

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137703.1 + phase: 0
(233 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
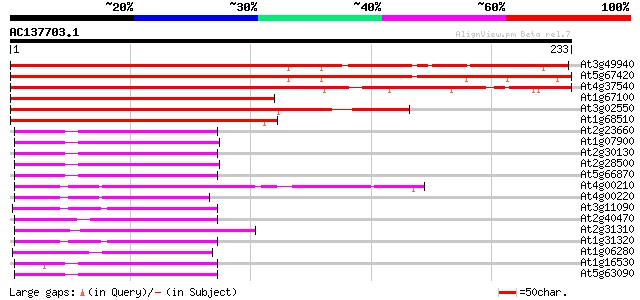
Score E
Sequences producing significant alignments: (bits) Value
At3g49940 unknown protein 254 2e-68
At5g67420 unknown protein 245 2e-65
At4g37540 unknown protein 241 2e-64
At1g67100 putative protein 173 6e-44
At3g02550 unknown protein 164 4e-41
At1g68510 unknown protein 158 3e-39
At2g23660 putative protein 54 7e-08
At1g07900 ASYMMETRIC LEAVES2-like protein 8 (ASL8) 54 9e-08
At2g30130 unknown protein 53 1e-07
At2g28500 putative protein 53 1e-07
At5g66870 unknown protein 51 5e-07
At4g00210 putative protein 50 1e-06
At4g00220 LOB DOMAIN 30 (LBD30) 49 2e-06
At3g11090 putative protein 48 4e-06
At2g40470 unknown protein 48 5e-06
At2g31310 putative protein 47 9e-06
At1g31320 putative protein 47 9e-06
At1g06280 putative protein 47 9e-06
At1g16530 putative protein 47 1e-05
At5g63090 LOBd (LOB) 46 2e-05
>At3g49940 unknown protein
Length = 247
Score = 254 bits (650), Expect = 2e-68
Identities = 145/252 (57%), Positives = 171/252 (67%), Gaps = 26/252 (10%)
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
MSCNGCRVLRKGCSE+CILRPC+QWIE+PEAQGHATVFVAKFFGRAGLMSFIS VPE Q
Sbjct: 1 MSCNGCRVLRKGCSENCILRPCIQWIESPEAQGHATVFVAKFFGRAGLMSFISAVPESQC 60
Query: 61 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTLRPLPELT----LL 116
PALFQSLL+EACGRTVNPVNGAVGLLWTGNW+VCQAAVETVLRGG+L+P+PEL
Sbjct: 61 PALFQSLLYEACGRTVNPVNGAVGLLWTGNWNVCQAAVETVLRGGSLKPIPELLNGGGFA 120
Query: 117 DAPLTATDEASE----------AEDGGADAMWKIREPNFGLASSRSSNKVCSGSKRRRSE 166
P +DEASE A+D G ++ F + SRS + ++R S
Sbjct: 121 GFPSPTSDEASEICTEMLNLRKADDSGDRNIY--HHCRFSSSRSRSRSTASPPKRKRLSS 178
Query: 167 EFVKVPAAINLDLRLTPIFQQKAVEERRHGSPSMTSEESVTTTACLETGIGDRW------ 220
E + P++ LDL L PI+ K + + +PSM SEESVTT + GDR+
Sbjct: 179 E--QQPSS-ELDLSLIPIYPIKTL-PFKEDTPSMYSEESVTTVSFQNNNAGDRYVRCGGG 234
Query: 221 SHGGDRKVLNLF 232
G K+LNLF
Sbjct: 235 GGGATTKLLNLF 246
>At5g67420 unknown protein
Length = 250
Score = 245 bits (625), Expect = 2e-65
Identities = 139/252 (55%), Positives = 167/252 (66%), Gaps = 21/252 (8%)
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
MSCNGCRVLRKGCSE+CILRPC+QWIET +AQGHATVFVAKFFGRAGLMSFIS VP+ QR
Sbjct: 1 MSCNGCRVLRKGCSENCILRPCIQWIETADAQGHATVFVAKFFGRAGLMSFISAVPDSQR 60
Query: 61 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTLRPLPELT-----L 115
PALFQSLL+EACGRTVNPVNGA+G+LWTGNW++CQAAVETVLRGG+LRP+PEL
Sbjct: 61 PALFQSLLYEACGRTVNPVNGAIGMLWTGNWNICQAAVETVLRGGSLRPIPELLTHGGGF 120
Query: 116 LDAPLTATDEASE-------AEDGGADAMWKIREPNFGLASSRSSNKVCSGSKRRRSEEF 168
P ++EASE + + F + SRS+ S +KR+R
Sbjct: 121 AGFPSPTSEEASEICTEMLNLQQNDSTDRNIYHHSRFSSSRSRSTMDSSSPTKRKRLSS- 179
Query: 169 VKVPAAINLDLRLTPIFQQK----AVEERRHGSPSMTSEES-VTTTACLETGIGDRWSHG 223
+ + LDL L P F K + RR +PSM SE+S TTT GD + +G
Sbjct: 180 -EDQPSSELDLSLIPNFPIKQATPSSTRRRSVTPSMNSEDSGTTTTTTAFCDKGDVYGNG 238
Query: 224 GDR--KVLNLFI 233
G K+LNLF+
Sbjct: 239 GGETTKLLNLFV 250
>At4g37540 unknown protein
Length = 240
Score = 241 bits (616), Expect = 2e-64
Identities = 143/249 (57%), Positives = 174/249 (69%), Gaps = 25/249 (10%)
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
MSCNGCRVLRKGCSE+CILRPCLQWIE+ E+QGHATVFVAKFFGRAGLMSFIS+VPE QR
Sbjct: 1 MSCNGCRVLRKGCSETCILRPCLQWIESAESQGHATVFVAKFFGRAGLMSFISSVPELQR 60
Query: 61 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTLRPLPELTLLDAPL 120
PALFQSLLFEACGRTVNPVNGAVG+LWT NWHVCQAAVETVLRGGTLRP+ +L + +
Sbjct: 61 PALFQSLLFEACGRTVNPVNGAVGMLWTRNWHVCQAAVETVLRGGTLRPISDLLESPSLM 120
Query: 121 TATDEASEA--EDGGADAMWKIREPNFGLASSRSSNKV-CSGSKRRRSEEFVKVPAAINL 177
+ DE+SE +D + R ++SRS+ ++ S R+R + + +N
Sbjct: 121 ISCDESSEIWHQDVSRNQTHHCR-----FSTSRSTTEMKDSLVNRKRLKSDSDLDLQVNH 175
Query: 178 DLRLT----------PIFQQKAVEERRHGSPSMTSEESVTTTACLETGI-GD--RWSHGG 224
L LT P K V+ R GSP SEESV TT+C E G+ GD + + G
Sbjct: 176 GLTLTAPAVPVPFLPPSSFCKVVKGDRPGSP---SEESV-TTSCWENGMRGDNKQKRNKG 231
Query: 225 DRKVLNLFI 233
++K+LNLF+
Sbjct: 232 EKKLLNLFV 240
>At1g67100 putative protein
Length = 276
Score = 173 bits (439), Expect = 6e-44
Identities = 74/110 (67%), Positives = 96/110 (87%)
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
MSCNGCRVLRKGCSE+C +RPCLQWI++ E+Q +ATVF+AKF+GRAGLM+ ++ P+ R
Sbjct: 46 MSCNGCRVLRKGCSENCSIRPCLQWIKSAESQANATVFLAKFYGRAGLMNLLNTGPDHLR 105
Query: 61 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTLRPL 110
PA+F+SLL+EACGR VNP+ G+VGLLW+GNWH+CQAAVE V+RG + P+
Sbjct: 106 PAIFRSLLYEACGRIVNPIYGSVGLLWSGNWHLCQAAVEAVMRGSPVTPI 155
>At3g02550 unknown protein
Length = 263
Score = 164 bits (415), Expect = 4e-41
Identities = 84/176 (47%), Positives = 114/176 (64%), Gaps = 18/176 (10%)
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
MSCNGCRVLRKGCSE C +RPCL WI++PEAQ +ATVF+AKF+GRAGLM+ I+ P R
Sbjct: 3 MSCNGCRVLRKGCSEDCSIRPCLAWIKSPEAQANATVFLAKFYGRAGLMNLINAGPNHLR 62
Query: 61 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTLRPL---------- 110
P +F+SLL EACGR VNP+ G+VGLLW+GNW +CQ AVE V++G ++ +
Sbjct: 63 PGIFRSLLHEACGRIVNPIYGSVGLLWSGNWQLCQDAVEAVMKGEPVKEIATDAATIGQG 122
Query: 111 PELTLLDAPLTATDEASEAEDGGADAMWKIREPNFGLASSRSSNKVCSGSKRRRSE 166
P L + D + D+ S A G+ + LA +R + +V + + + SE
Sbjct: 123 PPLKIYDIRHISKDDNSAAAATGS--------TDLKLAKTRRAKRVSTVAIQAESE 170
>At1g68510 unknown protein
Length = 233
Score = 158 bits (399), Expect = 3e-39
Identities = 70/113 (61%), Positives = 94/113 (82%), Gaps = 2/113 (1%)
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
+SCNGCRVLRKGC++ C +RPCLQWI++ ++Q +AT+F+AKF+GRAGL++ I + P+ R
Sbjct: 3 ISCNGCRVLRKGCNQDCTIRPCLQWIKSADSQANATLFLAKFYGRAGLLNLIESGPDHLR 62
Query: 61 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRG--GTLRPLP 111
PA+F+SLL+EACGR VNPV+G+VGL+W+GNW CQAAV+ VL G T PLP
Sbjct: 63 PAIFRSLLYEACGRIVNPVDGSVGLMWSGNWAQCQAAVDAVLNGLPITHTPLP 115
>At2g23660 putative protein
Length = 311
Score = 53.9 bits (128), Expect = 7e-08
Identities = 25/84 (29%), Positives = 43/84 (50%), Gaps = 5/84 (5%)
Query: 3 CNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQRPA 62
C C++LR+ C++ C+ P P +FV + FG + + ++++P QR
Sbjct: 6 CAACKLLRRKCTQECVFAPYF-----PPTNPQKFIFVHRVFGASNVTKILNDLPPDQRED 60
Query: 63 LFQSLLFEACGRTVNPVNGAVGLL 86
SL +EA R +P+ G VGL+
Sbjct: 61 TVNSLFYEAEARIRDPIYGCVGLI 84
>At1g07900 ASYMMETRIC LEAVES2-like protein 8 (ASL8)
Length = 190
Score = 53.5 bits (127), Expect = 9e-08
Identities = 25/85 (29%), Positives = 42/85 (49%), Gaps = 5/85 (5%)
Query: 3 CNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQRPA 62
C C++LR+ C+E C+L P P + FG + ++ F+ +PE QR
Sbjct: 34 CAACKILRRRCAERCVLAPYF-----PPTDPAKFTIAHRVFGASNIIKFLQELPESQRTD 88
Query: 63 LFQSLLFEACGRTVNPVNGAVGLLW 87
S+++EA R +PV G G ++
Sbjct: 89 AVNSMVYEAEARIRDPVYGCAGAIY 113
>At2g30130 unknown protein
Length = 193
Score = 53.1 bits (126), Expect = 1e-07
Identities = 28/84 (33%), Positives = 40/84 (47%), Gaps = 5/84 (5%)
Query: 3 CNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQRPA 62
C C++LR+ C++ CI P P H V K FG + + + +P QR
Sbjct: 9 CASCKLLRRRCAKDCIFAPYF-----PPDDPHKFAIVHKVFGASNVSKMLQELPVHQRAD 63
Query: 63 LFQSLLFEACGRTVNPVNGAVGLL 86
SL+FEA R +PV G VG +
Sbjct: 64 AVNSLVFEANARVRDPVYGCVGAI 87
>At2g28500 putative protein
Length = 229
Score = 53.1 bits (126), Expect = 1e-07
Identities = 24/85 (28%), Positives = 42/85 (49%), Gaps = 5/85 (5%)
Query: 3 CNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQRPA 62
C C++LR+ C++ C+L P P + FG + ++ F+ +PE QR
Sbjct: 53 CAACKILRRRCADKCVLAPYF-----PPTDPAKFTIAHRVFGASNIIKFLQELPESQRTD 107
Query: 63 LFQSLLFEACGRTVNPVNGAVGLLW 87
S+++EA R +PV G G ++
Sbjct: 108 AVNSMVYEAGARMRDPVYGCAGAIY 132
>At5g66870 unknown protein
Length = 313
Score = 51.2 bits (121), Expect = 5e-07
Identities = 27/84 (32%), Positives = 40/84 (47%), Gaps = 5/84 (5%)
Query: 3 CNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQRPA 62
C C+ LR+ C++ C+ P P Q FV K FG + + ++ + QR
Sbjct: 8 CAACKFLRRKCTQECVFAPYF-----PPDQPQKFAFVHKVFGASNVAKLLNELASNQRED 62
Query: 63 LFQSLLFEACGRTVNPVNGAVGLL 86
SL +EA R +PV G VGL+
Sbjct: 63 AVNSLFYEAEARLRDPVYGCVGLI 86
>At4g00210 putative protein
Length = 215
Score = 49.7 bits (117), Expect = 1e-06
Identities = 44/172 (25%), Positives = 75/172 (43%), Gaps = 15/172 (8%)
Query: 3 CNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQRPA 62
C C+ LR+ C C+ P + ++ E H T V K FG + + +P +R
Sbjct: 12 CGACKFLRRKCVADCVFAP---YFDSVEGTSHFTA-VHKVFGASNASKLLMMIPASRRLD 67
Query: 63 LFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTLRPLPELTLLDAPLTA 122
+L +EA R +PV G VG ++ A V+T L TL+ LP P +
Sbjct: 68 AVVTLTYEALARLRDPVYGCVGHIFALQHQAELAYVQTQL--STLQGLP------PPNSQ 119
Query: 123 TDEASEAEDGGADAMWKIREPNFGLASSRSSNKVCSGSKRRRSE--EFVKVP 172
+ +EA + + ++SS SS+ C ++ + + E ++VP
Sbjct: 120 NNSRTEAASSSNVPLISSVDSKDNMSSS-SSHIPCMSQQQEQEQPKEAIEVP 170
>At4g00220 LOB DOMAIN 30 (LBD30)
Length = 228
Score = 49.3 bits (116), Expect = 2e-06
Identities = 26/81 (32%), Positives = 40/81 (49%), Gaps = 4/81 (4%)
Query: 3 CNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQRPA 62
C C+ LR+ C CI P + ++ + H V K FG + + + +VPE +RP
Sbjct: 18 CGACKFLRRKCVAGCIFAP---YFDSEQGAAHFAA-VHKVFGASNVSKLLHHVPEHKRPD 73
Query: 63 LFQSLLFEACGRTVNPVNGAV 83
S+ FEA R +P+ G V
Sbjct: 74 AVVSICFEAQARLRDPIYGCV 94
>At3g11090 putative protein
Length = 165
Score = 48.1 bits (113), Expect = 4e-06
Identities = 27/85 (31%), Positives = 43/85 (49%), Gaps = 5/85 (5%)
Query: 2 SCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQRP 61
SC C++L++ C+ +CI P + + + A V K FG + + + VPE QR
Sbjct: 11 SCAACKLLKRRCTPTCIFAP---YFRSSDLITFAKVH--KVFGASNVSKLLGEVPEEQRQ 65
Query: 62 ALFQSLLFEACGRTVNPVNGAVGLL 86
SL +EA R +PV G +G +
Sbjct: 66 ETVNSLAYEAEVRLKDPVYGCIGAI 90
>At2g40470 unknown protein
Length = 224
Score = 47.8 bits (112), Expect = 5e-06
Identities = 27/84 (32%), Positives = 40/84 (47%), Gaps = 5/84 (5%)
Query: 3 CNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQRPA 62
C C++LR+ C++ C P E H V K FG + + + VPE QR
Sbjct: 46 CAACKLLRRRCAQECPFSPYFSPHEP-----HKFASVHKVFGASNVSKMLMEVPESQRAD 100
Query: 63 LFQSLLFEACGRTVNPVNGAVGLL 86
SL++EA R +PV G +G +
Sbjct: 101 AANSLVYEANVRLRDPVYGCMGAI 124
>At2g31310 putative protein
Length = 188
Score = 47.0 bits (110), Expect = 9e-06
Identities = 26/100 (26%), Positives = 45/100 (45%), Gaps = 4/100 (4%)
Query: 3 CNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQRPA 62
C GC+ LR+ C E C+ P + E + K FG + IS++P+ R
Sbjct: 8 CGGCKFLRRKCVEGCVFAPYFCY----EEGSSNFAAIHKVFGASNFSKLISHLPDHDRCD 63
Query: 63 LFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVL 102
+++ +EA R +P+ G V +++ V + VL
Sbjct: 64 AVRTISYEAHSRLHDPIYGCVSQIFSLQQQVVSLQAQVVL 103
>At1g31320 putative protein
Length = 172
Score = 47.0 bits (110), Expect = 9e-06
Identities = 25/84 (29%), Positives = 42/84 (49%), Gaps = 5/84 (5%)
Query: 3 CNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQRPA 62
C C++LR+ C++ C+ P + E Q A V + FG + + + +P QR
Sbjct: 14 CAACKLLRRRCAQDCVFSP---YFPADEPQKFANVH--RVFGASNVNKMLQELPIHQRGD 68
Query: 63 LFQSLLFEACGRTVNPVNGAVGLL 86
S+++EA R +PV G VG +
Sbjct: 69 AVSSMVYEANARVRDPVYGCVGAI 92
>At1g06280 putative protein
Length = 206
Score = 47.0 bits (110), Expect = 9e-06
Identities = 29/83 (34%), Positives = 39/83 (46%), Gaps = 5/83 (6%)
Query: 2 SCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQRP 61
+C C+ RK C+ CIL P +T E Q V K FG + + + V E R
Sbjct: 24 ACASCKHQRKKCNNECILSPYFPARKTKEFQA-----VHKVFGVSNVQKMVRTVREEDRT 78
Query: 62 ALFQSLLFEACGRTVNPVNGAVG 84
L SL +EA R +PV G+ G
Sbjct: 79 KLSDSLTWEALWRQKDPVLGSYG 101
>At1g16530 putative protein
Length = 165
Score = 46.6 bits (109), Expect = 1e-05
Identities = 25/85 (29%), Positives = 41/85 (47%), Gaps = 6/85 (7%)
Query: 3 CNGCRVLRKGC-SESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQRP 61
C GC++LR+ C +SC+ P P + + V K FG + + + + E R
Sbjct: 15 CAGCKLLRRKCVKDSCVFAPYF-----PAKEPYKFAIVHKIFGASNVNKMLQELSENHRS 69
Query: 62 ALFQSLLFEACGRTVNPVNGAVGLL 86
S+++EA R +PV G VG +
Sbjct: 70 DAVDSMVYEANARIQDPVYGCVGTI 94
>At5g63090 LOBd (LOB)
Length = 186
Score = 45.8 bits (107), Expect = 2e-05
Identities = 26/84 (30%), Positives = 37/84 (43%), Gaps = 5/84 (5%)
Query: 3 CNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQRPA 62
C C+ LR+ C CI P P + H V K FG + + ++ + QR
Sbjct: 12 CAACKFLRRKCMPGCIFAPYF-----PPEEPHKFANVHKIFGASNVTKLLNELLPHQRED 66
Query: 63 LFQSLLFEACGRTVNPVNGAVGLL 86
SL +EA R +PV G VG +
Sbjct: 67 AVNSLAYEAEARVRDPVYGCVGAI 90
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.134 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,461,760
Number of Sequences: 26719
Number of extensions: 220335
Number of successful extensions: 569
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 36
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 500
Number of HSP's gapped (non-prelim): 46
length of query: 233
length of database: 11,318,596
effective HSP length: 96
effective length of query: 137
effective length of database: 8,753,572
effective search space: 1199239364
effective search space used: 1199239364
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC137703.1


