

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138012.9 - phase: 0
(672 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
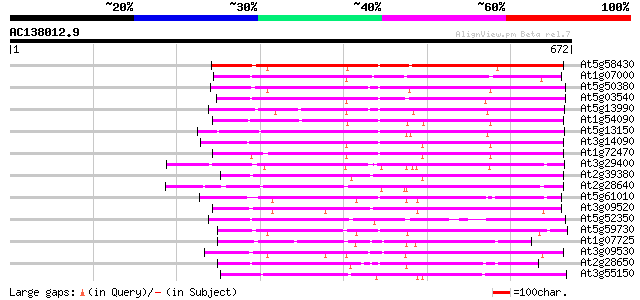
Score E
Sequences producing significant alignments: (bits) Value
At5g58430 leucine zipper-containing protein 302 3e-82
At1g07000 putative protein 233 3e-61
At5g50380 putative protein 197 2e-50
At5g03540 unknown protein (At5g03540) 186 5e-47
At5g13990 leucine zipper protein-like 179 4e-45
At1g54090 unknown protein 177 2e-44
At5g13150 putative protein 175 8e-44
At3g14090 unknown protein 173 3e-43
At1g72470 unknown protein 169 5e-42
At3g29400 unknown protein (At3g29400) 142 5e-34
At2g39380 hypothetical protein 142 6e-34
At2g28640 unknown protein 142 8e-34
At5g61010 putative protein 134 2e-31
At3g09520 hypothetical protein 132 5e-31
At5g52350 putative protein 130 2e-30
At5g59730 unknown protein 129 5e-30
At1g07725 putative protein 128 1e-29
At3g09530 hypothetical protein 127 2e-29
At2g28650 unknown protein 126 3e-29
At3g55150 unknown protein 123 4e-28
>At5g58430 leucine zipper-containing protein
Length = 624
Score = 302 bits (774), Expect = 3e-82
Identities = 174/434 (40%), Positives = 264/434 (60%), Gaps = 20/434 (4%)
Query: 242 DFEMDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFVLPEAKIN 301
D +D LPS +++LHE+ K ML AG+ K CS VY S R+ L++ + L K++
Sbjct: 193 DLIIDALPSATINDLHEMAKRMLGAGFGKACSHVYSSCRREFLEESMSR----LGLQKLS 248
Query: 302 TERERE---RYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQEA 358
E + + L+ RW+ A+++A +LFP E++ CD VF GFSSA F+E+C+ +
Sbjct: 249 IEEVHKMPWQELEDEIDRWIKAANVALRILFPSERRLCDRVFFGFSSAADLSFMEVCRGS 308
Query: 359 TFQLLNFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPNS----LVNEAIAVRNR 414
T QLLNFAD IA GS S RLFK++D+F + +L+P+FES+F + L NEA+ + R
Sbjct: 309 TIQLLNFADAIAIGSRSPERLFKVLDVFETMRDLMPEFESVFSDQFCSVLRNEAVTIWKR 368
Query: 415 LGDASRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILEEY 474
LG+A R +FM++ N I R PA V G H +T VM+Y+ +ACR R+ LEQ+ EE
Sbjct: 369 LGEAIRGIFMELENLIRRDPAKAAVPG--GGLHPITRYVMNYLRAACRSRQTLEQVFEES 426
Query: 475 PEVHNEVEASSFFLKQMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKFSR 534
V + + S+ QM IM +L+ L VKS+ KD AL ++F++NN +I K
Sbjct: 427 NGVPS--KDSTLLTVQMSWIMELLESNLEVKSKVYKDPALCYVFLMNNGRYIVQKVKDGD 484
Query: 535 LETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDNNESITKEL---LKEKIHLFN 591
L + G+DW + + K++Q Y+RS+W++++ LK+DN + L +KEK+ FN
Sbjct: 485 LGLLLGDDWIRKHNVKVKQYHMNYQRSSWNKMLGLLKVDNTAAGMNGLGKTMKEKLKQFN 544
Query: 592 NRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLH--GILGNQAYKYIKYG 649
+F+ IC+V S W ++ QL+ E+ S+ +L+PAYG F+GR G +G A KYIKYG
Sbjct: 545 IQFDEICKVHSTWVVFDEQLKEELKISLARLLVPAYGSFIGRFQNLGDIGKNADKYIKYG 604
Query: 650 MIEIQDLLNHLFLG 663
+ +I+ +N LF G
Sbjct: 605 VEDIEARINELFKG 618
>At1g07000 putative protein
Length = 599
Score = 233 bits (594), Expect = 3e-61
Identities = 147/427 (34%), Positives = 236/427 (54%), Gaps = 20/427 (4%)
Query: 245 MDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFVLPEAKINTER 304
++ L S + +L+ I M+ G+ KECS VY S R+ L++ L +++ + + +
Sbjct: 178 IEALQSSVIGDLNAIAVRMVAGGFAKECSRVYSSRRREFLEESL-SRLHLRGLSMEEVQE 236
Query: 305 ERERYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGF--SSATSHCFIEICQEATFQL 362
+ L+ RW+ A + V FP E+ CD VFS SS T F+E+C+ T QL
Sbjct: 237 SPWQDLEDEIDRWIKAVTLIFHVFFPSERLLCDRVFSDLPVSSVTDLSFMEVCRGTTTQL 296
Query: 363 LNFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFP----NSLVNEAIAVRNRLGDA 418
LNFAD IA GS RLFK+VD++ + +L+PK E+LF + L +EA+A+ RLG+A
Sbjct: 297 LNFADAIALGSRLPERLFKVVDLYEAMQDLIPKMETLFSDRYCSPLRHEALAIHKRLGEA 356
Query: 419 SRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILEEYPEVH 478
R +FM++ N I R P + G H +T VM+Y+ +AC+ R+ LEQIL+ +
Sbjct: 357 IRGIFMELENLIRRDP--PKTAFPGGGIHPITRYVMNYLRAACKSRQSLEQILD---QTG 411
Query: 479 NEVEASSFFLK-QMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKFSRLET 537
NE + + L Q+ ++ +L+ L K +D +L +FM+NN +I K + L
Sbjct: 412 NETGSDTRPLSVQIIWVLELLESNLEGKKRTYRDPSLCFLFMMNNDKYILDKAKDNELGL 471
Query: 538 IFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDNNESITKELLKEKIHLFNNRFEAI 597
+ G DW + AK++Q Y+RS+W++V+ L+ D L E + LF ++F+ +
Sbjct: 472 VLGEDWIVKHAAKLRQYHSNYRRSSWNQVVGLLRTDG----PYPKLVENLRLFKSQFDEV 527
Query: 598 CRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLH---GILGNQAYKYIKYGMIEIQ 654
C+VQS W + QLR E+ SSV I+ PAY F+ RL I G + +I Y + +++
Sbjct: 528 CKVQSQWVVSDGQLREELRSSVAGIVSPAYSNFIRRLKESPEINGRRGEPFIPYTVEDVE 587
Query: 655 DLLNHLF 661
++ LF
Sbjct: 588 FIIKRLF 594
>At5g50380 putative protein
Length = 637
Score = 197 bits (501), Expect = 2e-50
Identities = 139/450 (30%), Positives = 226/450 (49%), Gaps = 30/450 (6%)
Query: 241 CDFEMDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFVLPEAKI 300
CD +D + + +L EI + M+ AGYEKEC VY S R+ L L+ +L K+
Sbjct: 185 CDLWVDLINPTAVEDLKEIAERMIRAGYEKECVQVYSSVRRDALDDCLM----ILGVEKL 240
Query: 301 NTERERE---RYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQE 357
+ E ++ + +D ++W+ A I VL E+K CD +FS S+ CF E +
Sbjct: 241 SIEEVQKIDWKSMDEKMKKWIQAVKITVRVLLVGEKKICDEIFSSSESSKEVCFNETTKS 300
Query: 358 ATFQLLNFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPNSLV-NEAIAVRNRLG 416
QLLNF + +A G S +LF+++D++ L N++ E + + V NE V LG
Sbjct: 301 CVMQLLNFGEAVAIGRRSSEKLFRILDMYDALANVLQTLEVMVTDCFVCNETKGVLEALG 360
Query: 417 DASRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILEEYPE 476
DA+R F++ N + R +K+ ++ G+ H M VM+Y+ L +LE
Sbjct: 361 DAARGTFVEFENNV-RNETSKRPTTN-GEVHPMIRYVMNYMKLIVDYAVTLNSLLESNES 418
Query: 477 V----HNEVEASSFFLKQMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKF 532
+ E S K++ ++ L+ L KS+ +D L+H+FM+NN +I K
Sbjct: 419 SGVSGDDSTEEMSPLAKRILGLITSLESNLEDKSKLYEDGGLQHVFMMNNIYYIVQKVKD 478
Query: 533 SRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDN--------------NESI 578
S L + G+DW + + +I+Q Y R++W V+ L+ ++ + +
Sbjct: 479 SELGKLLGDDWVRKRRGQIRQYATGYLRASWSRVLSALRDESMGGSSSGSPSYGQRSNNS 538
Query: 579 TKELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGIL 638
+K LKE+ FN FE + R+Q+AW + QLR E+ S+ ++PAY F GR L
Sbjct: 539 SKMALKERFRGFNASFEELYRLQTAWKVPDPQLREELRISISEKVIPAYRAFFGRNRSQL 598
Query: 639 --GNQAYKYIKYGMIEIQDLLNHLFLGNKM 666
G A KYIKY +++ L LF GN++
Sbjct: 599 EGGRHAGKYIKYTPDDLESYLPDLFEGNQL 628
>At5g03540 unknown protein (At5g03540)
Length = 638
Score = 186 bits (471), Expect = 5e-47
Identities = 122/438 (27%), Positives = 220/438 (49%), Gaps = 25/438 (5%)
Query: 248 LPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFVLPEAKINTERERE 307
+PS L LH++ + M+ AG++++ +Y R +L++ L K+ V +K + +R +
Sbjct: 202 IPSRVLPLLHDLAQQMVQAGHQQQLLQIYRDTRSFVLEESL-KKLGVEKLSKEDVQRMQW 260
Query: 308 RYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQEATFQLLNFAD 367
L+ W+ IA +LF E++ CD +F GF S + CF E+ + LL+F D
Sbjct: 261 EVLEAKIGNWIHFMRIAVKLLFAGERQVCDQIFRGFDSLSDQCFAEVTVSSVSMLLSFGD 320
Query: 368 VIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPN----SLVNEAIAVRNRLGDASRVLF 423
IA S +LF ++D++ + L + E++F + + A + RL ++ F
Sbjct: 321 AIARSKRSPEKLFVLLDMYEIMRELHTEIETIFKGKACLEIRDSATGLTKRLAQTAQETF 380
Query: 424 MKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILEEYPEVHNEVEA 483
+ + V+ G H +T V++YV + L+Q+ E+ N ++
Sbjct: 381 GDFEEAVEKDATKTAVLD--GTVHPLTSYVINYVKFLFDYQTTLKQLFLEF---GNGDDS 435
Query: 484 SSFFLKQMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKFSRLETIFGNDW 543
+S +IM+ LQ L KS+ KD AL H+F++NN ++ + S + + G+DW
Sbjct: 436 NSQLASVTMRIMQALQNNLDGKSKQYKDPALTHLFLMNNIHYMVRSVRRSEAKDLLGDDW 495
Query: 544 FQNNKAKIQQNLDLYKRSAWDEVMD-------------FLKLDNNESITKELLKEKIHLF 590
Q ++ +QQ+ + YKR AW +++ L+ N+ +++ LLKE+ +F
Sbjct: 496 VQRHRRIVQQHANQYKRVAWTKILQSSSAQGLTSSGGGSLEGGNSSGVSRGLLKERFKMF 555
Query: 591 NNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGIL--GNQAYKYIKY 648
N +F+ + + QS W + ++LR + +V +LLPAY F+ R ++ G KYIKY
Sbjct: 556 NMQFDELHQRQSQWTVPDTELRESLRLAVAEVLLPAYRSFLKRFGPLVESGKNPQKYIKY 615
Query: 649 GMIEIQDLLNHLFLGNKM 666
+++ LL LF G M
Sbjct: 616 TAEDLERLLGELFEGKSM 633
>At5g13990 leucine zipper protein-like
Length = 695
Score = 179 bits (454), Expect = 4e-45
Identities = 128/444 (28%), Positives = 216/444 (47%), Gaps = 26/444 (5%)
Query: 239 GKCDFEMDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFVLPEA 298
G + E P + + L +I + M GY EC VY+ R+ +L + L
Sbjct: 247 GDQEIEYPGYPEDVVVVLRKIAEKMKAGGYGWECREVYLVGRRNILMRTLKQDCEF---E 303
Query: 299 KINTERERERYLDTMFQR---WMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEIC 355
K++ + ++ DT+ + W +++ FP E K + +F G + F +
Sbjct: 304 KVSIDEVQKMSWDTLEREIPIWNKTFKDCSSLFFPGELKLAERIFPGDEG---NLFCIVT 360
Query: 356 QEATFQLLNFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFP----NSLVNEAIAV 411
Q L FA+ +A S +LFK++DI+ L + P E LFP + L NE +
Sbjct: 361 HGLAIQFLGFAEAVAMTRRSTEKLFKILDIYETLRDSFPAMEELFPEELRSELRNEVTSA 420
Query: 412 RNRLGDASRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQIL 471
R+RLG+ + +F + + I + ++K V G H +T M+Y+ +C + LEQ+
Sbjct: 421 RSRLGETAIHIFCDLEHSI-KSDSSKTPVPG-GAVHPLTRYTMNYLKYSCEYKDTLEQVF 478
Query: 472 EEYPEVHNEVE-----ASSFFLKQMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHI 526
+ + ++ E E +S F Q+ +IM +L L KS+ KD L IFM+NN +I
Sbjct: 479 KSHSKMEREEEEPVESGNSAFASQLMRIMELLDGNLETKSKQYKDIPLSCIFMMNNGRYI 538
Query: 527 EAMNKFS-RLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLK---LDNNESITKEL 582
K S + + G+ W + ++++ Y+R W +++ FL L +N I K
Sbjct: 539 VQKIKGSAEIHEVMGDTWCRRRSSELRNYHKNYQRETWGKLLGFLGHEGLMHNGKIVKPN 598
Query: 583 LKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGIL--GN 640
LKE+ FN F+ I + Q+ W + QL+ E+ S+ +++PAY F+ R L G
Sbjct: 599 LKERFKSFNATFDEIHKTQTTWVVNDEQLQSELRVSITAVMIPAYRAFMARFGQYLDPGR 658
Query: 641 QAYKYIKYGMIEIQDLLNHLFLGN 664
Q KY+KY +I+DL++ LF GN
Sbjct: 659 QTEKYVKYQPEDIEDLIDQLFEGN 682
>At1g54090 unknown protein
Length = 622
Score = 177 bits (448), Expect = 2e-44
Identities = 122/448 (27%), Positives = 222/448 (49%), Gaps = 33/448 (7%)
Query: 244 EMDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFVLPEAKINTE 303
E+D + E + +L I + M+ AGY +EC VY + RK ++ + ++ ++ + +
Sbjct: 169 EIDLVTPEAVSDLRSIAQRMIGAGYSRECVQVYGNVRKSAMEM-IFKQLGIVKLGIGDVQ 227
Query: 304 RERERYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQEATFQLL 363
R ++ +RW+ A+ + V+F E++ C+ +F G T CF+EI + + QL
Sbjct: 228 RLDWEAVEVKIRRWIRAAKVCVRVVFASEKRLCEQIFEGTMEET--CFMEIVKTSALQLF 285
Query: 364 NFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPNS----LVNEAIAVRNRLGDAS 419
NF + I+ S +LFK++D+ + +L+P E +F +S ++ +A +++RL +A+
Sbjct: 286 NFPEAISISRRSPEKLFKILDLHDAITDLLPDMEEIFDSSSSESILVQATEIQSRLAEAA 345
Query: 420 RVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILEEYP---- 475
R + + N +FR P+ V G H +T VM+Y++ R L ++ P
Sbjct: 346 RGILTEFENAVFREPSVVPVPG--GTIHPLTRYVMNYLNLISDYRETLIDLVMTKPCRGL 403
Query: 476 EVHNEVEASSFFLKQMEQI----------MRMLQRKLIVKSENCKDRALRHIFMLNNRSH 525
+ N+ + ++E I M MLQ L KS + +D L HIF++NN +
Sbjct: 404 KCTNDRNDPDMDISELEGISPLALHMIWTMVMLQFNLEEKSLHYRDEPLSHIFVMNNVHY 463
Query: 526 IEAMNKFS-RLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDN-------NES 577
I K S L + G+ + + +Q Y+R+ W V++ L+ + +
Sbjct: 464 IVQKVKSSPELMELIGDKYLRKLTGIFRQAATKYQRATWVRVLNSLRDEGLHVSGSFSSG 523
Query: 578 ITKELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGI 637
++K L+E+ FN FE + R+QS W + +QLR E+ S+ L+PAY F+GR G
Sbjct: 524 VSKSALRERFKAFNTMFEEVHRIQSTWSVPDTQLREELRISLSEHLIPAYRSFLGRFRGH 583
Query: 638 L--GNQAYKYIKYGMIEIQDLLNHLFLG 663
+ G Y+KY + ++ + F G
Sbjct: 584 IESGRHPENYLKYSVENLETAVLDFFEG 611
>At5g13150 putative protein
Length = 653
Score = 175 bits (443), Expect = 8e-44
Identities = 130/467 (27%), Positives = 220/467 (46%), Gaps = 32/467 (6%)
Query: 225 SLNIIDR-LMEEYNQGKCDFEMDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVL 283
S N DR +++++ + + + D P E + L +I M+ AGYE EC Y R+
Sbjct: 184 STNDSDRCVLQDHEEAEEESFHDFSP-ESISTLKKIAGAMISAGYEAECCMSYEMSRRHA 242
Query: 284 LQKGLLNKIFVLPEAKINTERERERYLDTMFQRWMTASDIATTVLFPIEQKFCDLVF--S 341
++ L F + + +R L+ W++ +TVLFP E C+ VF
Sbjct: 243 FKEELTEVGFEGINVE-DVQRIGWESLEGEIASWISIVRRCSTVLFPGELSLCNAVFPDQ 301
Query: 342 GFSSATSHCFIEICQEATFQLLNFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFP 401
SS F + T + L+F+ + S +LFK +D++ L +L+P E
Sbjct: 302 DHSSVRKRLFTGLVSAVTIRFLDFSGAVVLTKRSSEKLFKFLDMYETLRDLIPAVEQS-D 360
Query: 402 NSLVNEAIAVRNRLGDASRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSAC 461
+ L+ E + RLG+A+ +F ++ I V S G H +T M+Y+ AC
Sbjct: 361 SDLIQEIKLAQTRLGEAAVTIFGELEKSIKSDNGRTPVPS--GAVHPLTRYTMNYLKYAC 418
Query: 462 RKRRKLEQILEEY----------PEVH--------NEVEASSFFLKQMEQIMRMLQRKLI 503
+ L+Q+ + Y PE +E S F +QM ++M +L L
Sbjct: 419 EYKETLDQVFQHYEANQTDNKPEPETKPRQQQREDDEEYKVSAFARQMIRVMELLDANLE 478
Query: 504 VKSENCKDRALRHIFMLNNRSHIEAMNKFS-RLETIFGNDWFQNNKAKIQQNLDLYKRSA 562
+KS +D +LR IF++NN +I K S + + G W + +++Q Y+R
Sbjct: 479 IKSRLYRDPSLRFIFLMNNGRYILQKIKGSIEIRDLMGQSWTRKRSTELRQYHKSYQRET 538
Query: 563 WDEVMDFLK---LDNNESITKELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSV 619
W +V+ + L N ++K +LKE+ +FN F+ I + QS W + Q++ E+ S+
Sbjct: 539 WGKVLQCMNQEGLQVNGKVSKPVLKERFKIFNAMFDEIHKTQSTWIVSDEQMQSELRVSI 598
Query: 620 GNILLPAYGIFVGRL--HGILGNQAYKYIKYGMIEIQDLLNHLFLGN 664
++++PAY F GR H G Q KY+KY +I+ ++ LF GN
Sbjct: 599 SSLVIPAYRSFFGRYKQHLDSGKQTDKYVKYQPEDIESFIDDLFDGN 645
>At3g14090 unknown protein
Length = 623
Score = 173 bits (438), Expect = 3e-43
Identities = 110/460 (23%), Positives = 227/460 (48%), Gaps = 28/460 (6%)
Query: 229 IDRLMEEYNQGKCDFEMDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGL 288
+ R Y EMD + E + +L IV+ M+ AGY +EC VY + RK ++ +
Sbjct: 157 LTRRRSSYRSTSSIREMDLISPEAVSDLRSIVQRMVAAGYSRECIQVYGTVRKSAME-AI 215
Query: 289 LNKIFVLPEAKINTERERERYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATS 348
++ ++ + + +R ++ ++W+ A+ + V+F E++ C+ +F G +A
Sbjct: 216 FKQLGIVKISIGDVQRLEWEVVEGKIRKWIRAAKVCIRVVFSSEKRLCEQLFDGICTAMD 275
Query: 349 H-CFIEICQEATFQLLNFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPN----S 403
CF+E + + +L F + I+ S +LFK++D+ L +++P E++F + +
Sbjct: 276 ETCFMETVKASALRLFTFPEAISISRRSPEKLFKILDLHDALTDMLPDIEAIFDSDSSDA 335
Query: 404 LVNEAIAVRNRLGDASRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRK 463
+ +A+ +++RL +A+R + + N + R P+ V G H +T VM+Y+
Sbjct: 336 IRAQAVEIQSRLAEAARGILSEFENAVLREPSIVPVPG--GTIHPLTRYVMNYIVMISDY 393
Query: 464 RRKLEQILEEYPEVHNEVEASSFFLKQMEQ----------IMRMLQRKLIVKSENCKDRA 513
++ L+ ++ P ++ +++ ++ +L L KS++ +D +
Sbjct: 394 KQTLDDLIMSNPSTGSDPNTPDMDFTELDSKSPLDLHLIWLIVVLHFNLEEKSKHYRDTS 453
Query: 514 LRHIFMLNNRSHI-EAMNKFSRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKL 572
L HIF++NN +I + + + L + G+ + + + Y+R+ W V++ L+
Sbjct: 454 LAHIFIMNNIHYIVQKVKRSPELREMIGDHYLRKLTGIFRHAATNYQRATWVRVLNSLRD 513
Query: 573 DN-------NESITKELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLP 625
+ + +++ L+E+ FN FE + R QS W + +QLR E+ S+ L+P
Sbjct: 514 EGLHVSGSFSSGVSRSALRERFKAFNTMFEEVHRTQSTWSVPDAQLREELRISLSEHLIP 573
Query: 626 AYGIFVGRLHGIL--GNQAYKYIKYGMIEIQDLLNHLFLG 663
AY F+GR G + G Y+KY + +I+ ++ F G
Sbjct: 574 AYRSFLGRFRGHIESGRHPENYLKYSVEDIETIVLDFFEG 613
>At1g72470 unknown protein
Length = 633
Score = 169 bits (428), Expect = 5e-42
Identities = 131/452 (28%), Positives = 218/452 (47%), Gaps = 39/452 (8%)
Query: 244 EMDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGL----LNKIFVLPEAK 299
E++ +P E + +L I + M+ AGY +EC VY S RK + + K+ + +
Sbjct: 177 EIELIPIESVIHLSWIARRMVSAGYLRECIQVYGSVRKSAVDSSFRRLGIEKLSIGDVQR 236
Query: 300 INTERERERYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSH--CFIEICQE 357
+N E L+ +RW+ A+ I V+F E+ C+ VF + H CF+E +
Sbjct: 237 LNWEA-----LEQKIRRWIRAAKICVRVVFASEKLLCEHVFESVGAVNIHEACFMETVKG 291
Query: 358 ATFQLLNFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFP----NSLVNEAIAVRN 413
QL NFA+ I+ S +LFK++D+ L L+P ES+F +S+ +A + +
Sbjct: 292 PAIQLFNFAEAISISRRSPEKLFKILDLHDALIELLPDIESVFDLKSSDSIRVQAAEILS 351
Query: 414 RLGDASRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILEE 473
RL +A+R + + N + R P+ V G H +T VM+Y+S R L ++
Sbjct: 352 RLAEAARGILSEFENAVLREPSRVPVPG--GTIHPLTRYVMNYISLISEYRPTLIDLIMS 409
Query: 474 YPEVH-NEVEASSFFLKQMEQ-----------IMRMLQRKLIVKSENCKDRALRHIFMLN 521
P + + F ++E I+ +LQ L KS+ K+ AL H+F++N
Sbjct: 410 KPSRNATDSNTPDFDFSELENNKGPLALHLIWIIVILQFNLEGKSKYYKNAALSHLFIMN 469
Query: 522 NRSHIEAMNKFS-RLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDN------ 574
N +I K S L + G+ + + K +Q Y+R+AW +V+ L+ +
Sbjct: 470 NAHYIVQKIKGSPELREMIGDLYLRKLTGKFRQAATYYQRAAWVKVLYCLRDEGLHTKGS 529
Query: 575 -NESITKELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGR 633
+ +++ L+E+ FN FE + RVQS W + SQLR E+ S+ L PAY F+GR
Sbjct: 530 FSSGVSRSALRERFKSFNALFEEVHRVQSQWLVPDSQLREELKISILEKLSPAYRSFLGR 589
Query: 634 L--HGILGNQAYKYIKYGMIEIQDLLNHLFLG 663
H G YIK + +++ + LF G
Sbjct: 590 FRSHIESGKHPENYIKISVEDLETEVLDLFEG 621
>At3g29400 unknown protein (At3g29400)
Length = 658
Score = 142 bits (359), Expect = 5e-34
Identities = 135/521 (25%), Positives = 231/521 (43%), Gaps = 60/521 (11%)
Query: 189 ENSVVIEMDSLLRPSNDIYSGMMQQLTTCINALQQDSLNIIDRLMEEYNQGKCDFEMDTL 248
EN + E++ S + G+ ++ A + L + N G D +D +
Sbjct: 143 ENRLPFELEHSSFRSIEADHGVEEESMASFGAASTEDLILGSNNDSRRNSG--DVVVDLV 200
Query: 249 PSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFVLPEAKINTE---RE 305
+ + +L I M+ +GY++EC V RK L + L N K++ E R
Sbjct: 201 NPDVILDLKNIANTMIASGYDRECIQVCTMVRKDALDEFLYNH----EVEKLSIEDVLRM 256
Query: 306 RERYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQEATFQLLNF 365
L+T ++W+ V E+ + +F + CF++ + QLLNF
Sbjct: 257 DWATLNTNIKKWVRVMRDIVQVYLLSEKSLDNQIFGDLNEIGLTCFVDTVKAPMMQLLNF 316
Query: 366 ADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLF---PNSLVN-EAIAVRNRLGDASRV 421
+ ++ G +L ++++++ + L+P+ ++LF P S V E V RLGD +R
Sbjct: 317 GEAVSLGPRQPEKLLRILEMYELASELLPEIDALFLDHPGSSVRTEYREVMRRLGDCART 376
Query: 422 LFMKMHNFIFRVPAAKQVVSSY----GQHHQMTIQVMSYVSSACRKRRKLEQILEEY--- 474
F++ + I AA VSS+ G H +T VM+Y+ + + L+ +L E+
Sbjct: 377 TFLEFKSAI----AAD--VSSHPFPGGAVHPLTNYVMNYLMALTDFKHTLDSLLMEHDDA 430
Query: 475 --------PEVHNEV---EASSF-----------FLKQMEQIMRMLQRKLIVKSENCKDR 512
P++ N V E S++ + I +L+ L KS+ KD
Sbjct: 431 EDLTIPPSPDIINPVMVEEESTYENSSSPEKFLAMTRHFYSITSVLEANLQEKSKLYKDV 490
Query: 513 ALRHIFMLNNRSHIEAMNKFSRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKL 572
+L+HIF+LNN ++ S L IFG+ W + + K QQ Y+R+ W V+ FLK
Sbjct: 491 SLQHIFLLNNIHYMTRKVLKSELRHIFGDKWNRKHTWKFQQQATEYERATWLPVLSFLKD 550
Query: 573 D--------NNESITKELL-KEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNIL 623
D + S +K L +E+ FN FE + + Q+ W I LR ++ + +
Sbjct: 551 DGSGSGPGSGSGSGSKNLRPRERFQGFNTAFEEVYKAQTGWLISDEGLREDVRTKASMWV 610
Query: 624 LPAYGIFVGRLHGILGNQAYKYIKYGMIEIQDLLNHLFLGN 664
+ AY F R + +YIKY +I+ LL LF G+
Sbjct: 611 IQAYWTFYSRHKNSVSE---RYIKYTTDDIERLLLDLFAGS 648
>At2g39380 hypothetical protein
Length = 637
Score = 142 bits (358), Expect = 6e-34
Identities = 108/432 (25%), Positives = 201/432 (46%), Gaps = 25/432 (5%)
Query: 253 LHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFVLPEAKINTERERE-RYLD 311
+ +L I + M+ GY KEC Y RK ++ +GL + + + KI+ + L+
Sbjct: 183 MSDLKVIAETMISCGYGKECIKSYKLIRKSIVDEGL--HLLGIEKCKISRFNRMDWDVLE 240
Query: 312 TMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQEATFQLLNFADVIAY 371
M + W+ A+ I L E+ CD VFS S+ CF EI EA L F +++A
Sbjct: 241 HMIKNWIKAAKIGVITLLRGEKLLCDHVFSASSTIRESCFYEIVNEAGINLFKFPELVAE 300
Query: 372 GSLSKWRLFKMVDIFVKLNNLVPKFESLFPNSLVNEA----IAVRNRLGDASRVLFMKMH 427
S R+F+++D++ +++L P E +F V ++ +L D+ M+
Sbjct: 301 KKPSPERIFRLMDLYAAISDLRPDIELIFHFDSVAAVKTLVLSSLKKLKDSIYTSLMEFE 360
Query: 428 NFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILEEYPEVHNEVEASSFF 487
+ I + + + +++ G H++T MS++SS R L +IL E+P N S+F
Sbjct: 361 STIQK--DSSKALTAGGGIHKLTRSTMSFISSLSEYSRVLSEILAEHPLKKNTRMLESYF 418
Query: 488 LKQMEQ--------------IMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKFS 533
+ + ++ + KL +K+E+ KD +L ++F++NN + + +
Sbjct: 419 TAPILEDEHNNHAVSVHLAWLILIFLCKLDIKAESYKDVSLSYLFLVNNIQFVVDTVRST 478
Query: 534 RLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDNNESITKELLKEKIHLFNNR 593
L + G+DW ++AK++ Y+ +AW V L + ++ E K F+
Sbjct: 479 HLRNLLGDDWLTKHEAKLRSYAANYEIAAWANVYISLPEKTSSRLSPEEAKTHFKRFHAV 538
Query: 594 FEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGILGNQ--AYKYIKYGMI 651
FE QS+ I ++LR E+ S+ ++P Y F G+ L + + +
Sbjct: 539 FEEAYMKQSSCVITDAKLRNELKVSIAKKIVPEYREFYGKYLPTLSKERNIEMLVSFKPD 598
Query: 652 EIQDLLNHLFLG 663
+++ L+ LF G
Sbjct: 599 NLENYLSDLFHG 610
>At2g28640 unknown protein
Length = 605
Score = 142 bits (357), Expect = 8e-34
Identities = 127/498 (25%), Positives = 227/498 (45%), Gaps = 36/498 (7%)
Query: 187 RGENSVVIEMDSLLRPSNDIYSGMMQQLTT-CINALQQDSLNIIDRLMEEYNQGKCDFEM 245
R N V I M L + I + L ++ S N +++ Y+Q E
Sbjct: 96 RAHNLVTIAMKQLEKEFYRILKSNRRNLDPESVSVRSSPSFNARNKV-SIYSQVPKSEEA 154
Query: 246 DTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFV-LPEAKINTER 304
D + K+ I M+ +GYE EC +Y R ++ + L N F L KI ++
Sbjct: 155 DVMTDLKM-----IADCMISSGYENECIKIYKKIRGSIMVEALSNLGFENLSFGKI--QK 207
Query: 305 ERERYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQEATFQLLN 364
++ ++W+ A+ + T LF E+ D VFS S CF EI ++ L
Sbjct: 208 LDWDSMEKNIKKWLEATKVLITNLFEGERILSDHVFSPSVSVAESCFTEITLDSALTLFI 267
Query: 365 FADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPNSLVNEAIAVRNRLGDASRVLFM 424
F +A + ++F +DI+ ++ L+P+ E +F + AVR + D+ L
Sbjct: 268 FPVSVARCKKTVEKIFLTLDIYQTISQLMPQIEEIFS---YDSTTAVRLQAADSLNNLGE 324
Query: 425 KMHNFIFRVPAAKQVVSSY-----GQHHQMTIQVMSYVSSACRKRRKLEQIL-------- 471
++++ + A+ SS G HQ+T VM+++ L +L
Sbjct: 325 EINSMVAEFEASITKESSKTPIRGGSVHQLTRYVMNFIVFLADYHECLAGVLTESTLPLP 384
Query: 472 EEY----PEVHNEVE--ASSFFLKQMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSH 525
E+Y E +NE E +SS ++ ++ +L K+ KS D AL ++F+ NN +
Sbjct: 385 EDYFGNNDEDNNEGETGSSSTVTTRIAWLILVLLCKIDTKSRMYNDMALSYLFLANNLHY 444
Query: 526 IEAMNKFSRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDNNESITKELLKE 585
+ + + S + + G++W N++ K+ Q L+ Y++ AW EV+ L N E + + + KE
Sbjct: 445 VISKVRTSNMRVVLGDEWVTNHEGKVTQYLEKYEKIAWGEVITSLFDSNEEMLEEHVAKE 504
Query: 586 KIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGILGNQAYKY 645
++ FN FE + QS W + S+LR ++ SV L F + H + ++
Sbjct: 505 RLMRFNEGFEEAFQKQSEWVVPDSKLRDDLKDSVTEKLTTVTTTFYEKYH----VENWEE 560
Query: 646 IKYGMIEIQDLLNHLFLG 663
+K+ ++ + L+ LFLG
Sbjct: 561 VKFAPEDLDNYLSDLFLG 578
>At5g61010 putative protein
Length = 639
Score = 134 bits (336), Expect = 2e-31
Identities = 113/458 (24%), Positives = 206/458 (44%), Gaps = 43/458 (9%)
Query: 228 IIDRLMEEYNQGKCDF-EMDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQK 286
I++ E ++ DF D + L ++ I M Y++ +I ++ L+
Sbjct: 186 IVEASSHEDDEQISDFYNSDLVDPIVLPHIKAIANAMFACEYDQPFCEAFIGVQREALE- 244
Query: 287 GLLNKIFVLPEAKINTERERERYLDTM----------FQRWMTASDIATTVLFPIEQKFC 336
E + E ER +D + ++W I T V E++ C
Sbjct: 245 ----------EYMVTLEMERFSCVDVLRMDWEDLNGAMRKWTKVVKIITQVYLASEKQLC 294
Query: 337 DLVFSGFSSATSHCFIEICQEATFQLLNFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKF 396
D + F S ++ CFIEI ++A LLNF + + S L + + ++ ++
Sbjct: 295 DQILGDFESISTACFIEISKDAILSLLNFGEAVVLRSCKPEMLERFLSMYEVSAEILVDV 354
Query: 397 ESLFPNSLVNEA-IAVRN---RLGDASRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQ 452
++LFP+ + IA N +L D + F+K + I + + G H +T
Sbjct: 355 DNLFPDETGSSLRIAFHNLSKKLADHTTTTFLKFKDAIASDESTRPFHG--GGIHHLTRY 412
Query: 453 VMSYVSSACRKRRKLEQILEEY------PEVHNEVEASSFF---LKQMEQIMRMLQRKLI 503
VM+Y+ L +L+ PE E S F + + I+ L+ L
Sbjct: 413 VMNYLKLLPEYTDSLNSLLQNIHVDDSIPEKTGEDVLPSTFSPMARHLRSIVTTLESSLE 472
Query: 504 VKSENCKDRALRHIFMLNNRSHIEAMNKFSRLETIFGNDWFQNNKAKIQQNLDLYKRSAW 563
K++ D AL+ IF++NN ++ K S L +FG++W + + A Q N+ Y+RS W
Sbjct: 473 RKAQLYADEALKSIFLMNNFRYMVQKVKGSELRRLFGDEWIRKHIASYQCNVTNYERSTW 532
Query: 564 DEVMDFLKLDNNESITKELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNIL 623
++ L+ DNN+S+ L+E+ LF+ F+ + + Q+ W + S+LR ++ S +
Sbjct: 533 SSILALLR-DNNDSV--RTLRERCRLFSLAFDDVYKNQTRWSVPDSELRDDLHISTSVKV 589
Query: 624 LPAYGIFVGRLHGILGNQAYKYIKYGMIEIQDLLNHLF 661
+ +Y F+GR +G K+I+Y +I+++L LF
Sbjct: 590 VQSYRGFLGRNAVRIGE---KHIRYTCEDIENMLLDLF 624
>At3g09520 hypothetical protein
Length = 628
Score = 132 bits (333), Expect = 5e-31
Identities = 106/448 (23%), Positives = 205/448 (45%), Gaps = 36/448 (8%)
Query: 244 EMDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFVLPEAKINTE 303
E++ + + + +L I + M+ +GY KEC ++Y S RK ++ +G I+ L K +T
Sbjct: 163 ELEEVSTNVMTDLKSIAECMIGSGYAKECLSIYKSIRKSIIDEG----IYRLEVEKTSTG 218
Query: 304 RERERYLDTM---FQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQEATF 360
+ ++ + M + W+ A ++ LF E+ CD VF + CF +I ++
Sbjct: 219 KVKKMSWEVMELKIRSWLKAVKVSMETLFKGEKILCDHVFESSDAIRESCFSDISRDGAL 278
Query: 361 QLLNFADVIAYGSLSKW----RLFKMVDIFVKLNNLVPKFESLFPNSLVN--EAIAVRN- 413
L F ++I + K ++F+++D++ + ES+F ++ ++A+++
Sbjct: 279 LLFGFPEIINTKTSKKHSPPEKVFRLLDMYTAIAGNWQAIESIFSFDSISVVRSLALKSL 338
Query: 414 -RLGDASRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILE 472
L ++ R L ++ + I + + +VV G H +TI VM ++S L IL
Sbjct: 339 ISLSESIRSLLVEFESGIQK--DSSKVVVPGGGVHPLTISVMDHLSLLADYSNVLVDILA 396
Query: 473 EYPEVHNEVEASSFF-------------LKQMEQIMRMLQRKLIVKSENCKDRALRHIFM 519
P + S+F + I+ +L K+ KS + KD +++++F+
Sbjct: 397 GSPPPDRSLLPESYFNVSESDDSPSSELTIRFAWIILVLLCKIDRKSIHYKDFSIQYLFL 456
Query: 520 LNNRSHIEAMNKFSRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDNNESIT 579
NN H+ + + S L+ + G DW + AK++Q YKR AW V+ L + +T
Sbjct: 457 TNNLQHVVSRARSSNLKNLLGEDWITRHFAKMRQFAGSYKRLAWGPVVATLPENRTVEMT 516
Query: 580 KELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGIL- 638
E +KE+ F+ FE S + +R EI S+ L+P Y F ++
Sbjct: 517 PEEVKERFEKFSESFENAYSKHSVCVVADPNIRDEIKVSISRKLVPIYREFYNTRGSVIL 576
Query: 639 -----GNQAYKYIKYGMIEIQDLLNHLF 661
+++ +I++ L+ LF
Sbjct: 577 GEGDGARNLNSVVRFTPEDIENYLSDLF 604
>At5g52350 putative protein
Length = 547
Score = 130 bits (328), Expect = 2e-30
Identities = 113/436 (25%), Positives = 194/436 (43%), Gaps = 48/436 (11%)
Query: 239 GKCDFEMDTL-PSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFVLPE 297
GK D+ + T+ P L LH++ + M+ AG+++E Y R+ +L + L K+ V
Sbjct: 146 GKTDYTVPTIIPPTVLPVLHDLAQQMVKAGHQQELFKTYRDIRRAVLAQSL-EKLGVERH 204
Query: 298 AKINTERERERYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQE 357
+K + ER + + W+ I+ +LF E++ C + G F EI
Sbjct: 205 SKYDVERMNQDVFEAKIMNWIHYIRISVKLLFAAEKEICHQILDGVEPFRDQSFAEITTI 264
Query: 358 ATFQLLNFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPNSLVNEAIAVRNRLGD 417
+ LL+F IA S ++F ++D++ + L P+FE +F + E ++ +
Sbjct: 265 SFGMLLSFGYAIAISRRSPEKVFVILDMYEIMIELQPEFELIFGSKPCTE---MKEDALN 321
Query: 418 ASRVLFMKMHNFI--FRVPA---AKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILE 472
+++L + I F V A + V G H +T V YV L Q+ +
Sbjct: 322 LTKLLAQTVKETIADFEVAIEMDATETVVMDGSVHALTSYVARYVKFLFDYEPTLRQLFQ 381
Query: 473 EYPEVHNEVEASSFFLKQMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKF 532
E+ N + + M IMR L+ L KS +D AL +F++NN +I
Sbjct: 382 EF----NSNDPDTKLKSVMTGIMRALRNNLDGKSRQFEDAALTQLFLMNNVYYI------ 431
Query: 533 SRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDNNESITKELLKEKIHLFNN 592
I Q++K+ L NESI K L+KEK FN+
Sbjct: 432 -----ILQCITVQSSKS---------------------GLIKNESIKKTLVKEKFKTFNS 465
Query: 593 RFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGIL--GNQAYKYIKYGM 650
+FE + + Q W + +LR + ++ +LLPAYG F+ R ++ G + KYI++
Sbjct: 466 QFEELHQRQCQWSVSDVELRESLRLAIAEVLLPAYGSFLKRFGPMIESGKNSQKYIRFTP 525
Query: 651 IEIQDLLNHLFLGNKM 666
+++ +LN F G +
Sbjct: 526 EDLERMLNDFFQGKNL 541
>At5g59730 unknown protein
Length = 634
Score = 129 bits (324), Expect = 5e-30
Identities = 115/451 (25%), Positives = 207/451 (45%), Gaps = 43/451 (9%)
Query: 250 SEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFVLPEAKINTERERE-- 307
S+ + +L I M+ GY KEC VY + RK ++ + L N L + N + ++
Sbjct: 164 SDAMDDLKMIADCMISTGYAKECVRVYKTVRKSIVDETLHN----LQMERFNLHQVQKMD 219
Query: 308 -RYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQEATFQLLNFA 366
L++ + W+ A +A LF E+ D VFS F EI QE L F
Sbjct: 220 WEILESKIKTWLKAVKLAVRKLFFGERILADHVFSSSGLIVESSFTEITQEGALILFTFP 279
Query: 367 DVIA-YGSLSKWRLFKMVDIFVKLNNLVPKFESLFPNSLVNEAIAVRN-------RLGDA 418
+ + L+ ++F+ +D++ L NL + ES+F A AVR+ RLGDA
Sbjct: 280 EYASKIKKLTPEKMFRFLDMYEALANLYVEIESIF---YFESAAAVRSQVINSLARLGDA 336
Query: 419 SRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILEEYP--- 475
+R++ + I + + ++ G H +T VM+Y+S + I E +
Sbjct: 337 TRLMMTDFESAIQKETSKTPIIG--GGVHPLTRYVMNYLSFLADYSDSIAAIFENWKLSV 394
Query: 476 --------------EVHNEVEASSFFLKQMEQIMRMLQRKLIVKSENCKDRALRHIFMLN 521
E + E SS ++ ++ + K+ K++ KD AL ++F+ N
Sbjct: 395 PTPLPDSLYISGGDEANPEDLYSSPVSVRIAWVILLTLCKIDGKAQPYKDVALSYLFLAN 454
Query: 522 NRSHIEAMNKFSRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDNNESITKE 581
N ++ + S L+ + G+DW ++ K++ D +++ AW +V+D L + I+ E
Sbjct: 455 NLQYVVVKVRSSTLKVLLGDDWVFRHEEKVKLYADKFEKLAWGKVLDLLPEIPTDEISPE 514
Query: 582 LLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGR---LHGIL 638
K + FN+ FE R Q++W I +LR +I ++ L+ F +G++
Sbjct: 515 EAKVLVARFNDEFETSYRKQTSWVIPDPKLRDQIKITLSQKLMLVCTEFYRMNRFAYGMV 574
Query: 639 G-NQAYKYIKYGMIEIQDLLNHLFLGNKMYG 668
G N+A +Y +I + L+ L+ G++ G
Sbjct: 575 GDNEAIS--RYTPEDIGNYLSDLYFGSRGSG 603
>At1g07725 putative protein
Length = 566
Score = 128 bits (321), Expect = 1e-29
Identities = 99/397 (24%), Positives = 183/397 (45%), Gaps = 34/397 (8%)
Query: 250 SEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFVLPEAKINTERERERY 309
S+ + +L I M+ +GYEK+C +Y +RK ++ L + F E +T+ ++ +
Sbjct: 174 SDAIVDLKMIANCMISSGYEKDCVKIYKKFRKKIIVDTLSHLGF---EKLTSTQMQKLEW 230
Query: 310 --LDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQEATFQLLNFAD 367
L+ + W+ + +A T LF E+ D +FS SS CF++I ++ L F+
Sbjct: 231 EILEKKIKIWVIVARVAITTLFNGERILSDHIFS--SSVAESCFVDITLQSALNLFIFSL 288
Query: 368 VIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPNSLVNEAIAVR-------NRLGDASR 420
+A + ++F +D++ + L PK + +F + AVR +L ++
Sbjct: 289 TVAKSRKTAEKIFPTLDVYQTILQLTPKIDQIFS---YDSTAAVRLQANESLEKLSESVN 345
Query: 421 VLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILEE----YPE 476
+ + + I + ++K +S G H Q+T VM+++ L IL+E PE
Sbjct: 346 AMMTEFQSSITK-ESSKSAISGGGVH-QLTRYVMNFIVFLADYSDSLATILKESSLPLPE 403
Query: 477 VHNEVEAS--------SFFLKQMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEA 528
+ S ++ ++ +L K+ KS D AL ++F+ NN ++
Sbjct: 404 DYFSSSGEENPGSGDRSPMAARLAWLILVLLCKIDAKSRLYNDSALSYLFLANNLHYVVT 463
Query: 529 MNKFSRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDNNESITKELLKEKIH 588
+ S L + G+DW N++ K+ Q L+ Y++ AW +V+ L D+ E E +
Sbjct: 464 KVRTSNLRLVLGDDWVANHEVKVNQYLEKYEKMAWGDVIASLPGDSTAGTEAE---ESLR 520
Query: 589 LFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLP 625
FN FE + W + LR EI +S+ L+P
Sbjct: 521 RFNEAFEEAYKKHKTWVVPDPNLRDEIQASIARKLMP 557
>At3g09530 hypothetical protein
Length = 637
Score = 127 bits (319), Expect = 2e-29
Identities = 109/461 (23%), Positives = 202/461 (43%), Gaps = 37/461 (8%)
Query: 234 EEYNQGKCDFEMDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIF 293
E + G E++ + L I M+ AG KEC+ Y S RK ++ + I+
Sbjct: 148 ESRSAGDSIIEVEEVSKNSRTELKSIADCMIAAGCAKECATTYKSIRKSIVDES----IY 203
Query: 294 VLPEAKINTERERE---RYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHC 350
L I++ + ++ ++ RW+ A ++ LF E+ CD +F S C
Sbjct: 204 RLGVENISSSKAKKMPCEVVELKMNRWIEAVKVSMKTLFNGEKTLCDEIFESSVSLREFC 263
Query: 351 FIEICQEATFQLLNFADVIAYGSLSK---WRLFKMVDIFVKLNNLVPKFESLFP----NS 403
F +I +E L F + I ++F ++D++ + + + E++F ++
Sbjct: 264 FRDISKEGALLLFGFPETITLRDKKNPHPEKIFPLLDMYCTITDNLLAIEAIFSFPSISN 323
Query: 404 LVNEAIAVRNRLGDASRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRK 463
+ +A + +RL ++ M + I R ++K VV G H MTI M+++S
Sbjct: 324 VRTQAHSSLSRLSESILAHLMDFESQI-RKDSSKTVVRGGGVH-PMTISAMNHISRLAEY 381
Query: 464 RRKLEQILE--------------EYPEVHNEVEASSFFLK-QMEQIMRMLQRKLIVKSEN 508
L IL+ Y V E+ LK + ++ +L K+ K+E
Sbjct: 382 SNALINILKGSSSSSSAKALLPKSYFNVSESEESPVSELKARFAWMILVLLCKIDGKAEM 441
Query: 509 CKDRALRHIFMLNNRSHIEAMNKFSRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMD 568
KD +++++F+ NN H+ + + + ++ + GNDW N K++Q Y+R AW +
Sbjct: 442 YKDFSMQYLFLANNLQHVVSRARSTNVKHVLGNDWIAKNSEKVRQFARSYERLAWGPLAS 501
Query: 569 FL-KLDNNESI--TKELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLP 625
+ +E++ + E + FN FE+ C QS + +L E+ S+G LLP
Sbjct: 502 MCPAISTSEAVEMSPEEAMMQFKKFNETFESTCEAQSECIVLDPKLLDEMRISIGRKLLP 561
Query: 626 AYGIFVGRLHG---ILGNQAYKYIKYGMIEIQDLLNHLFLG 663
Y F + G + ++Y +I + L+ LF G
Sbjct: 562 VYRDFYNAHRNAVMLAGTEGQWNVRYNPEDIGNHLSELFSG 602
>At2g28650 unknown protein
Length = 573
Score = 126 bits (317), Expect = 3e-29
Identities = 101/395 (25%), Positives = 187/395 (46%), Gaps = 21/395 (5%)
Query: 250 SEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFVLPEAKINTERERERY 309
S + +L I M+ +GY KEC +Y RK ++ + L N++ ++
Sbjct: 146 SNTMDDLKMIADCMISSGYSKECFKIYKRIRKSIINEAL-NQLGFENLTFSQIQKLEWEV 204
Query: 310 LDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSH-CFIEICQEATFQLLNFADV 368
++ ++W+ + A LF EQ D VFS SS F EI + L F +
Sbjct: 205 MEKKIRKWLRTTTRAVNTLFSGEQILSDHVFSSSSSTIRESAFAEITSQTALALFTFPEK 264
Query: 369 IAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPNSLVN------EAIAVRNRLGDASRVL 422
+A S ++F +D++ + +L+PK LF + + + V R G S +
Sbjct: 265 MAKCRKSPEKIFLTLDVYQTIVDLLPKINELFSSDSTSTVRSQVDLTLVNLREGVVSMI- 323
Query: 423 FMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILEE----YPEVH 478
+ + I + ++K ++S G HQ+T VM+++ L I+ + PE
Sbjct: 324 -DEFESSISK-ESSKSLISG-GGIHQLTRYVMNFIVFLADYSDTLSDIISKPSLPSPEEE 380
Query: 479 NEVEASSFFLKQMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKFSRLETI 538
+ SS ++ +++ L K+ KS D AL ++F++NN +++ + S L+T+
Sbjct: 381 KDSGDSSPVKSRISRLILFLLCKIDAKSRLYNDVALSYLFLINNVNYVVVKVRSSNLKTV 440
Query: 539 FGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDNNESITKELLKEKIHLFNNRFEAIC 598
DW + ++AK+++ + ++ W E+M L ++ ++T E E I F++ FE
Sbjct: 441 LSEDWVKKHEAKVKKYVAKFEEIVWGEMM--TSLSDDVTMTAE---EGIKRFSDGFEEAY 495
Query: 599 RVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGR 633
+ Q+ W + S+LR EI SVG +++P Y F R
Sbjct: 496 KRQTGWIVPDSKLRDEIKRSVGMMIIPRYSGFCER 530
>At3g55150 unknown protein
Length = 636
Score = 123 bits (308), Expect = 4e-28
Identities = 108/438 (24%), Positives = 202/438 (45%), Gaps = 30/438 (6%)
Query: 253 LHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGL-LNKIFVLPEAKINTERERERYLD 311
+ +L I + M+ GY KEC +Y RK ++ +GL L I + ++ + R L+
Sbjct: 184 MSDLKAIAESMISCGYGKECVKIYKRVRKSIVDEGLSLLGIEIYKGSRFH--RTDWVTLE 241
Query: 312 TMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQEATFQLLNFADVIAY 371
M + W+ A+ I LF E+ CD VFS +S CF EI EA L F + +A
Sbjct: 242 HMIKNWIKAAKIGIATLFRGEKLLCDHVFSASNSTRESCFYEIANEAATNLFKFPEFVAK 301
Query: 372 GSLSKWRLFKMVDIFVKLNNLVPKFESLFPNSLVNEAIAVRNRLGDASRVLFMKMHNFIF 431
S R+F ++D+ +++L E +F V V+++ + + L + +H+ +
Sbjct: 302 EKKSHERIFPLMDLQAAISDLWQDIEMIFHFDAV---AGVKSQALTSLQKLKVSIHSALT 358
Query: 432 RVPAAKQ-----VVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILEEYPEVHNEVEASSF 486
+ Q ++ G H++T M+++SS + L +IL ++P N S+
Sbjct: 359 DFESIIQKDTTKALTPGGGIHKLTRSTMNFISSLSKYSGVLSEILADHPLPRNTRLLESY 418
Query: 487 F---LKQMEQ-----------IMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKF 532
+ + EQ ++ +L KL K+E+ KD +L ++F+ NN I
Sbjct: 419 VRAPISEDEQHNHALSVHFAWLILVLLCKLDTKAEHYKDVSLSYLFLANNLQIIIETVGS 478
Query: 533 SRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKL-DNNESITKELLKEKIHLFN 591
+ L + G+DW ++ K+ Y+ +AW V F+ L + ++ E K F+
Sbjct: 479 TPLRNLLGDDWLNKHEDKLCAYAGNYEIAAWSNV--FMSLPEEPTDLSPEEAKIYFRRFH 536
Query: 592 NRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGILGNQ--AYKYIKYG 649
FE QS+ + ++LR E+ S+ L+P Y F + +LG + +++
Sbjct: 537 TAFEEAYMKQSSRVVPNAKLRDELKVSIAKKLVPEYREFYRKYLPMLGQERNIEILVRFK 596
Query: 650 MIEIQDLLNHLFLGNKMY 667
+++ ++ LF G ++
Sbjct: 597 PDNLENYISDLFHGTPIH 614
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.327 0.139 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,963,714
Number of Sequences: 26719
Number of extensions: 572644
Number of successful extensions: 2100
Number of sequences better than 10.0: 31
Number of HSP's better than 10.0 without gapping: 24
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 1978
Number of HSP's gapped (non-prelim): 37
length of query: 672
length of database: 11,318,596
effective HSP length: 106
effective length of query: 566
effective length of database: 8,486,382
effective search space: 4803292212
effective search space used: 4803292212
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC138012.9


