

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137670.5 + phase: 0
(409 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
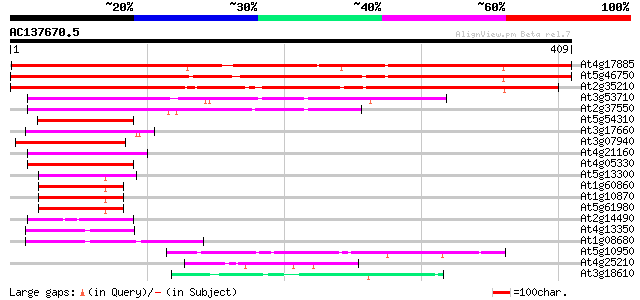
Score E
Sequences producing significant alignments: (bits) Value
At4g17885 unknown protein 500 e-142
At5g46750 zinc finger protein Glo3-like 474 e-134
At2g35210 unknown protein 448 e-126
At3g53710 ARF GAP-like zinc finger-containing protein ZIGA2 123 2e-28
At2g37550 Asp1 (asp1) 121 8e-28
At5g54310 unknown protein 91 1e-18
At3g17660 unknown protein 89 4e-18
At3g07940 putative GTPase activating protein 84 2e-16
At4g21160 unknown protein 83 2e-16
At4g05330 unknown protein 80 2e-15
At5g13300 putative protein 72 7e-13
At1g60860 GCN4-complementing like protein 71 9e-13
At1g10870 BRCA1-associated RING domain protein isolog 71 9e-13
At5g61980 GCN4-complementing protein - like 71 1e-12
At2g14490 unknown protein 55 5e-08
At4g13350 unknown protein 53 3e-07
At1g08680 unknown protein 53 3e-07
At5g10950 putative protein 45 6e-05
At4g25210 unknown protein 42 5e-04
At3g18610 unknown protein 42 8e-04
>At4g17885 unknown protein
Length = 413
Score = 500 bits (1287), Expect = e-142
Identities = 263/418 (62%), Positives = 327/418 (77%), Gaps = 19/418 (4%)
Query: 2 ASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISF 61
++D+ TDKN VFRKLK+KSENK CFDC+AKNPTWASVTYGIFLCIDCSA HR+LGVHISF
Sbjct: 5 SADNLTDKNIVFRKLKSKSENKVCFDCSAKNPTWASVTYGIFLCIDCSATHRNLGVHISF 64
Query: 62 VRSTNLDSWTPEQLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQLLSKEV 121
VRSTNLDSW+PEQL+ M FGGN+RAQVFF+QHGW GK+EAKYTSRAA+LY+Q+L+KEV
Sbjct: 65 VRSTNLDSWSPEQLRTMMFGGNNRAQVFFKQHGWTDGGKIEAKYTSRAADLYRQILAKEV 124
Query: 122 AKSMSEE--AALSAPPAASSQSAQGTNGLPDVKTNEVPIEKTVEKTVEKPEKTESSSSPR 179
AK+++EE + L + P A+SQ + +NG+ E E + K E T ++SSP+
Sbjct: 125 AKAIAEETNSGLLSSPVATSQLPEVSNGVSSYSVKE-------ELPLSKHEATSATSSPK 177
Query: 180 A-YTAVSNNLKKPIGAKKTGKSGGLGARKLTRKPSESLYEQKPEELPAPVSSSTITKNNL 238
A T V + KKPIGAK+TGK+GGLGARKLT KP ++LYEQKPEE+ APV + + NN
Sbjct: 178 ASNTVVPSTFKKPIGAKRTGKTGGLGARKLTTKPKDNLYEQKPEEV-APVIPAVSSTNNG 236
Query: 239 PS----GPPLTSRFEYTEDVQSSELNSGGSNVTGHVSVPKSSSSFFSDFGMDSGFQKKSG 294
S G SRFEY +D+QS + GG+ V HV+ PK SSSFFSDFGMDS F KKS
Sbjct: 237 ESKSSAGSSFASRFEYNDDLQSGGQSVGGTQVLNHVAPPK-SSSFFSDFGMDSSFPKKSS 295
Query: 295 PSSSKVQIQESDEARKKFSNAKSISSSQFFGDQNK-ANADAQATLSKFSGSSAISSADLF 353
+SSK Q++ESDEARKKF+NAKSISS+Q+FGDQNK A+ +++ATL KF+GS++ISSAD +
Sbjct: 296 SNSSKSQVEESDEARKKFTNAKSISSAQYFGDQNKNADLESKATLQKFAGSASISSADFY 355
Query: 354 GDSSD--NVDLAASDLINRISFQAQQDISSLKNIAGETGKKLTSLASSLMTDLQDRIL 409
G D N+D+ ASDLINR+SFQAQQD+SSL NIAGET KKL +LAS + +D+QDR+L
Sbjct: 356 GHDQDDSNIDITASDLINRLSFQAQQDLSSLVNIAGETKKKLGTLASGIFSDIQDRML 413
>At5g46750 zinc finger protein Glo3-like
Length = 402
Score = 474 bits (1219), Expect = e-134
Identities = 249/412 (60%), Positives = 314/412 (75%), Gaps = 13/412 (3%)
Query: 1 MASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 60
MA+++ TDKN VFRKLK+KSENK CFDC+AKNPTWASV YGIFLCIDCSAVHRSLGVHIS
Sbjct: 1 MATENLTDKNVVFRKLKSKSENKVCFDCSAKNPTWASVPYGIFLCIDCSAVHRSLGVHIS 60
Query: 61 FVRSTNLDSWTPEQLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQLLSKE 120
FVRSTNLDSW+PEQL+ M FGGN+RAQVFF+QHGWN GK+EAKYTSRAA++Y+Q L+KE
Sbjct: 61 FVRSTNLDSWSPEQLRTMMFGGNNRAQVFFKQHGWNDGGKIEAKYTSRAADMYRQTLAKE 120
Query: 121 VAKSMSEEAALSAPPAASSQSAQGTNGLPDVKTNEVPIEKTVEKTVEKPEKTESSSSPRA 180
VAK+M+EE L P +S ++Q + T+E P E ++ K E SS +
Sbjct: 121 VAKAMAEETVL--PSLSSVATSQPVESSENGFTSESPKESSL-----KQEAAVVSSPKAS 173
Query: 181 YTAVSNNLKKPIGAKKTGKSGGLGARKLTRKPSESLYEQKPEELPAPVSSSTITKNNLPS 240
V++ KKP+ ++K+GK+GGLGARKLT K ++LYEQKPEE + +++ T + +
Sbjct: 174 QKVVASTFKKPLVSRKSGKTGGLGARKLTTKSKDNLYEQKPEEPVPVIPAASPTNDTSAA 233
Query: 241 GPPLTSRFEYTEDVQSSELNSGGSNVTGHVSVPKSSSSFFSDFGMDSGFQKKSGPSSSKV 300
G SRFEY +D QS G+ V HV+ PK SS+FF++FGMDS F KKS SSSK
Sbjct: 234 GSSFASRFEYFDDEQSG--GQSGTRVLSHVAPPK-SSNFFNEFGMDSAFPKKSSSSSSKA 290
Query: 301 QIQESDEARKKFSNAKSISSSQFFGDQNK-ANADAQATLSKFSGSSAISSADLFGDSSD- 358
Q++E+DEARKKFSNAKSISS+QFFG+QN+ A+ D++ATL KFSGS+AISS+DLFG D
Sbjct: 291 QVEETDEARKKFSNAKSISSAQFFGNQNRDADLDSKATLQKFSGSAAISSSDLFGHGPDD 350
Query: 359 -NVDLAASDLINRISFQAQQDISSLKNIAGETGKKLTSLASSLMTDLQDRIL 409
N+D+ ASDLINRISFQAQQD+SS+ N+A ET KL + ASS+ +DLQDR+L
Sbjct: 351 SNIDITASDLINRISFQAQQDMSSIANLAEETKNKLGTFASSIFSDLQDRML 402
>At2g35210 unknown protein
Length = 395
Score = 448 bits (1153), Expect = e-126
Identities = 249/404 (61%), Positives = 305/404 (74%), Gaps = 15/404 (3%)
Query: 1 MASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 60
MAS++ DK +VF+KLK KS+NK CFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS
Sbjct: 1 MASENLNDKISVFKKLKAKSDNKICFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 60
Query: 61 FVRSTNLDSWTPEQLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQLLSKE 120
FVRSTNLDSW+ EQLKMM +GGN+RAQVFF+Q+GW+ GK EAKYTSRAA+LYKQ+L+KE
Sbjct: 61 FVRSTNLDSWSSEQLKMMIYGGNNRAQVFFKQYGWSDGGKTEAKYTSRAADLYKQILAKE 120
Query: 121 VAKSMSEEAALSAPPAASSQSAQGTNGLPDVKTNEVPIEKTVEKTVEKPEKTESSSSPRA 180
VAKS +EE L PP + S Q NGL +KT+E E K EKP+ SPR
Sbjct: 121 VAKSKAEE-ELDLPP-SPPDSTQVPNGLSSIKTSEALKESNTLKQQEKPDVV--PVSPR- 175
Query: 181 YTAVSNNLKKPIGAKKTGKSGGLGARKLTRKPSESLYEQKPEELPAPVSSSTITKNNLPS 240
+S ++KKP+GAKKTGK+GGLGARKLT K S +LY+QKPEE ++S ++ + S
Sbjct: 176 ---ISRSVKKPLGAKKTGKTGGLGARKLTTKSSGTLYDQKPEESVIIQATSPVSAKSARS 232
Query: 241 GPPLTSRFEYTEDVQSSELNSGGSNVTGHVSVPKSSSSFFSDFGMDSG-FQKKSGPSSSK 299
+SRF+Y ++VQ+ E + V HV+ PKSS F + M+ G FQKK SSSK
Sbjct: 233 S--FSSRFDYADNVQNRE-DYMSPQVVSHVAPPKSSGFFEEELEMNGGRFQKKPITSSSK 289
Query: 300 VQIQESDEARKKFSNAKSISSSQFFG-DQNKANADAQATLSKFSGSSAISSADLFGDSSD 358
+QIQE+DEARKKF+NAKSISS+Q+FG D N A+ +A+++L KFSGSSAISSADLFGD
Sbjct: 290 LQIQETDEARKKFTNAKSISSAQYFGNDNNSADLEAKSSLKKFSGSSAISSADLFGDGDG 349
Query: 359 N--VDLAASDLINRISFQAQQDISSLKNIAGETGKKLTSLASSL 400
+ +DL A DL+NR+S QAQQDISSLKN+A ET KKL S+ASSL
Sbjct: 350 DFPLDLTAGDLLNRLSLQAQQDISSLKNMAEETKKKLGSVASSL 393
>At3g53710 ARF GAP-like zinc finger-containing protein ZIGA2
Length = 459
Score = 123 bits (308), Expect = 2e-28
Identities = 89/323 (27%), Positives = 147/323 (44%), Gaps = 27/323 (8%)
Query: 14 RKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWTPE 73
R L+++ ENK C DC KNP WASV+YGIF+C++CS HR LGVHISFVRS +DSW+
Sbjct: 8 RTLQSQPENKVCVDCAQKNPQWASVSYGIFMCLECSGKHRGLGVHISFVRSVTMDSWSAI 67
Query: 74 QLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQLLSKEVAKSMSEEAALSA 133
Q+K M GGN R FF Q+G + + +KY S AA +Y+ + ++++E +
Sbjct: 68 QIKKMEAGGNERLNKFFAQYGIAKETDIISKYNSNAASVYRDRI-----QALAEGRPWND 122
Query: 134 PPAASSQS-----AQG--------TNGLPDVKTNEVPIEKTVEKTVEKPEKTESSSSPRA 180
PP + AQG NG D N+ + + + + S + A
Sbjct: 123 PPVVKEANKKPPLAQGGYGNNNNNNNGGWDSWDNDDSYKSDMRRNQSANDFRASGNREGA 182
Query: 181 YTAVSNNLKKPIGAKKTGKSGGLGARK-LTRKPSESLYEQKPEELPAPVSSSTITKNNLP 239
+ V + + I + ++ G R+ +E+ E KPE LP + +
Sbjct: 183 H--VKSKSSEDIYTRSQLEASAAGKESFFARRMAEN--ESKPEGLPPSQGGKYVGFGSSS 238
Query: 240 SGPPLTSRFEYTEDVQSSELNS----GGSNVTGHVSVPKSSSSFFSDFGMDSGFQKKSGP 295
+ PP ++ + V S S SV ++ + F+ + G+ K
Sbjct: 239 APPPRNNQQDDVFSVVSQGFGRLSLVAASAAQSAASVVQTGTKEFTSKVKEGGYDHKVSE 298
Query: 296 SSSKVQIQESDEARKKFSNAKSI 318
+ + V + ++ + + K +
Sbjct: 299 TVNVVANKTTEIGHRTWGIMKGV 321
>At2g37550 Asp1 (asp1)
Length = 456
Score = 121 bits (303), Expect = 8e-28
Identities = 81/254 (31%), Positives = 120/254 (46%), Gaps = 14/254 (5%)
Query: 14 RKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWTPE 73
R L+++ ENK C DC+ KNP WAS++YGIF+C++CS HR LGVHISFVRS +DSW+
Sbjct: 8 RTLQSQPENKVCVDCSQKNPQWASISYGIFMCLECSGKHRGLGVHISFVRSVTMDSWSEI 67
Query: 74 QLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYK---QLLSKE--------VA 122
Q+K M GGN R F Q+G + + + +KY S AA +Y+ Q L++ V
Sbjct: 68 QIKKMDAGGNERLNNFLAQYGISKETDIISKYNSNAASVYRDRIQALAEGRQWRDPPIVK 127
Query: 123 KSMSEEAALSAPPAASSQSAQGTNGLPDVKTNEVPIEKTVEKTVEKPEKTESSSSPRAYT 182
+S+ PP + NG D N+ T + + SS R
Sbjct: 128 ESVGGGLMNKKPPLSQGGGRDSGNGGWDNWDNDDSFRSTDMRRNQSAGDFRSSGG-RGAP 186
Query: 183 AVSNNLKKPIGAKKTGKSGGLGARKLTRKPSESLYEQKPEELPAPVSSSTITKNNLPSGP 242
A S + + + S ++ +E+ E KPE LP + + P
Sbjct: 187 AKSKSSEDIYSRSQLEASAANKESFFAKRMAEN--ESKPEGLPPSQGGKYVGFGSSPGPA 244
Query: 243 PLTSRFEYTEDVQS 256
P +++ DV S
Sbjct: 245 PRSNQQSGGGDVFS 258
>At5g54310 unknown protein
Length = 483
Score = 90.5 bits (223), Expect = 1e-18
Identities = 38/70 (54%), Positives = 48/70 (68%)
Query: 21 ENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWTPEQLKMMSF 80
EN+ C DC K P WASV GIF+C+ CS +HRSLGVHIS VRS LD+W PEQ+ +
Sbjct: 27 ENRECADCKTKGPRWASVNLGIFICMQCSGIHRSLGVHISKVRSATLDTWLPEQVAFIQS 86
Query: 81 GGNSRAQVFF 90
GN +A ++
Sbjct: 87 MGNDKANSYW 96
>At3g17660 unknown protein
Length = 117
Score = 89.0 bits (219), Expect = 4e-18
Identities = 43/100 (43%), Positives = 59/100 (59%), Gaps = 6/100 (6%)
Query: 12 VFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWT 71
+ L +N+ C DC +K P WASV GIF+C+ CS +HRSLGVHIS VRS LD+W
Sbjct: 18 ILEALLKHPDNRECADCRSKAPRWASVNLGIFICMQCSGIHRSLGVHISQVRSITLDTWL 77
Query: 72 PEQLKMMSFGGNSRAQVFFR----QH--GWNGDGKVEAKY 105
P+Q+ M GN++ ++ QH + D + AKY
Sbjct: 78 PDQVAFMKSTGNAKGNEYWESELPQHFERSSSDTFIRAKY 117
>At3g07940 putative GTPase activating protein
Length = 385
Score = 83.6 bits (205), Expect = 2e-16
Identities = 39/81 (48%), Positives = 51/81 (62%), Gaps = 1/81 (1%)
Query: 5 SFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRS 64
S +D KL + NK C DC + P W S++ G+F+CI CS VHRSLGVHIS V S
Sbjct: 42 SSSDPRDRLEKLLKQPGNKYCADCGSPEPKWVSLSLGVFICIKCSGVHRSLGVHISKVLS 101
Query: 65 TNLDSWTPEQLKMM-SFGGNS 84
LD WT +Q+ M+ +GGN+
Sbjct: 102 VKLDEWTDDQVDMLVGYGGNT 122
>At4g21160 unknown protein
Length = 337
Score = 83.2 bits (204), Expect = 2e-16
Identities = 39/88 (44%), Positives = 52/88 (58%), Gaps = 1/88 (1%)
Query: 14 RKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWTPE 73
R L T+S+N+ C DC A +P WAS G+F+C+ C VHRSLG HIS V S LD W+ E
Sbjct: 19 RDLLTQSDNRVCADCGAPDPKWASANIGVFICLKCCGVHRSLGSHISKVLSVTLDEWSDE 78
Query: 74 QL-KMMSFGGNSRAQVFFRQHGWNGDGK 100
++ M+ GGN+ A + G K
Sbjct: 79 EVDSMIEIGGNASANSIYEAFIPEGSSK 106
>At4g05330 unknown protein
Length = 336
Score = 80.1 bits (196), Expect = 2e-15
Identities = 35/78 (44%), Positives = 48/78 (60%), Gaps = 1/78 (1%)
Query: 14 RKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWTPE 73
R L + +N+ C DC A +P WAS G+F+C+ C VHRSLG HIS V S LD W+ E
Sbjct: 19 RDLLNQPDNRVCADCGASDPKWASANIGVFICLKCCGVHRSLGTHISKVLSVTLDEWSDE 78
Query: 74 QL-KMMSFGGNSRAQVFF 90
++ M+ GGN+ A +
Sbjct: 79 EVDSMIEIGGNASANSIY 96
>At5g13300 putative protein
Length = 750
Score = 71.6 bits (174), Expect = 7e-13
Identities = 33/73 (45%), Positives = 43/73 (58%), Gaps = 2/73 (2%)
Query: 22 NKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLD--SWTPEQLKMMS 79
N C DC A P WAS+ G+ +CI+CS VHR+LGVHIS VRS LD W P + +
Sbjct: 436 NDKCADCGAPEPDWASLNLGVLVCIECSGVHRNLGVHISKVRSLTLDVKVWEPSVISLFQ 495
Query: 80 FGGNSRAQVFFRQ 92
GN+ A + +
Sbjct: 496 ALGNTFANTVWEE 508
>At1g60860 GCN4-complementing like protein
Length = 776
Score = 71.2 bits (173), Expect = 9e-13
Identities = 33/64 (51%), Positives = 42/64 (65%), Gaps = 2/64 (3%)
Query: 22 NKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLD--SWTPEQLKMMS 79
N +C +CNA +P WAS+ G+ +CI+CS VHR+LGVHIS VRS LD W P L +
Sbjct: 479 NNTCAECNAPDPDWASLNLGVLMCIECSGVHRNLGVHISKVRSLTLDVKVWEPTILDLFR 538
Query: 80 FGGN 83
GN
Sbjct: 539 NLGN 542
>At1g10870 BRCA1-associated RING domain protein isolog
Length = 531
Score = 71.2 bits (173), Expect = 9e-13
Identities = 34/64 (53%), Positives = 41/64 (63%), Gaps = 2/64 (3%)
Query: 22 NKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLD--SWTPEQLKMMS 79
N +C +CNA P WAS+ G+ LCI CS VHR+LGVHIS VRS +LD W P L +
Sbjct: 235 NNACAECNAPEPDWASLNLGVLLCIQCSGVHRNLGVHISKVRSLSLDVKVWEPTILDLFR 294
Query: 80 FGGN 83
GN
Sbjct: 295 NLGN 298
>At5g61980 GCN4-complementing protein - like
Length = 850
Score = 70.9 bits (172), Expect = 1e-12
Identities = 32/64 (50%), Positives = 40/64 (62%), Gaps = 2/64 (3%)
Query: 22 NKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLD--SWTPEQLKMMS 79
N+ C DC A P WAS+ G+ +CI+CS +HR+LGVHIS VRS LD W P L +
Sbjct: 532 NERCADCGAPEPDWASLNLGVLICIECSGIHRNLGVHISKVRSLTLDVKVWEPSVLTLFQ 591
Query: 80 FGGN 83
GN
Sbjct: 592 SLGN 595
>At2g14490 unknown protein
Length = 119
Score = 55.5 bits (132), Expect = 5e-08
Identities = 32/78 (41%), Positives = 41/78 (52%), Gaps = 3/78 (3%)
Query: 14 RKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWTPE 73
R L + +N+ C DC A P WA I L C VHRSLG HI V S LD W+ E
Sbjct: 19 RDLLNQPDNRVCADCGASVPKWAKYQ-SIHLLKSCG-VHRSLGTHILKVLSVTLDEWSDE 76
Query: 74 QL-KMMSFGGNSRAQVFF 90
++ M+ GGN+ A +
Sbjct: 77 EVDSMIETGGNASANSIY 94
>At4g13350 unknown protein
Length = 602
Score = 52.8 bits (125), Expect = 3e-07
Identities = 24/81 (29%), Positives = 44/81 (53%), Gaps = 4/81 (4%)
Query: 12 VFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWT 71
+ R L ENK C +CN+ P + T+ F+C +CS +HR V+S ++ +T
Sbjct: 14 IIRSLLKLPENKRCINCNSLGPQYVCTTFWTFVCTNCSGIHREF---THRVKSISMAKFT 70
Query: 72 PEQLKMMSFGGNSRAQ-VFFR 91
+++ + GGN A+ ++F+
Sbjct: 71 SQEVTALKEGGNQHAKDIYFK 91
>At1g08680 unknown protein
Length = 159
Score = 52.8 bits (125), Expect = 3e-07
Identities = 30/130 (23%), Positives = 62/130 (47%), Gaps = 6/130 (4%)
Query: 12 VFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWT 71
+ R L N+ C +CN+ P + T+ F+C+ CS +HR V+S ++ +T
Sbjct: 14 IIRGLMKLPPNRRCINCNSLGPQYVCTTFWTFVCMACSGIHREF---THRVKSVSMSKFT 70
Query: 72 PEQLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQLLSKEVAKSMSEEAAL 131
++++++ GGN RA+ + + N D + + + AE ++ + + A
Sbjct: 71 SKEVEVLQNGGNQRAREIYLK---NWDHQRQRLPENSNAERVREFIKNVYVQKKYAGAND 127
Query: 132 SAPPAASSQS 141
+ P+ SQ+
Sbjct: 128 ADKPSKDSQA 137
>At5g10950 putative protein
Length = 395
Score = 45.4 bits (106), Expect = 6e-05
Identities = 60/259 (23%), Positives = 109/259 (41%), Gaps = 20/259 (7%)
Query: 115 QLLSKEVAKSMSEEAALSAPPAASSQSAQGTNGLPDVKTNEVPIEKTVEKTVEKPEKTES 174
+L+ ++ A+S S+E +L A +S++GT P K +T E +K +K +S
Sbjct: 87 ELIEEDDAESESDEISLQEESAG--ESSEGTPEPPKGKGKASLKGRTDEAMPKKKQKIDS 144
Query: 175 SSSPRAYTAVSNNLKKPIGAKKTGKSGGLGARKLTRKPSESLYEQKPEELPAPVSSSTIT 234
SS +A V KP +K GK G +G + + S + + +S
Sbjct: 145 SSKSKA-KEVEKKASKP-ETEKNGKMGDIGGKIVAAASRMSERFRSKGNVDQKETSKASK 202
Query: 235 KNNLPSGPPLTSRFEYTEDVQSSELNSGGSNVTGHVSVPK------SSSSFFSDFGMDSG 288
K + S LT R ++T+D E + + + PK S++ ++ D
Sbjct: 203 KPKMSS--KLTKR-KHTDDQDEDEEAGDDIDTSSEEAKPKVLKSCNSNADEVAENSSDED 259
Query: 289 FQKKSGPSSSKVQIQESDEARKKFSN------AKSISSSQFFGDQNKANADAQATLSKFS 342
K ++SK E +E + + K +S+ G+ N ++ + + T+SK +
Sbjct: 260 EPKVLKTNNSKADKDEDEEENETSDDEAEPKALKLSNSNSDNGENNSSDDEKEITISKIT 319
Query: 343 GSSAISSADLFGDSSDNVD 361
S I S ++ DN D
Sbjct: 320 -SKKIKSNTADEENGDNED 337
>At4g25210 unknown protein
Length = 368
Score = 42.4 bits (98), Expect = 5e-04
Identities = 38/148 (25%), Positives = 68/148 (45%), Gaps = 24/148 (16%)
Query: 128 EAALSAPPAASSQSAQGTNGLPDVKTNEVPIEKTVEKTVEKPE-------KTESSSSPRA 180
E + +PP +S + G++G + EVP K VE + +KPE ++ESSS P
Sbjct: 7 EEVVESPPVSSEEEESGSSGEESESSAEVP--KKVESS-QKPESDSEGESESESSSGPEP 63
Query: 181 YTAVSNNLK-KPIGAKKTGKSGGLGA---------RKLTRKPSESLYEQK---PEELPAP 227
+ + +K KP+G K ++ G A R L E++ +QK E + P
Sbjct: 64 ESEPAKTIKLKPVGTKPIPETSGSAATVPESSTAKRPLKEAAPEAIKKQKTSDTEHVKKP 123
Query: 228 VSSSTITKNNLPSGPPLTSR-FEYTEDV 254
+++ + K + + R F T+++
Sbjct: 124 ITNDEVKKISSEDAKKMFQRLFSETDEI 151
>At3g18610 unknown protein
Length = 636
Score = 41.6 bits (96), Expect = 8e-04
Identities = 49/201 (24%), Positives = 79/201 (38%), Gaps = 24/201 (11%)
Query: 119 KEVAKSMSEEAALSAPPAASSQSAQGTNGLPDVKTNEVPIEKTVEKTVEKPEKTESSSSP 178
+E + A L A SS S G++ ++E P E V K + ++ SSS
Sbjct: 191 EETVPMKKQTAVLEKAKAESSSSDDGSS------SDEEPTPAKKEPIVVKKDSSDESSSD 244
Query: 179 RAYTAVSNNLKKPIGAKKTGKSGGLGARKLTRKPSESLYEQKPEELPAPVSSSTITKNNL 238
V KKP K K+ + S S E ++ P P T+ KN
Sbjct: 245 EETPVVK---KKPTTVVKDAKA----------ESSSSEEESSSDDEPTPAKKPTVVKNAK 291
Query: 239 PSGPPLTSRFEYTEDVQSSELN--SGGSNVTGHVSVPKSSSSFFSD-FGMDSGFQKKSGP 295
P+ +S E +++ +S + + + V+ S +SSS SD + +K P
Sbjct: 292 PAAKDSSSSEEDSDEEESDDEKPPTKKAKVSSKTSKQESSSDESSDESDKEESKDEKVTP 351
Query: 296 SSSKVQIQESDEARKKFSNAK 316
++ D +K SNAK
Sbjct: 352 KKKDSDVEMVDAEQK--SNAK 370
Score = 29.6 bits (65), Expect = 3.1
Identities = 51/265 (19%), Positives = 95/265 (35%), Gaps = 13/265 (4%)
Query: 127 EEAALSAPPAASSQSAQGTNGLPDVKTNEVPIEKTVEKTVEKPEKTESSSSPRAYTAVSN 186
E+A + P S S+ ++ + KT E P + E + E E+ +SSS ++
Sbjct: 52 EKAEKTVPKKVESSSSDASDSDEEEKTKETPSKLKDESSSE--EEDDSSSDEE----IAP 105
Query: 187 NLKKPIGAKKTGKSGGLGARKLTRKPSESLYEQKP---EELPAPVSSSTITKNNLPSGPP 243
K+P KK T + +++P E+ SSS ++ P
Sbjct: 106 AKKRPEPIKKAKVESSSSDDDSTSDEETAPVKKQPAVLEKAKVESSSSDDDSSSDEETVP 165
Query: 244 LTSRFEYTEDVQSSELNSGGSNVTGHVSVPKSSSSFFSDFGMDSGFQKKSGPSSSKVQIQ 303
+ + E + +S + + +VP + + G SS +
Sbjct: 166 VKKQPAVLEKAKIESSSSDDDSSSDEETVPMKKQTAVLEKAKAESSSSDDGSSSD----E 221
Query: 304 ESDEARKKFSNAKSISSSQFFGDQNKANADAQATLSKFSGSSAISSADLFGDSSDNVDLA 363
E A+K+ K SS + D+ + T + SS++ S D A
Sbjct: 222 EPTPAKKEPIVVKKDSSDESSSDEETPVVKKKPTTVVKDAKAESSSSEEESSSDDEPTPA 281
Query: 364 ASDLINRISFQAQQDISSLKNIAGE 388
+ + + A +D SS + + E
Sbjct: 282 KKPTVVKNAKPAAKDSSSSEEDSDE 306
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.307 0.123 0.332
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,763,686
Number of Sequences: 26719
Number of extensions: 376032
Number of successful extensions: 1561
Number of sequences better than 10.0: 120
Number of HSP's better than 10.0 without gapping: 29
Number of HSP's successfully gapped in prelim test: 91
Number of HSP's that attempted gapping in prelim test: 1472
Number of HSP's gapped (non-prelim): 152
length of query: 409
length of database: 11,318,596
effective HSP length: 102
effective length of query: 307
effective length of database: 8,593,258
effective search space: 2638130206
effective search space used: 2638130206
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC137670.5


