

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137996.10 + phase: 0 /pseudo
(1594 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
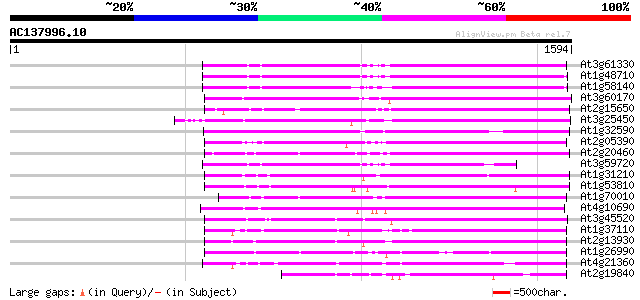
Score E
Sequences producing significant alignments: (bits) Value
At3g61330 copia-type polyprotein 684 0.0
At1g48710 hypothetical protein 681 0.0
At1g58140 hypothetical protein 674 0.0
At3g60170 putative protein 650 0.0
At2g15650 putative retroelement pol polyprotein 647 0.0
At3g25450 hypothetical protein 602 e-172
At1g32590 hypothetical protein, 5' partial 595 e-170
At2g05390 putative retroelement pol polyprotein 551 e-156
At2g20460 putative retroelement pol polyprotein 543 e-154
At3g59720 copia-type reverse transcriptase-like protein 534 e-151
At1g31210 putative reverse transcriptase 533 e-151
At1g53810 523 e-148
At1g70010 hypothetical protein 517 e-146
At4g10690 retrotransposon like protein 514 e-145
At3g45520 copia-like polyprotein 501 e-141
At1g37110 499 e-141
At2g13930 putative retroelement pol polyprotein 486 e-137
At1g26990 polyprotein, putative 483 e-136
At4g21360 putative transposable element 433 e-121
At2g19840 copia-like retroelement pol polyprotein 421 e-117
>At3g61330 copia-type polyprotein
Length = 1352
Score = 684 bits (1765), Expect = 0.0
Identities = 381/1039 (36%), Positives = 578/1039 (54%), Gaps = 42/1039 (4%)
Query: 549 REKQRSWYLDSGCSRHMTGEKALFLTLTMKDGGEVKFGGNQTGKIIGTGTI-----GNSS 603
+++ WYLDSG S HM G K++F L G V G ++ G G I
Sbjct: 329 QKENHKWYLDSGASNHMCGRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRLKNGDH 388
Query: 604 ISINNVWLVDGLKHNLLSISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKIN 663
I+NV+ + +K N+LS+ Q + GYD+ N ++ +++ IT ++ +N
Sbjct: 389 QFISNVYYIPSMKTNILSLGQLLEKGYDIRLKDNNLSIRDQESNLITKVPMSKNRMFVLN 448
Query: 664 F-SDLADQKVVCLLSMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGA 722
+D+A +C ++ W+WH R GH N+ + +S+ ++V+GLP I+ H + +C
Sbjct: 449 IRNDIAQCLKMCY---KEESWLWHLRFGHLNFGGLELLSRKEMVRGLPCIN-HPNQVCEG 504
Query: 723 CQKGKIVKSSFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWVKF 782
C GK K SF + +PLEL+H D+ GP+ SL S Y L+ +DD+SR TWV F
Sbjct: 505 CLLGKQFKMSFPKESSSRAQKPLELIHTDVCGPIKPKSLGKSNYFLLFIDDFSRKTWVYF 564
Query: 783 IKSKDYACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRT 842
+K K E+F F ++ E L I +RSD GGEF ++ F +CE +GI + + PR+
Sbjct: 565 LKEKSEVFEIFKKFKAHVEKESGLVIKTMRSDRGGEFTSKEFLKYCEDNGIRRQLTVPRS 624
Query: 843 PQQNGVVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYELF 902
PQQNGVVERKNRT+ EMAR+M+ L K WAEAV + Y+ NR + + KT E +
Sbjct: 625 PQQNGVVERKNRTILEMARSMLKSKRLPKELWAEAVACAVYLLNRSPTKSVSGKTPQEAW 684
Query: 903 KGRRPNISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVEE 962
GR+P +S+ FG + + K D K+++ IF+GY SK Y++YN +T+
Sbjct: 685 SGRKPGVSHLRVFGSIAHAHVPDEKRSKLDDKSEKYIFIGYDNNSKGYKLYNPDTKKTII 744
Query: 963 SMHVKFDDREPGSKTSEQSESNAGTTDSEDASESDQPSDSEKYTEVESSPEAEYNSEAEP 1022
S ++ FD+ S + + N ED E P+ E +E ++P P
Sbjct: 745 SRNIVFDEEGEWDWNSNEEDYNFFPHFEEDEPE---PTREEPPSEEPTTPPTS------P 795
Query: 1023 SPKVQNESASEDFQDNTQQVIQPKFKHKSSHPEELIIGSKDSPRRTRSHFRQEESLIGLL 1082
+ ES+SE P+F+ +EL +++ T L L
Sbjct: 796 TSSQIEESSSER---------TPRFRSI----QELYEVTENQENLT---------LFCLF 833
Query: 1083 SIIEPKTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTLLEQNGYSKTS*MNKE 1142
+ EP ++A+ W AM EE+ + + P + K +K
Sbjct: 834 AECEPMDFQKAIEKKTWRNAMDEEIKSIQKNDTWELTSLPNGHKAIGVKWVYKAKKNSKG 893
Query: 1143 K*PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQMDVKSAFLN 1202
+ K VA+GYSQ+ GIDY E FAPVARLE +RL++S A + ++QMDVKSAFLN
Sbjct: 894 EVERYKARLVAKGYSQRVGIDYDEVFAPVARLETVRLIISLAAQNKWKIHQMDVKSAFLN 953
Query: 1203 GVIEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDFERGQVD 1262
G +EEEVY++QP G+ D V +LKK LYGLKQAPRAW R+ + + DF + +
Sbjct: 954 GDLEEEVYIEQPQGYIVKGEEDKVLRLKKVLYGLKQAPRAWNTRIDKYFKEKDFIKCPYE 1013
Query: 1263 TTLFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELKFFLGIQIN 1322
L+ + K+DILI +YVDD+IF N S+ + F K M EFEM+ +G + ++LGI++
Sbjct: 1014 HALYIKIQKEDILIACLYVDDLIFTGNNPSIFEEFKKEMTKEFEMTDIGLMSYYLGIEVK 1073
Query: 1323 QSKEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGTVVDQKLYRGMIGSL 1382
Q G+++ Q Y KE+LKKFK++D + TPM LSK++ G VD ++ ++GSL
Sbjct: 1074 QEDNGIFITQEGYAKEVLKKFKIDDSNPVCTPMECGIKLSKKEEGEGVDPTTFKSLVGSL 1133
Query: 1383 LYLTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGLLYRKSLDYKLIGF 1442
YLT +RP IL++V + +R+ P +H A KRI RY+KGT N GL Y + DYKL+G+
Sbjct: 1134 RYLTCTRPDILYAVGVVSRYMEHPTTTHFKAAKRILRYIKGTVNFGLHYSTTSDYKLVGY 1193
Query: 1443 CDADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEYISAASCCTQLLWMK 1502
D+D+ GD +RKSTSG ++G+ +W SK+Q + +ST EAEY++A SC +W++
Sbjct: 1194 SDSDWGGDVDDRKSTSGFVFYIGDTAFTWMSKKQPIVTLSTCEAEYVAATSCVCHAIWLR 1253
Query: 1503 HQLEDYQI-NANSIPIYCDNTAAICLSKNPILHSRAKHIEIKHHFIRDYVQKGILDIQFI 1561
+ L++ + I+ DN +AI L+KNP+ H R+KHI+ ++H+IR+ V K + ++++
Sbjct: 1254 NLLKELSLPQEEPTKIFVDNKSAIALAKNPVFHDRSKHIDTRYHYIRECVSKKDVQLEYV 1313
Query: 1562 DIEHQWADIFTKPLSVERF 1580
Q AD FTKPL E F
Sbjct: 1314 KTHDQVADFFTKPLKRENF 1332
>At1g48710 hypothetical protein
Length = 1352
Score = 681 bits (1758), Expect = 0.0
Identities = 383/1043 (36%), Positives = 580/1043 (54%), Gaps = 44/1043 (4%)
Query: 549 REKQRSWYLDSGCSRHMTGEKALFLTLTMKDGGEVKFGGNQTGKIIGTGTI-----GNSS 603
+E+ WYLDSG S HM G K++F L G V G ++ G G I
Sbjct: 329 QEENHKWYLDSGASNHMCGRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRLKNGDH 388
Query: 604 ISINNVWLVDGLKHNLLSISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKIN 663
I+NV+ + +K N+LS+ Q + GYD+ N ++ +++ IT ++ +N
Sbjct: 389 QFISNVYYIPSMKTNILSLGQLLEKGYDIRLKDNNLSIRDQESNLITKVPMSKNRMFVLN 448
Query: 664 F-SDLADQKVVCLLSMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGA 722
+D+A +C ++ W+WH R GH N+ + +S+ ++V+GLP I+ H + +C
Sbjct: 449 IRNDIAQCLKMCY---KEESWLWHLRFGHLNFGGLELLSRKEMVRGLPCIN-HPNQVCEG 504
Query: 723 CQKGKIVKSSFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWVKF 782
C GK K SF + + LEL+H D+ GP+ SL S Y L+ +DD+SR TWV F
Sbjct: 505 CLLGKQFKMSFPKESSSRAQKSLELIHTDVCGPIKPKSLGKSNYFLLFIDDFSRKTWVYF 564
Query: 783 IKSKDYACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRT 842
+K K E+F F ++ E L I +RSD GGEF ++ F +CE +GI + + PR+
Sbjct: 565 LKEKSEVFEIFKKFKAHVEKESGLVIKTMRSDRGGEFTSKEFLKYCEDNGIRRQLTVPRS 624
Query: 843 PQQNGVVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYELF 902
PQQNGV ERKNRT+ EMAR+M+ L K WAEAV + Y+ NR + + KT E +
Sbjct: 625 PQQNGVAERKNRTILEMARSMLKSKRLPKELWAEAVACAVYLLNRSPTKSVSGKTPQEAW 684
Query: 903 KGRRPNISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVEE 962
GR+ +S+ FG + + K D K+++ IF+GY SK Y++YN +T+
Sbjct: 685 SGRKSGVSHLRVFGSIAHAHVPDEKRSKLDDKSEKYIFIGYDNNSKGYKLYNPDTKKTII 744
Query: 963 SMHVKFDDREPGSKTSEQSESNAGTTDSEDASESDQPSDSEKYTEVESSPEAEYNSEAEP 1022
S ++ FD+ S + + N ED E P+ E +E ++P P
Sbjct: 745 SRNIVFDEEGEWDWNSNEEDYNFFPHFEEDEPE---PTREEPPSEEPTTPPTS------P 795
Query: 1023 SPKVQNESASEDFQDNTQQVIQPKFKHKSSHPEELIIGSKDSPRRTRSHFRQEESLIGLL 1082
+ ES+SE P+F+ +EL +++ T L L
Sbjct: 796 TSSQIEESSSER---------TPRFRSI----QELYEVTENQENLT---------LFCLF 833
Query: 1083 SIIEPKTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTLLEQNGYSKTS*MNKE 1142
+ EP +EA+ W AM EE+ + + P + K +K
Sbjct: 834 AECEPMDFQEAIEKKTWRNAMDEEIKSIQKNDTWELTSLPNGHKTIGVKWVYKAKKNSKG 893
Query: 1143 K*PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQMDVKSAFLN 1202
+ K VA+GY Q+ GIDY E FAPVARLE +RL++S A + ++QMDVKSAFLN
Sbjct: 894 EVERYKARLVAKGYIQRAGIDYDEVFAPVARLETVRLIISLAAQNKWKIHQMDVKSAFLN 953
Query: 1203 GVIEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDFERGQVD 1262
G +EEEVY++QP G+ D V +LKK+LYGLKQAPRAW R+ + + DF + +
Sbjct: 954 GDLEEEVYIEQPQGYIVKGEEDKVLRLKKALYGLKQAPRAWNTRIDKYFKEKDFIKCPYE 1013
Query: 1263 TTLFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELKFFLGIQIN 1322
L+ + K+DILI +YVDD+IF N S+ + F K M EFEM+ +G + ++LGI++
Sbjct: 1014 HALYIKIQKEDILIACLYVDDLIFTGNNPSMFEEFKKEMTKEFEMTDIGLMSYYLGIEVK 1073
Query: 1323 QSKEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGTVVDQKLYRGMIGSL 1382
Q G+++ Q Y KE+LKKFK++D + TPM LSK++ G VD ++ ++GSL
Sbjct: 1074 QEDNGIFITQEGYAKEVLKKFKMDDSNPVCTPMECGIKLSKKEEGEGVDPTTFKSLVGSL 1133
Query: 1383 LYLTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGLLYRKSLDYKLIGF 1442
YLT +RP IL++V + +R+ P +H A KRI RY+KGT N GL Y + DYKL+G+
Sbjct: 1134 RYLTCTRPDILYAVGVVSRYMEHPTTTHFKAAKRILRYIKGTVNFGLHYSTTSDYKLVGY 1193
Query: 1443 CDADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEYISAASCCTQLLWMK 1502
D+D+ GD +RKSTSG ++G+ +W SK+Q + +ST EAEY++A SC +W++
Sbjct: 1194 SDSDWGGDVDDRKSTSGFVFYIGDTAFTWMSKKQPIVVLSTCEAEYVAATSCVCHAIWLR 1253
Query: 1503 HQLEDYQI-NANSIPIYCDNTAAICLSKNPILHSRAKHIEIKHHFIRDYVQKGILDIQFI 1561
+ L++ + I+ DN +AI L+KNP+ H R+KHI+ ++H+IR+ V K + ++++
Sbjct: 1254 NLLKELSLPQEEPTKIFVDNKSAIALAKNPVFHDRSKHIDTRYHYIRECVSKKDVQLEYV 1313
Query: 1562 DIEHQWADIFTKPLSVERFDFIK 1584
Q ADIFTKPL +R DFIK
Sbjct: 1314 KTHDQVADIFTKPL--KREDFIK 1334
>At1g58140 hypothetical protein
Length = 1320
Score = 674 bits (1738), Expect = 0.0
Identities = 383/1043 (36%), Positives = 569/1043 (53%), Gaps = 76/1043 (7%)
Query: 549 REKQRSWYLDSGCSRHMTGEKALFLTLTMKDGGEVKFGGNQTGKIIGTGTI-----GNSS 603
+E+ WYLDSG S HM G K++F L G V G ++ G G I
Sbjct: 329 QEENHKWYLDSGASNHMCGRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRLKNGDH 388
Query: 604 ISINNVWLVDGLKHNLLSISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKIN 663
I+NV+ + +K N+LS+ Q + GYD+ N ++ +++ IT ++ +N
Sbjct: 389 QFISNVYYIPSMKTNILSLGQLLEKGYDIRLKDNNLSIRDQESNLITKVPMSKNRMFVLN 448
Query: 664 F-SDLADQKVVCLLSMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGA 722
+D+A +C ++ W+WH R GH N+ + +S+ ++V+GLP I+ H + +C
Sbjct: 449 IRNDIAQCLKMCY---KEESWLWHLRFGHLNFGGLELLSRKEMVRGLPCIN-HPNQVCEG 504
Query: 723 CQKGKIVKSSFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWVKF 782
C GK K SF + +PLEL+H D+ GP+ SL S Y L+ +DD+SR TWV F
Sbjct: 505 CLLGKQFKMSFPKESSSRAQKPLELIHTDVCGPIKPKSLGKSNYFLLFIDDFSRKTWVYF 564
Query: 783 IKSKDYACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRT 842
+K K E+F F ++ E L I +RSD GGEF ++ F +CE +GI + + PR+
Sbjct: 565 LKEKSEVFEIFKKFKAHVEKESGLVIKTMRSDRGGEFTSKEFLKYCEDNGIRRQLTVPRS 624
Query: 843 PQQNGVVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYELF 902
PQQNGV ERKNRT+ EMAR+M+ L K WAEAV + Y+ NR + + KT E +
Sbjct: 625 PQQNGVAERKNRTILEMARSMLKSKRLPKELWAEAVACAVYLLNRSPTKSVSGKTPQEAW 684
Query: 903 KGRRPNISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVEE 962
GR+P +S+ FG + + K D K+++ IF+GY SK Y++YN +T+
Sbjct: 685 SGRKPGVSHLRVFGSIAHAHVPDEKRSKLDDKSEKYIFIGYDNNSKGYKLYNPDTKKTII 744
Query: 963 SMHVKFDDREPGSKTSEQSESNAGTTDSEDASESDQPSDSEKYTEVESSPEAEYNSEAEP 1022
S ++ FD+ E D S+ E Y P E E +P
Sbjct: 745 SRNIVFDEE----------------------GEWDWNSNEEDYNFF---PHFE---EDKP 776
Query: 1023 SPKVQNESASEDFQDNTQQVIQPKFKHKSSHPEELIIGSKDSPRRTRSHFRQEESLIGLL 1082
P + E SE+ P SS EE
Sbjct: 777 EP-TREEPPSEE-------PTTPPTSPTSSQIEEKC------------------------ 804
Query: 1083 SIIEPKTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTLLEQNGYSKTS*MNKE 1142
EP +EA+ W AM EE+ + + P + K +K
Sbjct: 805 ---EPMDFQEAIEKKTWRNAMDEEIKSIQKNDTWELTSLPNGHKAIGVKWVYKAKKNSKG 861
Query: 1143 K*PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQMDVKSAFLN 1202
+ K VA+GYSQ+ GIDY E FAPVARLE +RL++S A + ++QMDVKSAFLN
Sbjct: 862 EVERYKARLVAKGYSQRAGIDYDEVFAPVARLETVRLIISLAAQNKWKIHQMDVKSAFLN 921
Query: 1203 GVIEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDFERGQVD 1262
G +EEEVY++QP G+ D V +LKK+LYGLKQAPRAW R+ + + DF + +
Sbjct: 922 GDLEEEVYIEQPQGYIVKGEEDKVLRLKKALYGLKQAPRAWNTRIDKYFKEKDFIKCPYE 981
Query: 1263 TTLFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELKFFLGIQIN 1322
L+ + K+DILI +YVDD+IF N S+ + F K M EFEM+ +G + ++LGI++
Sbjct: 982 HALYIKIQKEDILIACLYVDDLIFTGNNPSMFEEFKKEMTKEFEMTDIGLMSYYLGIEVK 1041
Query: 1323 QSKEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGTVVDQKLYRGMIGSL 1382
Q G+++ Q Y KE+LKKFK++D + TPM LSK++ G VD ++ ++GSL
Sbjct: 1042 QEDNGIFITQEGYAKEVLKKFKMDDSNPVCTPMECGIKLSKKEEGEGVDPTTFKSLVGSL 1101
Query: 1383 LYLTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGLLYRKSLDYKLIGF 1442
YLT +RP IL++V + +R+ P +H A KRI RY+KGT N GL Y + DYKL+G+
Sbjct: 1102 RYLTCTRPDILYAVGVVSRYMEHPTTTHFKAAKRILRYIKGTVNFGLHYSTTSDYKLVGY 1161
Query: 1443 CDADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEYISAASCCTQLLWMK 1502
D+D+ GD +RKSTSG ++G+ +W SK+Q + +ST EAEY++A SC +W++
Sbjct: 1162 SDSDWGGDVDDRKSTSGFVFYIGDTAFTWMSKKQPIVTLSTCEAEYVAATSCVCHAIWLR 1221
Query: 1503 HQLEDYQI-NANSIPIYCDNTAAICLSKNPILHSRAKHIEIKHHFIRDYVQKGILDIQFI 1561
+ L++ + I+ DN +AI L+KNP+ H R+KHI+ ++H+IR+ V K + ++++
Sbjct: 1222 NLLKELSLPQEEPTKIFVDNKSAIALAKNPVFHDRSKHIDTRYHYIRECVSKKDVQLEYV 1281
Query: 1562 DIEHQWADIFTKPLSVERFDFIK 1584
Q ADIFTKPL +R DFIK
Sbjct: 1282 KTHDQVADIFTKPL--KREDFIK 1302
>At3g60170 putative protein
Length = 1339
Score = 650 bits (1676), Expect = 0.0
Identities = 376/1061 (35%), Positives = 577/1061 (53%), Gaps = 44/1061 (4%)
Query: 555 WYLDSGCSRHMTGEKALFLTLTMKDGGEVKFGGNQTGKIIGTGTI-----GNSSISINNV 609
W+LDSGCS HMTG K F L VK G + ++G G++ G + + I V
Sbjct: 300 WFLDSGCSNHMTGSKEWFSELEEGFNRTVKLGNDTRMSVVGKGSVKVKVNGVTQV-IPEV 358
Query: 610 WLVDGLKHNLLSISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKINFSDLAD 669
+ V L++NLLS+ Q + G + C + + +I ++ + +
Sbjct: 359 YYVPELRNNLLSLGQLQERGLAILIRDGTCKVYHPSKGAIMETNMSGNRMFFL-LASKPQ 417
Query: 670 QKVVCLLS---MNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGACQKG 726
+ +CL + M+ + +WH R GH N + ++ ++V GLP + + +C C G
Sbjct: 418 KNSLCLQTEEVMDKENHLWHCRFGHLNQEGLKLLAHKKMVIGLPILKATKE-ICAICLTG 476
Query: 727 KIVKSSFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWVKFIKSK 786
K + S K +S L+L+H D+ GP+ S G +Y L +DD++R TWV F+ K
Sbjct: 477 KQHRESMSKKTSWKSSTQLQLVHSDICGPITPISHSGKRYILSFIDDFTRKTWVYFLHEK 536
Query: 787 DYACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRTPQQN 846
A F F ++ E + +R+D GGEF + F FC HGI + ++ TPQQN
Sbjct: 537 SEAFATFKIFKASVEKEIGAFLTCLRTDRGGEFTSNEFGEFCRSHGISRQLTAAFTPQQN 596
Query: 847 GVVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYELFKGRR 906
GV ERKNRT+ R+M+ E + K FW+EA S +IQNR + T E + GR+
Sbjct: 597 GVAERKNRTIMNAVRSMLSERQVPKMFWSEATKWSVHIQNRSPTAAVEGMTPEEAWSGRK 656
Query: 907 PNISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVEESMHV 966
P + YF FGC Y+ K D K+++ +FLG SE SKA+R+Y+ + + S V
Sbjct: 657 PVVEYFRVFGCIGYVHIPDQKRSKLDDKSKKCVFLGVSEESKAWRLYDPVMKKIVISKDV 716
Query: 967 KFDDREPGSKTSEQSESNAGTTDSEDASESDQPSDSEKYTEVESSPEAEYNSEAEPSPKV 1026
FD+ + E+ T + D D EK +EV EP
Sbjct: 717 VFDEDKSWDWDQADVEAKEVTLECGD-------EDDEKNSEV-----------VEPIAVA 758
Query: 1027 QNESASEDFQDNTQQVIQPKFKHKSSHPEELIIGSKDSPRRTRSHFRQEES--------- 1077
D ++ ++ P S P + + P + + E
Sbjct: 759 SPNHVGSDNNVSSSPILAPSSPAPS--PVAAKVTRERRPPGWMADYETGEGEEIEENLSV 816
Query: 1078 -LIGLLSIIEPKTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTLLEQNGYSKT 1136
L+ +++ +P ++A+ D W AM+ E+ + P T + KT
Sbjct: 817 MLLMMMTEADPIQFDDAVKDKIWREAMEHEIESIVKNNTWELTTLPKGFTPIGVKWVYKT 876
Query: 1137 S*MNKEK*PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQMDV 1196
+ + K VA+GY+Q GIDYTE FAPVARL+ +R +L+ + ++Q+DV
Sbjct: 877 KLNEDGEVDKYKARLVAKGYAQCYGIDYTEVFAPVARLDTVRTILAISSQFNWEIFQLDV 936
Query: 1197 KSAFLNGVIEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDF 1256
KSAFL+G ++EEVYV+QP GF + VYKL+K+LYGLKQAPRAWY R+ + +K +F
Sbjct: 937 KSAFLHGELKEEVYVRQPEGFIREGEEEKVYKLRKALYGLKQAPRAWYSRIEAYFLKEEF 996
Query: 1257 ERGQVDTTLFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELKFF 1316
ER + TLF +T +ILIV +YVDD+IF ++ ++C F K M EFEMS +G++K F
Sbjct: 997 ERCPSEHTLFTKTRVGNILIVSLYVDDLIFTGSDKAMCDEFKKSMMLEFEMSDLGKMKHF 1056
Query: 1317 LGIQINQSKEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGTVVDQKLYR 1376
LGI++ QS G+++ Q +Y +E+L +F +++ + P+ P L+K++ G VD+ +++
Sbjct: 1057 LGIEVKQSDGGIFICQRRYAREVLARFGMDESNAVKNPIVPGTKLTKDENGEKVDETMFK 1116
Query: 1377 GMIGSLLYLTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGLLY--RKS 1434
++GSL+YLT +RP +++ VCL +RF S+PR SH A KRI RYLKGT LG+ Y RK+
Sbjct: 1117 QLVGSLMYLTVTRPDLMYGVCLISRFMSNPRMSHWLAAKRILRYLKGTVELGIFYRRRKN 1176
Query: 1435 LDYKLIGFCDADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEYISAASC 1494
KL+ F D+DYAGD +R+STSG + I WASK+Q +A+ST EAEYI+AA C
Sbjct: 1177 RSLKLMAFTDSDYAGDLNDRRSTSGFVFLMASGAICWASKKQPVVALSTTEAEYIAAAFC 1236
Query: 1495 CTQLLWMKHQLEDYQINANSIP-IYCDNTAAICLSKNPILHSRAKHIEIKHHFIRDYVQK 1553
Q +W++ LE S I CDN++ I LSK+P+LH ++KHIE++ H++RD V
Sbjct: 1237 ACQCVWLRKVLEKLGAEEKSATVINCDNSSTIQLSKHPVLHGKSKHIEVRFHYLRDLVNG 1296
Query: 1554 GILDIQFIDIEHQWADIFTKPLSVERFDFIKKNLNMHFVSD 1594
++ +++ E Q ADIFTKPL +E+F+ ++ L M +S+
Sbjct: 1297 DVVKLEYCPTEDQVADIFTKPLKLEQFEKLRALLGMVNMSE 1337
>At2g15650 putative retroelement pol polyprotein
Length = 1347
Score = 647 bits (1670), Expect = 0.0
Identities = 373/1046 (35%), Positives = 571/1046 (53%), Gaps = 40/1046 (3%)
Query: 555 WYLDSGCSRHMTGEKALFLTLTMKDGGEVKFGGNQTGKIIGTGTIGNSSIS-------IN 607
W +DSGC+ HMT E+ F + ++ + G I+ T G+ ++ I
Sbjct: 326 WLVDSGCTNHMTKEERYFSNINKSIKVPIRV---RNGDIVMTAGKGDITVMTRHGKRIIK 382
Query: 608 NVWLVDGLKHNLLSISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKINFSDL 667
NV+LV GL+ NLLS+ Q +GY V F C + + + K I + + +KI S +
Sbjct: 383 NVFLVPGLEKNLLSVPQIISSGYWVRFQDKRCIIQDANGKEI-MNIEMTDKSFKIKLSSV 441
Query: 668 ADQKVVCLLSMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGACQKGK 727
++ + + + WHKRLGH + + + ++ +LV GLP + C AC GK
Sbjct: 442 EEEAMTANVQTEE---TWHKRLGHVSNKRLQQMQDKELVNGLPRFKVTKET-CKACNLGK 497
Query: 728 IVKSSFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWVKFIKSKD 787
+ SF + T LE++H D+ GP+ S+ GS+Y ++ +DDY+ WV F+K K
Sbjct: 498 QSRKSFPKESQTKTREKLEIVHTDVCGPMQHQSIDGSRYYVLFLDDYTHMCWVYFLKQKS 557
Query: 788 YACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRTPQQNG 847
F F ++ + I +R P E+FCE GI + + P +PQQNG
Sbjct: 558 ETFATFKKFKALVEKQSNCSIKTLR----------PMEVFCEDEGINRQVTLPYSPQQNG 607
Query: 848 VVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTT-YELFKGRR 906
ERKNR+L EMAR+M+ E +L WAEAV TS Y+QNR+ + + + T E + G +
Sbjct: 608 AAERKNRSLVEMARSMLVEQDLPLKLWAEAVYTSAYLQNRLPSKAIEDDVTPMEKWCGHK 667
Query: 907 PNISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVEESMHV 966
PN+S+ FG CY+ +K DAKA+ GI +GYS ++K YRV+ E + VE S V
Sbjct: 668 PNVSHLRIFGSICYVHIPDQKRRKLDAKAKCGILIGYSNQTKGYRVFLLEDEKVEVSRDV 727
Query: 967 KFDDREPGSKTSEQSESNAGTTDSEDASESDQPSDSEKYTEVESSPEAEYNSEAEPSPKV 1026
F + + ++ D ES ++ + + A N E E S V
Sbjct: 728 VFQEDKKWDWDKQEEVKKTFVMSINDIQESRDQQETSSHDLSQIDDHAN-NGEGETSSHV 786
Query: 1027 QNESASEDFQDNTQQVIQPKFKHKSSHPEELIIGSKDSPRRTRSHFRQEESLIGLLSIIE 1086
++ ++ ++ ++ PK K+KS +E++ + +PR Q L++ E
Sbjct: 787 LSQVNDQEERETSES---PK-KYKSM--KEIL---EKAPRMENDEAAQGIEAC-LVANEE 836
Query: 1087 PKTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTLLEQNGYSKTS*MNKEK*PE 1146
P+T +EA D W AM EE+ + P ++ ++ K +
Sbjct: 837 PQTYDEARGDKEWEEAMNEEIKVIEKNRTWKLVDKPEKKNVISVKWIYKIKTDASGNHVK 896
Query: 1147 TKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQMDVKSAFLNGVIE 1206
K VA+G+SQ+ GIDY ETFAPV+R + IR LL+YA LYQMDVKSAFLNG +E
Sbjct: 897 HKARLVARGFSQEYGIDYLETFAPVSRYDTIRALLAYAAQMKWRLYQMDVKSAFLNGELE 956
Query: 1207 EEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDFERGQVDTTLF 1266
EEVYV QPPGF + V +L K+LYGLKQAPRAWY+R+ ++ I+N F R D L+
Sbjct: 957 EEVYVTQPPGFVIEGKEEKVLRLYKALYGLKQAPRAWYERIDSYFIQNGFARSMNDAALY 1016
Query: 1267 RRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELKFFLGIQINQSKE 1326
+ +D+LIV +YVDD+I N L F K M+DEFEM+ +G L +FLG+++NQ
Sbjct: 1017 SKKKGEDVLIVSLYVDDLIITGNNTHLINTFKKNMKDEFEMTDLGLLNYFLGMEVNQDDS 1076
Query: 1327 GVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLS--KEDTGTVVDQKLYRGMIGSLLY 1384
G+++ Q KY +L+ KF +++ K ++TP+ P + D D YR ++G LLY
Sbjct: 1077 GIFLSQEKYANKLIDKFGMKESKSVSTPLTPQGKRKGVEGDDKEFADPTKYRRIVGGLLY 1136
Query: 1385 LTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGLLYRKSLDYKLIGFCD 1444
L ASRP ++++ +R+ S P H KR+ RY+KGT+N G+L+ +L+G+ D
Sbjct: 1137 LCASRPDVMYASSYLSRYMSSPSIQHYQEAKRVLRYVKGTSNFGVLFTSKETPRLVGYSD 1196
Query: 1445 ADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEYISAASCCTQLLWMKHQ 1504
+D+ G ++KST+G LG + W S +Q T+A STAEAEYI+ + Q +W++
Sbjct: 1197 SDWGGSLEDKKSTTGYVFTLGLAMFCWQSCKQQTVAQSTAEAEYIAVCAATNQAIWLQRL 1256
Query: 1505 LEDYQIN-ANSIPIYCDNTAAICLSKNPILHSRAKHIEIKHHFIRDYVQKGILDIQFIDI 1563
ED+ + IPI CDN +AI + +NP+ H R KHIEIK+HF+R+ KG++ +++
Sbjct: 1257 FEDFGLKFKEGIPILCDNKSAIAIGRNPVQHRRTKHIEIKYHFVREAEHKGLIQLEYCKG 1316
Query: 1564 EHQWADIFTKPLSVERFDFIKKNLNM 1589
E Q AD+ TK LSV RF+ +++ L +
Sbjct: 1317 EDQLADVLTKALSVSRFEGLRRKLGV 1342
>At3g25450 hypothetical protein
Length = 1343
Score = 602 bits (1552), Expect = e-172
Identities = 370/1152 (32%), Positives = 619/1152 (53%), Gaps = 80/1152 (6%)
Query: 469 LNQNLKSQNKRIRRTKQLLLLRKQYQKVSNLKY*MIRSHSAFTLRYKGGKVKPPKLTQKD 528
+N+ +K+++ I R +L+ L++Q +K + +H A +L + L +K
Sbjct: 229 INKEIKAKSHVIDRLLKLIRLQEQKEKEED------DTHEAESLMMH----EVVYLNEK- 277
Query: 529 P*RYGYLNLNWPKMQVCLRAREKQRSWYLDSGCSRHMTGEKALFLTLTMKDGGEVKFGGN 588
N+ +++ C+ +WYLD+G S HMTG +A F L G+V+FG +
Sbjct: 278 -------NIRPTELESCIN-----NAWYLDNGASNHMTGNRAWFCKLDEMITGKVRFGDD 325
Query: 589 QTGKIIGTGTI-----GNSSISINNVWLVDGLKHNLLSISQFCDNGYDVTFSKTNCTLVN 643
I G G+I G + +V+ + LK N+LS+ Q ++G D+ + TL +
Sbjct: 326 SCINIKGKGSIPFISKGGERKILFDVYYIPDLKSNILSLGQATESGCDIRMREDYLTLHD 385
Query: 644 KDDKSITFKGKRVEN-VYKINFSDLADQKVVCLLSMNDKKWVWHKRLGHANWRLISKISK 702
++ ++ K +R N +YK++ ++ + K + L + N+ +WH RLGH ++ I + K
Sbjct: 386 REG-NLLIKAQRSRNRLYKVSL-EVENSKCLQLTTTNEST-IWHARLGHISFETIKAMIK 442
Query: 703 LQLVKGLPNIDYHSDALCGACQKGKIVKSSFKSKDIVSTSRPLELLHIDLFGPVNTASLY 762
+LV G+ + CG+C GK + SF ++ LEL+H DL GP++ ++
Sbjct: 443 KELVIGISSSVPQEKETCGSCLFGKQARHSFPKATSYRAAQVLELIHGDLCGPISPSTAA 502
Query: 763 GSKYGLVIVDDYSRWTWVKFIKSKDYACEVFSSFCTQIQSEKELKILKVRSDYGGEFENE 822
+Y V++DD+SR+ W +K K A F F ++ E I R+D GGEF +
Sbjct: 503 KKRYVFVLIDDHSRYMWSILLKEKSEAFGKFKEFKALVEQECGAIIKTFRTDRGGEFLSH 562
Query: 823 PFELFCEKHGILHEFSSPRTPQQNGVVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSC 882
F+ FC K GI ++P TPQQNGVVER+NRTL M R+++ N+ + W EAV S
Sbjct: 563 EFQEFCAKEGINRHLTAPYTPQQNGVVERRNRTLLGMTRSILKHMNMPNYLWGEAVRHST 622
Query: 883 YIQNRIYIRPMLEKTTYELFKGRRPNISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLG 942
Y+ NR+ R + +T YE+FK ++PN+ + FGC Y LKK D +++ ++LG
Sbjct: 623 YLINRVGTRSLSNQTPYEVFKHKKPNVEHLRVFGCVSYAKVEVPNLKKLDDRSRMLVYLG 682
Query: 943 YSERSKAYRVYNSETQCVEESMHVKFD--------------DREPGSKTSEQSE-SNAGT 987
SKAYR+ + + + S V FD D+E G+ T SE N G
Sbjct: 683 TEPGSKAYRLLDPTKRRIFVSRDVVFDENRSWMWQESSSETDKESGTFTITLSEFGNNGV 742
Query: 988 TDSEDASESDQPSDSEKYTEVESSPEAEYNSEAEPSPKVQNESASEDFQDNTQQVIQPKF 1047
T+++ ++E ++ ++E E E+ E EAE Q++ + + + +QVI+P +
Sbjct: 743 TENDISTEPEETEEAEINGEDENIIE-----EAETEEHDQSQEEPQPVRRSQRQVIRPNY 797
Query: 1048 KHK-----SSHPEELIIGSKDSPRRTRSHFRQEESLIGLLSIIEPKTVEEALSDDGWILA 1102
E L++ D EP +EA W A
Sbjct: 798 LKDYVLCAEIEAEHLLLAVND----------------------EPWDFKEANKSKEWRDA 835
Query: 1103 MQEELINFKGMMCGIWYPYPFRRTLLEQNGYSKTS*MNKEK*PETKPDFVAQGYSQQEGI 1162
+EE+ + + P + K + + K VA+GY Q+ G+
Sbjct: 836 CKEEIQSIEKNRTWSLVDLPVGSKAIGVKWVFKLKHNSDGSINKYKARLVAKGYVQRHGV 895
Query: 1163 DYTETFAPVARLEAIRLLLSYAINHGIILYQMDVKSAFLNGVIEEEVYVKQPPGFEDLKH 1222
D+ E FAPVAR+E +RL+++ A ++G ++ +DVK+AFL+G + E+VYV QP GF + +
Sbjct: 896 DFEEVFAPVARIETVRLIIALAASNGWEIHHLDVKTAFLHGELREDVYVSQPEGFTNKES 955
Query: 1223 PDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDFERGQVDTTLFRRTLKKDILIVQIYVD 1282
+ VYKL K+LYGL+QAPRAW +L+ L + FE+ + +L+R+ ++IL+V +YVD
Sbjct: 956 KEKVYKLHKALYGLRQAPRAWNTKLNEILKELKFEKCHKEPSLYRKQEGENILVVAVYVD 1015
Query: 1283 DIIFGSTNASLCK*FSKLMQDEFEMSMMGELKFFLGIQINQSKEGVYVHQTKYTKELLKK 1342
D++ +N + F K M +FEMS +G+L ++LGI++ QSK+G+ + Q +Y K++L++
Sbjct: 1016 DLLVTGSNLDIILNFKKGMVGKFEMSDLGKLTYYLGIEVLQSKDGITLKQERYAKKILEE 1075
Query: 1343 FKLEDCKVMNTPMHPTCTLSKEDTGTVVDQKLYRGMIGSLLYLTASRPAILFSVCLCARF 1402
+ C +NTPM + LSK +D+ YR IG L YL +RP + ++V + +R+
Sbjct: 1076 AGMSKCNTVNTPMIASLELSKAQDEKRIDETDYRRNIGCLRYLLHTRPDLSYNVGILSRY 1135
Query: 1403 QSDPRESHLTAVKRIFRYLKGTTNLGLLYRKSLDYKLIGFCDADYAGDRIERKSTSGNCQ 1462
+PRESH A+K+I RYL+GTT+ GL ++K + LIG+ D+ + D + KST G+
Sbjct: 1136 LQEPRESHGAALKQILRYLQGTTSHGLYFKKGENAGLIGYSDSSHNVDLDDGKSTGGHIF 1195
Query: 1463 FLGENLISWASKRQATIAMSTAEAEYISAASCCTQLLWMKHQL-EDYQINANSIPIYCDN 1521
+L + I+W S++Q + +S+ EAE+++A Q +W++ L E + I DN
Sbjct: 1196 YLNDCPITWCSQKQQVVTLSSCEAEFMAATEAAKQAIWLQELLAEVIGTECEKVTIRVDN 1255
Query: 1522 TAAICLSKNPILHSRAKHIEIKHHFIRDYVQKGILDIQFIDIEHQWADIFTKPLSVERFD 1581
+AI L+KNP+ H R+KHI ++HFIR+ V+ G ++++ + Q ADI TK L +F
Sbjct: 1256 KSAIALTKNPVFHGRSKHIHRRYHFIRECVENGQIEVEHVPGVRQKADILTKALGKIKFL 1315
Query: 1582 FIKKNLNMHFVS 1593
+++ + + VS
Sbjct: 1316 EMRELIGVQGVS 1327
>At1g32590 hypothetical protein, 5' partial
Length = 1263
Score = 595 bits (1535), Expect = e-170
Identities = 343/1054 (32%), Positives = 562/1054 (52%), Gaps = 62/1054 (5%)
Query: 550 EKQRSWYLDSGCSRHMTGEKALFLTLTMKDGGEVKFGGNQTGKIIGTGTI-----GNSSI 604
E+++ W+LDSGCS HM G + FL L V+ G ++ + G G + G +
Sbjct: 253 EEKQIWFLDSGCSNHMCGTREWFLELDSGFKQNVRLGDDRRMAVEGKGKLRLEVDGRIQV 312
Query: 605 SINNVWLVDGLKHNLLSISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKINF 664
I++V+ V GLK+NL S+ Q G C + +K +K + +N + F
Sbjct: 313 -ISDVYFVPGLKNNLFSVGQLQQKGLRFIIEGDVCEVWHKTEKRMVMHSTMTKNRMFVVF 371
Query: 665 SDLADQKVV----CLLSMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDY-HSDAL 719
+ + K CL + +WHKR GH N + + +++ ++VKGLP D +A+
Sbjct: 372 AAVKKSKETEETRCLQVIGKANNMWHKRFGHLNHQGLRSLAEKEMVKGLPKFDLGEEEAV 431
Query: 720 CGACQKGKIVKSSFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTW 779
C C KGK ++ S + +++ L+L+H D+ GP+N AS G +Y L +DD+SR W
Sbjct: 432 CDICLKGKQIRESIPKESAWKSTQVLQLVHTDICGPINPASTSGKRYILNFIDDFSRKCW 491
Query: 780 VKFIKSKDYACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSS 839
+ K + F F +++ E K++ +RSD GGE+ + F+ +C++ GI + ++
Sbjct: 492 TYLLSEKSETFQFFKEFKAEVERESGKKLVCLRSDRGGEYNSREFDEYCKEFGIKRQLTA 551
Query: 840 PRTPQQNGVVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTY 899
TPQQNGV ERKNR++ M R M+ E ++ + FW EAV + YI NR + + + T
Sbjct: 552 AYTPQQNGVAERKNRSVMNMTRCMLMEMSVPRKFWPEAVQYAVYILNRSPSKALNDITPE 611
Query: 900 ELFKGRRPNISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQC 959
E + +P++ + FG Y L K D K+ + + G S+ SKAYR+Y+ T
Sbjct: 612 EKWSSWKPSVEHLRIFGSLAYALVPYQKRIKLDEKSIKCVMFGVSKESKAYRLYDPATGK 671
Query: 960 VEESMHVKFDDREPGSKTSEQSESNAGTTDSEDASESDQPSDSEKYTEVESSPEAEYNSE 1019
+ S V+FD+ E G + ++S D+ D + E PE +N +
Sbjct: 672 ILISRDVQFDE-ERGWEWEDKSLEEELVWDNSDHEPAG-----------EEGPEINHNGQ 719
Query: 1020 AEPSP-KVQNESASEDFQDNTQQVIQPKFKHKSSHP--EELIIGSKDSPRRTRSHFRQEE 1076
+ + + E+ +E N V + + ++ ++G+ R + ++E
Sbjct: 720 QDQEETEEEEETVAETVHQNLPAVGTGGVRQRQQPVWMKDYVVGNA---RVLITQDEEDE 776
Query: 1077 SLIGLLSIIEPKTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTLLEQNGYSKT 1136
L + +P EEA + W AM+ E+ + + P ++ KT
Sbjct: 777 VLALFIGPDDPVCFEEAAQLEVWRKAMEAEITSIEENNTWELVELPEEAKVIGLKWIFKT 836
Query: 1137 S*MNKEK*PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQMDV 1196
K + + K VA+GY Q+ G+D+ E FAPVA+ + IRL+L A G ++Q+DV
Sbjct: 837 KFNEKGEVDKFKARLVAKGYHQRYGVDFYEVFAPVAKWDTIRLILGLAAEKGWSVFQLDV 896
Query: 1197 KSAFLNGVIEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDF 1256
KSAFL+G ++E+V+V+QP GFE + VYKLKK+LYGLKQAPRAWY R+ F K F
Sbjct: 897 KSAFLHGDLKEDVFVEQPKGFEVEEESSKVYKLKKALYGLKQAPRAWYSRIEEFFGKEGF 956
Query: 1257 ERGQVDTTLFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELKFF 1316
E+ + TLF + + D L+V +YVDD+I+ ++ + + F M +EF M+ +G++K+F
Sbjct: 957 EKCYCEHTLFVKKERSDFLVVSVYVDDLIYTGSSMEMIEGFKNSMMEEFAMTDLGKMKYF 1016
Query: 1317 LGIQINQSKEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGTVVDQKLYR 1376
LG+++ Q + G++++Q KY E++KK+ +E C + P+ P L+K G V
Sbjct: 1017 LGVEVIQDERGIFINQRKYAAEIIKKYGMEGCNSVKNPIVPGQKLTK--AGAV------- 1067
Query: 1377 GMIGSLLYLTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGLLYRKSLD 1436
+R+ P E HL AVKRI RY++GT +LG+ Y +
Sbjct: 1068 -----------------------SRYMESPNEQHLLAVKRILRYVQGTLDLGIQYERGGA 1104
Query: 1437 YKLIGFCDADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEYISAASCCT 1496
+L+GF D+DYAGD +RKSTSG LG I+WASK+Q + +ST EAE++SA+
Sbjct: 1105 TELVGFVDSDYAGDVDDRKSTSGYVFMLGGGAIAWASKKQPIVTLSTTEAEFVSASYGAC 1164
Query: 1497 QLLWMKHQLEDYQI-NANSIPIYCDNTAAICLSKNPILHSRAKHIEIKHHFIRDYVQKGI 1555
Q +W+++ LE+ ++CDN++ I LSKNP+LH R+KHI +++HF+R+ V++G
Sbjct: 1165 QAVWLRNVLEEIGCRQEGGTLVFCDNSSTIKLSKNPVLHGRSKHIHVRYHFLRELVKEGT 1224
Query: 1556 LDIQFIDIEHQWADIFTKPLSVERFDFIKKNLNM 1589
+ + + Q ADI TK + E F+ ++ + +
Sbjct: 1225 IRLDYCTTTDQVADIMTKAVKREVFEELRGRMGV 1258
>At2g05390 putative retroelement pol polyprotein
Length = 1307
Score = 551 bits (1420), Expect = e-156
Identities = 335/1047 (31%), Positives = 558/1047 (52%), Gaps = 70/1047 (6%)
Query: 554 SWYLDSGCSRHMTGEKALFLTLTMKDGGEVKFGGNQTGKIIGTGTI-----GNSSISINN 608
SWYLD+G S HMTG F L G+V+FG + I G G+I G ++ +
Sbjct: 279 SWYLDNGASNHMTGNLQWFSKLNEMITGKVRFGDDSRIDIKGKGSIVLITKGGIRKTLTD 338
Query: 609 VWLVDGLKHNLLSISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKINFSDLA 668
V+ + LK N++S+ Q + G DV TL +++ + + +YK+ DL
Sbjct: 339 VYFIPDLKSNIISLGQATEAGCDVRMKDDQLTLHDREGCLLLRATRSRNRLYKV---DLN 395
Query: 669 DQKVVCLLSMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGACQKGKI 728
+ V CL +L A + + +LV G+ NI + CG+C GK
Sbjct: 396 VENVKCL------------QLEAAT------MVRKELVIGISNIPKEKET-CGSCLLGKQ 436
Query: 729 VKSSFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWVKFIKSKDY 788
+ F S+ LEL+H DL GP+ ++ +Y LV++DD++R+ W +K K
Sbjct: 437 ARQPFPKATTYRASQVLELVHGDLCGPITQSTTAKKRYILVLIDDHTRYMWSMLLKEKSE 496
Query: 789 ACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRTPQQNGV 848
A E F F T+++ E +KI R+D GGEF ++ F+ FC K GI ++P TPQQNGV
Sbjct: 497 AFEKFRDFKTKVEQESGVKIKTFRTDKGGEFVSQEFQDFCAKEGINRHLTAPYTPQQNGV 556
Query: 849 VERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYELFKGRRPN 908
VER+NRTL M R+++ + + W EAV S YI NR+ R + +T YE+FK R+PN
Sbjct: 557 VERRNRTLLGMTRSILKHMKMPNYLWGEAVRHSTYIINRVGTRSLQNQTPYEVFKQRKPN 616
Query: 909 ISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYN-------------S 955
+ + FGC Y +L+K D +++ ++LG SKAYR+ + S
Sbjct: 617 VEHLRVFGCIGYAKIEGPHLRKLDDRSKMLVYLGTEPGSKAYRLLDPTNRKIIKWNNSDS 676
Query: 956 ETQCVEESMHVKFDD-REPGSKTSEQSESNAGTTDSEDASESDQPSDSEKYTEVESSPEA 1014
ET+ + + + + G + S+ E+ +SE++ E + ++ + ++++ E
Sbjct: 677 ETRDISGTFSLTLGEFGNNGIQESDDIETEKNGEESENSHEEEGENEHNEQEQIDA--EE 734
Query: 1015 EYNSEAEPSPKVQNESASEDFQDNTQQVIQPKFKHKSSHPEELIIGSKDSPRRTRSHFRQ 1074
S A P P ++ +T+QV +P + ++ ++ +
Sbjct: 735 TQPSHATPLPTLRR---------STRQVGKPNYL------DDYVL---------MAEIEG 770
Query: 1075 EESLIGLLSIIEPKTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTLLEQNGYS 1134
E+ L+ + EP +EA W A +EE+++ + P RR ++
Sbjct: 771 EQVLLAIND--EPWDFKEANKLKEWRDACKEEILSIEKNKTWSLIDLPVRRKVIGLKWVF 828
Query: 1135 KTS*MNKEK*PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQM 1194
K + + K VA+GY Q+ GIDY E FA VAR+E IR++++ A ++G ++ +
Sbjct: 829 KIKRNSDGSINKYKARLVAKGYVQRHGIDYDEVFAHVARIETIRVIIALAASNGWEVHHL 888
Query: 1195 DVKSAFLNGVIEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKN 1254
DVK+AFL+G + E+VYV QP GF + + VYKL K+LYGLKQAPRAW +L+ L +
Sbjct: 889 DVKTAFLHGELREDVYVTQPEGFTNKDNEGKVYKLHKALYGLKQAPRAWNTKLNKILQEL 948
Query: 1255 DFERGQVDTTLFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELK 1314
+F + + +++RR +K +LIV IYVDD++ ++ L F K M +FEMS +G+L
Sbjct: 949 NFVKCSKEPSVYRRQEEKKLLIVAIYVDDLLVTGSSLDLILCFKKDMAGKFEMSDLGQLT 1008
Query: 1315 FFLGIQINQSKEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGTVVDQKL 1374
++LGI++ K G+ + Q +Y +++++ + +C + PM L K + ++
Sbjct: 1009 YYLGIEVLHRKNGIILRQERYAMKIIEEAGMSNCNPVLIPMAAGLELCKAQEEKCITERD 1068
Query: 1375 YRGMIGSLLYLTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGLLYRKS 1434
YR MIG L Y+ +RP + + V + +R+ PRESH A+K++ RYLKGT + GL ++
Sbjct: 1069 YRRMIGCLRYIVHTRPDLSYCVGVLSRYLQQPRESHGNALKQVLRYLKGTMSHGLYLKRG 1128
Query: 1435 LDYKLIGFCDADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEYISAASC 1494
L+G+ D+ ++ D + KST+G+ +L + I+W S++Q +A+S+ EAE+++A
Sbjct: 1129 FKSGLVGYSDSSHSADLDDGKSTAGHIFYLHQCPITWCSQKQQVVALSSCEAEFMAATEA 1188
Query: 1495 CTQLLWMKHQL-EDYQINANSIPIYCDNTAAICLSKNPILHSRAKHIEIKHHFIRDYVQK 1553
Q +W++ E + + I DN +AI L+KN + H R+KHI ++HFIR+ V+
Sbjct: 1189 AKQAIWLQDLFAEVCGTTSEKVMIRVDNKSAIALTKNLVFHGRSKHIHRRYHFIRECVEN 1248
Query: 1554 GILDIQFIDIEHQWADIFTKPLSVERF 1580
++++ + Q ADI TKPL +F
Sbjct: 1249 NLVEVDHVPGVEQRADILTKPLGRIKF 1275
>At2g20460 putative retroelement pol polyprotein
Length = 1461
Score = 543 bits (1399), Expect = e-154
Identities = 344/1045 (32%), Positives = 549/1045 (51%), Gaps = 41/1045 (3%)
Query: 554 SWYLDSGCSRHMTGEKALFLTLTMKDGGEVKFGGNQTG---KIIGTGTIG-NSSISINNV 609
+W +DSG + H++ ++ LF TL D V F TG +I G GT+ N I + NV
Sbjct: 441 TWVIDSGATHHVSHDRKLFQTL---DTSIVSFVNLPTGPNVRISGVGTVLINKDIILQNV 497
Query: 610 WLVDGLKHNLLSISQFC-DNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKINFSDLA 668
+ + NL+SIS D G V F + C + + +GKR+ N+Y ++ A
Sbjct: 498 LFIPEFRLNLISISSLTTDLGTRVIFDPSCCQIQDLTKGLTLGEGKRIGNLYVLDTQSPA 557
Query: 669 DQKVVCLLSMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGACQKGKI 728
+ + ++ D VWHKRLGH ++ S++ L V G A C C K
Sbjct: 558 ----ISVNAVVDVS-VWHKRLGHPSF---SRLDSLSEVLGTTRHKNKKSAYCHVCHLAKQ 609
Query: 729 VKSSFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWVKFIKSKDY 788
K SF S + + S ELLHID++GP + ++ G KY L IVDD+SR TW+ +KSK
Sbjct: 610 KKLSFPSANNICNST-FELLHIDVWGPFSVETVEGYKYFLTIVDDHSRATWIYLLKSKSD 668
Query: 789 ACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRTPQQNGV 848
VF +F ++++ + ++ VRSD E F F + GI+ S P TP+QN V
Sbjct: 669 VLTVFPAFIDLVENQYDTRVKSVRSDNAKELA---FTEFYKAKGIVSFHSCPETPEQNSV 725
Query: 849 VERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYELFKGRRPN 908
VERK++ + +AR ++ ++N++ +W + V T+ ++ NR + KT +E+ G+ P+
Sbjct: 726 VERKHQHILNVARALMFQSNMSLPYWGDCVLTAVFLINRTPSALLSNKTPFEVLTGKLPD 785
Query: 909 ISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVEESMHVKF 968
S FGC CY + KF +++ +FLGY K Y++ + E+ V S +V+F
Sbjct: 786 YSQLKTFGCLCYSSTSSKQRHKFLPRSRACVFLGYPFGFKGYKLLDLESNVVHISRNVEF 845
Query: 969 DDREPGSKTSEQSESNAGTTDSEDASESDQPSDSEKYTEVESSPEAEYNSEAEPSPKVQN 1028
+ +S+QS TT S+ + D S T SP+ PS ++
Sbjct: 846 HEELFPLASSQQS----ATTASDVFTPMDPLSSGNSITSHLPSPQIS------PSTQISK 895
Query: 1029 ESASEDFQDNTQQVIQPKFKHKSSHPEELIIGSKDSPRRTRSHFRQEESLIGLLSIIEPK 1088
++ F + Q SHP I S + + SH + + I P+
Sbjct: 896 RRITK-FPAHLQDYHCYFVNKDDSHP---ISSSLSYSQISPSHMLY---INNISKIPIPQ 948
Query: 1089 TVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTLLEQNGYSKTS*MNKEK*PETK 1148
+ EA W A+ +E+ + P + + K
Sbjct: 949 SYHEAKDSKEWCGAIDQEIGAMERTDTWEITSLPPGKKAVGCKWVFTVKFHADGSLERFK 1008
Query: 1149 PDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQMDVKSAFLNGVIEEE 1208
VA+GY+Q+EG+DYTETF+PVA++ ++LLL + + L Q+D+ +AFLNG +EE
Sbjct: 1009 ARIVAKGYTQKEGLDYTETFSPVAKMATVKLLLKVSASKKWYLNQLDISNAFLNGDLEET 1068
Query: 1209 VYVKQPPGFEDLKH----PDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDFERGQVDTT 1264
+Y+K P G+ D+K P+ V +LKKS+YGLKQA R W+ + SN L+ FE+ D T
Sbjct: 1069 IYMKLPDGYADIKGTSLPPNVVCRLKKSIYGLKQASRQWFLKFSNSLLALGFEKQHGDHT 1128
Query: 1265 LFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELKFFLGIQINQS 1324
LF R + + +++ +YVDDI+ ST + ++ ++ F++ +G LK+FLG+++ ++
Sbjct: 1129 LFVRCIGSEFIVLLVYVDDIVIASTTEQAAQSLTEALKASFKLRELGPLKYFLGLEVART 1188
Query: 1325 KEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGTVVDQKLYRGMIGSLLY 1384
EG+ + Q KY ELL + DCK + PM P LSK D + D+++YR ++G L+Y
Sbjct: 1189 SEGISLSQRKYALELLTSADMLDCKPSSIPMTPNIRLSKNDGLLLEDKEMYRRLVGKLMY 1248
Query: 1385 LTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGLLYRKSLDYKLIGFCD 1444
LT +RP I F+V +F S PR +HL AV ++ +Y+KGT GL Y D L G+ D
Sbjct: 1249 LTITRPDITFAVNKLCQFSSAPRTAHLAAVYKVLQYIKGTVGQGLFYSAEDDLTLKGYTD 1308
Query: 1445 ADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEYISAASCCTQLLWMKHQ 1504
AD+ R+ST+G F+G +LISW SK+Q T++ S+AEAEY + A ++ W+
Sbjct: 1309 ADWGTCPDSRRSTTGFTMFVGSSLISWRSKKQPTVSRSSAEAEYRALALASCEMAWLSTL 1368
Query: 1505 LEDYQINANSIPIYCDNTAAICLSKNPILHSRAKHIEIKHHFIRDYVQKGILDIQFIDIE 1564
L ++++ +Y D+TAA+ ++ NP+ H R KHIEI H +R+ + G L + + +
Sbjct: 1369 LLALRVHSGVPILYSDSTAAVYIATNPVFHERTKHIEIDCHTVREKLDNGQLKLLHVKTK 1428
Query: 1565 HQWADIFTKPLSVERFDFIKKNLNM 1589
Q ADI TKPL +F + +++
Sbjct: 1429 DQVADILTKPLFPYQFAHLLSKMSI 1453
>At3g59720 copia-type reverse transcriptase-like protein
Length = 1272
Score = 534 bits (1376), Expect = e-151
Identities = 317/898 (35%), Positives = 471/898 (52%), Gaps = 67/898 (7%)
Query: 549 REKQRSWYLDSGCSRHMTGEKALFLTLTMKDGGEVKFGGNQTGKIIGTGTI-----GNSS 603
+E+ WYLDSG S HM G K++F L G V G ++ G G I
Sbjct: 329 QEENHKWYLDSGASNHMCGRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRLKNGDH 388
Query: 604 ISINNVWLVDGLKHNLLSISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKIN 663
I+NV+ + +K N+LS+ Q + GYD+ N ++ +K+ IT ++ +N
Sbjct: 389 QFISNVYYIPSMKTNILSLGQLLEKGYDIRLKDNNLSIRDKESNLITKVPMSKNRMFVLN 448
Query: 664 F-SDLADQKVVCLLSMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGA 722
+D+A +C ++ W+WH R GH N+ + +S+ ++V+GLP I+ H + +C
Sbjct: 449 IRNDIAQCLKMCY---KEESWLWHLRFGHLNFGGLELLSRKEMVRGLPCIN-HPNQVCEG 504
Query: 723 CQKGKIVKSSFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWVKF 782
C G K SF + +PLEL+H D+ GP+ SL S Y L+ +DD+SR TWV F
Sbjct: 505 CLLGNQFKMSFPKESSSRAQKPLELIHTDVCGPIKPKSLGKSNYFLLFIDDFSRKTWVYF 564
Query: 783 IKSKDYACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRT 842
+K K E+F F ++ E L I +RSD GGEF ++ F +CE +GI + + PR+
Sbjct: 565 LKEKSEVFEIFKKFKAHVEKESGLVIKTMRSDSGGEFTSKEFLKYCEDNGIRRQLTVPRS 624
Query: 843 PQQNGVVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYELF 902
PQQNGV ERKNRT+ EMAR+M+ L K WAEAV + Y+ NR + + KT E +
Sbjct: 625 PQQNGVAERKNRTILEMARSMLKSKRLPKELWAEAVACAVYLLNRSPTKSVSGKTPQEAW 684
Query: 903 KGRRPNISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVEE 962
GR+P +S+ FG + + K D K+++ IF+GY SK Y++YN +T+
Sbjct: 685 SGRKPGVSHLRVFGSIAHAHVPDEKRNKLDDKSEKYIFIGYDNNSKGYKLYNPDTKKTII 744
Query: 963 SMHVKFDDREPGSKTSEQSESNAGTTDSEDASESDQPSDSEKYTEVESSPEAEYNSEAEP 1022
S ++ FD+ S + + N ED E P+ E +E ++P P
Sbjct: 745 SRNIVFDEEGEWDWNSNEEDYNFFPHFEEDKPE---PTREEPPSEEPTTPPTS------P 795
Query: 1023 SPKVQNESASEDFQDNTQQVIQPKFKHKSSHPEELIIGSKDSPRRTRSHFRQEESLIGLL 1082
+ ES+SE P+F+ +EL +++ T L L
Sbjct: 796 TSSQIEESSSER---------TPRFRSI----QELYEVTENQENLT---------LFCLF 833
Query: 1083 SIIEPKTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTLLEQNGYSKTS*MNKE 1142
+ EP +EA+ W AM EE+ + + P + K +K
Sbjct: 834 AECEPMDFQEAIEKKTWRNAMDEEIKSIQKNDTWELTSLPNGHKAIGVKWVYKAKKNSKG 893
Query: 1143 K*PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQMDVKSAFLN 1202
+ K VA+GYSQ+ GIDY E FAPVARLE +RL++S A + ++QMDVKSAFLN
Sbjct: 894 EVERYKARLVAKGYSQRAGIDYDEIFAPVARLETVRLIISLAAQNKWKIHQMDVKSAFLN 953
Query: 1203 GVIEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDFERGQVD 1262
G +EEEVY++QP G+ D V +LKK LYGLKQAPRAW R+ + + DF + +
Sbjct: 954 GDLEEEVYIEQPQGYIVKGEEDKVLRLKKVLYGLKQAPRAWNTRIDKYFKEKDFIKCPYE 1013
Query: 1263 TTLFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELKFFLGIQIN 1322
L+ + K+DILI +YVDD+IF N S+ + F K M EFEM+ +G + ++LGI++
Sbjct: 1014 HALYIKIQKEDILIACLYVDDLIFTGNNPSMFEEFKKEMTKEFEMTDIGLMSYYLGIEVK 1073
Query: 1323 QSKEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGTVVDQKLYRGMIGSL 1382
Q G+++ Q Y KE+LKKFK++D ++GSL
Sbjct: 1074 QEDNGIFITQEGYAKEVLKKFKMDDSN--------------------------PSLVGSL 1107
Query: 1383 LYLTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGLLYRKSLDYKLI 1440
YLT +RP IL++V + +R+ P +H A KRI RY+KGT N GL Y + DYKL+
Sbjct: 1108 RYLTCTRPDILYAVGVVSRYMEHPTTTHFKAAKRILRYIKGTVNFGLHYSTTSDYKLV 1165
>At1g31210 putative reverse transcriptase
Length = 1415
Score = 533 bits (1373), Expect = e-151
Identities = 341/1066 (31%), Positives = 543/1066 (49%), Gaps = 53/1066 (4%)
Query: 553 RSWYLDSGCSRHMTGEKALFLTLTMKDGGEVKFGGNQTGKII---GTGTI--GNSSISIN 607
+ W+ DS + H+T + T +G + G+ T I G+ TI N I +N
Sbjct: 320 KEWHPDSAATAHVTSSTNGLQSATEYEGDDAVLVGDGTYLPITHTGSTTIKSSNGKIPLN 379
Query: 608 NVWLVDGLKHNLLSISQFCDN-GYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKINFSD 666
V +V ++ +LLS+S+ CD+ V F +++ + + G R +Y
Sbjct: 380 EVLVVPNIQKSLLSVSKLCDDYPCGVYFDANKVCIIDLQTQKVVTTGPRRNGLYV----- 434
Query: 667 LADQKVVCLLSMND---KKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGAC 723
L +Q+ V L S + VWH RLGHAN + + LQ K + + +C C
Sbjct: 435 LENQEFVALYSNRQCAATEEVWHHRLGHANSKALQH---LQNSKAIQINKSRTSPVCEPC 491
Query: 724 QKGKIVKSSFKSKDIVSTSR---PLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWV 780
Q GK + F ++S SR PL+ +H DL+GP S G KY + VDDYSR++W
Sbjct: 492 QMGKSSRLPF----LISDSRVLHPLDRIHCDLWGPSPVVSNQGLKYYAIFVDDYSRYSWF 547
Query: 781 KFIKSKDYACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSP 840
+ +K VF SF ++++ KI +SD GGEF + + +HGI H S P
Sbjct: 548 YPLHNKSEFLSVFISFQKLVENQLNTKIKVFQSDGGGEFVSNKLKTHLSEHGIHHRISCP 607
Query: 841 RTPQQNGVVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYE 900
TPQQNG+ ERK+R L E+ +M+ ++ + FW E+ T+ YI NR+ + + YE
Sbjct: 608 YTPQQNGLAERKHRHLVELGLSMLFHSHTPQKFWVESFFTANYIINRLPSSVLKNLSPYE 667
Query: 901 LFKGRRPNISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCV 960
G +P+ S FG CY KFD ++ + +FLGY+ + K YR + T V
Sbjct: 668 ALFGEKPDYSSLRVFGSACYPCLRPLAQNKFDPRSLQCVFLGYNSQYKGYRCFYPPTGKV 727
Query: 961 EESMHVKFDDREPGSKTSEQSESNAGTTDSEDASESDQPSDSE----------KYTEVES 1010
S +V F++ E K QS +T A + ++ S+ K ++ +
Sbjct: 728 YISRNVIFNESELPFKEKYQSLVPQYSTPLLQAWQHNKISEISVPAAPVQLFSKPIDLNT 787
Query: 1011 SPEAEYNSE-AEPSPKVQNESASEDFQDNTQQVIQPKFKHKSSHPEELIIGSKDSPRRTR 1069
++ + +P P NE + E+ +++ + +E +I S R++
Sbjct: 788 YAGSQVTEQLTDPEPTSNNEGSDEEVNPVAEEI---------AANQEQVINSHAMTTRSK 838
Query: 1070 SHFRQEESLIGLLS----IIEPKTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRR 1125
+ ++ + L++ EPKT+ A+ GW A+ EE+ + P
Sbjct: 839 AGIQKPNTRYALITSRMNTAEPKTLASAMKHPGWNEAVHEEINRVHMLHTWSLVPPTDDM 898
Query: 1126 TLLEQNGYSKTS*MNKEK*PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAI 1185
+L KT + K VA+G+ Q+EG+DY ETF+PV R IRL+L +
Sbjct: 899 NILSSKWVFKTKLHPDGSIDKLKARLVAKGFDQEEGVDYLETFSPVVRTATIRLVLDVST 958
Query: 1186 NHGIILYQMDVKSAFLNGVIEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYD 1245
+ G + Q+DV +AFL+G ++E V++ QP GF D + P HV +L K++YGLKQAPRAW+D
Sbjct: 959 SKGWPIKQLDVSNAFLHGELQEPVFMYQPSGFIDPQKPTHVCRLTKAIYGLKQAPRAWFD 1018
Query: 1246 RLSNFLIKNDFERGQVDTTLFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEF 1305
SNFL+ F + D +LF IL + +YVDDI+ ++ SL + + +++ F
Sbjct: 1019 TFSNFLLDYGFVCSKSDPSLFVCHQDGKILYLLLYVDDILLTGSDQSLLEDLLQALKNRF 1078
Query: 1306 EMSMMGELKFFLGIQINQSKEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKED 1365
M +G ++FLGIQI G+++HQT Y ++L++ + DC M TP+ L +
Sbjct: 1079 SMKDLGPPRYFLGIQIEDYANGLFLHQTAYATDILQQAGMSDCNPMPTPLPQ--QLDNLN 1136
Query: 1366 TGTVVDQKLYRGMIGSLLYLTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTT 1425
+ + +R + G L YLT +RP I F+V + P S +KRI RY+KGT
Sbjct: 1137 SELFAEPTYFRSLAGKLQYLTITRPDIQFAVNFICQRMHSPTTSDFGLLKRILRYIKGTI 1196
Query: 1426 NLGLLYRKSLDYKLIGFCDADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAE 1485
+GL +++ L + D+D+AG + R+ST+G C LG NLISW++KRQ T++ S+ E
Sbjct: 1197 GMGLPIKRNSTLTLSAYSDSDHAGCKNTRRSTTGFCILLGSNLISWSAKRQPTVSNSSTE 1256
Query: 1486 AEYISAASCCTQLLWMKHQLEDYQINANSIP--IYCDNTAAICLSKNPILHSRAKHIEIK 1543
AEY + ++ W+ L D I +P +YCDN +A+ LS NP LH+R+KH +
Sbjct: 1257 AEYRALTYAAREITWISFLLRDLGI-PQYLPTQVYCDNLSAVYLSANPALHNRSKHFDTD 1315
Query: 1544 HHFIRDYVQKGILDIQFIDIEHQWADIFTKPLSVERFDFIKKNLNM 1589
+H+IR+ V G+++ Q I Q AD+FTK L F ++ L +
Sbjct: 1316 YHYIREQVALGLIETQHISATFQLADVFTKSLPRRAFVDLRSKLGV 1361
>At1g53810
Length = 1522
Score = 523 bits (1346), Expect = e-148
Identities = 355/1117 (31%), Positives = 537/1117 (47%), Gaps = 92/1117 (8%)
Query: 555 WYLDSGCSRHMTGEKALFLTLTMKDGGE---VKFGGNQTGKIIGTGTIGNSS--ISINNV 609
W DS S H+T + + G + V G G+G+I +SS I + V
Sbjct: 326 WIPDSAASAHVTNNRHVLQQSQPYHGSDSIMVADGNFLPITHTGSGSIASSSGKIPLKEV 385
Query: 610 WLVDGLKHNLLSISQFC-DNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKINFSDLA 668
+ + +LLS+S+ D V F + + +K K + G+ + +Y + L
Sbjct: 386 LVCPDIVKSLLSVSKLTSDYPCSVEFDADSVRINDKATKKLLVMGRNRDGLYSLEEPKL- 444
Query: 669 DQKVVCLLSMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDALCGACQKGKI 728
Q + + VWH+RLGHAN ++ +++ K + I+ +C AC GK
Sbjct: 445 -QVLYSTRQNSASSEVWHRRLGHANAEVLHQLASS---KSIIIINKVVKTVCEACHLGKS 500
Query: 729 VKSSFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWVKFIKSKDY 788
+ F + SRPLE +H DL+GP T+S+ G +Y +V +D YSR+TW +K K
Sbjct: 501 TRLPFMLSTF-NASRPLERIHCDLWGPSPTSSVQGFRYYVVFIDHYSRFTWFYPLKLKSD 559
Query: 789 ACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRTPQQNGV 848
F F ++++ KI + D GGEF + F + HGI S P TPQQNG+
Sbjct: 560 FFSTFVMFQKLVENQLGHKIKIFQCDGGGEFISSQFLKHLQDHGIQQNMSCPYTPQQNGM 619
Query: 849 VERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPM-LEKTTYELFKGRRP 907
ERK+R + E+ +MI ++ L +W E+ T+ ++ N + + ++ Y+ G+ P
Sbjct: 620 AERKHRHIVELGLSMIFQSKLPLKYWLESFFTANFVINLLPTSSLDNNESPYQKLYGKAP 679
Query: 908 NISYFHQFGCTCYILNTKDYLK-KFDAKAQRGIFLGYSERSKAYRVYNSETQCVEESMHV 966
S FGC CY +DY KFD ++ + +FLGY+E+ K YR T + S HV
Sbjct: 680 EYSALRVFGCACYP-TLRDYASTKFDPRSLKCVFLGYNEKYKGYRCLYPPTGRIYISRHV 738
Query: 967 KFDDR-----------EPGSKT------------------------------------SE 979
FD+ P KT S
Sbjct: 739 VFDENTHPFESIYSHLHPQDKTPLLEAWFKSFHHVTPTQPDQSRYPVSSIPQPETTDLSA 798
Query: 980 QSESNAGTTDSEDASESDQPSDSEKYTEVESSPEA-----------EYNS----EAEPSP 1024
S A T +AS+ D D+E + V SPE Y+S + PSP
Sbjct: 799 APASVAAETAGPNASD-DTSQDNETISVVSGSPERTTGLDSASIGDSYHSPTADSSHPSP 857
Query: 1025 KVQNESASEDFQDNTQQVIQPKFKHKSSHPEELIIGSKDSPRRTRSHFRQEESLIGLLSI 1084
++ AS Q + + ++ ++ K+ + + L +SI
Sbjct: 858 -ARSSPASSPQGSPIQMAPAQQVQAPVTNEHAMVTRGKEGISKPNKRY---VLLTHKVSI 913
Query: 1085 IEPKTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTLLEQNGYSKTS*MNKEK* 1144
EPKTV EAL GW AMQEE+ N K PY +L +T
Sbjct: 914 PEPKTVTEALKHPGWNNAMQEEMGNCKETETWTLVPYSPNMNVLGSMWVFRTKLHADGSL 973
Query: 1145 PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQMDVKSAFLNGV 1204
+ K VA+G+ Q+EGIDY ET++PV R +RL+L A L QMDVK+AFL+G
Sbjct: 974 DKLKARLVAKGFKQEEGIDYLETYSPVVRTPTVRLILHVATVLKWELKQMDVKNAFLHGD 1033
Query: 1205 IEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDFERGQVDTT 1264
+ E VY++QP GF D PDHV L KSLYGLKQ+PRAW+DR SNFL++ F D +
Sbjct: 1034 LTETVYMRQPAGFVDKSKPDHVCLLHKSLYGLKQSPRAWFDRFSNFLLEFGFICSLFDPS 1093
Query: 1265 LFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELKFFLGIQINQS 1324
LF + D++++ +YVDD++ N+ + EF M MG++ +FLGIQI
Sbjct: 1094 LFVYSSNNDVILLLLYVDDMVITGNNSQSLTHLLAALNKEFRMKDMGQVHYFLGIQIQTY 1153
Query: 1325 KEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGTVVDQKLYRGMIGSLLY 1384
G+++ Q KY ++LL + +C M TP+ D +R + G L Y
Sbjct: 1154 DGGLFMSQQKYAEDLLITASMANCSPMPTPLPLQLDRVSNQDEVFSDPTYFRSLAGKLQY 1213
Query: 1385 LTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGLLYRKS---------L 1435
LT +RP I F+V + P S +KRI RY+KGT ++G+ Y +
Sbjct: 1214 LTLTRPDIQFAVNFVCQKMHQPSVSDFNLLKRILRYIKGTVSMGIQYNSNSSSVVSAYES 1273
Query: 1436 DYKLIGFCDADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEYISAASCC 1495
DY L + D+DYA + R+S G C F+G+N+ISW+SK+Q T++ S+ EAEY S +
Sbjct: 1274 DYDLSAYSDSDYANCKETRRSVGGYCTFMGQNIISWSSKKQPTVSRSSTEAEYRSLSETA 1333
Query: 1496 TQLLWMKHQLEDYQINANSIP-IYCDNTAAICLSKNPILHSRAKHIEIKHHFIRDYVQKG 1554
+++ WM L + ++ P ++CDN +A+ L+ NP H+R KH ++ HH+IR+ V
Sbjct: 1334 SEIKWMSSILREIGVSLPDTPELFCDNLSAVYLTANPAFHARTKHFDVDHHYIRERVALK 1393
Query: 1555 ILDIQFIDIEHQWADIFTKPLSVERFDFIKKNLNMHF 1591
L ++ I Q ADIFTK L E F ++ L + F
Sbjct: 1394 TLVVKHIPGHLQLADIFTKSLPFEAFTRLRFKLGVDF 1430
>At1g70010 hypothetical protein
Length = 1315
Score = 517 bits (1332), Expect = e-146
Identities = 338/1013 (33%), Positives = 527/1013 (51%), Gaps = 51/1013 (5%)
Query: 593 IIGTGTIGNSSISINNVWLVDGLKHNLLSISQFCDN-GYDVTFSKTNCTLVNKDDKSITF 651
I G+ +G I +N+V + K NLLS+S + G + F +T+C L + + +
Sbjct: 313 ISGSVHLGRHLI-LNDVLFIPQFKFNLLSVSSLTKSMGCRIWFDETSCVLQDATRELMVG 371
Query: 652 KGKRVENVYKINFSDLA----DQKV-VCLLSMNDKKWVWHKRLGHANWRLISKISKL-QL 705
GK+V N+Y ++ L+ D + V ++ +D +WHKRLGH + + + +S L
Sbjct: 372 MGKQVANLYIVDLDSLSHPGTDSSITVASVTSHD---LWHKRLGHPSVQKLQPMSSLLSF 428
Query: 706 VKGLPNIDYHSDALCGACQKGKIVKSSFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSK 765
K N D+H C C K F S + S SRP +L+HID +GP + + G +
Sbjct: 429 PKQKNNTDFH----CRVCHISKQKHLPFVSHNNKS-SRPFDLIHIDTWGPFSVQTHDGYR 483
Query: 766 YGLVIVDDYSRWTWVKFIKSKDYACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFE 825
Y L IVDDYSR TWV +++K V +F T ++++ E I VRSD E F
Sbjct: 484 YFLTIVDDYSRATWVYLLRNKSDVLTVIPTFVTMVENQFETTIKGVRSDNAPELN---FT 540
Query: 826 LFCEKHGILHEFSSPRTPQQNGVVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQ 885
F GI+ S P TPQQN VVERK++ + +AR++ ++++ +W + + T+ Y+
Sbjct: 541 QFYHSKGIVPYHSCPETPQQNSVVERKHQHILNVARSLFFQSHIPISYWGDCILTAVYLI 600
Query: 886 NRIYIRPMLEKTTYELFKGRRPNISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSE 945
NR+ + +K +E+ P + FGC CY + KF +A+ F+GY
Sbjct: 601 NRLPAPILEDKCPFEVLTKTVPTYDHIKVFGCLCYASTSPKDRHKFSPRAKACAFIGYPS 660
Query: 946 RSKAYRVYNSETQCVEESMHVKFDDRE---PGSKTSEQSES---NAGTTDSEDASESDQP 999
K Y++ + ET + S HV F + GS S++ ++ + T SD
Sbjct: 661 GFKGYKLLDLETHSIIVSRHVVFHEELFPFLGSDLSQEEQNFFPDLNPTPPMQRQSSDHV 720
Query: 1000 SDSEKYTEVESSPEAE-YNSEAEPSPKVQNESASEDFQDNTQQVIQPKFKHK--SSHPEE 1056
+ S+ + VE P A N+ EPS + + A + +Q + H SS P E
Sbjct: 721 NPSDSSSSVEILPSANPTNNVPEPSVQTSHRKAKKP------AYLQDYYCHSVVSSTPHE 774
Query: 1057 LIIGSKDSPRRTRSHFRQEESLIGLLSII----EPKTVEEALSDDGWILAMQEELINFKG 1112
+ R+ S+ R + + L+ + EP EA W AM E +G
Sbjct: 775 I--------RKFLSYDRINDPYLTFLACLDKTKEPSNYTEAEKLQVWRDAMGAEFDFLEG 826
Query: 1113 MMCGIWYPYPFRRTLLEQNGYSKTS*MNKEK*PETKPDFVAQGYSQQEGIDYTETFAPVA 1172
P + + K + K VAQGY+Q+EGIDY ETF+PVA
Sbjct: 827 THTWEVCSLPADKRCIGCRWIFKIKYNSDGSVERYKARLVAQGYTQKEGIDYNETFSPVA 886
Query: 1173 RLEAIRLLLSYAINHGIILYQMDVKSAFLNGVIEEEVYVKQPPGFE----DLKHPDHVYK 1228
+L +++LLL A + L Q+D+ +AFLNG ++EE+Y++ P G+ D P+ V +
Sbjct: 887 KLNSVKLLLGVAARFKLSLTQLDISNAFLNGDLDEEIYMRLPQGYASRQGDSLPPNAVCR 946
Query: 1229 LKKSLYGLKQAPRAWYDRLSNFLIKNDFERGQVDTTLFRRTLKKDILIVQIYVDDIIFGS 1288
LKKSLYGLKQA R WY + S+ L+ F + D T F + L V +Y+DDII S
Sbjct: 947 LKKSLYGLKQASRQWYLKFSSTLLGLGFIQSYCDHTCFLKISDGIFLCVLVYIDDIIIAS 1006
Query: 1289 TNASLCK*FSKLMQDEFEMSMMGELKFFLGIQINQSKEGVYVHQTKYTKELLKKFKLEDC 1348
N + M+ F++ +GELK+FLG++I +S +G+++ Q KY +LL + C
Sbjct: 1007 NNDAAVDILKSQMKSFFKLRDLGELKYFLGLEIVRSDKGIHISQRKYALDLLDETGQLGC 1066
Query: 1349 KVMNTPMHPTCTLSKEDTGTVVDQKLYRGMIGSLLYLTASRPAILFSVCLCARFQSDPRE 1408
K + PM P+ + + G V+ YR +IG L+YL +RP I F+V A+F PR+
Sbjct: 1067 KPSSIPMDPSMVFAHDSGGDFVEVGPYRRLIGRLMYLNITRPDITFAVNKLAQFSMAPRK 1126
Query: 1409 SHLTAVKRIFRYLKGTTNLGLLYRKSLDYKLIGFCDADYAGDRIERKSTSGNCQFLGENL 1468
+HL AV +I +Y+KGT GL Y + + +L + +ADY R R+STSG C FLG++L
Sbjct: 1127 AHLQAVYKILQYIKGTIGQGLFYSATSELQLKVYANADYNSCRDSRRSTSGYCMFLGDSL 1186
Query: 1469 ISWASKRQATIAMSTAEAEYISAASCCTQLLWMKHQLEDYQIN-ANSIPIYCDNTAAICL 1527
I W S++Q ++ S+AEAEY S + +L+W+ + L++ Q+ + ++CDN AAI +
Sbjct: 1187 ICWKSRKQDVVSKSSAEAEYRSLSVATDELVWLTNFLKELQVPLSKPTLLFCDNEAAIHI 1246
Query: 1528 SKNPILHSRAKHIEIKHHFIRDYVQKGILDIQFIDIEHQWADIFTKPLSVERF 1580
+ N + H R KHIE H +R+ + KG+ ++ I+ E Q AD FTKPL F
Sbjct: 1247 ANNHVFHERTKHIESDCHSVRERLLKGLFELYHINTELQIADPFTKPLYPSHF 1299
>At4g10690 retrotransposon like protein
Length = 1515
Score = 514 bits (1323), Expect = e-145
Identities = 346/1107 (31%), Positives = 541/1107 (48%), Gaps = 83/1107 (7%)
Query: 542 MQVCLRAREKQRSWYLDSGCSRHMTGEK-ALFLTLTMKDGGEVKFGGNQTGKIIGTGTI- 599
M+V + + W DS + H+T L + T V G I GTI
Sbjct: 311 MRVSDQNQASSHEWLPDSAATAHITNTTDGLQNSQTYSGDDSVIVGNGDFLPITHIGTIP 370
Query: 600 ---GNSSISINNVWLVDGLKHNLLSISQFCDNGYDVTFSKTNCTLVNKDDKS--ITFKGK 654
++ + +V + G+ +LLS+S+ D+ Y +F+ + ++V KD ++ + +G
Sbjct: 371 LNISQGTLPLEDVLVCPGITKSLLSVSKLTDD-YPCSFTFDSDSVVIKDKRTQQLLTQGN 429
Query: 655 RVENVYKINFSDLADQKVVCLLSMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDY 714
+ + +Y + D+ Q + VWH+RLGH N ++ + K + + ++
Sbjct: 430 KHKGLYVLK--DVPFQTYYSTRQQSSDDEVWHQRLGHPNKEVLQHLIKTKAIV----VNK 483
Query: 715 HSDALCGACQKGKIVKSSFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDY 774
S +C ACQ GK+ + F + + VS SRPLE +H DL+GP S G +Y ++ +D+Y
Sbjct: 484 TSSNMCEACQMGKVCRLPFVASEFVS-SRPLERIHCDLWGPAPVTSAQGFQYYVIFIDNY 542
Query: 775 SRWTWVKFIKSKDYACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGIL 834
SR+TW +K K VF F ++++ + KI + D GGEF + F GI
Sbjct: 543 SRFTWFYPLKLKSDFFSVFVLFQQLVENQYQHKIAMFQCDGGGEFVSYKFVAHLASCGIK 602
Query: 835 HEFSSPRTPQQNGVVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPML 894
S P TPQQNG+ ER++R L E+ +++ + + W EA TS ++ N + +
Sbjct: 603 QLISCPHTPQQNGIAERRHRYLTELGLSLMFHSKVPHKLWVEAFFTSNFLSNLLPSSTLS 662
Query: 895 E-KTTYELFKGRRPNISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVY 953
+ K+ YE+ G P + FG CY KFD K+ +FLGY+ + K YR
Sbjct: 663 DNKSPYEMLHGTPPVYTALRVFGSACYPYLRPYAKNKFDPKSLLCVFLGYNNKYKGYRCL 722
Query: 954 NSETQCVEESMHVKFDDRE-PGSKTSEQSESNAGT------------------------- 987
+ T V HV FD+R+ P S Q ++ +G+
Sbjct: 723 HPPTGKVYICRHVLFDERKFPYSDIYSQFQTISGSPLFTAWQKGFSSTALSRETPSTNVE 782
Query: 988 -------TDSEDASESDQPSDSEKYTEVESSPEAEYNSEAEPSPKVQN-------ESASE 1033
T S P+ +E T + A ++ PSP ES S+
Sbjct: 783 DIIFPSATVSSSVPTGCAPNIAETATAPDVDVAAAHDMVVPPSPITSTSLPTQPEESTSD 842
Query: 1034 DFQDNT------------QQVIQPKFKHKSSHPEELIIGSKDSP--------RRTRSHFR 1073
+T Q + F+ P + +I S + R +S
Sbjct: 843 QNHYSTDSETAISSAMTPQSINVSLFEDSDFPPLQSVISSTTAAPETSHPMITRAKSGIT 902
Query: 1074 QEESLIGLLSII----EPKTVEEALSDDGWILAMQEELINFKGMMCGIWYPYPFRRTLLE 1129
+ L S+ EPK+V+EAL D+GW AM EE+ P LL
Sbjct: 903 KPNPKYALFSVKSNYPEPKSVKEALKDEGWTNAMGEEMGTMHETDTWDLVPPEMVDRLLG 962
Query: 1130 QNGYSKTS*MNKEK*PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGI 1189
KT + K VA+GY Q+EG+DY ET++PV R +R +L A +
Sbjct: 963 CKWVFKTKLNSDGSLDRLKARLVARGYEQEEGVDYVETYSPVVRSATVRSILHVATINKW 1022
Query: 1190 ILYQMDVKSAFLNGVIEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLSN 1249
L Q+DVK+AFL+ ++E V++ QPPGFED PD+V KLKK++Y LKQAPRAW+D+ S+
Sbjct: 1023 SLKQLDVKNAFLHDELKETVFMTQPPGFEDPSRPDYVCKLKKAIYDLKQAPRAWFDKFSS 1082
Query: 1250 FLIKNDFERGQVDTTLFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSM 1309
+L+K F D +LF +D++ + +YVDD+I N L + ++ EF M
Sbjct: 1083 YLLKYGFICSFSDPSLFVYLKGRDVMFLLLYVDDMILTGNNDVLLQQLLNILSTEFRMKD 1142
Query: 1310 MGELKFFLGIQINQSKEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGTV 1369
MG L +FLGIQ + +G+++ Q KYT +LL + DC M TP+ L + +
Sbjct: 1143 MGALHYFLGIQAHYHNDGLFLSQEKYTSDLLVNAGMSDCSSMPTPLQ--LDLLQGNNKPF 1200
Query: 1370 VDQKLYRGMIGSLLYLTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGL 1429
+ +R + G L YLT +RP I F+V + P S +KRI YLKGT +G+
Sbjct: 1201 PEPTYFRRLAGKLQYLTLTRPDIQFAVNFVCQKMHAPTMSDFHLLKRILHYLKGTMTMGI 1260
Query: 1430 LYRKSLDYKLIGFCDADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEYI 1489
+ D L + D+D+AG + R+ST G C FLG N+ISW++KR T++ S+ EAEY
Sbjct: 1261 NLSSNTDSVLRCYSDSDWAGCKDTRRSTGGFCTFLGYNIISWSAKRHPTVSKSSTEAEYR 1320
Query: 1490 SAASCCTQLLWMKHQLEDYQINANSIP-IYCDNTAAICLSKNPILHSRAKHIEIKHHFIR 1548
+ + +++ W+ L++ + IP +YCDN +A+ LS NP LHSR+KH ++ ++++R
Sbjct: 1321 TLSFAASEVSWIGFLLQEIGLPQQQIPEMYCDNLSAVYLSANPALHSRSKHFQVDYYYVR 1380
Query: 1549 DYVQKGILDIQFIDIEHQWADIFTKPL 1575
+ V G L ++ I Q ADIFTK L
Sbjct: 1381 ERVALGALTVKHIPASQQLADIFTKSL 1407
>At3g45520 copia-like polyprotein
Length = 1363
Score = 501 bits (1291), Expect = e-141
Identities = 322/1062 (30%), Positives = 541/1062 (50%), Gaps = 50/1062 (4%)
Query: 555 WYLDSGCSRHMTGEKALFLTLTMKDGGEVKFGGNQTGKIIGTGTI-----GNSSISINNV 609
W +D+GC HMT ++ + GG V+ G ++ G GT+ ++++ NV
Sbjct: 316 WIMDTGCIYHMTHKREWLEDFDEEAGGSVRMGNKSISRVKGVGTVRIVNDNGLTVTLQNV 375
Query: 610 WLVDGLKHNLLSISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKINFSDLAD 669
+ + NLLS+ F G+ F N L K + +G+R + +Y ++ D
Sbjct: 376 RYIPDMDRNLLSLGTFEKAGHK--FESENGMLRIKSGNQVLLEGRRYDTLYILHGKPATD 433
Query: 670 QKVVCLLSMNDKKWVWHKRLGHANWRLISKISKLQLV--KGLPNIDYHSDALCGACQKGK 727
+ + + ND +WH+RL H + + +S + K + K + +D D + G +K
Sbjct: 434 ESLA-VARANDDTVLWHRRLCHMSQKNMSLLIKKGFLDKKKVSMLDTCEDCIYGRAKK-- 490
Query: 728 IVKSSFKSKDIVSTSRPLELLHIDLFG-PVNTASLYGSKYGLVIVDDYSRWTWVKFIKSK 786
+ + D T + LE +H DL+G P SL +Y + +DDY+R WV F+K+K
Sbjct: 491 -IGFNLAQHD---TKKKLEYVHSDLWGAPTVPMSLGNCQYFISFIDDYTRKVWVYFLKTK 546
Query: 787 DYACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRTPQQN 846
D A E F S+ + ++++ ++ +R+D G EF N F+ FCE+ G + TPQQN
Sbjct: 547 DEAFEKFVSWISLVENQSGERVKTLRTDNGLEFCNRMFDGFCEEKGFQRHRTCAYTPQQN 606
Query: 847 GVVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYELFKGRR 906
GVVER NRT+ E R+M+ ++ L K FWAEA +T+ + N+ + + + + G+
Sbjct: 607 GVVERMNRTIMEKVRSMLCDSGLPKRFWAEATHTAVLLINKTPCSAINFEFPDKRWSGKA 666
Query: 907 PNISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVEESMHV 966
P SY ++GC ++ K + +A++G+ +GY K Y+V+ E + S +V
Sbjct: 667 PIYSYLRRYGCVTFVHTDGG---KLNLRAKKGVLIGYPSGVKGYKVWLIEEKKCVVSRNV 723
Query: 967 KFDDREPGSKTSEQSESNAGTTDSEDASESDQPSDSEKYTEVESSPEAEYNSEAEPSPKV 1026
F + ++ E + D S D +++K ++S E S+A+ +P
Sbjct: 724 SFQENAVYKDLMQRKEQVSCEEDDHAGSYIDLDLEADK----DNSSGGE-QSQAQVTPAT 778
Query: 1027 QNESASEDFQDNTQQVIQPKFKHKSSHPEELIIGSKDSPRRTRSHFRQEESLIGLL---- 1082
+ S + T + + H+S L+ + R F E+ L
Sbjct: 779 RGAVTSTPPRYETDDIEETDV-HQSPLSYHLVRDRERREIRAPRRFDDEDYYAEALYTTE 837
Query: 1083 --SIIEPKTVEEALSD---DGWILAMQEELINFKGMMCGIWYPYPFRRTLLEQNG-YSKT 1136
+EP +EA+ D D W LAM EE+ + P ++ ++ Y
Sbjct: 838 DGDAVEPADYKEAVRDENWDKWRLAMNEEIESQLKNDTWTTVTRPEKQRIIGSRWIYKYK 897
Query: 1137 S*MNKEK*PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQMDV 1196
+ + P K VA+GY+Q+EG+DY E FAPV + +IR+LLS + L Q+DV
Sbjct: 898 QGIPGVEEPRFKARLVAKGYAQREGVDYHEIFAPVVKHVSIRILLSIVAQENLELEQLDV 957
Query: 1197 KSAFLNGVIEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDF 1256
K+AFL+G ++E++Y+ P G E L + V L KSLYGLKQAPR W ++ ++++ + F
Sbjct: 958 KTAFLHGELKEKIYMMPPEGCESLFKENEVCLLNKSLYGLKQAPRQWNEKFNHYMTEIGF 1017
Query: 1257 ERGQVDTTLFRRTLKKD-ILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELKF 1315
+R D+ + + L D + + YVDD++ + N K + +FEM +G K
Sbjct: 1018 KRSDYDSCAYTKKLSDDSTMYLLFYVDDMLVAANNMQAIDALKKELSIKFEMKDLGAAKK 1077
Query: 1316 FLGIQINQSKEG--VYVHQTKYTKELLKKFKLEDCKVMNTPM--HPTCTLSKEDTGTVVD 1371
LGI+I +E +++ Q Y ++LK F + + K TP+ H + E+ + +
Sbjct: 1078 ILGIEIIIDREAGVLWLSQESYLNKVLKTFNMLESKPALTPLGAHLKMKSATEEKLSTEE 1137
Query: 1372 QKL----YRGMIGSLLY-LTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTN 1426
+ + Y +GS++Y + +RP + + V + +RF S P + H VK + RY+KGT +
Sbjct: 1138 EYMNSVPYSSAVGSIMYAMIGTRPDLAYPVGVVSRFMSQPAKEHWLGVKWVLRYIKGTVD 1197
Query: 1427 LGLLYRKSLDYKLIGFCDADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEA 1486
L Y+++ D+ + G+CDADYA D +R+S +G LG N ISW S Q +A S+ E
Sbjct: 1198 TRLCYKRNSDFSICGYCDADYAADLDKRRSITGLVFTLGGNTISWKSGLQRVVAQSSTEC 1257
Query: 1487 EYISAASCCTQLLWMKHQLEDYQINANSIPIYCDNTAAICLSKNPILHSRAKHIEIKHHF 1546
EY+S + +W+K L+D+ ++ I+CD+ +AI LSKN + H R KHI++K HF
Sbjct: 1258 EYMSLTEAVKEAIWLKGLLKDFGYEQKNVEIFCDSQSAIALSKNNVHHERTKHIDVKFHF 1317
Query: 1547 IRDYVQKGILDIQFIDIEHQWADIFTKPLSVERF----DFIK 1584
IR+ + G +++ I E ADIFTK L V +F DF++
Sbjct: 1318 IREIIADGKVEVSKISTEKNPADIFTKVLPVNKFQTALDFLR 1359
>At1g37110
Length = 1356
Score = 499 bits (1286), Expect = e-141
Identities = 338/1081 (31%), Positives = 546/1081 (50%), Gaps = 100/1081 (9%)
Query: 555 WYLDSGCSRHMTGEKALFLTLTMKDGGEVKFGGNQTGKIIGTGTI-----GNSSISINNV 609
W LDSGC+ HMT + F++ K + G + + + G GTI G + + NV
Sbjct: 308 WILDSGCTSHMTSRRDWFISFQEKGNTTILLGDDHSVESQGQGTIRIDTHGGTIKILENV 367
Query: 610 WLVDGLKHNLLSISQFCDNGY-------DVTFSKTNCTLVNKDDKSITFKGKRVENVYKI 662
V L+ NL+S GY V + K N T + +G +Y +
Sbjct: 368 KYVPHLRRNLISTGTLDKLGYRHEGGEGKVRYFKNNKTAL---------RGSLSNGLYVL 418
Query: 663 NFSDLADQKVVCLLSMNDKKW-VWHKRLGHANWRLISKISKLQLV--KGLPNIDYHSDAL 719
+ S + + +C + K +WH RLGH + + ++ L+ K + +++
Sbjct: 419 DGSTVMSE--LCNAETDKVKTALWHSRLGHMSMNNLKVLAGKGLIDRKEINELEF----- 471
Query: 720 CGACQKGKIVKSSFKSKDIVSTSRPLELLHIDLFGPVN-TASLYGSKYGLVIVDDYSRWT 778
C C GK K SF S L +H DL+G N T S+ G +Y L I+DD +R
Sbjct: 472 CEHCVMGKSKKVSFNVGKHTSEDA-LSYVHADLWGSPNVTPSISGKQYFLSIIDDKTRKV 530
Query: 779 WVKFIKSKDYACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFS 838
W+ F+KSKD + F + + ++++ K+ +R+D G EF N F+ +C++HGI +
Sbjct: 531 WLYFLKSKDETFDKFCEWKSLVENQVNKKVKCLRTDNGLEFCNSRFDSYCKEHGIERHRT 590
Query: 839 SPRTPQQNGVVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTT 898
TPQQNGV ER NRT+ E R +++++ + + FWAEA T+ Y+ NR +
Sbjct: 591 CTYTPQQNGVAERMNRTIMEKVRCLLNKSGVEEVFWAEAAATAAYLINRSPASAINHNVP 650
Query: 899 YELFKGRRPNISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYN-SET 957
E++ R+P + +FG Y+ + LK +A +G FLGY +K Y+V+ E
Sbjct: 651 EEMWLNRKPGYKHLRKFGSIAYVHQDQGKLKP---RALKGFFLGYPAGTKGYKVWLLEEE 707
Query: 958 QCV-------EES-----MHVKFDDREPGSK---TSEQSESNAGTTDSEDASESDQPSDS 1002
+CV +ES + VK DD + ++ TS + E N S SDS
Sbjct: 708 KCVISRNVVFQESVVYRDLKVKEDDTDNLNQKETTSSEVEQNKFAEASGSGGVIQLQSDS 767
Query: 1003 EKYTEVESSP----EAEYNSEAEPSPKVQNESASEDFQDNTQQVIQP--KFKHKSSHPEE 1056
E TE E S E EY+ + + +PK + + +D ++ I P +F +SS
Sbjct: 768 EPITEGEQSSDSEEEVEYSEKTQETPKRTGLTTYKLARDRVRRNINPPTRFTEESSVTFA 827
Query: 1057 LIIGSKDSPRRTRSHFRQEESLIGLLSIIEPKTVEEALSD---DGWILAMQEELINFKGM 1113
L++ E+ I + EP++ +EA+ + W +A +E+ + M
Sbjct: 828 LVV---------------VENCI----VQEPQSYQEAMESQDCEKWDMATHDEMDSL--M 866
Query: 1114 MCGIW--YPYPFRRTLLEQNGYSKTS*MNKEK*PETKPD-----FVAQGYSQQEGIDYTE 1166
G W P R ++ K K P +P VA+GY+Q+EG+DY E
Sbjct: 867 KNGTWDLVDKPKDRKIIGCRWLFKL----KSGIPGVEPTRFKARLVAKGYTQREGVDYQE 922
Query: 1167 TFAPVARLEAIRLLLSYAINHGIILYQMDVKSAFLNGVIEEEVYVKQPPGFEDLKHPDHV 1226
FAPV + +IR+L+S ++ + L QMDVK+ FL+G +EEE+Y++QP GF + V
Sbjct: 923 IFAPVVKHVSIRILMSLVVDKDLELEQMDVKTTFLHGDLEEELYMEQPEGFVSDSSENKV 982
Query: 1227 YKLKKSLYGLKQAPRAWYDRLSNFLIKNDFERGQVDTTLF-RRTLKKDILIVQIYVDDII 1285
+LKKSLYGLKQ+PR W R F+ F R + D ++ + + D + + +YVDD++
Sbjct: 983 CRLKKSLYGLKQSPRQWNKRFDRFMSSQQFIRSEHDACVYVKHVSEHDFIYLLLYVDDML 1042
Query: 1286 FGSTNASLCK*FSKLMQDEFEMSMMGELKFFLGIQINQSKEG--VYVHQTKYTKELLKKF 1343
+ + + + EFEM MG LGI I + ++G + + Q Y +++L +F
Sbjct: 1043 IAGASKAEINRVKEQLSTEFEMKDMGGASRILGIDIYRDRKGGVLKLSQEIYIRKVLDRF 1102
Query: 1344 KLEDCKVMNTPMHPTCTLS---KEDTGTVVDQKLYRGMIGSLLY-LTASRPAILFSVCLC 1399
+ K+ N P+ L+ +ED D Y +GS++Y + +RP + +++CL
Sbjct: 1103 NMSGAKMTNAPVGAHFKLAAVREEDECVDTDVVPYSSAVGSIMYAMLGTRPDLAYAICLI 1162
Query: 1400 ARFQSDPRESHLTAVKRIFRYLKGTTNLGLLYRKSLDYKLIGFCDADYAGDRIERKSTSG 1459
+R+ S P H AVK + RYLKG +L L++ K D+ + G+CD++YA D R+S SG
Sbjct: 1163 SRYMSKPGSMHWEAVKWVMRYLKGAQDLNLVFTKEKDFTVTGYCDSNYAADLDRRRSISG 1222
Query: 1460 NCQFLGENLISWASKRQATIAMSTAEAEYISAASCCTQLLWMKHQLEDYQINANSIPIYC 1519
+G N +SW + Q +AMST EAEYI+ A + +W+K L+D + + + I+C
Sbjct: 1223 YVFTIGGNTVSWKASLQPVVAMSTTEAEYIALAEAAKEAMWIKGLLQDMGMQQDKVKIWC 1282
Query: 1520 DNTAAICLSKNPILHSRAKHIEIKHHFIRDYVQKGILDIQFIDIEHQWADIFTKPLSVER 1579
D+ +AICLSKN + H R KHI+++ ++IRD V+ G +D+ I D TK + V +
Sbjct: 1283 DSQSAICLSKNSVYHERTKHIDVRFNYIRDVVESGDVDVLKIHTSRNPVDALTKCIPVNK 1342
Query: 1580 F 1580
F
Sbjct: 1343 F 1343
>At2g13930 putative retroelement pol polyprotein
Length = 1335
Score = 486 bits (1250), Expect = e-137
Identities = 328/1069 (30%), Positives = 543/1069 (50%), Gaps = 71/1069 (6%)
Query: 555 WYLDSGCSRHMTGEKALFLTLTMKDGGEVKFGGNQTGKIIGTGTI-----GNSSISINNV 609
W LD+GCS HMT K F G VK G + + G G+I S + + +V
Sbjct: 285 WVLDTGCSFHMTPRKDWFKDFKELSSGYVKMGNDTYSPVKGIGSIKIRNSDGSQVILTDV 344
Query: 610 WLVDGLKHNLLSISQFCDNGYDVTFSKTNCTLVNKDDKSITFKGKRVENVYKINFSDLAD 669
+ + NL+S+ D G F + L S KG++ + +Y ++ +
Sbjct: 345 RYMPNMTRNLISLGTLEDRG--CWFKSQDGILKIVKGCSTILKGQKRDTLYILD-GVTEE 401
Query: 670 QKVVCLLSMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDA-LCGACQKGKI 728
+ + D+ +WH RLGH + K ++ + KG + + C C GK
Sbjct: 402 GESHSSAEVKDETALWHSRLGHMS----QKGMEILVKKGCLRREVIKELEFCEDCVYGKQ 457
Query: 729 VKSSFKSKDIVSTSRPLELLHIDLFG-PVNTASLYGSKYGLVIVDDYSRWTWVKFIKSKD 787
+ SF V T L +H DL+G P N ASL S+Y + VDDYSR W+ F++ KD
Sbjct: 458 HRVSFAPAQHV-TKEKLAYVHSDLWGSPHNPASLGNSQYFISFVDDYSRKVWIYFLRKKD 516
Query: 788 YACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRTPQQNG 847
A E F + ++++ + K+ K+R+D G E+ N FE FC++ GI+ + TPQQNG
Sbjct: 517 EAFEKFVEWKKMVENQSDRKVKKLRTDNGLEYCNHYFEKFCKEEGIVRHKTCAYTPQQNG 576
Query: 848 VVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYELFKGRRP 907
+ ER NRT+ + R+M+ + + K FWAEA +T+ Y+ NR + E + G P
Sbjct: 577 IAERLNRTIMDKVRSMLSRSGMEKKFWAEAASTAVYLINRSPSTAINFDLPEEKWTGALP 636
Query: 908 NISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVEESMHVK 967
++S +FGC YI + K + ++++GIF Y E K Y+V+ E + S +V
Sbjct: 637 DLSSLRKFGCLAYIHADQG---KLNPRSKKGIFTSYPEGVKGYKVWVLEDKKCVISRNVI 693
Query: 968 FDDREPGSKTSEQSESNAGTTDSEDASESDQPSDSE-------------------KYTEV 1008
F ++ S++ +D ED + +D E + V
Sbjct: 694 FREQVMFKDLKGDSQNTISESDLEDLRVNPDMNDQEFTDQGGATQDNSNPSEATTSHNPV 753
Query: 1009 ESSP--EAEYNSEAEPSPKVQNESASEDFQDNTQQVIQPKFKHKSSHPEELIIGSKDSPR 1066
+SP + SE E S V++ S + +D ++ I+ K+ S+ S+D +
Sbjct: 754 LNSPTHSQDEESEEEDSDAVEDLSTYQLVRDRVRRTIKANPKYNESNMVGFAYYSEDDGK 813
Query: 1067 RTRSHFRQEESLIGLLSIIEPKTVEEALSD---DGWILAMQEELINFKGMMCGIWYPYPF 1123
EPK+ +EAL D + W AM+EE+++ P
Sbjct: 814 P------------------EPKSYQEALLDPDWEKWNAAMKEEMVSMSKNHTWDLVTKPE 855
Query: 1124 RRTLLEQNG-YSKTS*MNKEK*PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLS 1182
+ L+ +++ + + + P VA+G++Q+EG+DY E F+PV + +IR LLS
Sbjct: 856 KVKLIGCRWVFTRKAGIPGVEAPRFIARLVAKGFTQKEGVDYNEIFSPVVKHVSIRYLLS 915
Query: 1183 YAINHGIILYQMDVKSAFLNGVIEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRA 1242
+++ + L QMDVK+AFL+G +EEE+Y+ QP GFE + + V LK+SLYGLKQ+PR
Sbjct: 916 MVVHYNMELQQMDVKTAFLHGFLEEEIYMAQPEGFEIKRGSNKVCLLKRSLYGLKQSPRQ 975
Query: 1243 WYDRLSNFLIKNDFERGQVDTTLFRRTLKKDILI-VQIYVDDIIFGSTNASLCK*FSKLM 1301
W R F+ + R D+ ++ + D I + +YVDD++ S N S +L+
Sbjct: 976 WNLRFDEFMRGIKYTRSAYDSCVYFKKCNGDTYIYLLLYVDDMLIASANKSEVNELKQLL 1035
Query: 1302 QDEFEMSMMGELKFFLGIQINQSKEG--VYVHQTKYTKELLKKFKLEDCKVMNTPMHPTC 1359
EFEM +G+ K LG++I++ ++ + + Q Y K++L+ F++++ K ++TP+
Sbjct: 1036 SREFEMKDLGDAKKILGMEISRDRDAGLLTLSQEGYVKKVLRSFQMDNAKPVSTPLGIHF 1095
Query: 1360 TLS----KEDTGTVVDQKL--YRGMIGSLLY-LTASRPAILFSVCLCARFQSDPRESHLT 1412
L KE K+ Y IGS++Y + +RP + +S+ + +RF S P + H
Sbjct: 1096 KLKAATDKEYEEQFERMKIVPYANTIGSIMYSMIGTRPDLAYSLGVISRFMSKPLKDHWQ 1155
Query: 1413 AVKRIFRYLKGTTNLGLLYRKSLDYKLIGFCDADYAGDRIERKSTSGNCQFLGENLISWA 1472
AVK + RY++GT L +RK D+ L G+CD+DY + R+S +G +G N ISW
Sbjct: 1156 AVKWVLRYMRGTEKKKLCFRKQEDFLLRGYCDSDYGSNFDTRRSITGYVFTVGGNTISWK 1215
Query: 1473 SKRQATIAMSTAEAEYISAASCCTQLLWMKHQLEDYQINANSIPIYCDNTAAICLSKNPI 1532
SK Q +A+S+ EAEY++ + LW+K + + + + ++ D+ +AI L+KN +
Sbjct: 1216 SKLQKVVAISSTEAEYMALTEAVKEALWLKGFAAELGHSQDYVEVHSDSQSAITLAKNSV 1275
Query: 1533 LHSRAKHIEIKHHFIRDYVQKGILDIQFIDIEHQWADIFTKPLSVERFD 1581
H R KHI+I+ HFIRD + G++ + I E A+IFTK + + +F+
Sbjct: 1276 HHERTKHIDIRLHFIRDIICAGLIKVVKIATECNPANIFTKTVPLAKFE 1324
>At1g26990 polyprotein, putative
Length = 1436
Score = 483 bits (1243), Expect = e-136
Identities = 344/1058 (32%), Positives = 528/1058 (49%), Gaps = 87/1058 (8%)
Query: 553 RSWYLDSGCSRHMTGEKALFLTLTMKDGGEVKFGGNQTGKIIGTGTIG-NSSISINNVWL 611
R+W +DSG S H+T E+ L+ T D V+ T KI GTG I ++S++NV
Sbjct: 419 RAWVIDSGASHHVTHERNLYHTYKALDRTFVRLPNGHTVKIEGTGFIQLTDALSLHNVLF 478
Query: 612 VDGLKHNLLSISQFCDNGYD-VTFSKTNCTLVNKDDKSITFKGKRVENVYKINFS-DLAD 669
+ K NLLS+S V+F+ C + + + KG +V N+Y +N L D
Sbjct: 479 IPEFKFNLLSVSVLTKTLQSKVSFTSDECMIQALTKELMLGKGSQVGNLYILNLDKSLVD 538
Query: 670 Q-----KVVCLLSMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHSDAL-CGAC 723
K VC N+ + +WHKRLGH ++ +KI L V LP + D+ C C
Sbjct: 539 VSSFPGKSVCSSVKNESE-MWHKRLGHPSF---AKIDTLSDVLMLPKQKINKDSSHCHVC 594
Query: 724 QKGKIVKSSFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSRWTWVKFI 783
K FKS + + + EL+HID +GP + ++ +Y L IVDD+SR TW+ +
Sbjct: 595 HLSKQKHLPFKSVNHIR-EKAFELVHIDTWGPFSVPTVDSYRYFLTIVDDFSRATWIYLL 653
Query: 784 KSKDYACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHEFSSPRTP 843
K K VF SF ++++ K+ VRSD E + ELF K GI + P TP
Sbjct: 654 KQKSDVLTVFPSFLKMVETQYHTKVCSVRSDNAHELKFN--ELFA-KEGIKADHPCPETP 710
Query: 844 QQNGVVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEKTTYELFK 903
+QN VVERK++ L +AR ++ ++ + +W + V T+ ++ NR+ + +T YE
Sbjct: 711 EQNFVVERKHQHLLNVARALMFQSGIPLEYWGDCVLTAVFLINRLLSPVINNETPYERLT 770
Query: 904 GRRPNISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSETQCVEES 963
+P+ S FGC CY + KFD +A+ IFLGY K Y++ + ET V S
Sbjct: 771 KGKPDYSSLKAFGCLCYCSTSPKSRTKFDPRAKACIFLGYPMGYKGYKLLDIETYSVSIS 830
Query: 964 MHVKF-DDREPGSKTSEQSESNAGTTDSEDASESDQPSDSEKYTEVESSPEAEYNSEAEP 1022
HV F +D P + S D P++ E V+SS +A +N +
Sbjct: 831 RHVIFYEDIFPFAS----SNITDAAKDFFPHIYLPAPNNDEHLPLVQSSSDAPHNHD--- 883
Query: 1023 SPKVQNESASEDFQDNTQQVIQPKFKHKSSHPEEL--IIGSKDSPRRTR----------S 1070
ES+S F + +PK + P L ++P T+ S
Sbjct: 884 ------ESSSMIFVPS-----EPKSTRQRKLPSHLQDFHCYNNTPTTTKTSPYPLTNYIS 932
Query: 1071 HFRQEESLIGLLSIIE----PKTVEEALSDDGWILAMQEELINFKGMMCGIWY--PYPFR 1124
+ E ++II P+ EA D W AM +E+ F + G W P
Sbjct: 933 YSYLSEPFGAFINIITATKLPQKYSEARLDKVWNDAMGKEISAF--VRTGTWSICDLPAG 990
Query: 1125 RTLLEQNGYSKTS*MNKEK*PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYA 1184
+ + + K VA+GY+QQEGID+ TF+PVA++ +++LLS A
Sbjct: 991 KVAVGCKWIITIKFLADGSIERHKARLVAKGYTQQEGIDFFNTFSPVAKMVTVKVLLSLA 1050
Query: 1185 INHGIILYQMDVKSAFLNGVIEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWY 1244
L+Q+D+ +A LNG +EEE+Y+K PPG+ +++ G + +P A
Sbjct: 1051 PKMKWYLHQLDISNALLNGDLEEEIYMKLPPGYSEIQ-------------GQEVSPNA-- 1095
Query: 1245 DRLSNFLIKNDFERGQVDTTLFRRTLKKDILIVQIYVDDIIFGSTNASLCK*FSKLMQDE 1304
+ D TLF + L+V +YVDDI+ ST + + +
Sbjct: 1096 -------------KCHGDHTLFVKAQDGFFLVVLVYVDDILIASTTEAASAELTSQLSSF 1142
Query: 1305 FEMSMMGELKFFLGIQINQSKEGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCTLSKE 1364
F++ +GE KFFLGI+I ++ +G+ + Q KY +LL DCK + PM P LSK
Sbjct: 1143 FQLRDLGEPKFFLGIEIARNADGISLCQRKYVLDLLASSDFSDCKPSSIPMEPNQKLSK- 1201
Query: 1365 DTGTVV-DQKLYRGMIGSLLYLTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKG 1423
DTGT++ D K YR ++G L YL +RP I F+V A++ S P + HL A+ +I RYLKG
Sbjct: 1202 DTGTLLEDGKQYRRILGKLQYLCLTRPDINFAVSKLAQYSSAPTDIHLQALHKILRYLKG 1261
Query: 1424 TTNLGLLYRKSLDYKLIGFCDADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAMST 1483
T GL Y ++ L GF D+D+ R+ +G F+G +L+SW SK+Q ++MS+
Sbjct: 1262 TIGQGLFYGADTNFDLRGFSDSDWQTCPDTRRCVTGFAIFVGNSLVSWRSKKQDVVSMSS 1321
Query: 1484 AEAEYISAASCCTQLLWMKHQLEDYQIN-ANSIPIYCDNTAAICLSKNPILHSRAKHIEI 1542
AEAEY + + +L+W+ + L ++I + +YCDN AA+ ++ N + H R KHIE
Sbjct: 1322 AEAEYRAMSVATKELIWLGYILTAFKIPFTHPAYLYCDNEAALHIANNSVFHERTKHIEN 1381
Query: 1543 KHHFIRDYVQKGILDIQFIDIEHQWADIFTKPLSVERF 1580
H +R+ ++ GIL F+ ++Q AD TKPL + F
Sbjct: 1382 DCHKVRECIEAGILKTIFVRTDNQLADTLTKPLYPKPF 1419
>At4g21360 putative transposable element
Length = 1308
Score = 433 bits (1113), Expect = e-121
Identities = 302/1060 (28%), Positives = 517/1060 (48%), Gaps = 91/1060 (8%)
Query: 549 REKQRSWYLDSGCSRHMTGEKALFLTLTMKDGGEVKFGGNQTGKIIGTGTI-----GNSS 603
+E + W +DSGC+ HMT F + + G + T + G+GT+ G S
Sbjct: 303 QEAKDKWVIDSGCTYHMTSRMDWFSEFNENETTMILLGDDHTVESKGSGTVKVNTHGGSI 362
Query: 604 ISINNVWLVDGLKHNLLSISQFCDNGYD-------VTFSKTNCTLVNKDDKSITFKGKRV 656
+ NV V L+ NL+S GY V F K N T + G V
Sbjct: 363 RVLKNVRFVPNLRRNLISTGTLDKLGYKHEGGDGKVRFYKENKTALC---------GNLV 413
Query: 657 ENVYKINFSDLADQKVVCLLSMNDKKWVWHKRLGHANWRLISKISKLQLVKGLPNIDYHS 716
+Y ++ + ++ + N+K +WH RLGH + + +++ L L D
Sbjct: 414 NGLYVLDGHTVVNENCN-VEGSNEKTELWHCRLGHMSLNNMKILAEKGL---LEKKDIKE 469
Query: 717 DALCGACQKGKIVKSSFKSKDIVSTSRPLELLHIDLFGPVNTASLYGSKYGLVIVDDYSR 776
+ C C GK K SF + T L +H DL +G +Y L I+DD SR
Sbjct: 470 LSFCENCVMGKSKKLSFNVGKHI-TDEVLGYIHADL---------WGKQYFLSIIDDKSR 519
Query: 777 WTWVKFIKSKDYACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKHGILHE 836
W+ F+K+KD E F + ++++ K+ +R+D G EF N F+ FC+++GI
Sbjct: 520 KVWLMFLKTKDETFERFCEWKELVENQVNKKVKILRTDNGLEFCNLKFDEFCKQNGIERH 579
Query: 837 FSSPRTPQQNGVVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIRPMLEK 896
+ TPQQNGV +R NRTL E R +++E+ L + FWAEA T+ Y+ NR +
Sbjct: 580 RTCTYTPQQNGVAKRMNRTLMEKVRCLLNESGLEEVFWAEAAATAAYLVNRSPASAVDHN 639
Query: 897 TTYELFKGRRPNISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYRVYNSE 956
EL+ ++P + +FGC Y+ + LK +A +G+FLGY + +K Y+V+ +
Sbjct: 640 VPEELWLDKKPGYKHLRRFGCIAYVHLDQGKLK---PRALKGVFLGYPQGTKGYKVWLLD 696
Query: 957 TQCVEESMHVKFDDREPGSKTSEQSESNAGTTDSEDASESDQPSDSE--KYTEVESSPEA 1014
+ S ++ F++ + E SE + + Q S E + ++
Sbjct: 697 EEKCVISRNIVFNENQVYKDIRESSEQSVKDISDLEGYNEFQVSVKEHGECSKTGGVTIE 756
Query: 1015 EYNSEAEPSPKVQNESASEDFQDNTQQVIQPKFKHKSSHPEELIIGSKDSPRRTRSHFRQ 1074
E + E++ V E + Q + + + + P++L +HF
Sbjct: 757 EIDQESDSENSVTQEPLIASIDLSNYQSARDRERRAPNPPQKL---------ADYTHFAL 807
Query: 1075 EESLIGLLSIIEPKTVEEALSDDGWIL---AMQEELINFKGMMCGIW--YPYPFRRTLLE 1129
+ + EP+ +A D WI M+EE+ + + G W +P + ++
Sbjct: 808 ALVMAEEIESEEPQCYHDAKKDKHWIKWNGGMKEEIDSL--LKNGTWDIVEWPKEQKVIS 865
Query: 1130 QNGYSKTS-*MNKEK*PETKPDFVAQGYSQQEGIDYTETFAPVARLEAIRLLLSYAINHG 1188
K + + K VA+G++QQ+GIDY E FAPV + +IR+L+S +
Sbjct: 866 CRWLFKLKPGIPGVEAQRYKARLVARGFTQQKGIDYEEVFAPVVKHISIRILMSAVVKDD 925
Query: 1189 IILYQMDVKSAFLNGVIEEEVYVKQPPGFEDLKHPDHVYKLKKSLYGLKQAPRAWYDRLS 1248
+ L QMDVK+ FL+G +++ +Y++QP GFE D V LKKSLYGLKQAPR W +
Sbjct: 926 MELEQMDVKTTFLHGELDQVLYMEQPEGFEVNPEKDQVCLLKKSLYGLKQAPRQWNKKFH 985
Query: 1249 NFLIKNDFERGQVDTTLFRRTLKK-DILIVQIYVDDIIFGSTNASLCK*FSKLMQDEFEM 1307
F++ F R + D+ ++ + + + + + +YVDD++ + + S + + +FEM
Sbjct: 986 AFMLSLQFARSEHDSCVYVKEVNPGEFVYLLLYVDDMLLAAKSKSEISKLKEALSLKFEM 1045
Query: 1308 SMMGELKFFLGIQI--NQSKEGVYVHQTKYTKELLKKFKLEDCKVMNTPM--HPTCTLSK 1363
MG LGI I N+ + + + QT+Y +++++F++ D KV++TPM H T
Sbjct: 1046 KDMGAASRILGIDIIRNRKEGTLRLSQTRYVDKVIQRFRMADAKVVSTPMGAHFKLTSLI 1105
Query: 1364 EDTGTVVDQKL-YRGMIGSLLY-LTASRPAILFSVCLCARFQSDPRESHLTAVKRIFRYL 1421
++ G+V + + Y +GS++Y + + P + +++ L +RF S P
Sbjct: 1106 DEIGSVDPEVVPYSSAVGSVMYAMIGTIPDVAYAMGLVSRFMSRP--------------- 1150
Query: 1422 KGTTNLGLLYRKSLDYKLIGFCDADYAGDRIERKSTSGNCQFLGENLISWASKRQATIAM 1481
+ ++ G+CD+D+A D +R+S SG +G N +SW S Q +A+
Sbjct: 1151 ------------GANLEVQGYCDSDHAADLDKRRSISGYVFTVGGNTVSWKSSLQHVVAL 1198
Query: 1482 STAEAEYISAASCCTQLLWMKHQLEDYQINANSIPIYCDNTAAICLSKNPILHSRAKHIE 1541
S+ +AE+I+ + +W++ LED + ++CD+ +AICLSKN H R KH+E
Sbjct: 1199 SSTQAEFIALTEAVKEAIWIRGLLEDMGLQPKPATVWCDSQSAICLSKNNAFHDRTKHVE 1258
Query: 1542 IKHHFIRDYVQKGILDIQFIDIEHQWADIFTKPLSVERFD 1581
+K +FIRD ++ G + ++ I AD+ TK + V++F+
Sbjct: 1259 VKFYFIRDIIEAGEVKVRKIHTSVNPADMLTKCIPVKKFE 1298
>At2g19840 copia-like retroelement pol polyprotein
Length = 1137
Score = 421 bits (1082), Expect = e-117
Identities = 278/852 (32%), Positives = 442/852 (51%), Gaps = 88/852 (10%)
Query: 772 DDYSRWTWVKFIKSKDYACEVFSSFCTQIQSEKELKILKVRSDYGGEFENEPFELFCEKH 831
+D+SR WV F+K+KD A F+ + ++++ E K+ +R+D G EF N F+ C+K
Sbjct: 319 NDWSRKVWVYFLKTKDEAFASFTEWKKMVETQSERKLKHLRTDNGLEFCNHKFDEVCKKE 378
Query: 832 GILHEFSSPRTPQQNGVVERKNRTLQEMARTMIHENNLAKHFWAEAVNTSCYIQNRIYIR 891
GI+ + TPQQNGV ER NRT+ R+M+ E+ L K FWA+A +T+ Y+ NR
Sbjct: 379 GIVRHRTCTYTPQQNGVAERLNRTIMNKVRSMLSESGLDKKFWAKAASTAVYLINRSPSS 438
Query: 892 PMLEKTTYELFKGRRPNISYFHQFGCTCYILNTKDYLKKFDAKAQRGIFLGYSERSKAYR 951
+ K EL+ PN S +FGC Y+ + + K D +A++G+F+GY K +R
Sbjct: 439 SIENKIPEELWTSAVPNFSGLKRFGCVVYVYSQEG---KLDPRAKKGVFVGYPNGVKGFR 495
Query: 952 VYNSETQCVEESMHVKFDDREPGSKTSEQSESNAGTTDSEDASESDQPS----DSEKYTE 1007
V+ E + S +V F RE ++S +G + + + PS + K E
Sbjct: 496 VWMIEEERCSISRNVVF--REDVMYKDILNQSTSGMSFDFPLATNRIPSFECAGNRKEDE 553
Query: 1008 VE-----SSPEAEYNSEAEPSPKVQNESASEDFQDNTQQVIQPKFKHKSSHPEELIIGSK 1062
+ S + + +SE P + S+ ++ T Q+ + K K ++ P++L
Sbjct: 554 ISVQGGVSDDDTKQSSEESP---ISTGSSGQNSGQRTYQIARDKPKRQTKIPDKL----- 605
Query: 1063 DSPRRTRSHFRQEESL---IGLLSII-------EPKTVEEALSDDG---WILAMQEE--- 1106
R + EE L G +I EP ++AL D W+ A+ EE
Sbjct: 606 ------RDYELNEEVLDEIAGYAYMITEDGGNPEPNDYQKALQDSDYKMWLKAVDEEIES 659
Query: 1107 --------LINFKGMMCGIWYPYPFRRTLLEQNGYSKTS*MNKEK*PETKPDFVAQGYSQ 1158
L+N I + F+R K+ + EK P K V +GYSQ
Sbjct: 660 LLKNNTWVLVNRDQFQKPIGCKWVFKR---------KSGIVGVEK-PRFKARLVVKGYSQ 709
Query: 1159 QEGIDYTETFAPVARLEAIRLLLSYAINHGIILYQMDVKSAFLNGVIEEEVYVKQPPGFE 1218
+EGIDY E F+PV + +IRLLLS + + L QMDVK+AFL+G ++E +Y++QP G+
Sbjct: 710 KEGIDYQEIFSPVVKHVSIRLLLSMVTHCDMELQQMDVKTAFLHGYLDETIYIEQPEGYV 769
Query: 1219 DLKHPDHVYKLKKSLYGLKQAPRAWYDRLSNFLIKNDFERGQVDTTLFRRTLKK-DILIV 1277
++PD V LK+SLYGL+Q+PR W +R + F+ K +ER + D+ ++ + L+ + + +
Sbjct: 770 HKRYPDKVCLLKRSLYGLRQSPRQWNNRFNEFMQKIGYERSKYDSCVYFKELQSGEYIYL 829
Query: 1278 QIYVDDIIFGSTNASLCK*FSKLMQDEFEMSMMGELKFFLGIQINQSKEG--VYVHQTKY 1335
+YVDDI+ S + L+ EFEM +G+ K LG++I + ++ + + Q Y
Sbjct: 830 LLYVDDILIASRDKRTVCDLKALLNSEFEMKDLGDAKKILGMEIVRDRKAGTMSISQEGY 889
Query: 1336 TKELLKKFKLEDCKVMNTPMHPTCTLSKEDTGTVVDQK------LYRGMIGSLLY-LTAS 1388
++L F ++ K + TPM L V+ Q Y+ +GSL+Y + +
Sbjct: 890 LLKVLGNFGMDQAKPVFTPMGAHFKLKPATDEEVMRQSEVMRAVPYQSAVGSLMYSMIGT 949
Query: 1389 RPAILFSVCLCARFQSDPRESHLTAVKRIFRYLKGTTNLGLLYRKSLDYKLIGFCDADYA 1448
RP + SV L RF S P + H AVK I RY++G+ + L Y+ + L G+CD+DYA
Sbjct: 950 RPDLAHSVGLVCRFMSKPLKEHWQAVKWILRYIRGSIDRKLCYKNEGELILEGYCDSDYA 1009
Query: 1449 GDRIERKSTSGNCQFLGENLISWASKRQATIAMSTAEAEYISAASCCTQLLWMKHQLEDY 1508
D+ R+STSG +A+S+ EAEY++ + +W+K + +
Sbjct: 1010 ADKEGRRSTSG----------------VKVVALSSTEAEYMALTDGAKEAIWLKGHVSEL 1053
Query: 1509 QINANSIPIYCDNTAAICLSKNPILHSRAKHIEIKHHFIRDYVQKGILDIQFIDIEHQWA 1568
++ I+CD+ +AI L+KN + H R KHI++K+HFIRD V G + + ID E A
Sbjct: 1054 GFVQKTVNIHCDSQSAIALAKNAVYHERTKHIDVKYHFIRDLVNNGEVQVLKIDTEDNPA 1113
Query: 1569 DIFTKPLSVERF 1580
DIFTK L V +F
Sbjct: 1114 DIFTKVLPVSKF 1125
Score = 39.3 bits (90), Expect = 0.018
Identities = 15/51 (29%), Positives = 25/51 (48%)
Query: 549 REKQRSWYLDSGCSRHMTGEKALFLTLTMKDGGEVKFGGNQTGKIIGTGTI 599
++ Q W +D+GCS HMT K + G+V+ N ++ G G +
Sbjct: 260 KDDQEEWIMDTGCSFHMTPRKEYLMDFVEAKSGKVRMANNSFSEVKGIGKV 310
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.332 0.143 0.445
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 33,611,666
Number of Sequences: 26719
Number of extensions: 1427326
Number of successful extensions: 6323
Number of sequences better than 10.0: 236
Number of HSP's better than 10.0 without gapping: 133
Number of HSP's successfully gapped in prelim test: 107
Number of HSP's that attempted gapping in prelim test: 5297
Number of HSP's gapped (non-prelim): 600
length of query: 1594
length of database: 11,318,596
effective HSP length: 113
effective length of query: 1481
effective length of database: 8,299,349
effective search space: 12291335869
effective search space used: 12291335869
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 67 (30.4 bits)
Medicago: description of AC137996.10


