

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137825.4 - phase: 0
(161 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
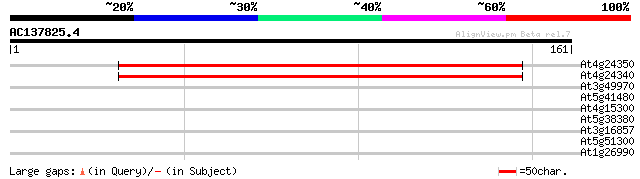
Score E
Sequences producing significant alignments: (bits) Value
At4g24350 putative protein 117 2e-27
At4g24340 unknown protein 109 7e-25
At3g49970 putative protein 29 0.97
At5g41480 folylpolyglutamate synthase-like protein 28 2.2
At4g15300 cytochrome P450 like protein 28 2.2
At5g38380 unknown protein 27 3.7
At3g16857 ARR1 protein, putative, 5' partial 27 3.7
At5g51300 unknown protein 26 8.2
At1g26990 polyprotein, putative 26 8.2
>At4g24350 putative protein
Length = 232
Score = 117 bits (294), Expect = 2e-27
Identities = 54/116 (46%), Positives = 74/116 (63%)
Query: 32 ISNINNEGPSIGIVVPNAYELNPLLHSSSFVPHNKFPYFDFAGRHFRIGELEKKKVIVVM 91
I +N GP IG+V A E N L S F P P+ D +GR FRIG++ KKV+ V
Sbjct: 38 IRKLNRGGPYIGLVTVIATEENAFLRSVDFRPDPTHPFLDLSGRRFRIGKIHGKKVVYVR 97
Query: 92 SGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQIGDVTIPKYWAHTGLWHW 147
G GM+N ATQ ++ +FN++G++H+GIAGN+N+ IGDV+IPK + GLW W
Sbjct: 98 CGRGMVNGAAATQQMIDVFNVKGIVHFGIAGNMNNSMSIGDVSIPKQITNAGLWDW 153
>At4g24340 unknown protein
Length = 338
Score = 109 bits (272), Expect = 7e-25
Identities = 52/116 (44%), Positives = 73/116 (62%)
Query: 32 ISNINNEGPSIGIVVPNAYELNPLLHSSSFVPHNKFPYFDFAGRHFRIGELEKKKVIVVM 91
I IN GP IG+V E + L S F P+ D +GR FRIG++ KKV+ V
Sbjct: 43 IRKINRRGPYIGLVTVFETEEDAFLGSVDFRLDPTHPFLDLSGRRFRIGKIHGKKVVYVR 102
Query: 92 SGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQIGDVTIPKYWAHTGLWHW 147
G GM+NA ATQ ++ +F+++G++H+GIAGN+N+ IGDV+IPK + GLW W
Sbjct: 103 CGIGMVNAAAATQQMIDVFSVKGIVHFGIAGNINNSMSIGDVSIPKQITNAGLWDW 158
>At3g49970 putative protein
Length = 526
Score = 29.3 bits (64), Expect = 0.97
Identities = 21/81 (25%), Positives = 35/81 (42%), Gaps = 1/81 (1%)
Query: 14 LGNSIYAYGAINQLSWREISNINNEGPSIGIVVPNAYELNPLLHS-SSFVPHNKFPYFDF 72
L +YA+ A N +SN ++ P+ + EL+ + S N++ +
Sbjct: 391 LSRDVYAHAAQNDRFQENLSNSDSPAPATAEKTLSPPELSSYKNELSKLNRENQYLKLEL 450
Query: 73 AGRHFRIGELEKKKVIVVMSG 93
+ ELEK+K VMSG
Sbjct: 451 LKVKMKFKELEKEKAFEVMSG 471
>At5g41480 folylpolyglutamate synthase-like protein
Length = 530
Score = 28.1 bits (61), Expect = 2.2
Identities = 21/82 (25%), Positives = 36/82 (43%), Gaps = 1/82 (1%)
Query: 15 GNSIYAYGAINQLSWREISNINNEGPSIGIVVPNAYELNPLLHSSSFVPHNKFPYFD-FA 73
G S+ Y + + LS +E + N E S + Y + P+L S + +F+
Sbjct: 120 GYSVGCYSSPHILSIKERISCNGEPVSASTLNDLFYSVKPILEQSIQEENGSLSHFEILT 179
Query: 74 GRHFRIGELEKKKVIVVMSGEG 95
G F + E E + V+ +G G
Sbjct: 180 GIAFSLFEKENVDIAVIEAGLG 201
>At4g15300 cytochrome P450 like protein
Length = 487
Score = 28.1 bits (61), Expect = 2.2
Identities = 14/37 (37%), Positives = 18/37 (47%), Gaps = 2/37 (5%)
Query: 124 LNSRFQIGDVTIPKYWAHTGLWHWQVHTSCFNYLDTF 160
L FQ+GD TIP W G H +H + Y D +
Sbjct: 369 LEHDFQVGDYTIPAGWTFMGYPH--IHFNSEKYEDPY 403
>At5g38380 unknown protein
Length = 361
Score = 27.3 bits (59), Expect = 3.7
Identities = 17/57 (29%), Positives = 35/57 (60%), Gaps = 3/57 (5%)
Query: 1 MEFLL-GVLVMIVLLGNSIYAYGAINQLSWREISNINNEGPSIGIVVPNAYELNPLL 56
M+F L G +V ++LG I+ ++ + ++ S++ +E PS GI+ N ++ +P+L
Sbjct: 305 MDFSLPGFIVCCLVLGFGIFEATSLERN--KKDSSLKSEDPSNGILGNNDFDTSPIL 359
>At3g16857 ARR1 protein, putative, 5' partial
Length = 569
Score = 27.3 bits (59), Expect = 3.7
Identities = 15/43 (34%), Positives = 22/43 (50%)
Query: 25 NQLSWREISNINNEGPSIGIVVPNAYELNPLLHSSSFVPHNKF 67
N++S R + N P V+P +Y HSSS +P+N F
Sbjct: 371 NRISERSGFSGRNNIPESSRVLPTSYTNLTTQHSSSSMPYNNF 413
>At5g51300 unknown protein
Length = 804
Score = 26.2 bits (56), Expect = 8.2
Identities = 17/59 (28%), Positives = 24/59 (39%), Gaps = 8/59 (13%)
Query: 24 INQLSWREISNINNEGPSIGIVV--------PNAYELNPLLHSSSFVPHNKFPYFDFAG 74
IN +R +N E I + P A P LH F+P +FP ++F G
Sbjct: 201 INTREYRARERLNRERQEIIAQIIKKNPAFKPPADYRPPKLHKKLFIPMKEFPGYNFIG 259
>At1g26990 polyprotein, putative
Length = 1436
Score = 26.2 bits (56), Expect = 8.2
Identities = 13/26 (50%), Positives = 15/26 (57%)
Query: 40 PSIGIVVPNAYELNPLLHSSSFVPHN 65
P I + PN E PL+ SSS PHN
Sbjct: 856 PHIYLPAPNNDEHLPLVQSSSDAPHN 881
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.142 0.448
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,799,314
Number of Sequences: 26719
Number of extensions: 158268
Number of successful extensions: 414
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 407
Number of HSP's gapped (non-prelim): 11
length of query: 161
length of database: 11,318,596
effective HSP length: 91
effective length of query: 70
effective length of database: 8,887,167
effective search space: 622101690
effective search space used: 622101690
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 56 (26.2 bits)
Medicago: description of AC137825.4


