

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137823.16 + phase: 0
(621 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
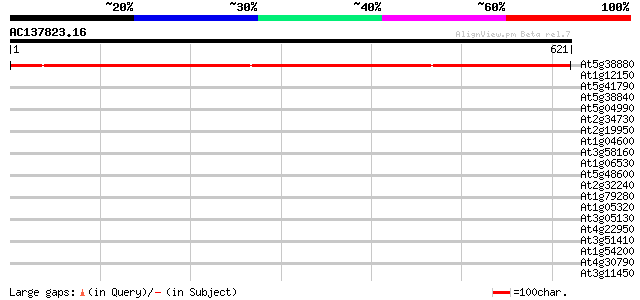
Score E
Sequences producing significant alignments: (bits) Value
At5g38880 putative protein 874 0.0
At1g12150 hypothetical protein 39 0.011
At5g41790 myosin heavy chain-like protein 36 0.056
At5g38840 kanadaptin - like protein 35 0.095
At5g04990 unknown protein 34 0.21
At2g34730 putative myosin heavy chain 34 0.21
At2g19950 hypothetical protein 34 0.28
At1g04600 putative myosin heavy chain 34 0.28
At3g58160 myosin heavy chain MYA3 33 0.36
At1g06530 hypothetical protein 33 0.36
At5g48600 chromosome condensation protein 33 0.47
At2g32240 putative myosin heavy chain 33 0.47
At1g79280 hypothetical protein 33 0.47
At1g05320 unknown protein 33 0.47
At3g05130 hypothetical protein 33 0.62
At4g22950 putative MADS Box / AGL protein 32 0.81
At3g51410 putative protein 32 1.1
At1g54200 unknown protein 32 1.1
At4g30790 putative protein 32 1.4
At3g11450 putative cell division related protein 32 1.4
>At5g38880 putative protein
Length = 775
Score = 874 bits (2258), Expect = 0.0
Identities = 441/620 (71%), Positives = 524/620 (84%), Gaps = 3/620 (0%)
Query: 1 MLEAYDRQCDEASRIFAEYHKRLCYYINQARDAQRSGVDSSVEMVNSFSAKNEKEAVYST 60
+LEAYD+QCDEA+RIFAEYHKRL Y+NQA DAQRS V+SS E+++S SA +E+EAVYST
Sbjct: 157 LLEAYDQQCDEATRIFAEYHKRLQVYVNQANDAQRS-VNSSNEVLSSLSANSEREAVYST 215
Query: 61 VKGSKSSDDVIVIETTREKNIRKACESLVAYMVDKIQSSFPAYEGSGVLSNPQAEAAKLG 120
VKG+KS+DDVI++ETTRE+NIR C+ L + M+++I++SFPAYEG+G+ S P+ E AKLG
Sbjct: 216 VKGTKSADDVILMETTRERNIRIVCDLLASRMIERIRNSFPAYEGNGICSLPELETAKLG 275
Query: 121 FDFDGQIPDEVRTVIVNCLKSPPLLLQAITAYTSHLKSQISREIEKIDVRADAEILRYKY 180
F++DG+I DE++TVIVN L+ PPLLLQAI AYT +K+ ISRE+EKIDVRADAE+LRYK+
Sbjct: 276 FEYDGEITDEMKTVIVNSLRGPPLLLQAIAAYTLRIKTLISREMEKIDVRADAEMLRYKF 335
Query: 181 ENNIVMDVSSSDGSSPLQYPLYGNGKLGADVPPGGSQNQLLERQKAHVQQFLATEDALNN 240
ENN V D SSSD SSPL Y GNGK+G D GS NQLLERQKAHVQQFLATEDALN
Sbjct: 336 ENNRVTDNSSSDVSSPLSYQFNGNGKIGTDTHFQGSNNQLLERQKAHVQQFLATEDALNK 395
Query: 241 AAEARDLCEKLMKRLHGGTDVTSRSIGIGATSQNVGSLRQLQLDVWAKEREVSGLKASLN 300
AAEARDLC K + RLHG D + S +G T+Q+ +LRQ +LDVW KERE +GL+ASLN
Sbjct: 396 AAEARDLCHKFINRLHGSADTATHSF-VGGTTQSGSNLRQFELDVWGKEREAAGLRASLN 454
Query: 301 TLMSEIQRLNKLCAERKEAEDSLKKKWKKIEEFDGRRSELESIYTALLKANTDAASFWSQ 360
TL+SEIQRLNKLCAERKEAEDSLKKKWKKIEEFD RRSELE+IYT LLKAN DA +FW+Q
Sbjct: 455 TLLSEIQRLNKLCAERKEAEDSLKKKWKKIEEFDARRSELETIYTTLLKANMDAVAFWNQ 514
Query: 361 QPSTAREYALSTIIPACSAVVETSNSAKDLIEKEVSAFYRSPDNSLYMLPSSPQALLEAI 420
QP AREYA +T+IPA VV+ SNSAKD IEKEVSAF++SPDNSLYMLP++PQ LLE++
Sbjct: 515 QPLAAREYASATVIPASEVVVDISNSAKDFIEKEVSAFFQSPDNSLYMLPATPQGLLESM 574
Query: 421 GSSGSSGQEAVANAEISAAILTARAGARDPSAIPSICRVSAALQYAAGGLEGSDAGLASI 480
G++GS+G EAVA AE +AA+LTARAGARDPSAIPSICR+SAALQY A GLEGSDA LAS+
Sbjct: 575 GANGSTGPEAVAYAEKNAALLTARAGARDPSAIPSICRISAALQYPA-GLEGSDASLASV 633
Query: 481 LESLEFCLKRRGSEASVLEDLLKAINLVHIRRDLVQSGHALLNHAYFVQQDYERTTNFSL 540
LESLEFCL+ RGSEA VLEDL KAI+LVHIR+DLV+SGH+LL+HA+ QQ YERTTN+ L
Sbjct: 634 LESLEFCLRVRGSEACVLEDLAKAIDLVHIRQDLVESGHSLLDHAFRAQQKYERTTNYCL 693
Query: 541 NLAAEQERAVMEKWLPELKTGVLNAQQSLEACKYVWGLLDEWWEQPASTVVDWATVDGSN 600
+LA+EQE + ++WLPEL+T V NAQ S E CKYV GLLDEWWEQPASTVVDW TVDG +
Sbjct: 694 DLASEQENTISDQWLPELRTAVQNAQASSEHCKYVRGLLDEWWEQPASTVVDWVTVDGQS 753
Query: 601 VAFWHNHVKKLLTCYDQELL 620
VA W NHVK+LL YD+E L
Sbjct: 754 VAAWQNHVKQLLAFYDKESL 773
>At1g12150 hypothetical protein
Length = 548
Score = 38.5 bits (88), Expect = 0.011
Identities = 68/307 (22%), Positives = 123/307 (39%), Gaps = 45/307 (14%)
Query: 216 SQNQLLERQKAHVQQFLATEDA--LNNAAEARDLCEKLMKRLHGGTDVTSRSIGIGATSQ 273
++ +LL +K + + T +A L +E L E+ MK+ H +++ + I ++
Sbjct: 252 AEKKLLVLRKEYEPELSRTLEAKLLETTSEIEVLREE-MKKAHE-SEMNTVKIITNELNE 309
Query: 274 NVGSLRQLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAERKEAEDSLKKKWKKIEEF 333
L++ D + V+ L+ L L E + L + AER E E++ KK+E
Sbjct: 310 ATMRLQEAADDECSLRSLVNSLRMELEDLRREREELQQKEAERLEIEET-----KKLEAL 364
Query: 334 DGRRSELESIYTALLKANTDAASFWSQQPSTAREYALSTIIPACSAVVETSNSAKDLIEK 393
+LE + T ++A +AA+ + S +E A E + +L+ +
Sbjct: 365 KQESLKLEQMKTEAIEARNEAANMNRKIESLKKE------TEAAMIAAEEAEKRLELVIR 418
Query: 394 EVSAFYRSPD---NSLYMLPSSPQALLEAIGSSGSSGQEAVANAEISAAILTARAGARDP 450
EV + + + M+ ++ + SSGS + + E
Sbjct: 419 EVEEAKSAEEKVREEMKMISQKQESKKQDEESSGSKIKITIQEFE--------------- 463
Query: 451 SAIPSICRVSAALQYAAGGLEGS-DAGLASILESLEFCLKRRGSEASVLEDLLKAINLVH 509
+L+ AG E + + LA+I LE KRR + LE LKAI +
Sbjct: 464 -----------SLKRGAGETEAAIEKKLATIAAELEEINKRRAEADNKLEANLKAIEEMK 512
Query: 510 IRRDLVQ 516
+L Q
Sbjct: 513 QATELAQ 519
>At5g41790 myosin heavy chain-like protein
Length = 1305
Score = 36.2 bits (82), Expect = 0.056
Identities = 48/197 (24%), Positives = 87/197 (43%), Gaps = 35/197 (17%)
Query: 146 LQAITAYTSHLKSQI-SREIEKIDVRADAEILRYKYE----NNIVMDVSSSDGSSPLQYP 200
+ A+T ++LK+++ S +++K + A+ E R K E +N + DV
Sbjct: 979 IMALTELINNLKNELDSLQVQKSETEAELE--REKQEKSELSNQITDVQK---------- 1026
Query: 201 LYGNGKLGADVPPGGSQNQLLERQKAHVQQFLATEDALNNAAEARDLCEKLMKRLHGGTD 260
A V + N L E K + F TE LN ++L++ G +
Sbjct: 1027 --------ALVEQEAAYNTLEEEHKQINELFKETEATLNKVTVDYKEAQRLLEER--GKE 1076
Query: 261 VTSRSIGIGATSQNVGSLRQLQLDVWAKERE-----VSGLKASLNTLMSEIQRLNKLCAE 315
VTSR IG + + SLR +L++ E E +S ++ L +++ ++ E
Sbjct: 1077 VTSRDSTIGVHEETMESLRN-ELEMKGDEIETLMEKISNIEVKLRLSNQKLRVTEQVLTE 1135
Query: 316 RKEAEDSLKKKWKKIEE 332
++EA K++ K +EE
Sbjct: 1136 KEEA--FRKEEAKHLEE 1150
>At5g38840 kanadaptin - like protein
Length = 730
Score = 35.4 bits (80), Expect = 0.095
Identities = 55/228 (24%), Positives = 88/228 (38%), Gaps = 32/228 (14%)
Query: 215 GSQNQLLERQKAHVQQFLATEDALNNAAEARD---LCEKLMKRLHGGTDVTSRSIGIGAT 271
GS+NQ +E V + D + EA++ L EK T+VTS
Sbjct: 386 GSENQTVET----VDSLVDKRDNVLKEIEAKNEQLLTEKSKMETENVTEVTSGDSLDALD 441
Query: 272 SQNVGSLRQLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAERKEAEDSLKKKWKKIE 331
+ G L D A+ ++ L+TL SE+ R+ L + +KK+ K +
Sbjct: 442 AYMTGLSTTLVQDKTAQ------IQQELSTLQSELSRILYLLKIADPTGEEVKKRELKSQ 495
Query: 332 EFDGRRSELESIYTALLKANTDAASFWSQQPSTAREYALSTIIPACSAVVETSNSAKDLI 391
E ++SE S+ + A ++ A++ S P E N A +
Sbjct: 496 ELKIKKSETPSVEKKINIPLKQADPNEHKEKEVAKDLVDSENKP------EVENKASETA 549
Query: 392 EKEVSAFYRSPDNSLYMLPSSPQALLEAIGSSGSSGQEAVANAEISAA 439
E++ + Y +PS PQ L GS+ + N EI AA
Sbjct: 550 EEKKTTVY---------VPSKPQWL----GSAANKAIIEEKNPEIVAA 584
>At5g04990 unknown protein
Length = 471
Score = 34.3 bits (77), Expect = 0.21
Identities = 55/217 (25%), Positives = 83/217 (37%), Gaps = 38/217 (17%)
Query: 280 QLQLDVWAK--EREVSGLKASLNTLMSEIQRLNKLCAERKEAEDSLKKKWKKIEEFDGR- 336
Q+Q+++ K ERE L+ + S Q +E K+ E + K ++E + +
Sbjct: 173 QVQVELLDKKMEREAKVLRQEIERKASAFQ------SELKKIESRTESLEKSVDEVNAKP 226
Query: 337 ---RSELESIYTALLKANTDAASFWSQQPSTAREYALSTI-----------------IPA 376
+ ELE IY L K N D ++F R YA + A
Sbjct: 227 WVTKDELERIYEELKKGNVDDSAFSEISIDELRAYARDIMEKEIEKHAADGLGRVDYALA 286
Query: 377 CSAVVETSNSAKDLIEKEVSAF----YRSPDNSLYMLPSSPQALLEAIGSSGSSG--QEA 430
+S L+ K S F R+ N++ ML S + GS G Q
Sbjct: 287 SGGAFVMEHSDPYLVGKGSSWFATTMRRAHTNAVKMLSPSFGEPGQCFPLKGSEGYVQIR 346
Query: 431 VANAEISAAIL---TARAGARDPSAIPSICRVSAALQ 464
+ I A A++ A D S+ P CRVS +LQ
Sbjct: 347 LRGPIIPEAFTLEHVAKSVAYDRSSAPKDCRVSGSLQ 383
>At2g34730 putative myosin heavy chain
Length = 829
Score = 34.3 bits (77), Expect = 0.21
Identities = 44/207 (21%), Positives = 80/207 (38%), Gaps = 15/207 (7%)
Query: 146 LQAITAYTSHLKSQISREIEKIDVRADAEILRYKYENNIVMDVSSSDGSSPLQYPLYG-- 203
L ++ LK +S EK+ + AE + + DV S + + +YG
Sbjct: 419 LDSLLLENRQLKDSLSDAAEKMSQLSQAEADHQELIRKLETDVEDSRNEASIYEDVYGCF 478
Query: 204 ----NGKLGADVPPGGSQNQLLERQKAHVQQFLATEDALNNAAEARDLCEKLMKRLHGGT 259
G++ ++ +L + + LA ++A + + D C K +
Sbjct: 479 VTEFVGQIKCTKQETDLEHSMLREAYELLLEDLARKEARKSKEDFEDSCVKSVM-----M 533
Query: 260 DVTSRSIGIGATSQNVGSLRQLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAERKE- 318
+ I A + + +L L V KE + L EI RL L E++
Sbjct: 534 EECCSVIYKEAVKEAHKKIVELNLHVTEKEGTLRSEMVDKERLKEEIHRLGCLVKEKENL 593
Query: 319 ---AEDSLKKKWKKIEEFDGRRSELES 342
AE++L + KKIE + ++L+S
Sbjct: 594 VQTAENNLATERKKIEVVSQQINDLQS 620
>At2g19950 hypothetical protein
Length = 713
Score = 33.9 bits (76), Expect = 0.28
Identities = 42/199 (21%), Positives = 87/199 (43%), Gaps = 18/199 (9%)
Query: 210 DVPPGGSQNQLLERQKAHVQQFLATEDALNNAAEARD--LCEKLMKRLHGGTDVT---SR 264
D P G+Q +L + K V+ D++ +A + + + + GT + +
Sbjct: 190 DAPKNGTQRELDDSSKRDVENL----DSVVHAPSVNEGNVAQSTGDEVKVGTSINLEKEQ 245
Query: 265 SIGIGATSQNVGSLRQLQLDVWAKE-----REVSGL-KASLNTLMSEIQRLNKLCAERKE 318
+ TS N+ + + D + + E GL KA+++T S+ RL ++CA
Sbjct: 246 EPKVPVTSTNLKREQDRRADTTSMKIQDQLEEAQGLLKATVSTGQSKEARLARVCAGLSS 305
Query: 319 AEDSLKKKWKKIEEFDGRRSELESIYTALLK-ANTDAASFWSQQPSTAREYALSTIIPAC 377
+K + ++EE EL Y A ++ D ++ ++ T E ++ + A
Sbjct: 306 RLQEIKAENAQLEELLTAEQELTKSYEASIRHLQKDLSA--AKSEVTKVESSMVEALAAK 363
Query: 378 SAVVETSNSAKDLIEKEVS 396
++ +ET SA D ++ + +
Sbjct: 364 NSEIETLVSAMDALKNQAA 382
>At1g04600 putative myosin heavy chain
Length = 1730
Score = 33.9 bits (76), Expect = 0.28
Identities = 19/69 (27%), Positives = 37/69 (53%), Gaps = 7/69 (10%)
Query: 288 KEREVSGLKASLNTLMSEIQRLNK-------LCAERKEAEDSLKKKWKKIEEFDGRRSEL 340
K +E+S L+++L + EI+ L+K L AE ++ ++S+ KI+E + + E+
Sbjct: 968 KSKEISDLQSALQDMQLEIEELSKGLEMTNDLAAENEQLKESVSSLQNKIDESERKYEEI 1027
Query: 341 ESIYTALLK 349
I +K
Sbjct: 1028 SKISEERIK 1036
>At3g58160 myosin heavy chain MYA3
Length = 1242
Score = 33.5 bits (75), Expect = 0.36
Identities = 39/167 (23%), Positives = 72/167 (42%), Gaps = 16/167 (9%)
Query: 272 SQNVGSLRQLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAERKEAEDSLKKKWKKIE 331
SQ +R +++ E E+ LKA L + CAE +E + ++ K++E
Sbjct: 1071 SQITSPIRDTEIESLTAEVEM--LKALLQVEKQRADISERKCAEARELGE---RRRKRLE 1125
Query: 332 EFDGRRSELESIYTALLKANTDAASFWSQQPSTAREYALSTIIPACSAVV--------ET 383
E + R +L+ LL + +D +SQ S R ++S A + VV E
Sbjct: 1126 ETERRVYQLQDSLNRLLYSMSDQ---FSQLKSILRSPSMSASTMASAPVVRDDLADSSEN 1182
Query: 384 SNSAKDLIEKEVSAFYRSPDNSLYMLPSSPQALLEAIGSSGSSGQEA 430
S ++ + A S DN P+ Q +++ + ++ + G E+
Sbjct: 1183 SEASSSDSDFTFPAPSPSSDNFSTFNPNQLQVIVQDLSTTEAKGTES 1229
>At1g06530 hypothetical protein
Length = 323
Score = 33.5 bits (75), Expect = 0.36
Identities = 20/58 (34%), Positives = 30/58 (51%), Gaps = 4/58 (6%)
Query: 288 KEREVSGLKASLNTLMSEIQR----LNKLCAERKEAEDSLKKKWKKIEEFDGRRSELE 341
KE+EV LK + +L S++ + L K E+ EDSLK KK+ + EL+
Sbjct: 202 KEKEVHDLKEKIKSLESDVAKGKTELQKWITEKMVVEDSLKDSEKKVVALESEIVELQ 259
>At5g48600 chromosome condensation protein
Length = 1241
Score = 33.1 bits (74), Expect = 0.47
Identities = 76/347 (21%), Positives = 129/347 (36%), Gaps = 39/347 (11%)
Query: 235 EDALNNAAEARDLCEKLMKRLHGGTDVTSRSIG----IGATSQNVGSLRQLQLDVWAKER 290
ED + + + L +KL K D+T S I +N+ L+++ LD K
Sbjct: 344 EDLKHVKQKIKKLEDKLEKDSSKIGDMTKESEDSSNLIPKLQENIPKLQKVLLDEEKKLE 403
Query: 291 EVSGL-KASLNTLMSEIQRLNKLCAERKEAEDSLKKKWKKIEEFDGRRSELESIYTALLK 349
E+ + K SE L K+ AE + E L K++ L + A LK
Sbjct: 404 EIKAIAKVETEGYRSE---LTKIRAELEPWEKDLIVHRGKLDVASSESELLSKKHEAALK 460
Query: 350 ANTDAASFWSQQPSTAREYALST-------------IIPACSAVVETSNSAKDLIEKEVS 396
A TDA S + +E A +T I A E+ + L+ +E +
Sbjct: 461 AFTDAQKQLSDISTRKKEKAAATTSWKADIKKKKQEAIEARKVEEESLKEQETLVPQEQA 520
Query: 397 AFYRSPD-NSLYMLPSSPQALLEAI----------GSSGSSGQEAVANAEISAAILTARA 445
A + + S S +L+A+ G G G +A+ AI TA A
Sbjct: 521 AREKVAELKSAMNSEKSQNEVLKAVLRAKENNQIEGIYGRMGDLGAIDAKYDVAISTACA 580
Query: 446 GARDPSAIPSICRVSAALQYAAGGLEGSDAGLAS--ILESLEFCLKRRGSEASVLEDLLK 503
G D + + A ++ L + G A+ ILE + + + ED+ +
Sbjct: 581 GL-DYIVVETTSSAQACVEL----LRKGNLGFATFMILEKQTDHIHKLKEKVKTPEDVPR 635
Query: 504 AINLVHIRRDLVQSGHALLNHAYFVQQDYERTTNFSLNLAAEQERAV 550
+LV ++ + ++ V +D ++ T + E R V
Sbjct: 636 LFDLVRVKDERMKLAFYAALGNTVVAKDLDQATRIAYGGNREFRRVV 682
>At2g32240 putative myosin heavy chain
Length = 1333
Score = 33.1 bits (74), Expect = 0.47
Identities = 33/121 (27%), Positives = 61/121 (50%), Gaps = 17/121 (14%)
Query: 235 EDALNNAAE-ARDLCEKL------MKRLHGGTDVTSRSIGIGATSQNVGSLRQLQLDVWA 287
E ALN A E ++L E L K+L D S I S++ L ++ ++
Sbjct: 689 EAALNIATENEKELTENLNAVTSEKKKLEATVDEYSVKI-----SESENLLESIRNELNV 743
Query: 288 KEREVSGLKASLNTL-MSEIQRLNKLCAERKEAEDSLKKKWKKIEEFDGRRSELESIYTA 346
+ ++ ++ L + E + + KL K AE+SL++K ++I+E +R ELE+++ +
Sbjct: 744 TQGKLESIENDLKAAGLQESEVMEKL----KSAEESLEQKGREIDEATTKRMELEALHQS 799
Query: 347 L 347
L
Sbjct: 800 L 800
Score = 32.7 bits (73), Expect = 0.62
Identities = 70/352 (19%), Positives = 135/352 (37%), Gaps = 45/352 (12%)
Query: 3 EAYDRQCDEASRIFAEYHKRLCYYINQARDAQRSGVDSSVEMVNSFSAKNEKEAVYSTVK 62
E R +EA + F E HK+ + + ++Q++ S + SAK +E + S +
Sbjct: 205 EGLKRSAEEAQK-FEELHKQSASHADS--ESQKALEFSELLKSTKESAKEMEEKMASLQQ 261
Query: 63 GSKSSDDVIVIETTREKNIRKACESLVAYMVDKIQSSFPAYEGSGVLSNPQAEAAKLGFD 122
K ++ + E ++ + L A + S E +S+ +A +L +
Sbjct: 262 EIKELNEKMSENEKVEAALKSSAGELAAVQEELALSKSRLLETEQKVSSTEALIDELTQE 321
Query: 123 FDGQIPDEVRTVIVNCLKSPPLLLQAITAYTSHLKSQISREIEKIDVRADAEILRYKYEN 182
+ + E R K +LQ + A T L++++S + E I+ + E+ +
Sbjct: 322 LEQKKASESR------FKEELSVLQDLDAQTKGLQAKLSEQ-EGINSKLAEEL-----KE 369
Query: 183 NIVMDVSSSDGSSPLQYPLYGNGKLGADVPPGGSQNQLLERQKAHVQQFLATEDALNNAA 242
+++ S D L+ N KL + + + LE A V +N A
Sbjct: 370 KELLESLSKDQEEKLRT---ANEKLAEVL----KEKEALEANVAEVT---------SNVA 413
Query: 243 EARDLCEKLMKRLHGGTDVTSRSIGIGATSQNVGSLRQLQLDVWAKEREVSGLKASLNTL 302
++C +L ++L + +N L + E+ SL L
Sbjct: 414 TVTEVCNELEEKLK-------------TSDENFSKTDALLSQALSNNSELEQKLKSLEEL 460
Query: 303 MSEIQRLNKLCAERK-EAEDSLKKKWKKIEEFDGRRSELESIYTALLKANTD 353
SE ++ E ED ++ + EE + ELE+ +TA + N +
Sbjct: 461 HSEAGSAAAAATQKNLELEDVVRSSSQAAEEAKSQIKELETKFTAAEQKNAE 512
Score = 32.0 bits (71), Expect = 1.1
Identities = 33/153 (21%), Positives = 65/153 (41%), Gaps = 9/153 (5%)
Query: 204 NGKLGADVPPGGSQNQLLERQKAHVQQFLATEDALNNAAEARDLCEKLMKRLHGGTDVTS 263
N KL ++ GS+ L+ + + ++ E N ++ E L K+L +
Sbjct: 1011 NLKLNLELANHGSEANELQTKLSALEA--EKEQTANELEASKTTIEDLTKQLTSEGEKLQ 1068
Query: 264 RSIGIGATSQNV------GSLRQLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAERK 317
I N + +LQ + E +++ + +TL+SEI++L + AE+
Sbjct: 1069 SQISSHTEENNQVNAMFQSTKEELQSVIAKLEEQLTVESSKADTLVSEIEKLRAVAAEKS 1128
Query: 318 EAEDSLKKKWKKIEEFDGRRSE-LESIYTALLK 349
E ++ K + E + E +E+ TA +K
Sbjct: 1129 VLESHFEELEKTLSEVKAQLKENVENAATASVK 1161
>At1g79280 hypothetical protein
Length = 2111
Score = 33.1 bits (74), Expect = 0.47
Identities = 32/125 (25%), Positives = 52/125 (41%), Gaps = 18/125 (14%)
Query: 235 EDALNNAAEARDLCEKLMKRLHGGTDV----------TSRSIGIGATSQNVGSLRQLQLD 284
E+ L DLC K M++L TD+ T R+I I ++ +RQL+
Sbjct: 1331 ENLLKTKQTELDLCMKEMEKLRMETDLHKKRVDELRETYRNIDIADYNRLKDEVRQLEEK 1390
Query: 285 VWAKEREVSGLKASLNTLMSEIQRLNK--------LCAERKEAEDSLKKKWKKIEEFDGR 336
+ AK+ K L ++I L K L K +D+ + + EF+ +
Sbjct: 1391 LKAKDAHAEDCKKVLLEKQNKISLLEKELTNCKKDLSEREKRLDDAQQAQATMQSEFNKQ 1450
Query: 337 RSELE 341
+ ELE
Sbjct: 1451 KQELE 1455
>At1g05320 unknown protein
Length = 841
Score = 33.1 bits (74), Expect = 0.47
Identities = 32/116 (27%), Positives = 57/116 (48%), Gaps = 18/116 (15%)
Query: 237 ALNNAAEARDLCEKLMKRLHGGTDVTSRSI---GIGATSQ-NVGSLRQLQLDVWAKEREV 292
AL AE D+ G T RS+ G+ TSQ + + D+ A + +
Sbjct: 234 ALQKGAEHEDI----------GNVSTKRSVELQGLFQTSQLKLEKAEEKLKDLEAIQVKN 283
Query: 293 SGLKASLNTLMSE----IQRLNKLCAERKEAEDSLKKKWKKIEEFDGRRSELESIY 344
S L+A+L+ M + + LN + + K +E+ L+K+ ++I+E R ELE+++
Sbjct: 284 SSLEATLSVAMEKERDLSENLNAVMEKLKSSEERLEKQAREIDEATTRSIELEALH 339
>At3g05130 hypothetical protein
Length = 634
Score = 32.7 bits (73), Expect = 0.62
Identities = 73/310 (23%), Positives = 120/310 (38%), Gaps = 25/310 (8%)
Query: 288 KEREVSGLKASLNTLMSEIQRLNKLCAERKEAEDSLKKKWKKIEEFDGRRSELESIYTAL 347
K+ E+ GLK + L+SE + + E+K + L++K K+ E ++ E + L
Sbjct: 248 KKNEIDGLKREIKVLLSEKNEMEIVKIEQKGVIEELERKLDKLNETVRSLTKEEKVLRDL 307
Query: 348 ---LKANTDAASFWSQQPSTAREYALSTIIPACSAVVETSNSAKDLIEKEVSAF-YRSPD 403
L+ N D S + AL + VE K+LIEK++ +S D
Sbjct: 308 VIGLEKNLD-ESMEKESGMMVEIDALGKERTIKESEVERLIGEKNLIEKQMEMLNVQSSD 366
Query: 404 NS--LYMLPSSPQALLEAIGSSGSSGQEAVANA-EISAAILTARAGARDPSAIP-----S 455
+ L L E I S E A E++ A+ + D + I
Sbjct: 367 KGKLIDQLSREKVELEERIFSRERKLVELNRKADELTHAVAVLQKNCDDQTKINGKLSCK 426
Query: 456 ICRVSAALQYAAGGLEGSDAGLASILESLE----FCLKRRGSEASVLEDLLKAINLVHIR 511
+ ++S AL E +D L + E LK A LE+L K V I
Sbjct: 427 VDQLSNALAQVELRREEADKALDEEKRNGEDLKAEVLKSEKMVAKTLEELEK----VKIE 482
Query: 512 RDLVQSGHALLNHAYFVQQDYERTTNFSLNLAAEQERAVMEKWLPELKTGVLNAQQSLEA 571
R + S L Q + ++ N L + R ME EL++ ++A++S+
Sbjct: 483 RKSLFSAKNDLES----QSESLKSENVKLEKELVELRKAMEALKTELESAGMDAKRSMVM 538
Query: 572 CKYVWGLLDE 581
K +L +
Sbjct: 539 LKSAASMLSQ 548
>At4g22950 putative MADS Box / AGL protein
Length = 219
Score = 32.3 bits (72), Expect = 0.81
Identities = 29/111 (26%), Positives = 55/111 (49%), Gaps = 10/111 (9%)
Query: 221 LERQKAHVQQFLATEDALNNAAEARDLCEKLMKRLHGGTDVTSRSIGIGATSQNVGSLRQ 280
+ER + +++ +N+ +ARD L K++ + +G G + ++ L+Q
Sbjct: 66 IERYQRRIKEIGNNHKRNDNSQQARDETSGLTKKIEQLEISKRKLLGEGIDACSIEELQQ 125
Query: 281 LQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAER----KEAEDSLKKKW 327
L+ + +R +S ++A L+ E + KL AE KE +D LK+KW
Sbjct: 126 LENQL---DRSLSRIRAKKYQLLRE--EIEKLKAEERNLVKENKD-LKEKW 170
>At3g51410 putative protein
Length = 255
Score = 32.0 bits (71), Expect = 1.1
Identities = 19/61 (31%), Positives = 34/61 (55%), Gaps = 3/61 (4%)
Query: 290 REVSGLKASLNTLMSEIQRLNKLCAERKEAEDSLKKKWKKIEEFDGRRSELESIYTALLK 349
R +SG S +++MSE+Q L+ + + +KK +++EE R ELE+ +L K
Sbjct: 182 RLISGGHRSSSSIMSELQNLDAVLRSDGDNSKEIKKMLERLEE---RTEELETALDSLFK 238
Query: 350 A 350
+
Sbjct: 239 S 239
>At1g54200 unknown protein
Length = 366
Score = 32.0 bits (71), Expect = 1.1
Identities = 19/84 (22%), Positives = 41/84 (48%), Gaps = 3/84 (3%)
Query: 28 NQARDAQRSGVDSSVEMVNSFSAKNEKEAVYSTVKGSKSSDDVIVIETTREKNIRKACES 87
N+ + R ++++ E++ S+ KN KE + +V+ + DD + T ++
Sbjct: 281 NRVMEENRRVIEAAKELLRSYQKKN-KEVIEVSVEDDEEDDDDDALSCTSSDLFE--LDN 337
Query: 88 LVAYMVDKIQSSFPAYEGSGVLSN 111
L A +D+ + P YE + + +N
Sbjct: 338 LSAIGIDRYREELPVYETTRLNTN 361
>At4g30790 putative protein
Length = 1148
Score = 31.6 bits (70), Expect = 1.4
Identities = 28/102 (27%), Positives = 45/102 (43%), Gaps = 5/102 (4%)
Query: 1 MLEAYDRQCDEASRIFAEYHKRLC-YYINQARDAQR--SGVDSSVEMVNSFSAK--NEKE 55
ML+ CD + E +RL Y+ R + SG++ E V+ + A+ ++ E
Sbjct: 648 MLKEKQMHCDSYEKRIRELEQRLSDEYLQGQRHNNKDVSGLNLMHEKVSEYKAEASSDVE 707
Query: 56 AVYSTVKGSKSSDDVIVIETTREKNIRKACESLVAYMVDKIQ 97
+ V GS+ D+V + K KA E + MVD Q
Sbjct: 708 GNKTHVSGSEPMDEVSCVSNLTSKQPCKAREGMDENMVDSSQ 749
>At3g11450 putative cell division related protein
Length = 663
Score = 31.6 bits (70), Expect = 1.4
Identities = 43/181 (23%), Positives = 77/181 (41%), Gaps = 45/181 (24%)
Query: 239 NNAAEARDLCEKLMKRLHGGTDVTSRSIGIGATSQNVGSLRQLQLDVWAKEREVSGLKAS 298
N+ AE++ EK+ K+ +GGT+ T+R + +++Q + W+KE
Sbjct: 446 NDEAESK---EKVSKKTNGGTEPTTRVSQLDSSTQ--------KKQPWSKE--------- 485
Query: 299 LNTLMSEIQRLNKLCAERKEAEDSLKKKWKKIEEFDGRRSELESIY----TALLKANTDA 354
EI L K + + ++W+ I E+ G +E I T LL+ A
Sbjct: 486 ------EIDMLRKGMIKYPK---GTSRRWEVISEYIGTGRSVEEILKATKTVLLQKPDSA 536
Query: 355 ASFWS----QQPSTAREYALST------IIPACSAVVETSNSAKDLIEKEVSAFYRSPDN 404
+F S ++PS + LST +P + S + ++ K S+ +S DN
Sbjct: 537 KAFDSFLEKRKPSASITSPLSTREELGESLPTMTTTTNAKPSKETVVGKSSSS--QSSDN 594
Query: 405 S 405
+
Sbjct: 595 N 595
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.314 0.129 0.361
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,997,649
Number of Sequences: 26719
Number of extensions: 532153
Number of successful extensions: 1591
Number of sequences better than 10.0: 52
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 43
Number of HSP's that attempted gapping in prelim test: 1560
Number of HSP's gapped (non-prelim): 72
length of query: 621
length of database: 11,318,596
effective HSP length: 105
effective length of query: 516
effective length of database: 8,513,101
effective search space: 4392760116
effective search space used: 4392760116
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 63 (28.9 bits)
Medicago: description of AC137823.16


