

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
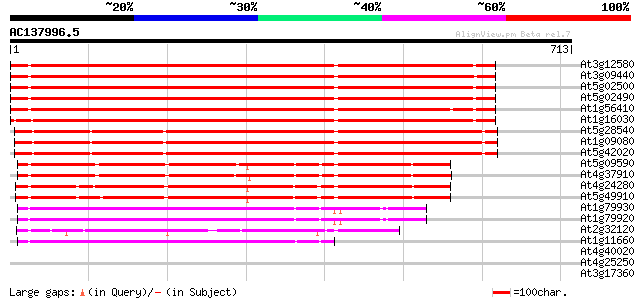
Query= AC137996.5 + phase: 0
(713 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At3g12580 putative protein 730 0.0
At3g09440 heat-shock protein (At-hsc70-3) 726 0.0
At5g02500 dnaK-type molecular chaperone hsc70.1 718 0.0
At5g02490 dnaK-type molecular chaperone hsc70.1 - like 709 0.0
At1g56410 hypothetical protein 709 0.0
At1g16030 unknown protein 703 0.0
At5g28540 luminal binding protein 574 e-164
At1g09080 putative luminal binding protein 572 e-163
At5g42020 luminal binding protein (dbj|BAA13948.1) 569 e-162
At5g09590 heat shock protein 70 (Hsc70-5) 421 e-118
At4g37910 heat shock protein 70 like protein 421 e-118
At4g24280 hsp 70-like protein 402 e-112
At5g49910 heat shock protein 70 (Hsc70-7) 400 e-111
At1g79930 putative heat-shock protein 248 6e-66
At1g79920 putative heat-shock protein (At1g79920) 248 1e-65
At2g32120 70kD heat shock protein 238 9e-63
At1g11660 heat-shock protein like 229 4e-60
At4g40020 putative protein 33 0.72
At4g25250 unknown protein 33 0.72
At3g17360 kinesin-like protein 33 0.72
>At3g12580 putative protein
Length = 650
Score = 730 bits (1885), Expect = 0.0
Identities = 388/621 (62%), Positives = 478/621 (76%), Gaps = 12/621 (1%)
Query: 1 MARKYEGRAVGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDYQRLIGNGA 60
MA K EG A+GIDLGTTYSCV VW H+RVEII NDQGNRTTPS VAFTD +RLIG+ A
Sbjct: 1 MAGKGEGPAIGIDLGTTYSCVGVW--QHDRVEIIANDQGNRTTPSYVAFTDSERLIGDAA 58
Query: 61 ENQSATNPENTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYKGQEKQF 120
+NQ A NP NTVFDAKRLIGR+YSDP VQ D WPF V SG +KPMI V +KG+EKQF
Sbjct: 59 KNQVAMNPTNTVFDAKRLIGRRYSDPSVQADKSHWPFKVVSGPGEKPMIVVNHKGEEKQF 118
Query: 121 CAEEISSAVLTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILN 180
AEEISS VL KMREIAEA+LGSPVKN VVTVPAYFNDSQR+AT DAG I+GLNV+RI+N
Sbjct: 119 SAEEISSMVLIKMREIAEAFLGSPVKNAVVTVPAYFNDSQRQATKDAGVISGLNVMRIIN 178
Query: 181 EPTAAAIAYGLDKRSNYDGKRNILVFDLGGGTFDVSILTIKGDVFDVKATAGNTHLGGED 240
EPTAAAIAYGLDK+++ G++N+L+FDLGGGTFDVS+LTI+ +F+VKATAG+THLGGED
Sbjct: 179 EPTAAAIAYGLDKKASSVGEKNVLIFDLGGGTFDVSLLTIEEGIFEVKATAGDTHLGGED 238
Query: 241 FDNRMVYYFVEEFKKKNKVEISGNPKSLRRLRTACERAKRILSFTFVTTVEVDALFMGID 300
FDNRMV +FV+EFK+KNK +I+GNP++LRRLRTACERAKR LS T TT+E+D+LF GID
Sbjct: 239 FDNRMVNHFVQEFKRKNKKDITGNPRALRRLRTACERAKRTLSSTAQTTIEIDSLFEGID 298
Query: 301 FSSSITRAKFEEINMDFFNECINIVDSCLRDSKIYKNDIDDVVLVGGSSRIPKVQDLLLE 360
F ++ITRA+FEE+NMD F +C+ V+ CLRD+K+ K+ + DVVLVGGS+RIPKVQ LL +
Sbjct: 299 FYTTITRARFEELNMDLFRKCMEPVEKCLRDAKMDKSSVHDVVLVGGSTRIPKVQQLLQD 358
Query: 361 FFKGKALFMNINPDEAIAYGAAVQAAILS-EDFKNVPNLVLRDVTPLSLGIATATYWENI 419
FF GK L +INPDEA+AYGAAVQAAILS E + V +L+L DVTPLSLG+ TA +
Sbjct: 359 FFNGKELCKSINPDEAVAYGAAVQAAILSGEGNEKVQDLLLLDVTPLSLGLETA---GGV 415
Query: 420 MDVVIPRNTSIPVKNTKGYCTAIDN-CGASIIVYEGERPRASDNNLLGLFTLSCL-PGPR 477
M V+IPRNT+IP K + + T DN G I VYEGER R DNNLLG F LS + P PR
Sbjct: 416 MTVLIPRNTTIPTKKEQIFSTYSDNQPGVLIQVYEGERARTKDNNLLGKFELSGIPPAPR 475
Query: 478 GQP-LEVCFSIDENGILTVSGKDITTGILNEITITNEKERLSKFEIEKKIEEAEKYRVED 536
G P + VCF ID NGIL VS +D TTG N+ITITN+K RLSK EIEK ++EAEKY+ ED
Sbjct: 476 GVPQITVCFDIDANGILNVSAEDKTTGQKNKITITNDKGRLSKEEIEKMVQEAEKYKAED 535
Query: 537 MKFLRKAKVISALDSCVYNIKSALNMKDVDLILSPQESEQINNAIIVAMNLLEMNKNNQH 596
+ +K +AL++ YN+++ + + + L + ++I +AI A+ L+ NQ
Sbjct: 536 EEHKKKVDAKNALENYAYNMRNTIKDEKIASKLDAADKKKIEDAIDQAIEWLD---GNQL 592
Query: 597 IEIDVLEHHLKEMNSMLNMLL 617
E D E +KE+ S+ N ++
Sbjct: 593 AEADEFEDKMKELESLCNPII 613
>At3g09440 heat-shock protein (At-hsc70-3)
Length = 649
Score = 726 bits (1873), Expect = 0.0
Identities = 385/621 (61%), Positives = 478/621 (75%), Gaps = 12/621 (1%)
Query: 1 MARKYEGRAVGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDYQRLIGNGA 60
MA K EG A+GIDLGTTYSCV VW H+RVEII NDQGNRTTPS VAFTD +RLIG+ A
Sbjct: 1 MAGKGEGPAIGIDLGTTYSCVGVW--QHDRVEIIANDQGNRTTPSYVAFTDSERLIGDAA 58
Query: 61 ENQSATNPENTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYKGQEKQF 120
+NQ A NP NTVFDAKRLIGR+++D VQ DI LWPF + SG +KPMI V YKG++K+F
Sbjct: 59 KNQVAMNPINTVFDAKRLIGRRFTDSSVQSDIKLWPFTLKSGPAEKPMIVVNYKGEDKEF 118
Query: 121 CAEEISSAVLTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILN 180
AEEISS +L KMREIAEAYLG+ +KN VVTVPAYFNDSQR+AT DAG IAGLNV+RI+N
Sbjct: 119 SAEEISSMILIKMREIAEAYLGTTIKNAVVTVPAYFNDSQRQATKDAGVIAGLNVMRIIN 178
Query: 181 EPTAAAIAYGLDKRSNYDGKRNILVFDLGGGTFDVSILTIKGDVFDVKATAGNTHLGGED 240
EPTAAAIAYGLDK++ G++N+L+FDLGGGTFDVS+LTI+ +F+VKATAG+THLGGED
Sbjct: 179 EPTAAAIAYGLDKKATSVGEKNVLIFDLGGGTFDVSLLTIEEGIFEVKATAGDTHLGGED 238
Query: 241 FDNRMVYYFVEEFKKKNKVEISGNPKSLRRLRTACERAKRILSFTFVTTVEVDALFMGID 300
FDNRMV +FV+EFK+KNK +ISGNP++LRRLRTACERAKR LS T TT+E+D+LF GID
Sbjct: 239 FDNRMVNHFVQEFKRKNKKDISGNPRALRRLRTACERAKRTLSSTAQTTIEIDSLFDGID 298
Query: 301 FSSSITRAKFEEINMDFFNECINIVDSCLRDSKIYKNDIDDVVLVGGSSRIPKVQDLLLE 360
F + ITRA+FEE+N+D F +C+ V+ CLRD+K+ KN IDDVVLVGGS+RIPKVQ LL++
Sbjct: 299 FYAPITRARFEELNIDLFRKCMEPVEKCLRDAKMDKNSIDDVVLVGGSTRIPKVQQLLVD 358
Query: 361 FFKGKALFMNINPDEAIAYGAAVQAAILS-EDFKNVPNLVLRDVTPLSLGIATATYWENI 419
FF GK L +INPDEA+AYGAAVQAAILS E + V +L+L DVTPLSLG+ TA +
Sbjct: 359 FFNGKELCKSINPDEAVAYGAAVQAAILSGEGNEKVQDLLLLDVTPLSLGLETA---GGV 415
Query: 420 MDVVIPRNTSIPVKNTKGYCTAIDN-CGASIIVYEGERPRASDNNLLGLFTLSCL-PGPR 477
M V+I RNT+IP K + + T DN G I VYEGER R DNNLLG F LS + P PR
Sbjct: 416 MTVLIQRNTTIPTKKEQVFSTYSDNQPGVLIQVYEGERARTKDNNLLGKFELSGIPPAPR 475
Query: 478 GQP-LEVCFSIDENGILTVSGKDITTGILNEITITNEKERLSKFEIEKKIEEAEKYRVED 536
G P + VCF ID NGIL VS +D TTG N+ITITN+K RLSK EIEK ++EAEKY+ ED
Sbjct: 476 GVPQITVCFDIDANGILNVSAEDKTTGQKNKITITNDKGRLSKDEIEKMVQEAEKYKSED 535
Query: 537 MKFLRKAKVISALDSCVYNIKSALNMKDVDLILSPQESEQINNAIIVAMNLLEMNKNNQH 596
+ +K +AL++ YN+++ + + + L+ + ++I ++I A+ LE NQ
Sbjct: 536 EEHKKKVDAKNALENYAYNMRNTIRDEKIGEKLAGDDKKKIEDSIEAAIEWLEA---NQL 592
Query: 597 IEIDVLEHHLKEMNSMLNMLL 617
E D E +KE+ S+ N ++
Sbjct: 593 AECDEFEDKMKELESICNPII 613
>At5g02500 dnaK-type molecular chaperone hsc70.1
Length = 651
Score = 718 bits (1853), Expect = 0.0
Identities = 380/621 (61%), Positives = 476/621 (76%), Gaps = 12/621 (1%)
Query: 1 MARKYEGRAVGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDYQRLIGNGA 60
M+ K EG A+GIDLGTTYSCV VW H+RVEII NDQGNRTTPS VAFTD +RLIG+ A
Sbjct: 1 MSGKGEGPAIGIDLGTTYSCVGVW--QHDRVEIIANDQGNRTTPSYVAFTDSERLIGDAA 58
Query: 61 ENQSATNPENTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYKGQEKQF 120
+NQ A NP NTVFDAKRLIGR++SD VQ D+ LWPF + +G DKPMI V+YKG+EK+F
Sbjct: 59 KNQVAMNPVNTVFDAKRLIGRRFSDSSVQSDMKLWPFKIQAGPADKPMIYVEYKGEEKEF 118
Query: 121 CAEEISSAVLTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILN 180
AEEISS VL KMREIAEAYLG +KN VVTVPAYFNDSQR+AT DAG IAGLNV+RI+N
Sbjct: 119 AAEEISSMVLIKMREIAEAYLGVTIKNAVVTVPAYFNDSQRQATKDAGVIAGLNVMRIIN 178
Query: 181 EPTAAAIAYGLDKRSNYDGKRNILVFDLGGGTFDVSILTIKGDVFDVKATAGNTHLGGED 240
EPTAAAIAYGLDK++ G++N+L+FDLGGGTFDVS+LTI+ +F+VKATAG+THLGGED
Sbjct: 179 EPTAAAIAYGLDKKATSVGEKNVLIFDLGGGTFDVSLLTIEEGIFEVKATAGDTHLGGED 238
Query: 241 FDNRMVYYFVEEFKKKNKVEISGNPKSLRRLRTACERAKRILSFTFVTTVEVDALFMGID 300
FDNRMV +FV+EFK+K+K +I+GNP++LRRLRT+CERAKR LS T TT+E+D+L+ GID
Sbjct: 239 FDNRMVNHFVQEFKRKSKKDITGNPRALRRLRTSCERAKRTLSSTAQTTIEIDSLYEGID 298
Query: 301 FSSSITRAKFEEINMDFFNECINIVDSCLRDSKIYKNDIDDVVLVGGSSRIPKVQDLLLE 360
F S+ITRA+FEE+NMD F +C+ V+ CLRD+K+ K+ + DVVLVGGS+RIPKVQ LL +
Sbjct: 299 FYSTITRARFEELNMDLFRKCMEPVEKCLRDAKMDKSTVHDVVLVGGSTRIPKVQQLLQD 358
Query: 361 FFKGKALFMNINPDEAIAYGAAVQAAILS-EDFKNVPNLVLRDVTPLSLGIATATYWENI 419
FF GK L +INPDEA+AYGAAVQ AILS E + V +L+L DVTPLSLG+ TA +
Sbjct: 359 FFNGKELCKSINPDEAVAYGAAVQGAILSGEGNEKVQDLLLLDVTPLSLGLETA---GGV 415
Query: 420 MDVVIPRNTSIPVKNTKGYCTAIDN-CGASIIVYEGERPRASDNNLLGLFTLSCL-PGPR 477
M +IPRNT+IP K + + T DN G I VYEGER R DNNLLG F LS + P PR
Sbjct: 416 MTTLIPRNTTIPTKKEQVFSTYSDNQPGVLIQVYEGERARTKDNNLLGKFELSGIPPAPR 475
Query: 478 GQP-LEVCFSIDENGILTVSGKDITTGILNEITITNEKERLSKFEIEKKIEEAEKYRVED 536
G P + VCF ID NGIL VS +D TTG N+ITITN+K RLSK EIEK ++EAEKY+ ED
Sbjct: 476 GVPQITVCFDIDANGILNVSAEDKTTGQKNKITITNDKGRLSKDEIEKMVQEAEKYKSED 535
Query: 537 MKFLRKAKVISALDSCVYNIKSALNMKDVDLILSPQESEQINNAIIVAMNLLEMNKNNQH 596
+ +K + +AL++ YN+++ + + + L + ++I ++I A+ LE NQ
Sbjct: 536 EEHKKKVEAKNALENYAYNMRNTIQDEKIGEKLPAADKKKIEDSIEQAIQWLE---GNQL 592
Query: 597 IEIDVLEHHLKEMNSMLNMLL 617
E D E +KE+ S+ N ++
Sbjct: 593 AEADEFEDKMKELESICNPII 613
>At5g02490 dnaK-type molecular chaperone hsc70.1 - like
Length = 653
Score = 709 bits (1830), Expect = 0.0
Identities = 376/621 (60%), Positives = 473/621 (75%), Gaps = 12/621 (1%)
Query: 1 MARKYEGRAVGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDYQRLIGNGA 60
MA K EG A+GIDLGTTYSCV VW H+RVEII NDQGNRTTPS VAFTD +RLIG+ A
Sbjct: 1 MAGKGEGPAIGIDLGTTYSCVGVW--QHDRVEIIANDQGNRTTPSYVAFTDSERLIGDAA 58
Query: 61 ENQSATNPENTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYKGQEKQF 120
+NQ A NP NTVFDAKRLIGR++SD VQ D LWPF + SG +KPMI V+YKG+EKQF
Sbjct: 59 KNQVAMNPVNTVFDAKRLIGRRFSDASVQSDRQLWPFTIISGTAEKPMIVVEYKGEEKQF 118
Query: 121 CAEEISSAVLTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILN 180
AEEISS VL KMREIAEA+LG+ VKN VVTVPAYFNDSQR+AT DAG IAGLNV+RI+N
Sbjct: 119 AAEEISSMVLIKMREIAEAFLGTTVKNAVVTVPAYFNDSQRQATKDAGVIAGLNVLRIIN 178
Query: 181 EPTAAAIAYGLDKRSNYDGKRNILVFDLGGGTFDVSILTIKGDVFDVKATAGNTHLGGED 240
EPTAAAIAYGLDK++ G++N+L+FDLGGGTFDVS+LTI+ +F+VKATAG+THLGGED
Sbjct: 179 EPTAAAIAYGLDKKATSVGEKNVLIFDLGGGTFDVSLLTIEEGIFEVKATAGDTHLGGED 238
Query: 241 FDNRMVYYFVEEFKKKNKVEISGNPKSLRRLRTACERAKRILSFTFVTTVEVDALFMGID 300
FDNRMV +FV+EFK+KNK +I+G P++LRRLRTACERAKR LS T TT+E+D+L+ G D
Sbjct: 239 FDNRMVNHFVQEFKRKNKQDITGQPRALRRLRTACERAKRTLSSTAQTTIEIDSLYGGAD 298
Query: 301 FSSSITRAKFEEINMDFFNECINIVDSCLRDSKIYKNDIDDVVLVGGSSRIPKVQDLLLE 360
F S ITRA+FEE+NMD F +C+ V+ CLRD+K+ K+ + ++VLVGGS+RIPKVQ LL +
Sbjct: 299 FYSPITRARFEEMNMDLFRKCMEPVEKCLRDAKMDKSTVHEIVLVGGSTRIPKVQQLLQD 358
Query: 361 FFKGKALFMNINPDEAIAYGAAVQAAILS-EDFKNVPNLVLRDVTPLSLGIATATYWENI 419
FF GK L +INPDEA+AYGAAVQAAILS E + V +L+L DVTPLSLG+ TA +
Sbjct: 359 FFNGKELCKSINPDEAVAYGAAVQAAILSGEGNEKVQDLLLLDVTPLSLGLETA---GGV 415
Query: 420 MDVVIPRNTSIPVKNTKGYCTAIDN-CGASIIVYEGERPRASDNNLLGLFTLSCL-PGPR 477
M +I RNT+IP K + + T DN G I V+EGER R DNNLLG F LS + P PR
Sbjct: 416 MTTLIQRNTTIPTKKEQVFSTYSDNQPGVLIQVFEGERARTKDNNLLGKFELSGIPPAPR 475
Query: 478 GQP-LEVCFSIDENGILTVSGKDITTGILNEITITNEKERLSKFEIEKKIEEAEKYRVED 536
G P + VCF ID NGIL VS +D TTG N+ITITN+K RLSK +IEK ++EAEKY+ ED
Sbjct: 476 GVPQITVCFDIDANGILNVSAEDKTTGKKNKITITNDKGRLSKEDIEKMVQEAEKYKSED 535
Query: 537 MKFLRKAKVISALDSCVYNIKSALNMKDVDLILSPQESEQINNAIIVAMNLLEMNKNNQH 596
+ +K + +AL++ YN+++ + + + L + +++ ++I A+ L+ NQ
Sbjct: 536 EEHKKKVEAKNALENYAYNMRNTIRDEKIGEKLPAADKKKVEDSIEEAIQWLD---GNQL 592
Query: 597 IEIDVLEHHLKEMNSMLNMLL 617
E D E +KE+ S+ N ++
Sbjct: 593 GEADEFEDKMKELESVCNPII 613
>At1g56410 hypothetical protein
Length = 617
Score = 709 bits (1830), Expect = 0.0
Identities = 384/621 (61%), Positives = 468/621 (74%), Gaps = 14/621 (2%)
Query: 1 MARKYEGRAVGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDYQRLIGNGA 60
MA K EG A+GIDLGTTYSCV VW H+RVEII NDQGNRTTPS VAFTD +RLIG+ A
Sbjct: 1 MAGKGEGPAIGIDLGTTYSCVGVW--QHDRVEIIANDQGNRTTPSYVAFTDSERLIGDAA 58
Query: 61 ENQSATNPENTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYKGQEKQF 120
+NQ A NP NTVFDAKRLIGR++SD VQ D+ WPF VT G DKPMI V YKG+EKQF
Sbjct: 59 KNQVAMNPVNTVFDAKRLIGRRFSDASVQSDMKFWPFKVTPGQADKPMIFVNYKGEEKQF 118
Query: 121 CAEEISSAVLTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILN 180
AEEISS VL KMREIAEAYLGS +KN VVTVPAYFNDSQR+AT DAG IAGLNV+RI+N
Sbjct: 119 AAEEISSMVLIKMREIAEAYLGSSIKNAVVTVPAYFNDSQRQATKDAGVIAGLNVLRIIN 178
Query: 181 EPTAAAIAYGLDKRSNYDGKRNILVFDLGGGTFDVSILTIKGDVFDVKATAGNTHLGGED 240
EPTAAAIAYGLDK++ G +N+L+FDLGGGTFDVS+LTI+ +F+VKATAG+THLGGED
Sbjct: 179 EPTAAAIAYGLDKKATSVGIKNVLIFDLGGGTFDVSLLTIEEGIFEVKATAGDTHLGGED 238
Query: 241 FDNRMVYYFVEEFKKKNKVEISGNPKSLRRLRTACERAKRILSFTFVTTVEVDALFMGID 300
FDNRMV +FV+EFK+KNK +ISG+ ++LRRLRTACERAKR LS T TTVEVD+LF GID
Sbjct: 239 FDNRMVNHFVQEFKRKNKKDISGDARALRRLRTACERAKRTLSSTAQTTVEVDSLFEGID 298
Query: 301 FSSSITRAKFEEINMDFFNECINIVDSCLRDSKIYKNDIDDVVLVGGSSRIPKVQDLLLE 360
F S ITRAKFEE+NMD F +C+ V CLRDSK+ K+ + DVVLVGGS+RIPKVQ LL +
Sbjct: 299 FYSPITRAKFEEMNMDLFRKCMEPVMKCLRDSKMDKSMVHDVVLVGGSTRIPKVQQLLQD 358
Query: 361 FFKGKALFMNINPDEAIAYGAAVQAAILS-EDFKNVPNLVLRDVTPLSLGIATATYWENI 419
FF GK L +INPDEA+AYGAAVQAAILS E + V +L+L DVTPLSLGI T +
Sbjct: 359 FFNGKELCKSINPDEAVAYGAAVQAAILSGEGNEKVQDLLLLDVTPLSLGIETI---GGV 415
Query: 420 MDVVIPRNTSIPVKNTKGYCTAIDNCGASII-VYEGERPRASDNNLLGLFTLSCL-PGPR 477
M +I RNT+IP K + + T +DN +I VYEGER R DNN+LG F LS + P PR
Sbjct: 416 MTTLIQRNTTIPAKKEQEFTTTVDNQPDVLIQVYEGERARTIDNNILGQFVLSGIPPAPR 475
Query: 478 GQP-LEVCFSIDENGILTVSGKDITTGILNEITITNEKERLSKFEIEKKIEEAEKYRVED 536
G P VCF ID NGIL VS +D TG N+ITITN+K RLSK +IEK ++EAEKY+ ED
Sbjct: 476 GIPQFTVCFDIDSNGILNVSAEDKATGKKNKITITNDKGRLSKDDIEKMVQEAEKYKSED 535
Query: 537 MKFLRKAKVISALDSCVYNIKSALNMKDVDLILSPQESEQINNAIIVAMNLLEMNKNNQH 596
+ +K + + L++ YN+ + L +D+ L + ++ ++I + L+ +NQ
Sbjct: 536 EEHKKKVEAKNGLENYAYNVGNTL--RDMGEKLPAADKKKFEDSIEEVIQWLD---DNQL 590
Query: 597 IEIDVLEHHLKEMNSMLNMLL 617
E D EH +KE+ S+ + ++
Sbjct: 591 AEADEFEHKMKELESVWSTII 611
Score = 33.5 bits (75), Expect = 0.42
Identities = 47/221 (21%), Positives = 91/221 (40%), Gaps = 25/221 (11%)
Query: 491 GILTVSGKDITTGILNEITITNEKERLSKFEIEKK------IEEAEKYRVEDMKFLRKAK 544
GI T+ G +TT I TI +KE+ ++ + + E E+ R D L +
Sbjct: 408 GIETIGGV-MTTLIQRNTTIPAKKEQEFTTTVDNQPDVLIQVYEGERARTIDNNILGQF- 465
Query: 545 VISALDSCVYNIKSALNMKDVDL--ILSPQESEQINNAIIVAMNLLEMNKNNQHIEIDVL 602
V+S + I D+D IL+ ++ N + + + + D +
Sbjct: 466 VLSGIPPAPRGIPQFTVCFDIDSNGILNVSAEDKATGK----KNKITITNDKGRLSKDDI 521
Query: 603 EHHLKEMNSMLNMLLKIVEDMKFLRKATVMSALDSCVKNITNALNTKDVNLIISPQESEK 662
E ++E + ED + +K + L++ N+ N L +D+ + + +K
Sbjct: 522 EKMVQEAEKYKS------EDEEHKKKVEAKNGLENYAYNVGNTL--RDMGEKLPAADKKK 573
Query: 663 INNAIFGAMNLLDKNKNNQQINIDVLEHHLKELTNMLKMLL 703
++I + LD +NQ D EH +KEL ++ ++
Sbjct: 574 FEDSIEEVIQWLD---DNQLAEADEFEHKMKELESVWSTII 611
>At1g16030 unknown protein
Length = 646
Score = 703 bits (1814), Expect = 0.0
Identities = 368/621 (59%), Positives = 474/621 (76%), Gaps = 13/621 (2%)
Query: 1 MARKYEGRAVGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDYQRLIGNGA 60
MA K E +A+GIDLGTTYSCV VW++D RVEII NDQGNRTTPS VAFTD +RLIG+ A
Sbjct: 1 MATKSE-KAIGIDLGTTYSCVGVWMND--RVEIIPNDQGNRTTPSYVAFTDTERLIGDAA 57
Query: 61 ENQSATNPENTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYKGQEKQF 120
+NQ A NP+NTVFDAKRLIGRK+SDP VQ DI+ WPF V SG +KPMI V YK +EKQF
Sbjct: 58 KNQVALNPQNTVFDAKRLIGRKFSDPSVQSDILHWPFKVVSGPGEKPMIVVSYKNEEKQF 117
Query: 121 CAEEISSAVLTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILN 180
EEISS VL KM+E+AEA+LG VKN VVTVPAYFNDSQR+AT DAGAI+GLNV+RI+N
Sbjct: 118 SPEEISSMVLVKMKEVAEAFLGRTVKNAVVTVPAYFNDSQRQATKDAGAISGLNVLRIIN 177
Query: 181 EPTAAAIAYGLDKRSNYDGKRNILVFDLGGGTFDVSILTIKGDVFDVKATAGNTHLGGED 240
EPTAAAIAYGLDK+ G++N+L+FDLGGGTFDVS+LTI+ VF+VKATAG+THLGGED
Sbjct: 178 EPTAAAIAYGLDKKGTKAGEKNVLIFDLGGGTFDVSLLTIEEGVFEVKATAGDTHLGGED 237
Query: 241 FDNRMVYYFVEEFKKKNKVEISGNPKSLRRLRTACERAKRILSFTFVTTVEVDALFMGID 300
FDNR+V +FV EF++K+K +I+GN ++LRRLRTACERAKR LS T TT+E+D+L GID
Sbjct: 238 FDNRLVNHFVAEFRRKHKKDIAGNARALRRLRTACERAKRTLSSTAQTTIEIDSLHEGID 297
Query: 301 FSSSITRAKFEEINMDFFNECINIVDSCLRDSKIYKNDIDDVVLVGGSSRIPKVQDLLLE 360
F ++I+RA+FEE+NMD F +C++ V+ L+D+K+ K+ + DVVLVGGS+RIPK+Q LL +
Sbjct: 298 FYATISRARFEEMNMDLFRKCMDPVEKVLKDAKLDKSSVHDVVLVGGSTRIPKIQQLLQD 357
Query: 361 FFKGKALFMNINPDEAIAYGAAVQAAILS-EDFKNVPNLVLRDVTPLSLGIATATYWENI 419
FF GK L +INPDEA+AYGAAVQAAIL+ E + V +L+L DV PLSLG+ TA +
Sbjct: 358 FFNGKELCKSINPDEAVAYGAAVQAAILTGEGSEKVQDLLLLDVAPLSLGLETA---GGV 414
Query: 420 MDVVIPRNTSIPVKNTKGYCTAIDN-CGASIIVYEGERPRASDNNLLGLFTLSCL-PGPR 477
M V+IPRNT++P K + + T DN G I VYEGER R DNNLLG F L + P PR
Sbjct: 415 MTVLIPRNTTVPCKKEQVFSTYADNQPGVLIQVYEGERARTRDNNLLGTFELKGIPPAPR 474
Query: 478 GQP-LEVCFSIDENGILTVSGKDITTGILNEITITNEKERLSKFEIEKKIEEAEKYRVED 536
G P + VCF ID NGIL VS +D T G+ N+ITITN+K RLSK EIEK +++AEKY+ ED
Sbjct: 475 GVPQINVCFDIDANGILNVSAEDKTAGVKNQITITNDKGRLSKEEIEKMVQDAEKYKAED 534
Query: 537 MKFLRKAKVISALDSCVYNIKSALNMKDVDLILSPQESEQINNAIIVAMNLLEMNKNNQH 596
+ +K + ++L++ YN+++ + + + L+ ++ ++I AI + +E NQ
Sbjct: 535 EQVKKKVEAKNSLENYAYNMRNTIKDEKLAQKLTQEDKQKIEKAIDETIEWIE---GNQL 591
Query: 597 IEIDVLEHHLKEMNSMLNMLL 617
E+D E+ LKE+ + N ++
Sbjct: 592 AEVDEFEYKLKELEGICNPII 612
Score = 32.7 bits (73), Expect = 0.72
Identities = 18/83 (21%), Positives = 41/83 (48%), Gaps = 3/83 (3%)
Query: 621 EDMKFLRKATVMSALDSCVKNITNALNTKDVNLIISPQESEKINNAIFGAMNLLDKNKNN 680
ED + +K ++L++ N+ N + + + ++ ++ +KI AI + ++ N
Sbjct: 533 EDEQVKKKVEAKNSLENYAYNMRNTIKDEKLAQKLTQEDKQKIEKAIDETIEWIE---GN 589
Query: 681 QQINIDVLEHHLKELTNMLKMLL 703
Q +D E+ LKEL + ++
Sbjct: 590 QLAEVDEFEYKLKELEGICNPII 612
>At5g28540 luminal binding protein
Length = 669
Score = 574 bits (1479), Expect = e-164
Identities = 326/620 (52%), Positives = 426/620 (68%), Gaps = 18/620 (2%)
Query: 7 GRAVGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDYQRLIGNGAENQSAT 66
G +GIDLGTTYSCV V+ + H VEII NDQGNR TPS V FTD +RLIG A+NQ+A
Sbjct: 35 GSVIGIDLGTTYSCVGVYKNGH--VEIIANDQGNRITPSWVGFTDSERLIGEAAKNQAAV 92
Query: 67 NPENTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYK-GQEKQFCAEEI 125
NPE TVFD KRLIGRK+ D VQKD L P+ + + + KP I VK K G+ K F EEI
Sbjct: 93 NPERTVFDVKRLIGRKFEDKEVQKDRKLVPYQIVNK-DGKPYIQVKIKDGETKVFSPEEI 151
Query: 126 SSAVLTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILNEPTAA 185
S+ +LTKM+E AEAYLG +K+ VVTVPAYFND+QR+AT DAG IAGLNV RI+NEPTAA
Sbjct: 152 SAMILTKMKETAEAYLGKKIKDAVVTVPAYFNDAQRQATKDAGVIAGLNVARIINEPTAA 211
Query: 186 AIAYGLDKRSNYDGKRNILVFDLGGGTFDVSILTIKGDVFDVKATAGNTHLGGEDFDNRM 245
AIAYGLDK+ G++NILVFDLGGGTFDVS+LTI VF+V +T G+THLGGEDFD+R+
Sbjct: 212 AIAYGLDKKG---GEKNILVFDLGGGTFDVSVLTIDNGVFEVLSTNGDTHLGGEDFDHRV 268
Query: 246 VYYFVEEFKKKNKVEISGNPKSLRRLRTACERAKRILSFTFVTTVEVDALFMGIDFSSSI 305
+ YF++ KKK++ +IS + K+L +LR CERAKR LS VE+++LF G+DFS +
Sbjct: 269 MEYFIKLIKKKHQKDISKDNKALGKLRRECERAKRALSSQHQVRVEIESLFDGVDFSEPL 328
Query: 306 TRAKFEEINMDFFNECINIVDSCLRDSKIYKNDIDDVVLVGGSSRIPKVQDLLLEFFKGK 365
TRA+FEE+N D F + + V + D+ + K+ ID++VLVGGS+RIPKVQ LL +FF+GK
Sbjct: 329 TRARFEELNNDLFRKTMGPVKKAMDDAGLQKSQIDEIVLVGGSTRIPKVQQLLKDFFEGK 388
Query: 366 ALFMNINPDEAIAYGAAVQAAILS-EDFKNVPNLVLRDVTPLSLGIATATYWENIMDVVI 424
+NPDEA+AYGAAVQ ILS E +++L DV PL+LGI T +M +I
Sbjct: 389 EPNKGVNPDEAVAYGAAVQGGILSGEGGDETKDILLLDVAPLTLGIETV---GGVMTKLI 445
Query: 425 PRNTSIPVKNTKGYCTAID-NCGASIIVYEGERPRASDNNLLGLFTLSCL-PGPRGQP-L 481
PRNT IP K ++ + T D SI V+EGER D LLG F L+ + P PRG P +
Sbjct: 446 PRNTVIPTKKSQVFTTYQDQQTTVSIQVFEGERSLTKDCRLLGKFDLNGIPPAPRGTPQI 505
Query: 482 EVCFSIDENGILTVSGKDITTGILNEITITNEKERLSKFEIEKKIEEAEKYRVEDMKFLR 541
EV F +D NGIL V +D +G +ITITNEK RLS+ EI++ ++EAE++ ED K
Sbjct: 506 EVTFEVDANGILNVKAEDKASGKSEKITITNEKGRLSQEEIDRMVKEAEEFAEEDKKVKE 565
Query: 542 KAKVISALDSCVYNIKSALNMKD-VDLILSPQESEQINNAIIVAMNLLEMNKNNQHIEID 600
K +AL++ VYN+K+ +N KD + L E E+I A A+ L+ N+N++ E D
Sbjct: 566 KIDARNALETYVYNMKNQVNDKDKLADKLEGDEKEKIEAATKEALEWLDENQNSEKEEYD 625
Query: 601 VLEHHLKEMNSMLNMLLKIV 620
LKE+ ++ N ++ V
Sbjct: 626 ---EKLKEVEAVCNPIITAV 642
Score = 37.0 bits (84), Expect = 0.038
Identities = 27/87 (31%), Positives = 44/87 (50%), Gaps = 4/87 (4%)
Query: 621 EDMKFLRKATVMSALDSCVKNITNALNTKD-VNLIISPQESEKINNAIFGAMNLLDKNKN 679
ED K K +AL++ V N+ N +N KD + + E EKI A A+ LD+N+N
Sbjct: 559 EDKKVKEKIDARNALETYVYNMKNQVNDKDKLADKLEGDEKEKIEAATKEALEWLDENQN 618
Query: 680 NQQINIDVLEHHLKELTNMLKMLLKIV 706
+++ D LKE+ + ++ V
Sbjct: 619 SEKEEYD---EKLKEVEAVCNPIITAV 642
>At1g09080 putative luminal binding protein
Length = 678
Score = 572 bits (1475), Expect = e-163
Identities = 322/619 (52%), Positives = 422/619 (68%), Gaps = 17/619 (2%)
Query: 7 GRAVGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDYQRLIGNGAENQSAT 66
G +GIDLGTTYSCV V+ + H VEII NDQGNR TPS VAFTD +RLIG A+NQ+A
Sbjct: 50 GTVIGIDLGTTYSCVGVYHNKH--VEIIANDQGNRITPSWVAFTDTERLIGEAAKNQAAK 107
Query: 67 NPENTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYKGQEKQFCAEEIS 126
NPE T+FD KRLIGRK+ DP VQ+DI P+ V + + KP I VK KG+EK F EEIS
Sbjct: 108 NPERTIFDPKRLIGRKFDDPDVQRDIKFLPYKVVNK-DGKPYIQVKVKGEEKLFSPEEIS 166
Query: 127 SAVLTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILNEPTAAA 186
+ +LTKM+E AEA+LG +K+ V+TVPAYFND+QR+AT DAGAIAGLNV+RI+NEPT AA
Sbjct: 167 AMILTKMKETAEAFLGKKIKDAVITVPAYFNDAQRQATKDAGAIAGLNVVRIINEPTGAA 226
Query: 187 IAYGLDKRSNYDGKRNILVFDLGGGTFDVSILTIKGDVFDVKATAGNTHLGGEDFDNRMV 246
IAYGLDK+ G+ NILV+DLGGGTFDVSILTI VF+V +T+G+THLGGEDFD+R++
Sbjct: 227 IAYGLDKKG---GESNILVYDLGGGTFDVSILTIDNGVFEVLSTSGDTHLGGEDFDHRVM 283
Query: 247 YYFVEEFKKKNKVEISGNPKSLRRLRTACERAKRILSFTFVTTVEVDALFMGIDFSSSIT 306
YF++ KKK +IS + K+L +LR CE AKR LS VE+++LF G+DFS +T
Sbjct: 284 DYFIKLVKKKYNKDISKDHKALGKLRRECELAKRSLSNQHQVRVEIESLFDGVDFSEPLT 343
Query: 307 RAKFEEINMDFFNECINIVDSCLRDSKIYKNDIDDVVLVGGSSRIPKVQDLLLEFFKGKA 366
RA+FEE+NMD F + + V L+D+ + K+DID++VLVGGS+RIPKVQ +L +FF GK
Sbjct: 344 RARFEELNMDLFKKTMEPVKKALKDAGLKKSDIDEIVLVGGSTRIPKVQQMLKDFFDGKE 403
Query: 367 LFMNINPDEAIAYGAAVQAAILS-EDFKNVPNLVLRDVTPLSLGIATATYWENIMDVVIP 425
NPDEA+AYGAAVQ +LS E + N++L DV PLSLGI T +M +IP
Sbjct: 404 PSKGTNPDEAVAYGAAVQGGVLSGEGGEETQNILLLDVAPLSLGIETV---GGVMTNIIP 460
Query: 426 RNTSIPVKNTKGYCTAID-NCGASIIVYEGERPRASDNNLLGLFTLS-CLPGPRGQP-LE 482
RNT IP K ++ + T D +I VYEGER DN LG F L+ LP PRG P +E
Sbjct: 461 RNTVIPTKKSQVFTTYQDQQTTVTINVYEGERSMTKDNRELGKFDLTGILPAPRGVPQIE 520
Query: 483 VCFSIDENGILTVSGKDITTGILNEITITNEKERLSKFEIEKKIEEAEKYRVEDMKFLRK 542
V F +D NGIL V +D ITITN+K RL++ EIE+ I EAE++ ED K
Sbjct: 521 VTFEVDANGILQVKAEDKVAKTSQSITITNDKGRLTEEEIEEMIREAEEFAEEDKIMKEK 580
Query: 543 AKVISALDSCVYNIKSALNMKD-VDLILSPQESEQINNAIIVAMNLLEMNKNNQHIEIDV 601
+ L++ VYN+KS + K+ + +S ++ E++ + A+ LE N N + + D
Sbjct: 581 IDARNKLETYVYNMKSTVADKEKLAKKISDEDKEKMEGVLKEALEWLEENVNAEKEDYD- 639
Query: 602 LEHHLKEMNSMLNMLLKIV 620
LKE+ + + ++K V
Sbjct: 640 --EKLKEVELVCDPVIKSV 656
>At5g42020 luminal binding protein (dbj|BAA13948.1)
Length = 668
Score = 569 bits (1467), Expect = e-162
Identities = 324/620 (52%), Positives = 425/620 (68%), Gaps = 18/620 (2%)
Query: 7 GRAVGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDYQRLIGNGAENQSAT 66
G +GIDLGTTYSCV V+ + H VEII NDQGNR TPS V FTD +RLIG A+NQ+A
Sbjct: 35 GSVIGIDLGTTYSCVGVYKNGH--VEIIANDQGNRITPSWVGFTDSERLIGEAAKNQAAV 92
Query: 67 NPENTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYK-GQEKQFCAEEI 125
NPE TVFD KRLIGRK+ D VQKD L P+ + + + KP I VK K G+ K F EEI
Sbjct: 93 NPERTVFDVKRLIGRKFEDKEVQKDRKLVPYQIVNK-DGKPYIQVKIKDGETKVFSPEEI 151
Query: 126 SSAVLTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILNEPTAA 185
S+ +LTKM+E AEAYLG +K+ VVTVPAYFND+QR+AT DAG IAGLNV RI+NEPTAA
Sbjct: 152 SAMILTKMKETAEAYLGKKIKDAVVTVPAYFNDAQRQATKDAGVIAGLNVARIINEPTAA 211
Query: 186 AIAYGLDKRSNYDGKRNILVFDLGGGTFDVSILTIKGDVFDVKATAGNTHLGGEDFDNRM 245
AIAYGLDK+ G++NILVFDLGGGTFDVS+LTI VF+V +T G+THLGGEDFD+R+
Sbjct: 212 AIAYGLDKKG---GEKNILVFDLGGGTFDVSVLTIDNGVFEVLSTNGDTHLGGEDFDHRI 268
Query: 246 VYYFVEEFKKKNKVEISGNPKSLRRLRTACERAKRILSFTFVTTVEVDALFMGIDFSSSI 305
+ YF++ KKK++ +IS + K+L +LR CERAKR LS VE+++LF G+D S +
Sbjct: 269 MEYFIKLIKKKHQKDISKDNKALGKLRRECERAKRALSSQHQVRVEIESLFDGVDLSEPL 328
Query: 306 TRAKFEEINMDFFNECINIVDSCLRDSKIYKNDIDDVVLVGGSSRIPKVQDLLLEFFKGK 365
TRA+FEE+N D F + + V + D+ + K+ ID++VLVGGS+RIPKVQ LL +FF+GK
Sbjct: 329 TRARFEELNNDLFRKTMGPVKKAMDDAGLQKSQIDEIVLVGGSTRIPKVQQLLKDFFEGK 388
Query: 366 ALFMNINPDEAIAYGAAVQAAILS-EDFKNVPNLVLRDVTPLSLGIATATYWENIMDVVI 424
+NPDEA+AYGAAVQ ILS E +++L DV PL+LGI T +M +I
Sbjct: 389 EPNKGVNPDEAVAYGAAVQGGILSGEGGDETKDILLLDVAPLTLGIETV---GGVMTKLI 445
Query: 425 PRNTSIPVKNTKGYCTAID-NCGASIIVYEGERPRASDNNLLGLFTLSCL-PGPRGQP-L 481
PRNT IP K ++ + T D SI V+EGER D LLG F L+ + P PRG P +
Sbjct: 446 PRNTVIPTKKSQVFTTYQDQQTTVSIQVFEGERSLTKDCRLLGKFDLTGVPPAPRGTPQI 505
Query: 482 EVCFSIDENGILTVSGKDITTGILNEITITNEKERLSKFEIEKKIEEAEKYRVEDMKFLR 541
EV F +D NGIL V +D +G +ITITNEK RLS+ EI++ ++EAE++ ED K
Sbjct: 506 EVTFEVDANGILNVKAEDKASGKSEKITITNEKGRLSQEEIDRMVKEAEEFAEEDKKVKE 565
Query: 542 KAKVISALDSCVYNIKSALNMKD-VDLILSPQESEQINNAIIVAMNLLEMNKNNQHIEID 600
K +AL++ VYN+K+ ++ KD + L E E+I A A+ L+ N+N++ E D
Sbjct: 566 KIDARNALETYVYNMKNQVSDKDKLADKLEGDEKEKIEAATKEALEWLDENQNSEKEEYD 625
Query: 601 VLEHHLKEMNSMLNMLLKIV 620
LKE+ ++ N ++ V
Sbjct: 626 ---EKLKEVEAVCNPIITAV 642
Score = 35.0 bits (79), Expect = 0.15
Identities = 26/87 (29%), Positives = 44/87 (49%), Gaps = 4/87 (4%)
Query: 621 EDMKFLRKATVMSALDSCVKNITNALNTKD-VNLIISPQESEKINNAIFGAMNLLDKNKN 679
ED K K +AL++ V N+ N ++ KD + + E EKI A A+ LD+N+N
Sbjct: 559 EDKKVKEKIDARNALETYVYNMKNQVSDKDKLADKLEGDEKEKIEAATKEALEWLDENQN 618
Query: 680 NQQINIDVLEHHLKELTNMLKMLLKIV 706
+++ D LKE+ + ++ V
Sbjct: 619 SEKEEYD---EKLKEVEAVCNPIITAV 642
>At5g09590 heat shock protein 70 (Hsc70-5)
Length = 682
Score = 421 bits (1083), Expect = e-118
Identities = 255/561 (45%), Positives = 352/561 (62%), Gaps = 31/561 (5%)
Query: 10 VGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAF-TDYQRLIGNGAENQSATNP 68
+GIDLGTT SCVAV + ++I N +G RTTPSVVAF T + L+G A+ Q+ TNP
Sbjct: 60 IGIDLGTTNSCVAVM--EGKNPKVIENAEGARTTPSVVAFNTKGELLVGTPAKRQAVTNP 117
Query: 69 ENTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYKGQEKQFCAEEISSA 128
NTV KRLIGRK+ DP QK++ + P+ + N + + +Q+ +I +
Sbjct: 118 TNTVSGTKRLIGRKFDDPQTQKEMKMVPYKIVRAPNGDAWV----EANGQQYSPSQIGAF 173
Query: 129 VLTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILNEPTAAAIA 188
+LTKM+E AEAYLG V VVTVPAYFND+QR+AT DAG IAGL+V RI+NEPTAAA++
Sbjct: 174 ILTKMKETAEAYLGKSVTKAVVTVPAYFNDAQRQATKDAGRIAGLDVERIINEPTAAALS 233
Query: 189 YGLDKRSNYDGKRNILVFDLGGGTFDVSILTIKGDVFDVKATAGNTHLGGEDFDNRMVYY 248
YG+ + I VFDLGGGTFDVS+L I VF+VKAT G+T LGGEDFDN ++ +
Sbjct: 234 YGMTNKEGL-----IAVFDLGGGTFDVSVLEISNGVFEVKATNGDTFLGGEDFDNALLDF 288
Query: 249 FVEEFKKKNKVEISGNPKSLRRLRTACERAKRILSFTFVTTVEVDALFMGID------FS 302
V EFK ++++ + +L+RLR A E+AK LS T T E++ F+ D F+
Sbjct: 289 LVNEFKTTEGIDLAKDRLALQRLREAAEKAKIELSSTSQT--EINLPFITADASGAKHFN 346
Query: 303 SSITRAKFEEINMDFFNECINIVDSCLRDSKIYKNDIDDVVLVGGSSRIPKVQDLLLEFF 362
++TR++FE + + +CL+D+ I ++D+V+LVGG +R+PKVQ ++ E F
Sbjct: 347 ITLTRSRFETLVNHLIERTRDPCKNCLKDAGISAKEVDEVLLVGGMTRVPKVQSIVAEIF 406
Query: 363 KGKALFMNINPDEAIAYGAAVQAAILSEDFKNVPNLVLRDVTPLSLGIATATYWENIMDV 422
GK+ +NPDEA+A GAA+Q IL D K L+L DVTPLSLGI T +
Sbjct: 407 -GKSPSKGVNPDEAVAMGAALQGGILRGDVK---ELLLLDVTPLSLGIETL---GGVFTR 459
Query: 423 VIPRNTSIPVKNTKGYCTAIDN-CGASIIVYEGERPRASDNNLLGLFTLSCL-PGPRGQP 480
+I RNT+IP K ++ + TA DN I V +GER A+DN LLG F L + P PRG P
Sbjct: 460 LITRNTTIPTKKSQVFSTAADNQTQVGIRVLQGEREMATDNKLLGEFDLVGIPPSPRGVP 519
Query: 481 -LEVCFSIDENGILTVSGKDITTGILNEITITNEKERLSKFEIEKKIEEAEKYRVEDMKF 539
+EV F ID NGI+TVS KD TTG + +ITI LS+ +I+K + EAE + +D +
Sbjct: 520 QIEVTFDIDANGIVTVSAKDKTTGKVQQITI-RSSGGLSEDDIQKMVREAELHAQKDKER 578
Query: 540 LRKAKVISALDSCVYNIKSAL 560
+ D+ +Y+I+ +L
Sbjct: 579 KELIDTKNTADTTIYSIEKSL 599
>At4g37910 heat shock protein 70 like protein
Length = 682
Score = 421 bits (1083), Expect = e-118
Identities = 250/562 (44%), Positives = 354/562 (62%), Gaps = 31/562 (5%)
Query: 10 VGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDY-QRLIGNGAENQSATNP 68
+GIDLGTT SCV+V + +I N +G+RTTPSVVA + L+G A+ Q+ TNP
Sbjct: 55 IGIDLGTTNSCVSVM--EGKTARVIENAEGSRTTPSVVAMNQKGELLVGTPAKRQAVTNP 112
Query: 69 ENTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYKGQEKQFCAEEISSA 128
NT+F +KRLIGR++ DP QK++ + P+ + N + + ++F +I +
Sbjct: 113 TNTIFGSKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWV----EANGQKFSPSQIGAN 168
Query: 129 VLTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILNEPTAAAIA 188
VLTKM+E AEAYLG + VVTVPAYFND+QR+AT DAG IAGL+V RI+NEPTAAA++
Sbjct: 169 VLTKMKETAEAYLGKSINKAVVTVPAYFNDAQRQATKDAGKIAGLDVQRIINEPTAAALS 228
Query: 189 YGLDKRSNYDGKRNILVFDLGGGTFDVSILTIKGDVFDVKATAGNTHLGGEDFDNRMVYY 248
YG++ + I VFDLGGGTFDVSIL I VF+VKAT G+T LGGEDFDN ++ Y
Sbjct: 229 YGMNNKEGV-----IAVFDLGGGTFDVSILEISSGVFEVKATNGDTFLGGEDFDNTLLEY 283
Query: 249 FVEEFKKKNKVEISGNPKSLRRLRTACERAKRILSFTFVTTVEVDALFMGIDFSS----- 303
V EFK+ + ++++ + +L+RLR A E+AK LS T T E++ F+ D S
Sbjct: 284 LVNEFKRSDNIDLTKDNLALQRLREAAEKAKIELSST--TQTEINLPFITADASGAKHLN 341
Query: 304 -SITRAKFEEINMDFFNECINIVDSCLRDSKIYKNDIDDVVLVGGSSRIPKVQDLLLEFF 362
++TR+KFE + + +CL+D+ + ++D+V+LVGG +R+PKVQ+++ E F
Sbjct: 342 ITLTRSKFEGLVGKLIERTRSPCQNCLKDAGVTIKEVDEVLLVGGMTRVPKVQEIVSEIF 401
Query: 363 KGKALFMNINPDEAIAYGAAVQAAILSEDFKNVPNLVLRDVTPLSLGIATATYWENIMDV 422
GK+ +NPDEA+A GAA+Q IL D K +L+L DV PLSLGI T +
Sbjct: 402 -GKSPCKGVNPDEAVAMGAAIQGGILRGDVK---DLLLLDVVPLSLGIETL---GAVFTK 454
Query: 423 VIPRNTSIPVKNTKGYCTAIDN-CGASIIVYEGERPRASDNNLLGLFTLSCL-PGPRGQP 480
+IPRNT+IP K ++ + TA DN I V +GER A+DN +LG F L + P PRG P
Sbjct: 455 LIPRNTTIPTKKSQVFSTAADNQMQVGIKVLQGEREMAADNKVLGEFDLVGIPPAPRGMP 514
Query: 481 -LEVCFSIDENGILTVSGKDITTGILNEITITNEKERLSKFEIEKKIEEAEKYRVEDMKF 539
+EV F ID NGI TVS KD TG ITI LS EI + ++EAE +D +
Sbjct: 515 QIEVTFDIDANGITTVSAKDKATGKEQNITI-RSSGGLSDDEINRMVKEAELNAQKDQEK 573
Query: 540 LRKAKVISALDSCVYNIKSALN 561
+ + ++ D+ +Y+++ +L+
Sbjct: 574 KQLIDLRNSADTTIYSVEKSLS 595
>At4g24280 hsp 70-like protein
Length = 718
Score = 402 bits (1032), Expect = e-112
Identities = 247/561 (44%), Positives = 344/561 (61%), Gaps = 26/561 (4%)
Query: 8 RAVGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDY-QRLIGNGAENQSAT 66
+ VGIDLGTT S VA + + I+ N +G RTTPSVVA+T RL+G A+ Q+
Sbjct: 79 KVVGIDLGTTNSAVAAM--EGGKPTIVTNAEGQRTTPSVVAYTKSGDRLVGQIAKRQAVV 136
Query: 67 NPENTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYKGQEKQFCAEEIS 126
NPENT F KR IGRK ++ V ++ + V N+ + ++ KQF AEEIS
Sbjct: 137 NPENTFFSVKRFIGRKMNE--VDEESKQVSYRVVRDENNN--VKLECPAINKQFAAEEIS 192
Query: 127 SAVLTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILNEPTAAA 186
+ VL K+ + A +L V V+TVPAYFNDSQR AT DAG IAGL V+RI+NEPTAA+
Sbjct: 193 AQVLRKLVDDASRFLNDKVTKAVITVPAYFNDSQRTATKDAGRIAGLEVLRIINEPTAAS 252
Query: 187 IAYGLDKRSNYDGKRNILVFDLGGGTFDVSILTIKGDVFDVKATAGNTHLGGEDFDNRMV 246
+AYG D+++N ILVFDLGGGTFDVS+L + VF+V +T+G+THLGG+DFD R+V
Sbjct: 253 LAYGFDRKAN----ETILVFDLGGGTFDVSVLEVGDGVFEVLSTSGDTHLGGDDFDKRVV 308
Query: 247 YYFVEEFKKKNKVEISGNPKSLRRLRTACERAKRILSFTFVTTVEVDALFMGID----FS 302
+ EFKK +++ + ++L+RL A E+AK LS T + + + D
Sbjct: 309 DWLAAEFKKDEGIDLLKDKQALQRLTEAAEKAKIELSSLTQTNMSLPFITATADGPKHIE 368
Query: 303 SSITRAKFEEINMDFFNECINIVDSCLRDSKIYKNDIDDVVLVGGSSRIPKVQDLLLEFF 362
+++TRAKFEE+ D + V++ LRD+K+ DID+V+LVGGS+RIP VQ+L+ +
Sbjct: 369 TTLTRAKFEELCSDLLDRVRTPVENSLRDAKLSFKDIDEVILVGGSTRIPAVQELVRK-V 427
Query: 363 KGKALFMNINPDEAIAYGAAVQAAILSEDFKNVPNLVLRDVTPLSLGIATATYWENIMDV 422
GK + +NPDE +A GAAVQA +L+ D V ++VL DVTPLS+G+ T +M
Sbjct: 428 TGKEPNVTVNPDEVVALGAAVQAGVLAGD---VSDIVLLDVTPLSIGLETL---GGVMTK 481
Query: 423 VIPRNTSIPVKNTKGYCTAID-NCGASIIVYEGERPRASDNNLLGLFTLSCL-PGPRGQP 480
+IPRNT++P ++ + TA D I V +GER DN LG F L + P PRG P
Sbjct: 482 IIPRNTTLPTSKSEVFSTAADGQTSVEINVLQGEREFVRDNKSLGSFRLDGIPPAPRGVP 541
Query: 481 -LEVCFSIDENGILTVSGKDITTGILNEITITNEKERLSKFEIEKKIEEAEKYRVEDMKF 539
+EV F ID NGIL+VS D TG +ITIT L K E+++ ++EAE++ +D +
Sbjct: 542 QIEVKFDIDANGILSVSAVDKGTGKKQDITITG-ASTLPKDEVDQMVQEAERFAKDDKEK 600
Query: 540 LRKAKVISALDSCVYNIKSAL 560
+ DS VY + L
Sbjct: 601 RDAIDTKNQADSVVYQTEKQL 621
>At5g49910 heat shock protein 70 (Hsc70-7)
Length = 718
Score = 400 bits (1028), Expect = e-111
Identities = 248/561 (44%), Positives = 341/561 (60%), Gaps = 26/561 (4%)
Query: 8 RAVGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDYQ-RLIGNGAENQSAT 66
+ VGIDLGTT S VA + + I+ N +G RTTPSVVA+T + RL+G A+ Q+
Sbjct: 79 KVVGIDLGTTNSAVAAM--EGGKPTIVTNAEGQRTTPSVVAYTKSKDRLVGQIAKRQAVV 136
Query: 67 NPENTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYKGQEKQFCAEEIS 126
NPENT F KR IGR+ ++ V ++ + V N + G KQF AEEIS
Sbjct: 137 NPENTFFSVKRFIGRRMNE--VAEESKQVSYRVIKDENGNVKLDCPAIG--KQFAAEEIS 192
Query: 127 SAVLTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILNEPTAAA 186
+ VL K+ + A +L V V+TVPAYFNDSQR AT DAG IAGL V+RI+NEPTAA+
Sbjct: 193 AQVLRKLVDDASRFLNDKVTKAVITVPAYFNDSQRTATKDAGRIAGLEVLRIINEPTAAS 252
Query: 187 IAYGLDKRSNYDGKRNILVFDLGGGTFDVSILTIKGDVFDVKATAGNTHLGGEDFDNRMV 246
+AYG +++SN ILVFDLGGGTFDVS+L + VF+V +T+G+THLGG+DFD R+V
Sbjct: 253 LAYGFERKSN----ETILVFDLGGGTFDVSVLEVGDGVFEVLSTSGDTHLGGDDFDKRVV 308
Query: 247 YYFVEEFKKKNKVEISGNPKSLRRLRTACERAKRILSFTFVTTVEVDALFMGID----FS 302
+ FKK +++ + ++L+RL A E+AK LS T + + + D
Sbjct: 309 DWLASTFKKDEGIDLLKDKQALQRLTEAAEKAKIELSSLTQTNMSLPFITATADGPKHIE 368
Query: 303 SSITRAKFEEINMDFFNECINIVDSCLRDSKIYKNDIDDVVLVGGSSRIPKVQDLLLEFF 362
+++TR KFEE+ D + V++ LRD+K+ DID+V+LVGGS+RIP VQDL+ +
Sbjct: 369 TTLTRGKFEELCSDLLDRVRTPVENSLRDAKLSFKDIDEVILVGGSTRIPAVQDLVRK-L 427
Query: 363 KGKALFMNINPDEAIAYGAAVQAAILSEDFKNVPNLVLRDVTPLSLGIATATYWENIMDV 422
GK +++NPDE +A GAAVQA +LS D V ++VL DVTPLSLG+ T +M
Sbjct: 428 TGKEPNVSVNPDEVVALGAAVQAGVLSGD---VSDIVLLDVTPLSLGLETL---GGVMTK 481
Query: 423 VIPRNTSIPVKNTKGYCTAID-NCGASIIVYEGERPRASDNNLLGLFTLSCL-PGPRGQP 480
+IPRNT++P ++ + TA D I V +GER DN +G F L + P PRG P
Sbjct: 482 IIPRNTTLPTSKSEVFSTAADGQTSVEINVLQGEREFVRDNKSIGSFRLDGIPPAPRGVP 541
Query: 481 -LEVCFSIDENGILTVSGKDITTGILNEITITNEKERLSKFEIEKKIEEAEKYRVEDMKF 539
+EV F ID NGIL+VS D TG +ITIT L K E++ ++EAE++ ED +
Sbjct: 542 QIEVKFDIDANGILSVSASDKGTGKKQDITITG-ASTLPKDEVDTMVQEAERFAKEDKEK 600
Query: 540 LRKAKVISALDSCVYNIKSAL 560
+ DS VY + L
Sbjct: 601 RDAIDTKNQADSVVYQTEKQL 621
>At1g79930 putative heat-shock protein
Length = 831
Score = 248 bits (634), Expect = 6e-66
Identities = 167/533 (31%), Positives = 272/533 (50%), Gaps = 20/533 (3%)
Query: 10 VGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDYQRLIGNGAENQSATNPE 69
VG D G VAV ++++ ND+ NR TP++V F D QR IG + NP+
Sbjct: 4 VGFDFGNENCLVAV--ARQRGIDVVLNDESNRETPAIVCFGDKQRFIGTAGAASTMMNPK 61
Query: 70 NTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYKGQEKQFCAEEISSAV 129
N++ KRLIGR++SDP +Q+DI PF VT G + P+I Y G+++ F ++ +
Sbjct: 62 NSISQIKRLIGRQFSDPELQRDIKSLPFSVTEGPDGYPLIHANYLGEKRAFTPTQVMGMM 121
Query: 130 LTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILNEPTAAAIAY 189
L+ ++ IAE L + V + + +P YF D QR+A +DA IAGL+ +R+++E TA A+AY
Sbjct: 122 LSNLKGIAEKNLNTAVVDCCIGIPVYFTDLQRRAVLDAATIAGLHPLRLIHETTATALAY 181
Query: 190 GLDKRSNYDGKR-NILVFDLGGGTFDVSILTIKGDVFDVKATAGNTHLGGEDFDNRMVYY 248
G+ K + + N+ D+G + V I K + + A + LGG DFD + +
Sbjct: 182 GIYKTDLPESDQLNVAFIDIGHASMQVCIAGFKKGQLKILSHAFDRSLGGRDFDEVLFNH 241
Query: 249 FVEEFKKKNKVEISGNPKSLRRLRTACERAKRILSFTFVTTVEVDALFMGIDFSSSITRA 308
F +FK + K+++S N K+ RLR CE+ K++LS + + ++ L D I R
Sbjct: 242 FAAKFKDEYKIDVSQNAKASLRLRATCEKLKKVLSANPLAPLNIECLMDEKDVRGVIKRE 301
Query: 309 KFEEINMDFFNECINIVDSCLRDSKIYKNDIDDVVLVGGSSRIPKVQDLLLEFFKGKALF 368
+FEEI++ ++ L D+ + D+ V ++G SR+P + +L EFF GK
Sbjct: 302 EFEEISIPILERVKRPLEKALSDAGLTVEDVHMVEVIGSGSRVPAMIKILTEFF-GKEPR 360
Query: 369 MNINPDEAIAYGAAVQAAILSEDFKNVPNLVLRDVTPLSLGIA---TATYWEN------I 419
+N E ++ G A+Q AILS FK V + + P S+ +A A+ +N
Sbjct: 361 RTMNASECVSRGCALQCAILSPTFK-VREFQVHESFPFSISLAWKGAASEAQNGGAENQQ 419
Query: 420 MDVVIPRNTSIP-VKNTKGYCTAIDNCGASII-VYEGERPRASDNNLLGLFTLSCLPGPR 477
+V P+ IP VK Y + + V + + P +G F S G R
Sbjct: 420 STIVFPKGNPIPSVKALTFYRSGTFSVDVQYSDVNDLQAPPKISTYTIGPFQSS--KGER 477
Query: 478 GQPLEVCFSIDENGILTVSGKDITTGILNEITITNE-KERLSKFEIEKKIEEA 529
+ L+V ++ +GI++V + E+ +T E E +K + +K EA
Sbjct: 478 AK-LKVKVRLNLHGIVSVESATLLEEEEVEVPVTKEHSEETTKMDSDKASAEA 529
>At1g79920 putative heat-shock protein (At1g79920)
Length = 736
Score = 248 bits (632), Expect = 1e-65
Identities = 167/533 (31%), Positives = 272/533 (50%), Gaps = 20/533 (3%)
Query: 10 VGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDYQRLIGNGAENQSATNPE 69
VG D G VAV ++++ ND+ NR TP++V F D QR IG + NP+
Sbjct: 4 VGFDFGNENCLVAV--ARQRGIDVVLNDESNRETPAIVCFGDKQRFIGTAGAASTMMNPK 61
Query: 70 NTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYKGQEKQFCAEEISSAV 129
N++ KRLIGR++SDP +Q+DI PF VT G + P+I Y G+ + F ++ +
Sbjct: 62 NSISQIKRLIGRQFSDPELQRDIKSLPFSVTEGPDGYPLIHANYLGEIRAFTPTQVMGMM 121
Query: 130 LTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILNEPTAAAIAY 189
L+ ++ IAE L + V + + +P YF D QR+A +DA IAGL+ + +++E TA A+AY
Sbjct: 122 LSNLKGIAEKNLNTAVVDCCIGIPVYFTDLQRRAVLDAATIAGLHPLHLIHETTATALAY 181
Query: 190 GLDKRSNYDGKR-NILVFDLGGGTFDVSILTIKGDVFDVKATAGNTHLGGEDFDNRMVYY 248
G+ K + + N+ D+G + V I K + + A + LGG DFD + +
Sbjct: 182 GIYKTDLPENDQLNVAFIDIGHASMQVCIAGFKKGQLKILSHAFDRSLGGRDFDEVLFNH 241
Query: 249 FVEEFKKKNKVEISGNPKSLRRLRTACERAKRILSFTFVTTVEVDALFMGIDFSSSITRA 308
F +FK + K+++S N K+ RLR CE+ K++LS + + ++ L D I R
Sbjct: 242 FAAKFKDEYKIDVSQNAKASLRLRATCEKLKKVLSANPMAPLNIECLMAEKDVRGVIKRE 301
Query: 309 KFEEINMDFFNECINIVDSCLRDSKIYKNDIDDVVLVGGSSRIPKVQDLLLEFFKGKALF 368
+FEEI++ ++ L D+ + D+ V +VG SR+P + +L EFF GK
Sbjct: 302 EFEEISIPILERVKRPLEKALSDAGLTVEDVHMVEVVGSGSRVPAMIKILTEFF-GKEPR 360
Query: 369 MNINPDEAIAYGAAVQAAILSEDFKNVPNLVLRDVTPLSLGIA---TATYWEN------I 419
+N E ++ G A+Q AILS FK V + + P S+ +A AT +N
Sbjct: 361 RTMNASECVSRGCALQCAILSPTFK-VREFQVHESFPFSISLAWKGAATDAQNGGTENQQ 419
Query: 420 MDVVIPRNTSIP-VKNTKGYCTAIDNCGASII-VYEGERPRASDNNLLGLFTLSCLPGPR 477
+V P+ IP VK Y + + V + + P +G F S G R
Sbjct: 420 STIVFPKGNPIPSVKALTFYRSGTFSIDVQYSDVNDLQAPPKISTYTIGPFQSS--KGER 477
Query: 478 GQPLEVCFSIDENGILTVSGKDITTGILNEITIT-NEKERLSKFEIEKKIEEA 529
+ L+V ++ +GI++V + E+++T ++ E +K + +K EA
Sbjct: 478 AK-LKVKVRLNLHGIVSVESATLLEEEEVEVSVTKDQSEETAKMDTDKASAEA 529
>At2g32120 70kD heat shock protein
Length = 563
Score = 238 bits (607), Expect = 9e-63
Identities = 172/504 (34%), Positives = 264/504 (52%), Gaps = 39/504 (7%)
Query: 9 AVGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDYQRLIGNGAENQSATNP 68
A+GID+GT+ +AVW + ++V I+ N + + S V F D + G NQ A
Sbjct: 30 ALGIDIGTSQCSIAVW--NGSQVHILRNTRNQKLIKSFVTFKD--EVPAGGVSNQLAHEQ 85
Query: 69 EN----TVFDAKRLIGRKYSDPIVQKDIMLWPFMV-TSGVNDKPMITVKYKGQEKQFCAE 123
E +F+ KRL+GR +DP+V L PF+V T + +P I + E
Sbjct: 86 EMLTGAAIFNMKRLVGRVDTDPVVHASKNL-PFLVQTLDIGVRPFIAALVNNAWRSTTPE 144
Query: 124 EISSAVLTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILNEPT 183
E+ + L ++R +AEA L PV+NVV+TVP F+ Q A A+AGL+V+R++ EPT
Sbjct: 145 EVLAIFLVELRLMAEAQLKRPVRNVVLTVPVSFSRFQLTRFERACAMAGLHVLRLMPEPT 204
Query: 184 AAAIAYGLDKR-SNYDG-----KRNILVFDLGGGTFDVSILTIKGDVFDVKATAGNTHLG 237
A A+ Y ++ + +D +R ++F++G G DV++ G V +KA AG+ +G
Sbjct: 205 AIALLYAQQQQMTTHDNMGSGSERLAVIFNMGAGYCDVAVTATAGGVSQIKALAGSP-IG 263
Query: 238 GEDFDNRMVYYFVEEFKKKNKVEISGNPKSLRRLRTACERAKRILSFTFVTTVEVDALFM 297
GED + + N ++ LR A + A L+ +EVD L
Sbjct: 264 GEDILQNTIRHIAPP-----------NEEASGLLRVAAQDAIHRLTDQENVQIEVD-LGN 311
Query: 298 GIDFSSSITRAKFEEINMDFFNECINIVDSCLRDSKIYKNDIDDVVLVGGSSRIPKVQDL 357
G S + R +FEE+N F EC +V CLRD+++ DIDD+++VGG S IPKV+ +
Sbjct: 312 GNKISKVLDRLEFEEVNQKVFEECERLVVQCLRDARVNGGDIDDLIMVGGCSYIPKVRTI 371
Query: 358 LLEFFKGKALFMNINPDEAIAYGAAVQAAILS---EDFKNVPNLVLRDVTPLSLGIATAT 414
+ K ++ +NP EA GAA++ A+ S + F ++ L ++ TPL++G+
Sbjct: 372 IKNVCKKDEIYKGVNPLEAAVRGAALEGAVTSGIHDPFGSLDLLTIQ-ATPLAVGVRAN- 429
Query: 415 YWENIMDVVIPRNTSIPVKNTKGYCTAIDN-CGASIIVYEGERPRASDNNLLGLFTLSCL 473
N VIPRNT +P + + T DN A II+YEGE +N+LLG F L +
Sbjct: 430 --GNKFIPVIPRNTMVPARKDLFFTTVQDNQKEALIIIYEGEGETVEENHLLGYFKLVGI 487
Query: 474 -PGPRGQP-LEVCFSIDENGILTV 495
P P+G P + VC ID + L V
Sbjct: 488 PPAPKGVPEINVCMDIDASNALRV 511
>At1g11660 heat-shock protein like
Length = 773
Score = 229 bits (584), Expect = 4e-60
Identities = 132/405 (32%), Positives = 223/405 (54%), Gaps = 6/405 (1%)
Query: 10 VGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDYQRLIGNGAENQSATNPE 69
VG D+G +AV ++++ ND+ NR P++V+F + QR +G A + +P+
Sbjct: 4 VGFDVGNENCVIAV--AKQRGIDVLLNDESNRENPAMVSFGEKQRFMGAAAAASATMHPK 61
Query: 70 NTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYKGQEKQFCAEEISSAV 129
+T+ KRLIGRK+ +P VQ D+ L+PF + + I ++Y G+ + F +I +
Sbjct: 62 STISQLKRLIGRKFREPDVQNDLRLFPFETSEDSDGGIQIRLRYMGEIQSFSPVQILGML 121
Query: 130 LTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILNEPTAAAIAY 189
L+ +++IAE L +PV + V+ +P+YF +SQR A +DA AIAGL +R++++ TA A+ Y
Sbjct: 122 LSHLKQIAEKSLKTPVSDCVIGIPSYFTNSQRLAYLDAAAIAGLRPLRLMHDSTATALGY 181
Query: 190 GLDKRS--NYDGKRNILVFDLGGGTFDVSILTIKGDVFDVKATAGNTHLGGEDFDNRMVY 247
G+ K I+ D+G V + + + V++ A + +LGG DFD +
Sbjct: 182 GIYKTDLVANSSPTYIVFIDIGHCDTQVCVASFESGSMRVRSHAFDRNLGGRDFDEVLFN 241
Query: 248 YFVEEFKKKNKVEISGNPKSLRRLRTACERAKRILSFTFVTTVEVDALFMGIDFSSSITR 307
+F EFK+K +++ N K+ RLR +CE+ K++LS + ++ L D S I R
Sbjct: 242 HFALEFKEKYNIDVYTNTKACVRLRASCEKVKKVLSANAEAQLNIECLMEEKDVRSFIKR 301
Query: 308 AKFEEINMDFFNECINIVDSCLRDSKIYKNDIDDVVLVGGSSRIPKVQDLLLEFFKGKAL 367
+FE+++ I L DS + + I V LVG SRIP + +L FK + L
Sbjct: 302 EEFEQLSAGLLERLIVPCQKALADSGLSLDQIHSVELVGSGSRIPAISKMLSSLFK-REL 360
Query: 368 FMNINPDEAIAYGAAVQAAILSEDFKNVPNLVLRDVTPLSLGIAT 412
+N E +A G A+Q A+LS F+ V + ++D P ++G ++
Sbjct: 361 GRTVNASECVARGCALQCAMLSPVFR-VRDYEVQDSYPFAIGFSS 404
>At4g40020 putative protein
Length = 615
Score = 32.7 bits (73), Expect = 0.72
Identities = 27/110 (24%), Positives = 55/110 (49%), Gaps = 5/110 (4%)
Query: 515 ERLSKFEIEK--KIEEAEKYRVEDMKFLRKAKVISALDSCVYNIKSALNMKDVDLILSPQ 572
E LSK E+ K+ + K + + ++A I+ ++ N+K AL K+ +L + +
Sbjct: 285 ENLSKKAKEENHKVRDILKQAINEANVAKEAAGIARAENS--NLKDALLDKEEELQFALK 342
Query: 573 ESEQIN-NAIIVAMNLLEMNKNNQHIEIDVLEHHLKEMNSMLNMLLKIVE 621
E E++ N + N+ ++ K IE+ + E + +N +M ++VE
Sbjct: 343 EIERVKVNEAVANDNIKKLKKMLSEIEVAMEEEKQRSLNRQESMPKEVVE 392
>At4g25250 unknown protein
Length = 199
Score = 32.7 bits (73), Expect = 0.72
Identities = 16/49 (32%), Positives = 29/49 (58%)
Query: 579 NAIIVAMNLLEMNKNNQHIEIDVLEHHLKEMNSMLNMLLKIVEDMKFLR 627
NA +V NLL+ K + E+ +L+ + EM ++ L + V +MK++R
Sbjct: 83 NATLVVSNLLQKAKAAKSHEVSILKDCVDEMKDTIDELKQAVAEMKYVR 131
>At3g17360 kinesin-like protein
Length = 2008
Score = 32.7 bits (73), Expect = 0.72
Identities = 37/191 (19%), Positives = 83/191 (43%), Gaps = 21/191 (10%)
Query: 477 RGQPLEVCFSIDENGILTVSGK---------DITTGILN-EITITNEKERLSKFEIEKKI 526
R + LE ++ + I T++ + D+ +L ++ IT+ E + + ++++ +
Sbjct: 1792 RVEELESLLAVKQKEICTLNTRIAAADSMTHDVIRDLLGVKMDITSYAELIDQHQVQRVV 1851
Query: 527 EEAEKYRVE----DMKFLRKAKVISALDSCVYNIKSALNMKDVDLILSPQESEQINNAII 582
E+A+++ E + + + + I L + S LN KD D++ + +Q+ +
Sbjct: 1852 EKAQQHAEEILSKEQEVMNLKRHIDYLFKDRESCMSELNKKDTDVLATQISLDQLQERVQ 1911
Query: 583 VAMNLLEMNKNNQH------IEIDVLEHHLKEMNSMLNMLLKIVEDMKFLRKATVMSALD 636
+ EM KN++ E+D H+ + N + K K L L+
Sbjct: 1912 LLSMQNEMLKNDKSNLLRKLAELDRTVHNAQASNHRVPQTTKDTASFK-LADTDYTKRLE 1970
Query: 637 SCVKNITNALN 647
+ K +++A N
Sbjct: 1971 NAQKLLSHANN 1981
Score = 29.3 bits (64), Expect = 8.0
Identities = 40/178 (22%), Positives = 78/178 (43%), Gaps = 32/178 (17%)
Query: 525 KIEEAEKYRVEDMKFLRKAKVISALDSCVYNIKSALNMKDVDLILSPQESEQINNAIIVA 584
K+EE E+ R E++KF K + +YN + + +L P +
Sbjct: 274 KMEEEER-RDENLKFSCKCSFLE-----IYN-------EQITDLLEPSSTN--------- 311
Query: 585 MNLLEMNKNNQHIEIDVLEHHLKEMNSMLNMLLKIVEDMKFLRKATVMSALDSCVKNITN 644
+ L E ++E +++EH+++ ++ +L +LL+ + K AT M++ S ++
Sbjct: 312 LQLREDLGKGVYVE-NLVEHNVRTVSDVLKLLLQGATNRKIA--ATRMNSESSRSHSVFT 368
Query: 645 ALNTKDVNLIISPQESEKINNAIFGAMNLLDKNKNNQQINIDVLEHHLKELTNMLKML 702
I S E + + + F +NL+D + +Q + LKE N+ K L
Sbjct: 369 CT-------IESLWEKDSLTRSRFARLNLVDLAGSERQKSSGAEGDRLKEAANINKSL 419
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.136 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,392,274
Number of Sequences: 26719
Number of extensions: 672763
Number of successful extensions: 2127
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 19
Number of HSP's successfully gapped in prelim test: 22
Number of HSP's that attempted gapping in prelim test: 1973
Number of HSP's gapped (non-prelim): 66
length of query: 713
length of database: 11,318,596
effective HSP length: 106
effective length of query: 607
effective length of database: 8,486,382
effective search space: 5151233874
effective search space used: 5151233874
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 64 (29.3 bits)
Medicago: description of AC137996.5


