

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138063.5 + phase: 0 /pseudo
(840 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
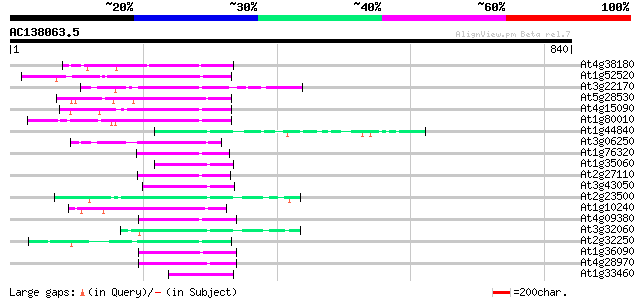
Score E
Sequences producing significant alignments: (bits) Value
At4g38180 unknown protein (At4g38180) 73 8e-13
At1g52520 F6D8.26 67 5e-11
At3g22170 far-red impaired response protein, putative 65 2e-10
At5g28530 far-red impaired response protein (FAR1) - like 65 2e-10
At4g15090 unknown protein 64 4e-10
At1g80010 hypothetical protein 60 4e-09
At1g44840 hypothetical protein 60 4e-09
At3g06250 unknown protein 59 1e-08
At1g76320 putative phytochrome A signaling protein 57 4e-08
At1g35060 hypothetical protein 55 2e-07
At2g27110 Mutator-like transposase 52 1e-06
At3g43050 putative protein 52 2e-06
At2g23500 Mutator-like transposase 52 2e-06
At1g10240 unknown protein 52 2e-06
At4g09380 putative protein 51 2e-06
At3g32060 hypothetical protein 51 3e-06
At2g32250 Mutator-like transposase 50 4e-06
At1g36090 hypothetical protein 50 4e-06
At4g28970 putative protein 50 5e-06
At1g33460 mutator transposase MUDRA, putative 49 9e-06
>At4g38180 unknown protein (At4g38180)
Length = 788
Score = 72.8 bits (177), Expect = 8e-13
Identities = 67/268 (25%), Positives = 120/268 (44%), Gaps = 18/268 (6%)
Query: 79 KKSKREGTGSRKCGCPFRLRGYFSATKLWSLNIVSGL---HNHKMEPKLEGHMLAG--RL 133
++ KR T +R GC L + W +VSG HNH++ P + H L ++
Sbjct: 137 REIKRPRTITR-VGCKASLSVKMQDSGKW---LVSGFVKDHNHELVPPDQVHCLRSHRQI 192
Query: 134 TAKESKIVGDMTRNLIKPKNILLDL----KGRWKDNRTVAKQIYNARHRYKLSIRGSR*E 189
+ ++ + + P+ I+ L G K T R+ + SI G
Sbjct: 193 SGPAKTLIDTLQAAGMGPRRIMSALIKEYGGISKVGFTEVDCRNYMRNNRQKSIEGEIQL 252
Query: 190 MQELFQKFEENNYVFNYRTVSGS--RTVQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQY 247
+ + ++ +N F Y +V GS ++V ++F+A P+++ F F + D+TY++N+Y
Sbjct: 253 LLDYLRQMNADNPNFFY-SVQGSEDQSVGNVFWADPKAIMDFTHFGDTVTFDTTYRSNRY 311
Query: 248 KMPLFEIVGLTSTEMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKKQMPKVIVTDRDTG 307
++P G+ G AF+ E E +F W + L P I TD D
Sbjct: 312 RLPFAPFTGVNHHGQPILFGCAFIINETEASFVWLF--NTWLAAMSAHPPVSITTDHDAV 369
Query: 308 LMNAVANIFPKSTALVCEFHILKNVRAR 335
+ A+ ++FP + C++HILK + +
Sbjct: 370 IRAAIMHVFPGARHRFCKWHILKKCQEK 397
>At1g52520 F6D8.26
Length = 703
Score = 66.6 bits (161), Expect = 5e-11
Identities = 77/324 (23%), Positives = 136/324 (41%), Gaps = 22/324 (6%)
Query: 18 EFFVTKEKKKYEDREEMINWARGQAIEAGFTLIIDKS-YSGSGRRKPKLVLA-----FER 71
EF ++E ++ N+ A E GF + + S + + K VL F+R
Sbjct: 81 EFDAPAVGMEFESYDDAYNYYNCYASEVGFRVRVKNSWFKRRSKEKYGAVLCCSSQGFKR 140
Query: 72 SGEYKGTKKSKREGTGSRKCGCPFRLRGYFSATKLWSLNIVSGLHNHKMEPKLEGHMLAG 131
+ +K R G CP +R +K W + V+ HNH + KL +
Sbjct: 141 INDVNRVRKETRTG-------CPAMIRMRQVDSKRWRVVEVTLDHNHLLGCKLYKSVKRK 193
Query: 132 RLTAKESKIVGDMTRNLIKPKNILLDLKGRWKDNRTVAKQIYNARHRYKLSI--RGSR*E 189
R S V D + + + ++D N T+ K+ N+ L RG
Sbjct: 194 RKCV--SSPVSD-AKTIKLYRACVVDNGSNVNPNSTLNKKFQNSTGSPDLLNLKRGDSAA 250
Query: 190 MQELFQKFEENNYVFNY-RTVSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQYK 248
+ F + + N F Y V+ ++++F+A S + F V+ +DS+Y + +++
Sbjct: 251 IYNYFCRMQLTNPNFFYLMDVNDEGQLRNVFWADAFSKVSCSYFGDVIFIDSSYISGKFE 310
Query: 249 MPLFEIVGLTSTEMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKKQMPKVIVTDRDTGL 308
+PL G+ T + FL E +++ W L+ L K+ P+ IVTDR L
Sbjct: 311 IPLVTFTGVNHHGKTTLLSCGFLAGETMESYHWLLKV---WLSVMKRSPQTIVTDRCKPL 367
Query: 309 MNAVANIFPKSTALVCEFHILKNV 332
A++ +FP+S HI++ +
Sbjct: 368 EAAISQVFPRSHQRFSLTHIMRKI 391
>At3g22170 far-red impaired response protein, putative
Length = 814
Score = 65.1 bits (157), Expect = 2e-10
Identities = 77/338 (22%), Positives = 147/338 (42%), Gaps = 45/338 (13%)
Query: 107 WSLNIVSGLHNHKMEPKLEGHMLAGRLTAKESKIVGDMTRNLIKPKNILL---DLKGRWK 163
W ++ HNH++ P A ++ + KI M + + K ++ D K ++
Sbjct: 171 WVIHSFVREHNHELLP-------AQAVSEQTRKIYAAMAKQFAEYKTVISLKSDSKSSFE 223
Query: 164 DNRTVAKQIYNARHRYKLSIRGSR*EMQELFQKFEENNYVFNYRTVSGS-RTVQDLFFAH 222
RT++ + + +K+ + + + + N F Y G + V+++F+
Sbjct: 224 KGRTLSVETGD----FKILL--------DFLSRMQSLNSNFFYAVDLGDDQRVKNVFWVD 271
Query: 223 PESVKLFNTFPTVLVMDSTYKTNQYKMPLFEIVGLTSTEMTYNVGFAFLTMEKEDNFTWG 282
+S + +F V+ +D+TY N+YKMPL VG+ +G A ++ E ++W
Sbjct: 272 AKSRHNYGSFCDVVSLDTTYVRNKYKMPLAIFVGVNQHYQYMVLGCALISDESAATYSWL 331
Query: 283 LQESANLLKCKKQMPKVIVTDRDTGLMNAVANIFPKSTALVCEFHILKNVRARCILDCKV 342
++ L Q PKV++T+ D + + V IFP + + +H+L KV
Sbjct: 332 ME--TWLRAIGGQAPKVLITELDVVMNSIVPEIFPNTRHCLFLWHVL----------MKV 379
Query: 343 KDSKGKHVKSSDLVNTIMNAWEVIVSSPSEELYADAVLQFRKVCEQFPKF-LKYVESTI- 400
++ G+ VK D N + + I S +E +A RK + +F LK + I
Sbjct: 380 SENLGQVVKQHD--NFMPKFEKCIYKSGKDEDFA------RKWYKNLARFGLKDDQWMIS 431
Query: 401 LEPLKKKIVKAWTNRVMHFGCTTTNGVESAHGTLKGYL 438
L +KK + V+ G +T+ +S + Y+
Sbjct: 432 LYEDRKKWAPTYMTDVLLAGMSTSQRADSINAFFDKYM 469
>At5g28530 far-red impaired response protein (FAR1) - like
Length = 700
Score = 64.7 bits (156), Expect = 2e-10
Identities = 69/290 (23%), Positives = 125/290 (42%), Gaps = 34/290 (11%)
Query: 71 RSGEYKGTKKSKREGTGSRK---CGCPFRL---RGYFSATKLWSLNIVSGLHNHKMEPKL 124
RSG + KK+ E RK CGC +L + W ++ S +HNH++
Sbjct: 119 RSGFNQPRKKANVEHPRERKSVRCGCDGKLYLTKEVVDGVSHWYVSQFSNVHNHELLEDD 178
Query: 125 EGHMLAGRLTAKESKIVGDMTRNLIKPKN--------ILLDLKGRWKDNRT--VAKQIYN 174
+ +L ++S D R L+ K LL+L+ + + K + N
Sbjct: 179 QVRLLPAYRKIQQS----DQERILLLSKAGFPVNRIVKLLELEKGVVSGQLPFIEKDVRN 234
Query: 175 ARHRYKLSIR-----------GSR*EMQELFQKFEENNYVFNYRTVSG-SRTVQDLFFAH 222
K S++ E+ E + E + F Y S ++ V+++ +A+
Sbjct: 235 FVRACKKSVQENDAFMTEKRESDTLELLECCKGLAERDMDFVYDCTSDENQKVENIAWAY 294
Query: 223 PESVKLFNTFPTVLVMDSTYKTNQYKMPLFEIVGLTSTEMTYNVGFAFLTMEKEDNFTWG 282
+SV+ ++ F V+V D++Y++ Y + L G+ + +G L E +FTW
Sbjct: 295 GDSVRGYSLFGDVVVFDTSYRSVPYGLLLGVFFGIDNNGKAMLLGCVLLQDESCRSFTWA 354
Query: 283 LQESANLLKCKKQMPKVIVTDRDTGLMNAVANIFPKSTALVCEFHILKNV 332
LQ ++ + P+ I+TD DTGL +A+ P + +V HI+ +
Sbjct: 355 LQTFVRFMRGRH--PQTILTDIDTGLKDAIGREMPNTNHVVFMSHIVSKL 402
>At4g15090 unknown protein
Length = 768
Score = 63.9 bits (154), Expect = 4e-10
Identities = 66/279 (23%), Positives = 120/279 (42%), Gaps = 31/279 (11%)
Query: 75 YKGTKKSKREGTGSR-----KCGCPFRLRGYFSATKLWSLNIVSGLHNHKMEPKLEGHML 129
Y T +S+ G+ SR K C + W ++ HNH++ P L H
Sbjct: 43 YGVTPESESSGSSSRRSTVKKTDCKASMHVKRRPDGKWIIHEFVKDHNHELLPALAYHFR 102
Query: 130 AGR---------------LTAKESKIVGDMTRNLIKPKNILLDLKGRWKDNRTVAKQIYN 174
R ++ + K+ +M+R KNI L+ V+ Q+
Sbjct: 103 IQRNVKLAEKNNIDILHAVSERTKKMYVEMSRQSGGYKNIGSLLQ------TDVSSQV-- 154
Query: 175 ARHRYKLSIRGSR*EMQELFQKFEENNYVFNYRT-VSGSRTVQDLFFAHPESVKLFNTFP 233
+ RY G + E F++ ++ N F Y ++ + +++LF+A +S + +F
Sbjct: 155 DKGRYLALEEGDSQVLLEYFKRIKKENPKFFYAIDLNEDQRLRNLFWADAKSRDDYLSFN 214
Query: 234 TVLVMDSTYKTNQYKMPLFEIVGLTSTEMTYNVGFAFLTMEKEDNFTWGLQESANLLKCK 293
V+ D+TY K+PL +G+ +G A + E + F W ++ L
Sbjct: 215 DVVSFDTTYVKFNDKLPLALFIGVNHHSQPMLLGCALVADESMETFVWLIK--TWLRAMG 272
Query: 294 KQMPKVIVTDRDTGLMNAVANIFPKSTALVCEFHILKNV 332
+ PKVI+TD+D LM+AV+ + P + +H+L+ +
Sbjct: 273 GRAPKVILTDQDKFLMSAVSELLPNTRHCFALWHVLEKI 311
>At1g80010 hypothetical protein
Length = 696
Score = 60.5 bits (145), Expect = 4e-09
Identities = 66/326 (20%), Positives = 141/326 (43%), Gaps = 29/326 (8%)
Query: 27 KYEDREEMINWARGQAIEAGFTLIIDKSYSG-SGRRKPKLVLAFERSGEYKGTKK--SKR 83
++E ++ ++ A E GF + + S++ + + K VL G +K K S+R
Sbjct: 71 EFESYDDAYSFYNSYARELGFAIRVKSSWTKRNSKEKRGAVLCCNCQG-FKLLKDAHSRR 129
Query: 84 EGTGSRKCGCPFRLRGYFSATKLWSLNIVSGLHNHKMEPKLEGHMLAGRLTAKESKIVGD 143
+ T + GC +R W ++ V HNH +P+ + + + K S
Sbjct: 130 KET---RTGCQAMIRLRLIHFDRWKVDQVKLDHNHSFDPQRAHNSKSHK---KSSSSASP 183
Query: 144 MTRNLIKP------------KNILLD----LKGRWKDNRTVAKQIYNARHRYKLSIRGSR 187
T+ +P + + LD L T + + + +L +RG
Sbjct: 184 ATKTNPEPPPHVQVRTIKLYRTLALDTPPALGTSLSSGETSDLSLDHFQSSRRLELRGGF 243
Query: 188 *EMQELFQKFEENNYVFNY-RTVSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQ 246
+Q+ F + + ++ F Y ++ +++++F+ + ++ F VL+ D+T +N
Sbjct: 244 RALQDFFFQIQLSSPNFLYLMDLADDGSLRNVFWIDARARAAYSHFGDVLLFDTTCLSNA 303
Query: 247 YKMPLFEIVGLTSTEMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKKQMPKVIVTDRDT 306
Y++PL VG+ T +G L + + + W + A L + P++ +T++
Sbjct: 304 YELPLVAFVGINHHGDTILLGCGLLADQSFETYVWLFR--AWLTCMLGRPPQIFITEQCK 361
Query: 307 GLMNAVANIFPKSTALVCEFHILKNV 332
+ AV+ +FP++ + H+L N+
Sbjct: 362 AMRTAVSEVFPRAHHRLSLTHVLHNI 387
>At1g44840 hypothetical protein
Length = 926
Score = 60.5 bits (145), Expect = 4e-09
Identities = 86/429 (20%), Positives = 165/429 (38%), Gaps = 89/429 (20%)
Query: 218 LFFAHPESVKLFNTFPTVLVMDSTYKTNQYKMPLFEIVGLTSTEMTYNVGFAFLTMEKED 277
+F A S+ F +LV+D T+ +YK L G + Y +GFA + E ++
Sbjct: 557 MFLAFGASIAGFRHLRRILVVDGTHLKGKYKGVLLTSSGQDANFQVYPLGFAVVDSENDE 616
Query: 278 NFTWGLQESANLLKCKKQMPKVIVTDRDTGLMNAVANIFPKSTALVCEFHILKNVRARCI 337
++TW + ++ K + I++DR + ++ AV +FP++ C H+ +N++ +
Sbjct: 617 SWTWFFTKLERIIADSKTL--TILSDRHSSILVAVKRVFPQANHGACIIHLCRNIQTK-- 672
Query: 338 LDCKVKDSKGKHVKSSDLVNTIMNAWEVIVSSPSEELYADAVLQFRKVCEQFPKFLKYVE 397
K+ L + NA + +E Y Q + K+L +
Sbjct: 673 ------------YKNKALTQLVKNAGYAFTGTKFKEFYG----QIETTNQNCGKYLHDIG 716
Query: 398 STILEPLKKKIVKAWTN---RVMHFGCTTTNGVESAHGTLKGYLEDSKGDLAKGWKAINA 454
+ WT R F T+N E+ + L + + + + I +
Sbjct: 717 -----------MANWTRHYFRGQRFNLMTSNIAETLNKALN---KGRSSHIVELIRFIRS 762
Query: 455 MLTVQFTEIQATFGRNSTVLEHKFKGNKMYKSLEYNVSRTAMNYIYKQASRVKDCGSDSK 514
MLT F R L+HK + ++ +++T + ++
Sbjct: 763 MLTRWFN------ARRKKSLKHK---GPVPPEVDKQITKTMLT-------------TNGS 800
Query: 515 KCGCVIRWTHAL---------------PCTCLIYKKLNL---GSCIRLDEINIHWKKLCF 556
K G + W++ + CTC Y KL + + + + I +K L
Sbjct: 801 KVGRITNWSYEINGMLGGRNVVDLEKKQCTCKRYDKLKIPCSHALVAANSFKISYKAL-- 858
Query: 557 EVDDDVEESNRHVSIQNELNAIQERFNKADYNMKLIIKE-QMREMTFPHT--TSLTPPIQ 613
VDD + H + + A+ F +A + I++E + R M P+T S P +
Sbjct: 859 -VDDCFKP---HAWVSSYPGAV---FPEAIGRDEKILEELEHRSMLPPYTRRPSGRPKVA 911
Query: 614 KLKTKGAQK 622
++ + G K
Sbjct: 912 RIPSTGEYK 920
>At3g06250 unknown protein
Length = 764
Score = 58.5 bits (140), Expect = 1e-08
Identities = 54/231 (23%), Positives = 94/231 (40%), Gaps = 30/231 (12%)
Query: 91 CGCPFRLRGYFSATKLWSLNIVSGLHNHKMEPKLEGHMLAGRLTAKESKIVGDMTRNLIK 150
CG R++ S W ++ ++ HNH +EP G+ A KI D+T L
Sbjct: 251 CGAYMRIKRQDSGG--WIVDRLNKDHNHDLEP--------GKKNAGMKKITDDVTGGLDS 300
Query: 151 PKNILL-DLKGRWKDNR--TVAKQIYNARHRYKLSIRGSR*EMQELFQKFEENNYVFNYR 207
I L DL R T+ K+ Y + + FQ + + F Y
Sbjct: 301 VDLIELNDLSNHISSTRENTIGKEWYPV--------------LLDYFQSKQAEDMGFFYA 346
Query: 208 T-VSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQYKMPLFEIVGLTSTEMTYNV 266
+ + + +F+A S + F +V D++Y+ Y +P +G +
Sbjct: 347 IELDSNGSCMSIFWADSRSRFACSQFGDAVVFDTSYRKGDYSVPFATFIGFNHHRQPVLL 406
Query: 267 GFAFLTMEKEDNFTWGLQESANLLKCKKQMPKVIVTDRDTGLMNAVANIFP 317
G A + E ++ F+W Q + ++ P+ +V D+D + AVA +FP
Sbjct: 407 GGALVADESKEAFSWLFQTWLRAMSGRR--PRSMVADQDLPIQQAVAQVFP 455
>At1g76320 putative phytochrome A signaling protein
Length = 670
Score = 57.0 bits (136), Expect = 4e-08
Identities = 33/140 (23%), Positives = 71/140 (50%), Gaps = 2/140 (1%)
Query: 190 MQELFQKFEENNYVFNYRTVSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQYKM 249
++ L + EEN F S ++++F+ + ++ + +F V+ +++Y ++YK+
Sbjct: 167 LEFLMRMQEENPKFFFAVDFSEDHLLRNVFWVDAKGIEDYKSFSDVVSFETSYFVSKYKV 226
Query: 250 PLFEIVGLTSTEMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKKQMPKVIVTDRDTGLM 309
PL VG+ +G L + + W +Q + L+ Q PKV++TD++ +
Sbjct: 227 PLVLFVGVNHHVQPVLLGCGLLADDTVYTYVWLMQ--SWLVAMGGQKPKVMLTDQNNAIK 284
Query: 310 NAVANIFPKSTALVCEFHIL 329
A+A + P++ C +H+L
Sbjct: 285 AAIAAVLPETRHCYCLWHVL 304
>At1g35060 hypothetical protein
Length = 873
Score = 55.1 bits (131), Expect = 2e-07
Identities = 31/118 (26%), Positives = 56/118 (47%), Gaps = 2/118 (1%)
Query: 218 LFFAHPESVKLFNTFPTVLVMDSTYKTNQYKMPLFEIVGLTSTEMTYNVGFAFLTMEKED 277
+F A S++ F VLV+D T+ +YK L G + Y + FA + E +D
Sbjct: 508 MFLAFGASIQGFKHLRRVLVVDGTHLKGKYKGVLLTASGQDANFQVYPLAFAVVDSENDD 567
Query: 278 NFTWGLQESANLLKCKKQMPKVIVTDRDTGLMNAVANIFPKSTALVCEFHILKNVRAR 335
+TW + ++ + I++DR + V +FP++ C H+ +N++AR
Sbjct: 568 AWTWFFTKLERIIADNNTL--TILSDRHESIKVGVKKVFPQAHHGACIIHLCRNIQAR 623
>At2g27110 Mutator-like transposase
Length = 851
Score = 52.0 bits (123), Expect = 1e-06
Identities = 31/140 (22%), Positives = 63/140 (44%), Gaps = 3/140 (2%)
Query: 192 ELFQKFEENNYVFNYRT-VSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQYKMP 250
E F++ + N F Y + + ++F+A S + F + +D+ Y+ NQ+++P
Sbjct: 202 EYFKRMQAENPGFFYAVQLDEDNQMSNVFWADSRSRVAYTHFGDTVTLDTRYRCNQFRVP 261
Query: 251 LFEIVGLTSTEMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKKQMPKVIVTDRDTGLMN 310
G+ G A + E + +F W + L + Q P +VTD+D +
Sbjct: 262 FAPFTGVNHHGQAILFGCALILDESDTSFIWLFK--TFLTAMRDQPPVSLVTDQDRAIQI 319
Query: 311 AVANIFPKSTALVCEFHILK 330
A +FP + + ++ +L+
Sbjct: 320 AAGQVFPGARHCINKWDVLR 339
>At3g43050 putative protein
Length = 608
Score = 51.6 bits (122), Expect = 2e-06
Identities = 31/137 (22%), Positives = 64/137 (46%), Gaps = 2/137 (1%)
Query: 200 NNYVFNYRTVSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQYKMPLFEIVGLTS 259
N ++ + + Q LFFA ++ FN V+++D+T+ L +
Sbjct: 299 NEGTISFVELDEKKNFQYLFFALGACIEGFNAMRKVIIVDATHLKIAQGGVLVVALAQDL 358
Query: 260 TEMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKKQMPKVIVTDRDTGLMNAVANIFPKS 319
Y + F L EK+ +++W ++ +++ ++ V V+DR+ LM + ++P S
Sbjct: 359 EHHHYPIDFGVLDGEKDVSWSWFFEKLKSVIPDSSEL--VFVSDRNQSLMKSQRELYPLS 416
Query: 320 TALVCEFHILKNVRARC 336
C +H+ +NV+ C
Sbjct: 417 QHGCCIYHLCQNVKGAC 433
>At2g23500 Mutator-like transposase
Length = 784
Score = 51.6 bits (122), Expect = 2e-06
Identities = 82/387 (21%), Positives = 145/387 (37%), Gaps = 52/387 (13%)
Query: 68 AFERSGEYKGTKKSK-REGTGSRKCGCPFRLRGY-FSATKLWSLNIVSGLHN-------- 117
AF S Y K K R K C +RLR S T ++S+ + +H+
Sbjct: 219 AFANSFGYVIKKSDKERYVLKCAKESCSWRLRASNISTTDIFSIRRYNKMHSCTRLSKGS 278
Query: 118 -----HKMEPKLEGHMLAGRLTAKESKIVGDMTRNLIKPKNILLDLKGRWKDNRTVAKQI 172
K P+L +L + V + L++ K L +K + T +
Sbjct: 279 SRLRKRKGNPQLVAALLHDHFPGQLETPVPSIIMELVQTK---LGVKVSYS---TALRGK 332
Query: 173 YNARHRYKLSIRGSR*EMQ-ELFQKFEENNYVFNYRTVSGSRTVQDLFFAHPESVKLFNT 231
Y+A + K S S ++ L+ + N+ Y + + Q +F A S++ F
Sbjct: 333 YHAIYDLKGSPEESYKDINCYLYMLKKVNDGTVTYLKLDENDKFQYVFVALGASIEGFRV 392
Query: 232 FPTVLVMDSTYKTNQYKMPLFEIVGLTSTEMTYNVGFAFLTMEKEDNFTWGLQESANLLK 291
VL++D+T+ N Y L Y + FA L E + ++ W ++ ++
Sbjct: 393 MRKVLIVDATHLKNGYGGVLVFASAQDPNRHHYIIAFAVLDGENDASWEWFFEKLKTVVP 452
Query: 292 CKKQMPKVIVTDRDTGLMNAVANIFPKSTALVCEFHILKNVRARCILDCKVKDSKGKHVK 351
++ V +TDR+ L+ A+ N++ + C +H+ +NV+ H
Sbjct: 453 DTSEL--VFMTDRNASLIKAIRNVYTAAHHGYCIWHLSQNVKGH-----------ATHTN 499
Query: 352 SSDLVNTIMNAWEVIVSSPSEELYADAVLQFRKVCEQFPKFLKYVESTILEPLKKKIVKA 411
L V V + Y L ++PK KY+E T ++ +
Sbjct: 500 RDVLAWKFQELSRVYVVADFNRAYDGFKL-------RYPKATKYLEDTTVK-------EK 545
Query: 412 WTNRVM---HFGCTTTNGVESAHGTLK 435
W + T+N VES + K
Sbjct: 546 WARCCFPGERYNLDTSNCVESLNNVFK 572
>At1g10240 unknown protein
Length = 680
Score = 51.6 bits (122), Expect = 2e-06
Identities = 55/249 (22%), Positives = 108/249 (43%), Gaps = 15/249 (6%)
Query: 88 SRKCGCPFRLRGYFSATKL----WSLNIVSGLHNHKM-EPKLEGHMLAGRLTAKESK--- 139
S +CGC LR T+L W + + HNH++ EP + A R + K
Sbjct: 119 SSRCGCQAYLR-ISKLTELGSTEWRVTGFANHHNHELLEPNQVRFLPAYRSISDADKSRI 177
Query: 140 IVGDMTRNLIKPKNILLDLKGRWKDNRT--VAKQIYNARHRYK-LSIRGSR*EMQELFQK 196
++ T ++ LL+L+ + K + N +K L + + Q
Sbjct: 178 LMFSKTGISVQQMMRLLELEKCVEPGFLPFTEKDVRNLLQSFKKLDPEDENIDFLRMCQS 237
Query: 197 FEENNYVFNYR-TVSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQYKMPLFEIV 255
+E + F + T+ + ++++ +++ S++ + F +V D+T++ + +MPL V
Sbjct: 238 IKEKDPNFKFEFTLDANDKLENIAWSYASSIQSYELFGDAVVFDTTHRLSAVEMPLGIWV 297
Query: 256 GLTSTEMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKKQMPKVIVTDRDTGLMNAVANI 315
G+ + + G L E +++W LQ + K P+ I+TD + L A+A
Sbjct: 298 GVNNYGVPCFFGCVLLRDENLRSWSWALQAFTGFMNGK--APQTILTDHNMCLKEAIAGE 355
Query: 316 FPKSTALVC 324
P + +C
Sbjct: 356 MPATKHALC 364
>At4g09380 putative protein
Length = 960
Score = 51.2 bits (121), Expect = 2e-06
Identities = 34/148 (22%), Positives = 65/148 (42%), Gaps = 4/148 (2%)
Query: 193 LFQKFEENNYVFNYRTVSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQYKMPLF 252
L+ + N Y + G + + LF A ++ F V+V+D+T+ Y L
Sbjct: 368 LYMLEKVNPGTVTYVELEGEKKFKYLFIALGACIEGFRAMRKVIVVDATHLKTVYGGMLV 427
Query: 253 EIVGLTSTEMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKKQMPKVI-VTDRDTGLMNA 311
Y + F + EK+ ++ W L+ NL +P ++ ++DR + A
Sbjct: 428 IATAQDPNHHHYPLAFGIIDSEKDVSWIWFLE---NLKTVYSDVPGLVFISDRHQSIKKA 484
Query: 312 VANIFPKSTALVCEFHILKNVRARCILD 339
V ++P + C +H+ +N+R R +D
Sbjct: 485 VKTVYPNALHAACIWHLCQNMRDRVKID 512
>At3g32060 hypothetical protein
Length = 487
Score = 50.8 bits (120), Expect = 3e-06
Identities = 58/274 (21%), Positives = 108/274 (39%), Gaps = 33/274 (12%)
Query: 167 TVAKQIYNARHRYKLSIRGSR*EMQE-----LFQKFEENNYVFNYRTVSGSRTVQDLFFA 221
TV + Y+A + ++GSR E + L+ + N+ Y + + Q +F A
Sbjct: 197 TVLRGKYHAIY----DLKGSREESYKDINCYLYMLKKVNDGTVTYLKLDENDKFQYVFVA 252
Query: 222 HPESVKLFNTFPTVLVMDSTYKTNQYKMPLFEIVGLTSTEMTYNVGFAFLTMEKEDNFTW 281
S++ F VL++D+T+ N Y L Y + FA L E + ++ W
Sbjct: 253 LGASIEGFRVMRKVLIVDATHLKNGYGGVLVFASAQDPNRHHYIIAFAVLDGENDASWEW 312
Query: 282 GLQESANLLKCKKQMPKVIVTDRDTGLMNAVANIFPKSTALVCEFHILKNVRARCILDCK 341
++ ++ ++ V +TDR+ L+ A+ N++ + C +H+ +NV+
Sbjct: 313 FFEKLKTVVPDTSEL--VFMTDRNASLIKAIRNVYTAAHHGYCIWHLSQNVKGH------ 364
Query: 342 VKDSKGKHVKSSDLVNTIMNAWEVIVSSPSEELYADAVLQFRKVCEQFPKFLKYVESTIL 401
H L V V + Y L ++PK KY+E T +
Sbjct: 365 -----ATHTNRDVLAWKFQELSRVYVVADFNRAYDGFKL-------RYPKATKYLEDTTV 412
Query: 402 EPLKKKIVKAWTNRVMHFGCTTTNGVESAHGTLK 435
+ + A + T+N +ES + K
Sbjct: 413 KEKWARCCFAGE----RYNLDTSNCMESLNNVFK 442
>At2g32250 Mutator-like transposase
Length = 684
Score = 50.4 bits (119), Expect = 4e-06
Identities = 62/311 (19%), Positives = 116/311 (36%), Gaps = 46/311 (14%)
Query: 28 YEDREEMINWARGQAIEAGFTLIIDKSYSGSGRRKPKLVLAFERSGEYKGTKKSKREGTG 87
+E +E + R A GF + I S +R K + + GTK+ K
Sbjct: 44 FESKEAAYYFYREYARSVGFGITIKASRRS--KRSGKFIDVKIACSRF-GTKREKATAIN 100
Query: 88 SRKC---GCPFRLRGYFSATKLWSLNIVSGLHNHKMEPKLEGHMLAGRLTAKESKIVGDM 144
R C GC L + W + HNH+
Sbjct: 101 PRSCPKTGCKAGLHMKRKEDEKWVIYNFVKEHNHE------------------------- 135
Query: 145 TRNLIKPKNILLDLKGRWKDNRTVA--KQIYNARHRYKLSIRGSR*EMQELFQKFEENNY 202
I P + + ++G+ K +A K + A L + + E F + ++
Sbjct: 136 ----ICPDDFYVSVRGKNKPAGALAIKKGLQLALEEEDLKL------LLEHFMEMQDKQP 185
Query: 203 VFNYRT-VSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQYKMPLFEIVGLTSTE 261
F Y + V+++F+ ++ + +F V++ D+ Y N Y++P +G++
Sbjct: 186 GFFYAVDFDSDKRVRNVFWLDAKAKHDYCSFSDVVLFDTFYVRNGYRIPFAPFIGVSHHR 245
Query: 262 MTYNVGFAFLTMEKEDNFTWGLQESANLLKCKKQMPKVIVTDRDTGLMNAVANIFPKSTA 321
+G A + E ++W + L Q P V++TD+D L + V +FP
Sbjct: 246 QYVLLGCALIGEVSESTYSWLFR--TWLKAVGGQAPGVMITDQDKLLSDIVVEVFPDVRH 303
Query: 322 LVCEFHILKNV 332
+ C + +L +
Sbjct: 304 IFCLWSVLSKI 314
>At1g36090 hypothetical protein
Length = 645
Score = 50.4 bits (119), Expect = 4e-06
Identities = 33/148 (22%), Positives = 66/148 (44%), Gaps = 4/148 (2%)
Query: 193 LFQKFEENNYVFNYRTVSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQYKMPLF 252
L+ + N Y + G + + LF A ++ F V+V+D+T+ Y L
Sbjct: 328 LYMLEKVNPGTVTYVELEGEKKFKYLFIALGACIEGFRAMRKVIVVDATHLKTVYGGMLV 387
Query: 253 EIVGLTSTEMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKKQMPKVI-VTDRDTGLMNA 311
Y + F + EK+ ++ W L++ L +P+++ ++DR + A
Sbjct: 388 IATAHDPNHHHYPLAFGIIDSEKDVSWIWFLEK---LKTVYSDVPRLVFISDRHQSIKKA 444
Query: 312 VANIFPKSTALVCEFHILKNVRARCILD 339
V ++P + C +H+ +N+R R +D
Sbjct: 445 VKTVYPNALHAACIWHLCQNMRDRVKID 472
>At4g28970 putative protein
Length = 914
Score = 50.1 bits (118), Expect = 5e-06
Identities = 32/147 (21%), Positives = 63/147 (42%), Gaps = 2/147 (1%)
Query: 193 LFQKFEENNYVFNYRTVSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQYKMPLF 252
L+ + N Y + G + + LF A ++ F V+V+D+T+ Y L
Sbjct: 368 LYMLEKVNPGTVTYVELEGEKKFKYLFIALGACIEGFRAMRKVIVVDATHLKTVYGGMLV 427
Query: 253 EIVGLTSTEMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKKQMPKVIVTDRDTGLMNAV 312
Y + F + EK+ ++ W L++ + + V ++DR + AV
Sbjct: 428 IATAQDPNHHHYPLAFGIIDSEKDVSWIWFLEKLKTVYSDVPGL--VFISDRHQSIKKAV 485
Query: 313 ANIFPKSTALVCEFHILKNVRARCILD 339
++P + C +H+ +N+R R +D
Sbjct: 486 KTVYPNALHAACIWHLCQNMRDRVTID 512
>At1g33460 mutator transposase MUDRA, putative
Length = 826
Score = 49.3 bits (116), Expect = 9e-06
Identities = 26/98 (26%), Positives = 49/98 (49%)
Query: 238 MDSTYKTNQYKMPLFEIVGLTSTEMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKKQMP 297
+D T+ + K L + + Y V +A + E +N+ W L + + L+ K
Sbjct: 374 IDGTFLKHAVKGCLLTAIAHDANNQIYPVAWATVQFENAENWLWFLNQLKHDLELKDGSG 433
Query: 298 KVIVTDRDTGLMNAVANIFPKSTALVCEFHILKNVRAR 335
V+++DR G+++AV N P + C HI++N++ R
Sbjct: 434 YVVISDRCKGIISAVKNALPNAEHRPCVKHIVENLKKR 471
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.136 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,616,289
Number of Sequences: 26719
Number of extensions: 878704
Number of successful extensions: 2383
Number of sequences better than 10.0: 92
Number of HSP's better than 10.0 without gapping: 40
Number of HSP's successfully gapped in prelim test: 52
Number of HSP's that attempted gapping in prelim test: 2291
Number of HSP's gapped (non-prelim): 113
length of query: 840
length of database: 11,318,596
effective HSP length: 108
effective length of query: 732
effective length of database: 8,432,944
effective search space: 6172915008
effective search space used: 6172915008
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 64 (29.3 bits)
Medicago: description of AC138063.5


