

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137822.15 + phase: 0
(279 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
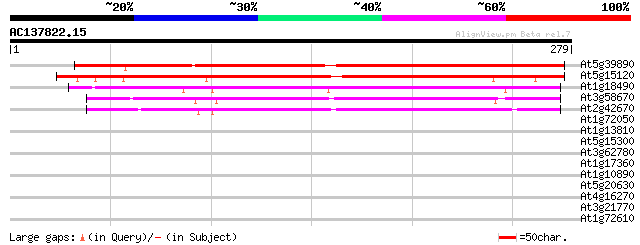
Score E
Sequences producing significant alignments: (bits) Value
At5g39890 unknown protein 239 9e-64
At5g15120 unknown protein 233 7e-62
At1g18490 unknown protein 208 3e-54
At3g58670 unknown protein 179 1e-45
At2g42670 unknown protein 167 4e-42
At1g72050 transcription factor IIIA (TFIIIA) 30 1.9
At1g13810 hypothetical protein 30 1.9
At5g15300 unknown protein 28 5.5
At3g62780 shock protein SRC2-like 28 5.5
At1g17360 COP1-Interacting Protein 7, putative 28 7.1
At1g10890 unknown protein 28 7.1
At5g20630 germin-like protein (GLP3b) 27 9.3
At4g16270 peroxidase like protein 27 9.3
At3g21770 putative peroxidase 27 9.3
At1g72610 germin-like protein 27 9.3
>At5g39890 unknown protein
Length = 276
Score = 239 bits (611), Expect = 9e-64
Identities = 123/246 (50%), Positives = 159/246 (64%), Gaps = 8/246 (3%)
Query: 33 RRVKKHVPKALQELFDSCKQTFKG--PGTVPSPRDVHKLCHILDNMKPEDVGLSRDLQFF 90
RR KK + +Q+LFD+CK+ F GTVPS ++ L +LD +KPEDVG++ + +F
Sbjct: 37 RRSKKTLICPVQKLFDTCKKVFADGKSGTVPSQENIEMLRAVLDEIKPEDVGVNPKMSYF 96
Query: 91 KPGNIIKENQRVTYTTVYKCDNFSLCIFFLPERGVIPLHNHPGMTVFSKLLLGQMHIKSY 150
+ + + VTY +Y C FS+CIF LP GVIPLHNHP MTVFSKLL G MHIKSY
Sbjct: 97 R-STVTGRSPLVTYLHIYACHRFSICIFCLPPSGVIPLHNHPEMTVFSKLLFGTMHIKSY 155
Query: 151 DWVDHEASHNLLQPSSKLRLAKLKANKTFTAPCDTSVLYPTTGGNIHEFTAITPCAVLDV 210
DWV QPSS RLAK+K + FTAPCDTS+LYP GGN+H FTA T CAVLDV
Sbjct: 156 DWVPDSP-----QPSSDTRLAKVKVDSDFTAPCDTSILYPADGGNMHCFTAKTACAVLDV 210
Query: 211 IGPPYSKEDGRDCSYYKDYPYNAFPNEEKIGEVKDKDDSYGLLEEIDMPENCQMDGIEYL 270
IGPPYS GR C+YY DYP+++F + + ++K+ L E + PE+ + + Y
Sbjct: 211 IGPPYSDPAGRHCTYYFDYPFSSFSVDGVVVAEEEKEGYAWLKEREEKPEDLTVTALMYS 270
Query: 271 GPPIDD 276
GP I +
Sbjct: 271 GPTIKE 276
>At5g15120 unknown protein
Length = 293
Score = 233 bits (595), Expect = 7e-62
Identities = 126/270 (46%), Positives = 171/270 (62%), Gaps = 22/270 (8%)
Query: 24 KKRNKPYNH------RRVKKHVPK----ALQELFDSCKQTFK--GPGTVPSPRDVHKLCH 71
KK+NK N RR K P A++ LF++CK+ F GPG +PS + +L
Sbjct: 29 KKKNKNKNKKMMMTWRRKKIDSPADGITAVRRLFNTCKEVFSNGGPGVIPSEDKIQQLRE 88
Query: 72 ILDNMKPEDVGLSRDLQFFKPGNII--KENQRVTYTTVYKCDNFSLCIFFLPERGVIPLH 129
ILD+MKPEDVGL+ + +F+P + + + + +TY +++CD FS+ IF LP GVIPLH
Sbjct: 89 ILDDMKPEDVGLTPTMPYFRPNSGVEARSSPPITYLHLHQCDQFSIGIFCLPPSGVIPLH 148
Query: 130 NHPGMTVFSKLLLGQMHIKSYDWVDHEASHNLLQPSSKLRLAKLKANKTFTAPCDTSVLY 189
NHPGMTVFSKLL G MHIKSYDWV + SK RLAKLK + TFTAPC+ S+LY
Sbjct: 149 NHPGMTVFSKLLFGTMHIKSYDWVVDAPMRD-----SKTRLAKLKVDSTFTAPCNASILY 203
Query: 190 PTTGGNIHEFTAITPCAVLDVIGPPYSKEDGRDCSYYKDYPYNAFPNEEK--IGEVKDKD 247
P GGN+H FTAIT CAVLDV+GPPY +GR C+Y+ ++P + +E+ + ++K+
Sbjct: 204 PEDGGNMHRFTAITACAVLDVLGPPYCNPEGRHCTYFLEFPLDKLSSEDDDVLSSEEEKE 263
Query: 248 DSYGLLEEIDMPE-NCQMDGIEYLGPPIDD 276
L E D PE + + G Y GP ++D
Sbjct: 264 GYAWLQERDDNPEDHTNVVGALYRGPKVED 293
>At1g18490 unknown protein
Length = 282
Score = 208 bits (529), Expect = 3e-54
Identities = 108/259 (41%), Positives = 153/259 (58%), Gaps = 15/259 (5%)
Query: 30 YNHRRVKKHVPKALQELFDSCKQTFKGPGTVPSPRDVHKLCHILDNMKPEDVGLSR---- 85
Y R ++ PK +QEL+D CK+TF G P+ + KLC +LD++ P DVGL
Sbjct: 23 YMASRNQEKSPK-VQELYDLCKETFTGKAPSPASMAIQKLCSVLDSVSPADVGLEEVSQD 81
Query: 86 DLQFFKPGNIIKEN------QRVTYTTVYKCDNFSLCIFFLPERGVIPLHNHPGMTVFSK 139
D + + + + N Q +T+ +++CD F++CIF P VIPLH+HP M VFSK
Sbjct: 82 DDRGYGVSGVSRFNRVGRWAQPITFLDIHECDTFTMCIFCFPTSSVIPLHDHPEMAVFSK 141
Query: 140 LLLGQMHIKSYDWVDHEA---SHNLLQPSSKLRLAKLKANKTFTAPCDTSVLYPTTGGNI 196
+L G +H+K+YDWV+ + S RLAKL ++K T + LYP TGGN+
Sbjct: 142 ILYGSLHVKAYDWVEPPCIITQDKGVPGSLPARLAKLVSDKVITPQSEIPALYPKTGGNL 201
Query: 197 HEFTAITPCAVLDVIGPPYSKEDGRDCSYYKDYPYNAFPNEEKIGEVKD-KDDSYGLLEE 255
H FTA+TPCAVLD++ PPY + GR CSYY DYP++ F E + +V + K+D Y L +
Sbjct: 202 HCFTALTPCAVLDILSPPYKESVGRSCSYYMDYPFSTFALENGMKKVDEGKEDEYAWLVQ 261
Query: 256 IDMPENCQMDGIEYLGPPI 274
ID P++ M Y GP I
Sbjct: 262 IDTPDDLHMRPGSYTGPTI 280
>At3g58670 unknown protein
Length = 242
Score = 179 bits (454), Expect = 1e-45
Identities = 100/246 (40%), Positives = 142/246 (57%), Gaps = 16/246 (6%)
Query: 39 VPKALQELFDSCKQTFKGPGTVPSPRDVHKLCHILDNMKPEDVGLSRDLQFFK----PGN 94
+P +Q LF++CK + G V S + K+ ++L+ +KP DVGL ++ Q + PGN
Sbjct: 1 MPYFIQRLFNTCKSSLSPNGPV-SEEALDKVRNVLEKIKPSDVGLEQEAQLVRNWPGPGN 59
Query: 95 IIKENQR----VTYTTVYKCDNFSLCIFFLPERGVIPLHNHPGMTVFSKLLLGQMHIKSY 150
N + Y +++CD+FS+ IF +P +IPLHNHPGMTV SKL+ G MH+KSY
Sbjct: 60 ERNGNHHSLPAIKYLQLHECDSFSIGIFCMPPGSIIPLHNHPGMTVLSKLVYGSMHVKSY 119
Query: 151 DWVDHEASHNLLQPSSKLRLAKLKANKTFTAPCDTSVLYPTTGGNIHEFTAITPCAVLDV 210
DW + + S L + R AKL + T+P + LYPTTGGNIH F AIT CA+ D+
Sbjct: 120 DWAEPDQSE--LDDPLQARPAKLVKDIDMTSPSPATTLYPTTGGNIHCFKAITHCAIFDI 177
Query: 211 IGPPYSKEDGRDCSYYKDYPYNAFPNEEKI--GEVKDKDDSYGLLEEIDMPENCQMDGIE 268
+ PPYS GR C+Y++ P P E ++ GEV + LEE P+N + +
Sbjct: 178 LSPPYSSTHGRHCNYFRKSPMLDLPGEIEVMNGEV---ISNVTWLEEYQPPDNFVIWRVP 234
Query: 269 YLGPPI 274
Y GP I
Sbjct: 235 YRGPVI 240
>At2g42670 unknown protein
Length = 241
Score = 167 bits (424), Expect = 4e-42
Identities = 91/244 (37%), Positives = 140/244 (57%), Gaps = 13/244 (5%)
Query: 39 VPKALQELFDSCKQTFKGPGTVPSPRDVHKLCHILDNMKPEDVGLSRDLQFFKP--GNII 96
+P Q L+++CK +F G + + K+ ++L+ +KP DVG+ +D Q + G +
Sbjct: 1 MPYFAQRLYNTCKASFSSDGPITEDA-LEKVRNVLEKIKPSDVGIEQDAQLARSRSGPLN 59
Query: 97 KEN------QRVTYTTVYKCDNFSLCIFFLPERGVIPLHNHPGMTVFSKLLLGQMHIKSY 150
+ N + Y +++CD+FS+ IF +P +IPLHNHPGMTV SKL+ G MH+KSY
Sbjct: 60 ERNGSNQSPPAIKYLHLHECDSFSIGIFCMPPSSMIPLHNHPGMTVLSKLVYGSMHVKSY 119
Query: 151 DWVDHEASHNLLQPSSKLRLAKLKANKTFTAPCDTSVLYPTTGGNIHEFTAITPCAVLDV 210
DW++ + + + S+ R AKL + TA + LYP +GGNIH F AIT CA+LD+
Sbjct: 120 DWLEPQLTEP--EDPSQARPAKLVKDTEMTAQSPVTTLYPKSGGNIHCFKAITHCAILDI 177
Query: 211 IGPPYSKEDGRDCSYYKDYPYNAFPNEEKIGEVKDKDDSYGLLEEIDMPENCQMDGIEYL 270
+ PPYS E R C+Y++ P E ++ D ++ LEE P++ + I Y
Sbjct: 178 LAPPYSSEHDRHCTYFRKSRREDLPGELEVDGEVVTDVTW--LEEFQPPDDFVIRRIPYR 235
Query: 271 GPPI 274
GP I
Sbjct: 236 GPVI 239
>At1g72050 transcription factor IIIA (TFIIIA)
Length = 412
Score = 29.6 bits (65), Expect = 1.9
Identities = 22/68 (32%), Positives = 30/68 (43%), Gaps = 3/68 (4%)
Query: 16 CYVKRVIAKKRNKPYNHRRVKKHVPKALQELFDSCKQTFKGPGTVPSPRDVHKLCHILDN 75
CYV A R K + +R + H K + ++CK F G V R V K H DN
Sbjct: 96 CYVDDCAASYRRKDHLNRHLLTHKGKLFKCPKENCKSEFSVQGNV--GRHVKKY-HSNDN 152
Query: 76 MKPEDVGL 83
++ GL
Sbjct: 153 RDKDNTGL 160
>At1g13810 hypothetical protein
Length = 303
Score = 29.6 bits (65), Expect = 1.9
Identities = 15/46 (32%), Positives = 22/46 (47%), Gaps = 1/46 (2%)
Query: 202 ITPCAVLDVIGPPYSKEDGRDCSYYKDYPYNAFPNEEKIGEVKDKD 247
+T C VL+V P+ D +K PYN P + + E+ D D
Sbjct: 162 VTSCGVLEV-KCPFDNRDNSKVYPWKKVPYNCVPQLQGLMEIVDTD 206
>At5g15300 unknown protein
Length = 514
Score = 28.1 bits (61), Expect = 5.5
Identities = 16/55 (29%), Positives = 24/55 (43%), Gaps = 1/55 (1%)
Query: 223 CSYYKDYPYNAFPNEEKIGEVKDKDDSYGLLEEIDMPENCQMDGIEYLGPPIDDT 277
C Y + + NE+ + KD+ Y LL I Q DG++ + DDT
Sbjct: 424 CKIYGNVELGKYANEKLLSMRKDESGDYVLLSNI-YASTGQWDGVQKVRKMFDDT 477
>At3g62780 shock protein SRC2-like
Length = 298
Score = 28.1 bits (61), Expect = 5.5
Identities = 15/53 (28%), Positives = 27/53 (50%)
Query: 142 LGQMHIKSYDWVDHEASHNLLQPSSKLRLAKLKANKTFTAPCDTSVLYPTTGG 194
+G++H++ + +DH + Q ++ K KA+ +FT V PT GG
Sbjct: 93 IGEVHVQVKELLDHLGNDKTGQRYVTYQIGKSKADISFTYSFTGPVEVPTGGG 145
>At1g17360 COP1-Interacting Protein 7, putative
Length = 1061
Score = 27.7 bits (60), Expect = 7.1
Identities = 19/56 (33%), Positives = 23/56 (40%), Gaps = 6/56 (10%)
Query: 216 SKEDGRDCSYYKDYPYNAFPNEEKIGEVKDKDDSYGLL-----EEIDMPENCQMDG 266
S E+G S P FP E + KD DD Y + +ID P N DG
Sbjct: 320 SNEEG-PLSTPSSIPDATFPKESEENSKKDDDDVYSTISDDSQNQIDKPGNFMTDG 374
>At1g10890 unknown protein
Length = 592
Score = 27.7 bits (60), Expect = 7.1
Identities = 20/75 (26%), Positives = 33/75 (43%), Gaps = 5/75 (6%)
Query: 67 HKLCHILDNMKP-EDVGLSRDLQFFKPGNIIKENQRVTYTTVYKCDNFSLCIFFLPERG- 124
HK+ +D+ P E+V + FF I E + + + ++F LC F ER
Sbjct: 378 HKILRFVDDFLPVENVVFGGNTYFFAKERYIFEGKGPEEIDITETEDFLLCFDFTAERFG 437
Query: 125 ---VIPLHNHPGMTV 136
+P H++ TV
Sbjct: 438 PRLPLPFHSYYAETV 452
>At5g20630 germin-like protein (GLP3b)
Length = 211
Score = 27.3 bits (59), Expect = 9.3
Identities = 18/43 (41%), Positives = 19/43 (43%), Gaps = 3/43 (6%)
Query: 94 NIIKENQRVTYTTVYKCDN---FSLCIFFLPERGVIPLHNHPG 133
NIIK + Y N SL L GVIPLH HPG
Sbjct: 65 NIIKAAVTPAFAPAYAGINGLGVSLARLDLAGGGVIPLHTHPG 107
>At4g16270 peroxidase like protein
Length = 362
Score = 27.3 bits (59), Expect = 9.3
Identities = 18/52 (34%), Positives = 22/52 (41%), Gaps = 3/52 (5%)
Query: 150 YDWVDHEASHNLLQPSSKLRLAKLKANKTFTAPCDTSVLYPTTGGNIHEFTA 201
Y WV+ L P L +L + F CD SVL T G + E TA
Sbjct: 81 YSWVETTV---LEDPRMAASLLRLHFHDCFVNGCDASVLLDDTEGLVGEKTA 129
>At3g21770 putative peroxidase
Length = 329
Score = 27.3 bits (59), Expect = 9.3
Identities = 15/42 (35%), Positives = 20/42 (46%)
Query: 154 DHEASHNLLQPSSKLRLAKLKANKTFTAPCDTSVLYPTTGGN 195
DH +H PS L ++ + F CD SVL +T GN
Sbjct: 47 DHIQNHIHNGPSLAAPLIRMHFHDCFVRGCDGSVLINSTSGN 88
>At1g72610 germin-like protein
Length = 208
Score = 27.3 bits (59), Expect = 9.3
Identities = 11/26 (42%), Positives = 16/26 (61%)
Query: 120 LPERGVIPLHNHPGMTVFSKLLLGQM 145
L +GVIP+H HPG + +L G +
Sbjct: 91 LAPKGVIPMHTHPGASEVLFVLTGSI 116
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.140 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,355,312
Number of Sequences: 26719
Number of extensions: 348135
Number of successful extensions: 673
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 651
Number of HSP's gapped (non-prelim): 15
length of query: 279
length of database: 11,318,596
effective HSP length: 98
effective length of query: 181
effective length of database: 8,700,134
effective search space: 1574724254
effective search space used: 1574724254
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC137822.15


