

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137826.3 - phase: 0 /pseudo
(1546 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
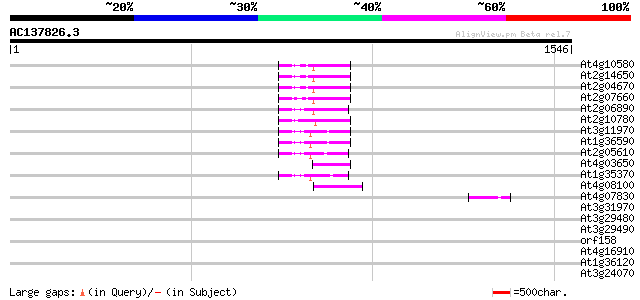
Score E
Sequences producing significant alignments: (bits) Value
At4g10580 putative reverse-transcriptase -like protein 77 6e-14
At2g14650 putative retroelement pol polyprotein 72 2e-12
At2g04670 putative retroelement pol polyprotein 71 4e-12
At2g07660 putative retroelement pol polyprotein 67 6e-11
At2g06890 putative retroelement integrase 67 1e-10
At2g10780 pseudogene 66 1e-10
At3g11970 hypothetical protein 63 2e-09
At1g36590 hypothetical protein 63 2e-09
At2g05610 putative retroelement pol polyprotein 60 8e-09
At4g03650 putative reverse transcriptase 57 6e-08
At1g35370 hypothetical protein 57 8e-08
At4g08100 putative polyprotein 47 1e-04
At4g07830 putative reverse transcriptase 46 2e-04
At3g31970 hypothetical protein 41 0.005
At3g29480 hypothetical protein 41 0.006
At3g29490 hypothetical protein 38 0.052
orf158 -mitochondrial genome- 37 0.089
At4g16910 retrotransposon like protein 37 0.089
At1g36120 putative reverse transcriptase gb|AAD22339.1 37 0.089
At3g24070 putative retroelement 33 1.7
>At4g10580 putative reverse-transcriptase -like protein
Length = 1240
Score = 77.4 bits (189), Expect = 6e-14
Identities = 67/217 (30%), Positives = 102/217 (46%), Gaps = 34/217 (15%)
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTK 799
D S +C DYR+LNRVT +N+Y LP ID+L +QL A+ +SK+ L T
Sbjct: 538 DGSFRLCIDYRELNRVTVKNRYPLPRIDELLDQLRGATC---FSKIDL----------TS 584
Query: 800 KGIQDPLWAL*VSSDIVWVDKCSFGCHEFD*LCDQMT-----------------LGFILI 842
Q P+ V + +G EF + +T L +I
Sbjct: 585 GYHQIPIAEADVRKTAF---RTRYGHFEFVVMPFGLTNAPAVFMRLMNSVFQEFLDEFVI 641
Query: 843 VCVDDILLFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP-IGNVVMNERIMV 901
+ +DDIL++ +S ++ HL V + QKL+A SF ++ +G++V E + V
Sbjct: 642 IFIDDILVYSKSPEEQEVHLRRVMEKLREQKLFAKLSKCSFWQREMGFLGHIVSAEGVSV 701
Query: 902 DLQNIEPVMNWV*PSFVIESRGFMGLASYYRGFEDAF 938
D + IE + +W P+ E R F+G A YYR F F
Sbjct: 702 DPEKIEAIRDWPRPTNATEIRSFLGWAGYYRRFVKGF 738
>At2g14650 putative retroelement pol polyprotein
Length = 1328
Score = 72.0 bits (175), Expect = 2e-12
Identities = 67/217 (30%), Positives = 99/217 (44%), Gaps = 34/217 (15%)
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTK 799
D S +C DYR LN VT +NKY LP ID+L +QL A+ +SK+ L T
Sbjct: 513 DGSFRLCIDYRGLNWVTVKNKYPLPRIDELLDQLRGATC---FSKIDL----------TS 559
Query: 800 KGIQDPLWAL*VSSDIVWVDKCSFGCHEFD*LCDQMT-----------------LGFILI 842
P+ V + +G EF + +T L +I
Sbjct: 560 GYHLIPIAEADVRKTAF---RTRYGHFEFVVMPFGLTNAPAAFMRLMNSVFQEVLDEFVI 616
Query: 843 VCVDDILLFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP-IGNVVMNERIMV 901
+ +DDIL++ +S + HL V + QKL+A SF ++ +G++V E + V
Sbjct: 617 IFIDDILVYSKSLEEHEVHLRRVMEKLREQKLFAKLSKCSFWQREMGFLGHIVSAEGVSV 676
Query: 902 DLQNIEPVMNWV*PSFVIESRGFMGLASYYRGFEDAF 938
D + IE + +W P+ E R F+GLA YYR F F
Sbjct: 677 DPEKIEAIRDWHTPTNATEIRSFLGLAGYYRRFVKGF 713
Score = 42.4 bits (98), Expect = 0.002
Identities = 34/121 (28%), Positives = 53/121 (43%), Gaps = 10/121 (8%)
Query: 1264 RLTKTSHFILVKKHYNAQQLSKVYMRR**GCTECPILSS*TVVPVYIYVLW-----ENA* 1318
RLTK++HF+ ++K A L+K Y+ P+ + W E
Sbjct: 1003 RLTKSAHFLAIRKTDGAAVLAKKYVSEIVKLHGVPVSIVSDRDSKFTSAFWRAFQAEMGT 1062
Query: 1319 EVGHVTHF*NNLPSSNGWTIRENNPSVRGHIEGMCDIDFRGHWDKFLTLCEFSYNNNYTL 1378
+V T + + TI+ +R MC +D+ GHW L+L EF+YNN+Y
Sbjct: 1063 KVQMSTAYHPQTYGQSERTIQTLEDMLR-----MCVLDWGGHWADHLSLVEFAYNNSYPA 1117
Query: 1379 S 1379
S
Sbjct: 1118 S 1118
>At2g04670 putative retroelement pol polyprotein
Length = 1411
Score = 71.2 bits (173), Expect = 4e-12
Identities = 66/217 (30%), Positives = 98/217 (44%), Gaps = 34/217 (15%)
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTK 799
D S +C DYR LN VT +NKY LP ID+L +QL A+ +SK+ L T
Sbjct: 564 DGSFRLCIDYRGLNWVTVKNKYPLPRIDELLDQLRGATC---FSKIDL----------TS 610
Query: 800 KGIQDPLWAL*VSSDIVWVDKCSFGCHEFD*LCDQMT-----------------LGFILI 842
Q P+ V + +G EF + +T L +I
Sbjct: 611 GYHQIPIAEADVRKTAF---RTRYGHFEFVVMPFALTNAPAAFMRLMNSVFQEFLDEFVI 667
Query: 843 VCVDDILLFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP-IGNVVMNERIMV 901
+ +DDIL++ +S + HL V + QKL+A SF ++ +G++V E + V
Sbjct: 668 IFIDDILVYSKSPEEHEVHLRRVMEKLREQKLFAKLSKCSFWQREIGFLGHIVSAEGVSV 727
Query: 902 DLQNIEPVMNWV*PSFVIESRGFMGLASYYRGFEDAF 938
D + IE + +W P+ E R F+ L YYR F F
Sbjct: 728 DPEKIEAIRDWPRPTNATEIRSFLRLTGYYRRFVKGF 764
Score = 43.5 bits (101), Expect = 0.001
Identities = 33/121 (27%), Positives = 54/121 (44%), Gaps = 10/121 (8%)
Query: 1264 RLTKTSHFILVKKHYNAQQLSKVYMRR**GCTECPILSS*TVVPVYIYVLW-----ENA* 1318
RLTK++HF+ ++K A L+K Y+ P+ + + W +
Sbjct: 1083 RLTKSAHFLAIRKTDGAAVLAKKYVSEIVKLHGVPVSIVSDRDSKFTFAFWRAFQAKMGT 1142
Query: 1319 EVGHVTHF*NNLPSSNGWTIRENNPSVRGHIEGMCDIDFRGHWDKFLTLCEFSYNNNYTL 1378
+V T + + TI+ +R MC +D+ GHW L+L EF+YNN+Y
Sbjct: 1143 KVQMSTAYHPQTDGQSERTIQTLEDMLR-----MCVLDWGGHWADHLSLVEFAYNNSYQA 1197
Query: 1379 S 1379
S
Sbjct: 1198 S 1198
>At2g07660 putative retroelement pol polyprotein
Length = 949
Score = 67.4 bits (163), Expect = 6e-11
Identities = 63/217 (29%), Positives = 97/217 (44%), Gaps = 34/217 (15%)
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTK 799
D S +C DYR LN+VT +NKY LP ID+L +QL A + +SK+ L +
Sbjct: 201 DGSFRLCIDYRGLNKVTVKNKYPLPRIDELMDQLGGAQW---FSKIDLASGYH------- 250
Query: 800 KGIQDPLWAL*VSSDIVWVDKCSFGCHEFD*LCDQMT-----------------LGFILI 842
Q P+ V + +G EF + +T L +I
Sbjct: 251 ---QIPIEPTDVRKTAF---RTRYGHFEFVVMPFGLTNAPAAFMKMMNGVFRDFLDEFVI 304
Query: 843 VCVDDILLFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP-IGNVVMNERIMV 901
+ +DDIL+ +S +HL V + +L+A SF + +G+V+ ++ + V
Sbjct: 305 IFIDDILVHSKSWEAHQEHLRAVLERLREHELFAKLSKFSFWQRSVGFLGHVISDQGVSV 364
Query: 902 DLQNIEPVMNWV*PSFVIESRGFMGLASYYRGFEDAF 938
D + I + W P E R F+GLA YYR F +F
Sbjct: 365 DPEKIRSIKEWPRPRNATEIRSFLGLAGYYRRFVMSF 401
Score = 37.0 bits (84), Expect = 0.089
Identities = 13/27 (48%), Positives = 21/27 (77%)
Query: 1353 CDIDFRGHWDKFLTLCEFSYNNNYTLS 1379
C +D+ G+W+K+L L EF+YNN++ S
Sbjct: 741 CVLDWGGNWEKYLRLIEFAYNNSFQAS 767
>At2g06890 putative retroelement integrase
Length = 1215
Score = 66.6 bits (161), Expect = 1e-10
Identities = 60/215 (27%), Positives = 99/215 (45%), Gaps = 38/215 (17%)
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTK 799
D S MC D R +N VT + + +P +DD+ ++L +S +SK+ L K
Sbjct: 466 DGSWRMCFDCRAINNVTVKYCHPIPRLDDMLDELHGSSI---FSKIDL-----------K 511
Query: 800 KGIQDPLWAL*VSSDIVWVD--KCSFGCHEFD*LCDQMT-----------------LGFI 840
G + ++ W K G +E+ + +T +G
Sbjct: 512 SGYHQ----IRMNEGDEWKTAFKTKHGLYEWLVMPFGLTHAPSTFMRLMNHVLRAFIGIF 567
Query: 841 LIVCVDDILLFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP-IGNVVMNERI 899
+IV DDIL++ ES + ++HL V +V +++LYA +FC L +G VV + +
Sbjct: 568 VIVYFDDILVYSESLREHIEHLDSVLNVLRKEELYANLKKCTFCTDNLVFLGFVVSADGV 627
Query: 900 MVDLQNIEPVMNWV*PSFVIESRGFMGLASYYRGF 934
VD + ++ + +W P V E R F GLA +YR F
Sbjct: 628 KVDEEKVKAIRDWPSPKTVGEVRSFHGLAGFYRRF 662
>At2g10780 pseudogene
Length = 1611
Score = 66.2 bits (160), Expect = 1e-10
Identities = 63/210 (30%), Positives = 96/210 (45%), Gaps = 20/210 (9%)
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTK 799
D S +C DYR LN+VT +NKY LP ID+L +QL A + +SK+ L A G +
Sbjct: 706 DGSFRLCIDYRGLNKVTVKNKYPLPRIDELMDQLGGAQW---FSKIDL------ASGYHQ 756
Query: 800 KGIQDP---LWAL*VSSDIVWVDKCSFGCHEFD*LCDQMTLGFI-------LIVCVDDIL 849
I+ A D FG +M G +I+ ++DIL
Sbjct: 757 IPIEPTDVRKTAFRTRYDHFEFVVMPFGLTNAPAAFMKMMNGVFRDFLDEFVIIFINDIL 816
Query: 850 LFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP-IGNVVMNERIMVDLQNIEP 908
++ +S +HL V + +L+A SF + +G+V+ ++ + VD + I
Sbjct: 817 VYSKSWEAHQEHLRAVLERLREHELFAKLSKCSFWQRSVGFLGHVISDQGVSVDPEKIRS 876
Query: 909 VMNWV*PSFVIESRGFMGLASYYRGFEDAF 938
+ W P E R F+GLA YYR F +F
Sbjct: 877 IKEWPRPRNATEIRSFLGLAGYYRRFVMSF 906
Score = 37.0 bits (84), Expect = 0.089
Identities = 13/27 (48%), Positives = 21/27 (77%)
Query: 1353 CDIDFRGHWDKFLTLCEFSYNNNYTLS 1379
C +D+ G+W+K+L L EF+YNN++ S
Sbjct: 1349 CVLDWGGNWEKYLRLVEFAYNNSFQAS 1375
>At3g11970 hypothetical protein
Length = 1499
Score = 62.8 bits (151), Expect = 2e-09
Identities = 61/224 (27%), Positives = 99/224 (43%), Gaps = 48/224 (21%)
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTK 799
D + +C DYR+LN +T ++ + +PLI+DL ++L A +SK+ L +
Sbjct: 648 DGTWRLCVDYRELNGMTVKDSFPIPLIEDLMDELGGAVI---FSKIDL-----------R 693
Query: 800 KGIQDPLWAL*VSSDIVWVDKCSFGCHE------------------FD*LCDQMTLGFI- 840
G V D + K +F H F L + + F+
Sbjct: 694 AGYHQ------VRMDPDDIQKTAFKTHSGHFEYLVMPFGLTNAPATFQGLMNFIFKPFLR 747
Query: 841 --LIVCVDDILLFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP----IGNVV 894
++V DDIL++ S + HL V +V KL+A +S C +P +G+ +
Sbjct: 748 KFVLVFFDDILVYSSSLEEHRQHLKQVFEVMRANKLFA---KLSKCAFAVPKVEYLGHFI 804
Query: 895 MNERIMVDLQNIEPVMNWV*PSFVIESRGFMGLASYYRGFEDAF 938
+ I D I+ V W P+ + + RGF+GLA YYR F +F
Sbjct: 805 SAQGIETDPAKIKAVKEWPQPTTLKQLRGFLGLAGYYRRFVRSF 848
Score = 33.5 bits (75), Expect = 0.98
Identities = 35/120 (29%), Positives = 52/120 (43%), Gaps = 14/120 (11%)
Query: 1264 RLTKTSHFILVKKHYNAQQLSKVYMRR**GCTECP--ILSS*TVVPVY-----IYVLWEN 1316
RL+K +HFI + Y+A ++ Y+ CP I+S VV + L
Sbjct: 1179 RLSKAAHFIALSHPYSALTVAHAYLDNVFKLHGCPTSIVSDRDVVFTSEFWREFFTLQGV 1238
Query: 1317 A*EVGHVTHF*NNLPSSNGWTIRENNPSVRGHIEGMCDIDFRGHWDKFLTLCEFSYNNNY 1376
A ++ H P S+G T N + ++ MC D W K+L L E+ YN NY
Sbjct: 1239 ALKLTSAYH-----PQSDGQT-EVVNRCLETYLRCMCH-DRPQLWSKWLALAEYWYNTNY 1291
>At1g36590 hypothetical protein
Length = 1499
Score = 62.8 bits (151), Expect = 2e-09
Identities = 61/224 (27%), Positives = 99/224 (43%), Gaps = 48/224 (21%)
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTK 799
D + +C DYR+LN +T ++ + +PLI+DL ++L A +SK+ L +
Sbjct: 648 DGTWRLCVDYRELNGMTVKDSFPIPLIEDLMDELGGAVI---FSKIDL-----------R 693
Query: 800 KGIQDPLWAL*VSSDIVWVDKCSFGCHE------------------FD*LCDQMTLGFI- 840
G V D + K +F H F L + + F+
Sbjct: 694 AGYHQ------VRMDPDDIQKTAFKTHSGHFEYLVMPFGLTNAPATFQGLMNFIFKPFLR 747
Query: 841 --LIVCVDDILLFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP----IGNVV 894
++V DDIL++ S + HL V +V KL+A +S C +P +G+ +
Sbjct: 748 KFVLVFFDDILVYSSSLEEHRQHLKQVFEVMRANKLFA---KLSKCAFAVPKVEYLGHFI 804
Query: 895 MNERIMVDLQNIEPVMNWV*PSFVIESRGFMGLASYYRGFEDAF 938
+ I D I+ V W P+ + + RGF+GLA YYR F +F
Sbjct: 805 SAQGIETDPAKIKAVKEWPQPTTLKQLRGFLGLAGYYRRFVRSF 848
Score = 34.3 bits (77), Expect = 0.58
Identities = 35/120 (29%), Positives = 53/120 (44%), Gaps = 14/120 (11%)
Query: 1264 RLTKTSHFILVKKHYNAQQLSKVYMRR**GCTECP--ILSS*TVVPVY-----IYVLWEN 1316
RL+K +HFI + Y+A +++ Y+ CP I+S VV + L
Sbjct: 1179 RLSKAAHFIALSHPYSALTVAQAYLDNVFKLHGCPTSIVSDRDVVFTSEFWREFFTLQGV 1238
Query: 1317 A*EVGHVTHF*NNLPSSNGWTIRENNPSVRGHIEGMCDIDFRGHWDKFLTLCEFSYNNNY 1376
A ++ H P S+G T N + ++ MC D W K+L L E+ YN NY
Sbjct: 1239 ALKLTSAYH-----PQSDGQT-EVVNRCLETYLRCMCH-DRPQLWSKWLALAEYWYNTNY 1291
>At2g05610 putative retroelement pol polyprotein
Length = 780
Score = 60.5 bits (145), Expect = 8e-09
Identities = 57/220 (25%), Positives = 97/220 (43%), Gaps = 48/220 (21%)
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTK 799
D + +C DYR+LN +T ++++ +PLI+DL ++L ++ YSK+ L +
Sbjct: 90 DGTWRLCVDYRELNGMTVKDRFPIPLIEDLMDELGGSNV---YSKIDL-----------R 135
Query: 800 KGIQDPLWAL*VSSDIVWVDKCSFGCHE------------------FD*LCDQMTLGFI- 840
G V D + + K +F H F L + F+
Sbjct: 136 AGYHQ------VRMDPLDIHKTAFKTHNGHYEYLVMPFGLTNAPASFQSLMNSFFKPFLR 189
Query: 841 --LIVCVDDILLFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP----IGNVV 894
++V DDIL++ S + HL V +V +++ F +S C +P +G+ +
Sbjct: 190 KFVLVFFDDILIYSTSMEEHKKHLEAVFEV---MRVHHLFAKMSKCAFAVPRVEYLGHFI 246
Query: 895 MNERIMVDLQNIEPVMNWV*PSFVIESRGFMGLASYYRGF 934
E I D I+ V +W P + + GF+GL YYR F
Sbjct: 247 SGEGIATDPAKIKAVQDWPVPVNLKQLCGFLGLTGYYRRF 286
>At4g03650 putative reverse transcriptase
Length = 839
Score = 57.4 bits (137), Expect = 6e-08
Identities = 37/106 (34%), Positives = 57/106 (52%), Gaps = 1/106 (0%)
Query: 834 QMTLGFILIVCVDDILLFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP-IGN 892
Q L +I+ +DDIL++ +S + HL V + QKL+A SF ++ +G+
Sbjct: 558 QEFLDEFVIIFIDDILVYSKSPEEHEVHLRRVMEKLREQKLFAKLSKCSFWQREMGFLGH 617
Query: 893 VVMNERIMVDLQNIEPVMNWV*PSFVIESRGFMGLASYYRGFEDAF 938
+V E + VD + IE + +W P+ E R F+GLA YYR F F
Sbjct: 618 IVSAEGVSVDPEKIEAIRDWPRPTNATEIRSFLGLAGYYRRFIKGF 663
>At1g35370 hypothetical protein
Length = 1447
Score = 57.0 bits (136), Expect = 8e-08
Identities = 55/216 (25%), Positives = 92/216 (42%), Gaps = 48/216 (22%)
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTK 799
D + +C DY +LN +T ++++ +PLI+DL ++L + +SK+ L +
Sbjct: 627 DGTWRLCVDYTELNGMTVKDRFLIPLIEDLMDELGGSVV---FSKIDL-----------R 672
Query: 800 KGIQDPLWAL*VSSDIVWVDKCSFGCHE------------------FD*LCDQMTLGFI- 840
G V D + K +F H F L + + F+
Sbjct: 673 AGYHQ------VRMDPDDIQKTAFKTHNGHFEYLVMLFGLTNAPATFQSLMNSVFRDFLR 726
Query: 841 --LIVCVDDILLFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLPIGNVVMNER 898
++V DDIL++ S + +HL +V +V KL+A +G+ +
Sbjct: 727 KFVLVFFDDILIYSSSIEEHKEHLRLVFEVMRLHKLFAKGSKEH-------LGHFISARE 779
Query: 899 IMVDLQNIEPVMNWV*PSFVIESRGFMGLASYYRGF 934
I D I+ V W P+ V + RGF+G A YYR F
Sbjct: 780 IETDPAKIQAVKEWPTPTTVKQVRGFLGFAGYYRRF 815
Score = 32.0 bits (71), Expect = 2.9
Identities = 32/118 (27%), Positives = 57/118 (48%), Gaps = 10/118 (8%)
Query: 1264 RLTKTSHFILVKKHYNAQQLSKVYMRR**GCTECP--ILSS*TVVPVYIYVLWENA*EVG 1321
RL+K +HF+ + Y+A +++ ++ CP I+S V+ + W+ ++
Sbjct: 1150 RLSKAAHFVALAHPYSALTVAQAFLDNVYKHHGCPTSIVSDRDVL--FTSDFWKEFFKLQ 1207
Query: 1322 HVTHF*NNL--PSSNGWTIRENNPSVRGHIEGMCDIDFRGH-WDKFLTLCEFSYNNNY 1376
V ++ P S+G T N + ++ MC R H W+K+L L E+ YN NY
Sbjct: 1208 GVELRMSSAYHPQSDGQT-EVVNRCLENYLRCMCHA--RPHLWNKWLPLAEYWYNTNY 1262
>At4g08100 putative polyprotein
Length = 1054
Score = 46.6 bits (109), Expect = 1e-04
Identities = 42/140 (30%), Positives = 69/140 (49%), Gaps = 5/140 (3%)
Query: 837 LGFILIVCVDDILLFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP-IGNVVM 895
+G +IV DDIL++ ++ V HL +V D+ ++KLYA +FC L +G VV
Sbjct: 617 IGVFVIVYFDDILVYSKNLEYHVMHLKLVLDLLRKEKLYANLKKCTFCTDNLVFLGFVVS 676
Query: 896 NERIMVDLQNIEPVMNWV*PSFVIESR-GFM---GLASYYRGFEDAFDHNTYLGTTSVT* 951
+ I VD + ++ + W P V E GF + ++ ++ +++ +
Sbjct: 677 ADGIKVDEEKVKAIREWPNPKNVSEKDIGFKWEDAQENAFQALKEKLTNSSVPILPNFMK 736
Query: 952 SLIINYYASQLILGVVLMQD 971
S I AS L +G VLMQD
Sbjct: 737 SFEIECDASGLGIGAVLMQD 756
>At4g07830 putative reverse transcriptase
Length = 611
Score = 45.8 bits (107), Expect = 2e-04
Identities = 35/121 (28%), Positives = 54/121 (43%), Gaps = 10/121 (8%)
Query: 1264 RLTKTSHFILVKKHYNAQQLSKVYMRR**GCTECPILSS*TVVPVYIYVLW-----ENA* 1318
RLTK++HF+ ++K A L+K Y+ P+ + W E
Sbjct: 268 RLTKSAHFLAIRKTDGAAVLAKKYVSEIVKLHGVPVSIVSDRDSKFTSAFWRAFQAEMGT 327
Query: 1319 EVGHVTHF*NNLPSSNGWTIRENNPSVRGHIEGMCDIDFRGHWDKFLTLCEFSYNNNYTL 1378
+V T + + TI+ +R MC +D+RGHW L+L EF+YNN+Y
Sbjct: 328 KVQMSTAYHPQTDGQSERTIQTLEDMLR-----MCVLDWRGHWADHLSLVEFAYNNSYQA 382
Query: 1379 S 1379
S
Sbjct: 383 S 383
>At3g31970 hypothetical protein
Length = 1329
Score = 41.2 bits (95), Expect = 0.005
Identities = 34/121 (28%), Positives = 52/121 (42%), Gaps = 10/121 (8%)
Query: 1264 RLTKTSHFILVKKHYNAQQLSKVYMRR**GCTECPILSS*TVVPVYIYVLW-----ENA* 1318
RLTK++HF+ ++K A L+K Y+ P+ + W E
Sbjct: 1001 RLTKSAHFLAIRKTDGAVLLAKKYVSEIVELHGVPVSIVSDRDSKFTSAFWRAFQGEMGT 1060
Query: 1319 EVGHVTHF*NNLPSSNGWTIRENNPSVRGHIEGMCDIDFRGHWDKFLTLCEFSYNNNYTL 1378
+V T + + TI+ +R MC +D GHW L+L EF+YNN+Y
Sbjct: 1061 KVQMSTAYHPQTDGQSERTIQTLEDMLR-----MCVLDRGGHWADHLSLVEFAYNNSYQA 1115
Query: 1379 S 1379
S
Sbjct: 1116 S 1116
Score = 39.7 bits (91), Expect = 0.014
Identities = 17/40 (42%), Positives = 24/40 (59%)
Query: 899 IMVDLQNIEPVMNWV*PSFVIESRGFMGLASYYRGFEDAF 938
+ VDL+ IE + +W P+ E R F+GLA YY+ F F
Sbjct: 659 VSVDLEKIEAIRDWPRPTNATEIRSFLGLAGYYKRFVKVF 698
>At3g29480 hypothetical protein
Length = 718
Score = 40.8 bits (94), Expect = 0.006
Identities = 18/40 (45%), Positives = 24/40 (60%)
Query: 899 IMVDLQNIEPVMNWV*PSFVIESRGFMGLASYYRGFEDAF 938
+ VDL+ IE + +W P+ E R F+GLA YYR F F
Sbjct: 416 VSVDLEKIEAIKDWPRPTNATEIRSFLGLAGYYRRFVKGF 455
>At3g29490 hypothetical protein
Length = 438
Score = 37.7 bits (86), Expect = 0.052
Identities = 14/25 (56%), Positives = 19/25 (76%)
Query: 1352 MCDIDFRGHWDKFLTLCEFSYNNNY 1376
MC +D+ GHW L+L EF+YNN+Y
Sbjct: 101 MCVLDWGGHWADHLSLVEFAYNNSY 125
>orf158 -mitochondrial genome-
Length = 158
Score = 37.0 bits (84), Expect = 0.089
Identities = 15/43 (34%), Positives = 23/43 (52%)
Query: 892 NVVMNERIMVDLQNIEPVMNWV*PSFVIESRGFMGLASYYRGF 934
+++ E + D +E ++ W P E RGF+GL YYR F
Sbjct: 37 HIISGEGVSADPAKLEAMVGWPEPKNTTELRGFLGLTGYYRRF 79
>At4g16910 retrotransposon like protein
Length = 687
Score = 37.0 bits (84), Expect = 0.089
Identities = 13/27 (48%), Positives = 21/27 (77%)
Query: 1353 CDIDFRGHWDKFLTLCEFSYNNNYTLS 1379
C +D+ G+W+K+L L EF+YNN++ S
Sbjct: 433 CVLDWGGNWEKYLRLVEFAYNNSFQAS 459
>At1g36120 putative reverse transcriptase gb|AAD22339.1
Length = 1235
Score = 37.0 bits (84), Expect = 0.089
Identities = 14/28 (50%), Positives = 20/28 (71%)
Query: 1352 MCDIDFRGHWDKFLTLCEFSYNNNYTLS 1379
MC +D+ GHW L+L +F+YNN+Y S
Sbjct: 1024 MCVLDWGGHWADHLSLVKFAYNNSYQAS 1051
Score = 34.3 bits (77), Expect = 0.58
Identities = 15/22 (68%), Positives = 18/22 (81%)
Query: 752 LNRVTTRNKYTLPLIDDLFNQL 773
LNRVT +NKY LP ID+L +QL
Sbjct: 576 LNRVTLKNKYPLPRIDELLDQL 597
>At3g24070 putative retroelement
Length = 262
Score = 32.7 bits (73), Expect = 1.7
Identities = 15/48 (31%), Positives = 26/48 (53%), Gaps = 1/48 (2%)
Query: 401 YRTFYNCGDFRHISRYCPKPRILDLMQK*TRAVVPASNDKNSRGHPRG 448
++ +NCGD H++R C P + D+ R++ P ++ NS G G
Sbjct: 85 FKFCFNCGDMNHLARNCLIPWV-DVPDPYERSLSPPPHESNSDGSAEG 131
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.359 0.161 0.583
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,907,365
Number of Sequences: 26719
Number of extensions: 1142332
Number of successful extensions: 4398
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 21
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 4314
Number of HSP's gapped (non-prelim): 71
length of query: 1546
length of database: 11,318,596
effective HSP length: 112
effective length of query: 1434
effective length of database: 8,326,068
effective search space: 11939581512
effective search space used: 11939581512
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 67 (30.4 bits)
Medicago: description of AC137826.3


