D DDV+ S+GG SA + G
Sbjct: 38 VSDEDDDDGSEGDDDDDEEEEEDD-DDDDDVQVLQ--SLGGPPVQ---SAEDEDEEGDED 91
Query: 69 GEGKERKEEGDVKLDGG-----NVQEGSSSGCDGMMDRSDHGMESRNSSGSSADKSNRRS 123
G G + ++GD D +V++ G + ++ + ++S +++
Sbjct: 92 GNGDDDDDDGDDDDDDDDDEDEDVEDEGDLGTEYLVRPVGRAEDEEDASDFEPEENGVEE 151
Query: 124 SLDVKEKDWNAFNVDSNGGAG 144
+D E D N D++GGAG
Sbjct: 152 DIDEGEDDEN----DNSGGAG 168
>At2g43900 putative inositol polyphosphate 5'-phosphatase
Length = 1305
Score = 38.5 bits (88), Expect = 0.027
Identities = 42/194 (21%), Positives = 79/194 (40%), Gaps = 20/194 (10%)
Query: 44 NLSIGGDDADVNASAFENSSGGGSGGEGKERKEEGDV------KLDGGNVQEGSSSGCDG 97
NL I + ++ + + + + G + K K +GD K DG + + S
Sbjct: 1098 NLRIDSNPSNSKSQSLKKNEGDSNSKSSK--KSDGDSNSKSSKKSDGDSNSKSSKKS--- 1152
Query: 98 MMDRSDHGMESRNSSGSSADKSNRRSSLDVKEKDWNAFNVDSNGGAGSESYSDFFSEF-- 155
D + S+ S G S KS+++S D K + DSN + +S D S+
Sbjct: 1153 --DGDSNSKSSKKSDGDSNSKSSKKSDGDSNSKSSKKSDGDSNSKSSKKSDGDSCSKSQK 1210
Query: 156 ---GDQNGKGYDH-DLNTEVKHANEIPGDQYAQTYNRDSNTEVKLGNEIPSDGMNASVDY 211
GD N K D ++ K + GD ++++ ++ ++ SDG ++S +
Sbjct: 1211 KSDGDTNSKSQKKGDGDSSSKSHKKNDGDSSSKSHKKNDGDSSSKSHK-KSDGDSSSKSH 1269
Query: 212 VQYQEGQSYDASAR 225
+ + S + R
Sbjct: 1270 KKSEGDSSLSLTRR 1283
Score = 30.0 bits (66), Expect = 9.8
Identities = 29/136 (21%), Positives = 48/136 (34%), Gaps = 10/136 (7%)
Query: 12 QDDEDFFDKLVEDDVGNVNDEANDSDDVKAFSNLSIGGDDADVNASAFENSSGGGSGGEG 71
+ D D K + G+ N +++ D + S S D D N+ + G G
Sbjct: 1175 KSDGDSNSKSSKKSDGDSNSKSSKKSDGDSCSK-SQKKSDGDTNSKS--QKKGDGDSSSK 1231
Query: 72 KERKEEGDVKLDGGNVQEGSSSGCDGMMDRSDHGMESRNSSGSSADKSNRRSSLDVKEKD 131
+K +GD +G SS H +SS S KS SSL + +
Sbjct: 1232 SHKKNDGDSSSKSHKKNDGDSSS-------KSHKKSDGDSSSKSHKKSEGDSSLSLTRRT 1284
Query: 132 WNAFNVDSNGGAGSES 147
++ G ++
Sbjct: 1285 MEILQAHTSRRVGRKT 1300
>At3g06130 unknown protein
Length = 473
Score = 38.1 bits (87), Expect = 0.036
Identities = 30/130 (23%), Positives = 50/130 (38%), Gaps = 15/130 (11%)
Query: 10 EDQDDEDFF----DKLVEDDVGNVNDEAND----SDDVKAFSNLSIGGDDADVNASAFEN 61
ED DD+DF D+ +DD +DE +D S+ +K + +NA N
Sbjct: 188 EDDDDDDFSDEFDDEFTDDDDDEFDDEFDDLPLPSNKMKPNMTMMPNAQQMMMNAQKNAN 247
Query: 62 SSGGGSGGEGKERKEEGDVKLDGGNVQEGSSSGCDGMMDR-------SDHGMESRNSSGS 114
+GG + GK G GG G + G G ++ H + +N G
Sbjct: 248 LAGGPAKNGGKGAPAAGGGGAGGGKGAGGGAKGGPGNQNQGGGKNGGGGHPQDGKNGGGG 307
Query: 115 SADKSNRRSS 124
+ ++ +
Sbjct: 308 GGPNAGKKGN 317
>At1g67120 hypothetical protein
Length = 5138
Score = 38.1 bits (87), Expect = 0.036
Identities = 38/187 (20%), Positives = 69/187 (36%), Gaps = 9/187 (4%)
Query: 10 EDQDDEDFFDKLVEDDVGNVNDEANDSDDVKAFSNLSIGGDDADVNASAFENSSGGGSGG 69
ED++DE D+ +++ +G+V +A +D+ K ++ D+ D + E + G S
Sbjct: 4357 EDKEDEGSEDEPLDNGIGDVGSDAEKADE-KPWNK-----DEEDEEENMNEKNESGPSIV 4410
Query: 70 EGKERKEEGDVKLDGGNVQEGSSSGCDGMMDRSDHGMESRNSSGSSADKSNRRSSLDVKE 129
+ R E K DG V+ D+ + G + D N + KE
Sbjct: 4411 DKDTRSRELRAKDDG--VETADEPEESNTSDKPEEGNDENVEQDDFDDTDNLEEKIQTKE 4468
Query: 130 KDWNAFNVD-SNGGAGSESYSDFFSEFGDQNGKGYDHDLNTEVKHANEIPGDQYAQTYNR 188
+ D N + D E ++ + + + KH E DQ
Sbjct: 4469 EALGGLTPDVDNEQIDDDMEMDKTEEVEKEDANQQEEPCSEDQKHPEEGENDQEETQEPS 4528
Query: 189 DSNTEVK 195
+ N E +
Sbjct: 4529 EENMEAE 4535
>At4g12770 auxilin-like protein
Length = 909
Score = 37.7 bits (86), Expect = 0.047
Identities = 38/167 (22%), Positives = 71/167 (41%), Gaps = 10/167 (5%)
Query: 2 ASNPPFHVEDQDDEDFFDKLVEDDVGNVNDEANDSDDVKAFSNLSIGGDD-ADVNASAFE 60
+S P + DDED F+ + E + + + ++ ++V FS++S N+S F+
Sbjct: 109 SSMPVYDKPVYDDEDVFESIPELKIPSTSSQSARFENV--FSSISSSPTKHRKQNSSPFD 166
Query: 61 NSSGGGSGGEGKERKEEGDVKLDGGNVQEGSSSGCDGMMDRSDHGMESRNSSGSSADKSN 120
+ G G + +R+E+G D G +S S+ +S+ +A+ S+
Sbjct: 167 DLMGNNLGKKESDREEKGSSIFDDLIPGFGRTSSPPAKRTTSETTSQSQKPPYRTAETSS 226
Query: 121 --RRSSLDVKEKDWNAFNVDSNGGAGSESYSDFFSEFGDQNGKGYDH 165
+ V E+ + S GG ++D E G N + DH
Sbjct: 227 NVKEDPFVVLEESTSTLREPSTGG-----FTDPLEEIGKFNSRKTDH 268
>At3g13460 unknown protein
Length = 608
Score = 37.7 bits (86), Expect = 0.047
Identities = 81/391 (20%), Positives = 135/391 (33%), Gaps = 81/391 (20%)
Query: 200 IPSDGMNASVDYVQ-----------YQEGQSY-DASARNSTSGEDVNSSQYWES---LYP 244
+PSD + SV YV Y GQ + D A + G D+NS Y E+ +YP
Sbjct: 2 LPSDAADPSVCYVPNPYNPYQYYNVYGSGQEWTDYPAYTNPEGVDMNSGIYGENGTVVYP 61
Query: 245 ---GWK-YDYNT-----------GQWYQVDEHNATAATQGSSEVNTAEVSYMQ-QTAQSA 288
G+ Y Y+ GQ Y ++ S ++ + Q + +
Sbjct: 62 QGYGYAAYPYSPATSPAPQLGGEGQLYGAQQYQYPNYFPNSGPYASSVATPTQPDLSANK 121
Query: 289 VAGTLAESAATETVPSWNQVSQGNNGY----PEHMIF----------------------D 322
AG A + V S +++G+NG P + D
Sbjct: 122 PAGVKTLPADSNNVASAAGITKGSNGSAPVKPTNQATLNTSSNLYGMGAPGGGLAAGYQD 181
Query: 323 PQYP--GWYYDTIAQEWRSLETYHSSIQYAVQGHG--NGHASSGTFSHNDNSLYRDYGQ- 377
P+Y G+Y W Y S +Q V G G + ++ S T + N YR
Sbjct: 182 PRYAYEGYYAPV---PWHDGSKY-SDVQRPVSGSGVASSYSKSSTVPSSRNQNYRSNSHY 237
Query: 378 ---------VGYYESQGVGSQAANNNWSGSYGINHQQDLDRHTTDTATKSGGSAYGGNQQ 428
GY +QG ++ N G YG + L ++ +++ G +
Sbjct: 238 TSVHQPSSVTGYGTAQGYYNRMYQNKLYGQYGSTGRSALGYGSSGYDSRTNGRGWAATDN 297
Query: 429 FDHSFGSSNSV---NKNQQNASSSFGSVPLYNKVNHGHGLVNGTVEVQRFAPSGNFGQHY 485
S+G NS N+N + + P + G ++ ++EV+ N +
Sbjct: 298 KYRSWGRGNSYYYGNENNVDGLNELNRGPRAKGTKNQKGNLDDSLEVKEQTGESNVTEVG 357
Query: 486 NYSNTQFDEQKNISNDYAESHQPFGYSNQSY 516
NT + Y + P Y+N +
Sbjct: 358 EADNTCVVPDR---EQYNKEDFPVDYANAMF 385
>At1g74250 putative heat shock protein
Length = 630
Score = 37.7 bits (86), Expect = 0.047
Identities = 47/187 (25%), Positives = 82/187 (43%), Gaps = 31/187 (16%)
Query: 10 EDQDDE-DFFDKLV---EDDVGNVNDEANDSDDVKAFSNLSIGGDDADVNASAFENSSGG 65
ED+DDE K+V ++ NV + D D+ + + I GD A+ S F+N
Sbjct: 414 EDEDDEMTLLKKMVSGQKNKQKNVVSKEEDEDE----TEVEIEGDTAEF--SEFDNQK-- 465
Query: 66 GSGGEGKERKEEGDVKLDGGNVQEGSSS----GCDGMMDRSDHGMESRNSSGSSADKSNR 121
S G KE KEE + + G ++ + +S G G D + + ES +SG+ AD
Sbjct: 466 -STGRNKEAKEERNKQNAGNDMADDTSKVQIPGEGGNPDENMNATES--ASGALADS--- 519
Query: 122 RSSLDVKEKDWNAFNVDSNGGAGSESYSDFFSEFGDQNGKGYDHDLNTEVKHANEIPGDQ 181
++ + N+ D+ G S + D+N +G ++ ++E + D
Sbjct: 520 ------QKDEANSMEYDNRKSTGRRRRS---KKGKDKNNQGELNEKSSEADDTQYVNRDM 570
Query: 182 YAQTYNR 188
+Q Y +
Sbjct: 571 ESQDYKK 577
>At3g28830 hypothetical protein
Length = 536
Score = 37.0 bits (84), Expect = 0.080
Identities = 34/121 (28%), Positives = 49/121 (40%), Gaps = 7/121 (5%)
Query: 44 NLSIGGDDADVNASAFENSSGGGSGGEGKERKEEGDVKLDGG---NVQEGSSSGCDGMMD 100
N++ G D DVN S+ GG + E K G V G +E SSSG
Sbjct: 177 NIASYGLDVDVNDSSIV---GGAASSESSSTKS-GSVSSSGSVSTKSKESSSSGSSASGS 232
Query: 101 RSDHGMESRNSSGSSADKSNRRSSLDVKEKDWNAFNVDSNGGAGSESYSDFFSEFGDQNG 160
+ ES S ++ K + S K K+ + + + +GS S S S G +G
Sbjct: 233 VATKSKESSGGSAATKSKESSGGSAATKSKESSGGSATTGKTSGSPSGSPKASPSGSVSG 292
Query: 161 K 161
K
Sbjct: 293 K 293
>At2g10350 hypothetical protein
Length = 863
Score = 37.0 bits (84), Expect = 0.080
Identities = 35/150 (23%), Positives = 64/150 (42%), Gaps = 20/150 (13%)
Query: 96 DGMMDRSDHGMESRNSSGSSADKSNRRSSLDVKEKDWNAFNVDSNGGAGSESYSDFFS-- 153
D MD ++S +S+ A++ N +DVK ++ N +VDS SE S
Sbjct: 375 DDPMDVFVTPLQSEHSNNDDANEGNPVYDIDVKYQNANEEDVDSQMQDASERVPAIHSGL 434
Query: 154 ------EFGDQNGKGYDHDLNTEVKHANE-IPGDQYAQTYNRDSNTEVKLGNEIPSDGMN 206
+ +Q D D+++++K A+E +P ++ N+E E+ ++
Sbjct: 435 DLSKEHNYEEQESNANDEDVDSQMKDASERVPATHSGLDLPKEHNSE-----ELQTNANK 489
Query: 207 ASVDYVQYQEGQSYDASARNSTSGEDVNSS 236
VD G+ D+ R S D++ S
Sbjct: 490 TDVD------GKMQDSLDREPASHSDIDLS 513
>At1g65440
Length = 1703
Score = 37.0 bits (84), Expect = 0.080
Identities = 28/88 (31%), Positives = 36/88 (40%), Gaps = 12/88 (13%)
Query: 48 GGDDADVNASAFENSSGGGSGG-----EGKERKEEGDVKL-------DGGNVQEGSSSGC 95
GG+ +D S S GGGSGG GK+ E+G D GN G S
Sbjct: 1609 GGNKSDGAGSWGSGSGGGGSGGWGNDSGGKKSSEDGGFGSGSGGGGSDWGNESGGKKSSA 1668
Query: 96 DGMMDRSDHGMESRNSSGSSADKSNRRS 123
DG G +S G + S+R+S
Sbjct: 1669 DGGWGSESGGKKSDGEGGWGNEPSSRKS 1696
Score = 34.3 bits (77), Expect = 0.52
Identities = 34/141 (24%), Positives = 50/141 (35%), Gaps = 13/141 (9%)
Query: 27 GNVNDEANDSDDVKAFSNLSIGGDDADVNASAFENSSGGGSGGEGKERKEEGDV-----K 81
G +D +D D N G +D + NS GGG G E +K G
Sbjct: 1551 GRRDDMNSDRQD----GNGDWGNNDTGTADGGWGNSGGGGWGSESAGKKTGGGSTGGWGS 1606
Query: 82 LDGGNVQEGSSSGCDGMMDRSDHGMESRNSSGSSADKSNRRSSLDVKEKDWNAFNVDSNG 141
GGN +G+ S G G + + S++ S DW + +G
Sbjct: 1607 ESGGNKSDGAGSWGSGSGGGGSGGWGNDSGGKKSSEDGGFGSGSGGGGSDWG----NESG 1662
Query: 142 GAGSESYSDFFSEFGDQNGKG 162
G S + + SE G + G
Sbjct: 1663 GKKSSADGGWGSESGGKKSDG 1683
>At3g05220 unknown protein
Length = 541
Score = 36.6 bits (83), Expect = 0.10
Identities = 36/165 (21%), Positives = 59/165 (34%), Gaps = 14/165 (8%)
Query: 7 FHVEDQDDEDFFDKLVEDDVGNVNDEANDSDDV-KAFSNLSIGGDDADVNASAFEN---- 61
F +D DDEDF + +DD +D+ +D D++ + GG+ N N
Sbjct: 148 FSEDDYDDEDFSEDDYDDD--EFDDDEDDDDEMGHGHGHGGGGGNHMPPNKMMMPNKMMP 205
Query: 62 ------SSGGGSGGEGKERKEEGDVKLDGGNVQEGSSSGCDGMMDRSDHGMESRNSSGSS 115
+GGG G + K GG ++G G + + G E +N
Sbjct: 206 QMGGHHGNGGGPKGPNEIMMMMNGFKGGGGGGKKGGGGGFEIPVQMKGMG-EGKNGKDGK 264
Query: 116 ADKSNRRSSLDVKEKDWNAFNVDSNGGAGSESYSDFFSEFGDQNG 160
K + + KE ++ +G FF + NG
Sbjct: 265 KGKGGEKGKKEGKENKGGGKTGKTDAKSGGGGLLGFFKKGKSGNG 309
>At3g22330 DEAD-Box RNA helicase like protein
Length = 616
Score = 36.2 bits (82), Expect = 0.14
Identities = 41/155 (26%), Positives = 63/155 (40%), Gaps = 30/155 (19%)
Query: 20 KLVEDDVGNVNDE----------ANDSDDVKAFSNLSIGGDDADVNASAFENSSGGGSGG 69
K++E +VG+ E A+ + + + S S GG D +S S GGG GG
Sbjct: 461 KIIEREVGSRFTELPSIAVERGSASMFEGIGSRSGGSFGGGMRDRGSSFGGRSGGGGYGG 520
Query: 70 EGKERKEEGDVKLDGGNVQEGSSSGCDGMMDRSDHG-MESRNSSGSSADKSNRRSSLDVK 128
GG GSS+ G DRS G R+ G +D+S++
Sbjct: 521 SSGGY---------GGGRSGGSSNRYSGDSDRSGFGSFGMRSPEGYGSDRSSQSGGR--- 568
Query: 129 EKDWNAFNVDSNGGAG---SESYSDFFSEFGDQNG 160
++F +GG+ S + DF S+ Q+G
Sbjct: 569 ----SSFGGGRSGGSSNNRSSGFGDFGSDRSSQSG 599
>At5g66540 unknown protein
Length = 524
Score = 35.8 bits (81), Expect = 0.18
Identities = 40/190 (21%), Positives = 76/190 (39%), Gaps = 37/190 (19%)
Query: 23 EDDVGNVNDEANDSDDVKAFSNLSIGGDDADVNASAFENSSGGGSGGEGKERKEEGDVKL 82
EDD+ ++ + DSDDV DD D + +S G E ++ +EE + +
Sbjct: 108 EDDIDEMDMDGFDSDDV----------DDEDKEIES-NDSEGEDEEEEEEDEEEEEEEEE 156
Query: 83 DGGNVQEGSSSGCDGMM-------------DRSDHGMESRNSSG-SSADKSNRRSSLDVK 128
+ ++G + G + + ++G++ +N G + K N D +
Sbjct: 157 EEEEEKDGDNEGIEDKFFKIKELEEFLEEGEAEEYGIDHKNKKGVAQRKKQNLSDDEDEE 216
Query: 129 EKDWNAFNVDSNGGAGSES----------YSDFFSEFGDQNGKGYDHDLNTEVKHANEIP 178
+ D +V+ + AG + Y DFF G + K DL+ + + E
Sbjct: 217 DDDDEEEDVEFDAFAGGDDEETDKLGKARYDDFFG--GKKETKMKLKDLSEDEEAEIENK 274
Query: 179 GDQYAQTYNR 188
G++ T+ R
Sbjct: 275 GNEKLSTHER 284
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.309 0.128 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,630,417
Number of Sequences: 26719
Number of extensions: 1667804
Number of successful extensions: 5871
Number of sequences better than 10.0: 147
Number of HSP's better than 10.0 without gapping: 16
Number of HSP's successfully gapped in prelim test: 142
Number of HSP's that attempted gapping in prelim test: 5489
Number of HSP's gapped (non-prelim): 410
length of query: 1399
length of database: 11,318,596
effective HSP length: 112
effective length of query: 1287
effective length of database: 8,326,068
effective search space: 10715649516
effective search space used: 10715649516
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 66 (30.0 bits)
Medicago: description of AC138087.1


