

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137546.15 + phase: 0 /pseudo
(355 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
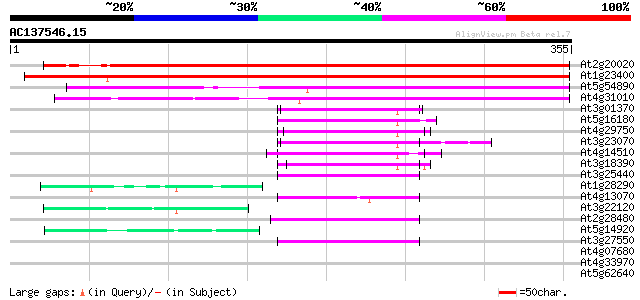
Score E
Sequences producing significant alignments: (bits) Value
At2g20020 CAF1-like protein 489 e-138
At1g23400 CAF2-like protein 410 e-115
At5g54890 putative protein 254 7e-68
At4g31010 unknown protein 240 7e-64
At3g01370 unknown protein 61 8e-10
At5g16180 putative protein 53 3e-07
At4g29750 putative protein 52 5e-07
At3g23070 hypothetical protein 52 5e-07
At4g14510 hypothetical protein 49 3e-06
At3g18390 unknown protein 49 3e-06
At3g25440 unknown protein 49 5e-06
At1g28290 proline-rich protein, putative 45 8e-05
At4g13070 putative protein 44 1e-04
At3g22120 unknown protein 43 2e-04
At2g28480 hypothetical protein 43 3e-04
At5g14920 unknown protein 42 4e-04
At3g27550 hypothetical protein 42 5e-04
At4g07680 hypothetical protein 40 0.001
At4g33970 extensin-like protein 39 0.006
At5g62640 unknown protein 38 0.007
>At2g20020 CAF1-like protein
Length = 701
Score = 489 bits (1258), Expect = e-138
Identities = 236/333 (70%), Positives = 271/333 (80%), Gaps = 15/333 (4%)
Query: 22 RPTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAPFE 81
+P PN+ +KP+V P S P L + PE VK+SEDG++YVI GAPFE
Sbjct: 108 KPDSGPNRPKNKPRV-PDSPPQL-------------DAKPE-VKLSEDGLTYVINGAPFE 152
Query: 82 FKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKK 141
FKYSYTETPK KP+++REP + PFGP TM RPWTGR PLP S+K +EFDSF LPP KK
Sbjct: 153 FKYSYTETPKVKPLKLREPAYAPFGPTTMGRPWTGRAPLPQSQKTPREFDSFRLPPVGKK 212
Query: 142 GVKPVQSPGPFLPGTSPRYVMSREEVLGEPLTKEEINELVRSTLKSSRQLNLGRDGFIHN 201
G+KPVQ PGPF PG PRYV S+EE+LGEPLTKEE+ ELV S LK++RQLN+GRDG HN
Sbjct: 213 GLKPVQKPGPFRPGVGPRYVYSKEEILGEPLTKEEVRELVTSCLKTTRQLNMGRDGLTHN 272
Query: 202 MLDNIHAHWKRRRVCKIKCIGVCTVDMDNVCQQLEEKTGGKVIYRRGGVIYLFRGRNYNH 261
ML+NIH WKRRRVCKIKC GVCTVDMDNVC+QLEEK GGKVIYRRGGV++LFRGRNYNH
Sbjct: 273 MLNNIHDLWKRRRVCKIKCKGVCTVDMDNVCEQLEEKIGGKVIYRRGGVLFLFRGRNYNH 332
Query: 262 KTRPRFPLMLWKPVPPVYPRLIQQVPEGLTLEEATEMRQKGRTLTPICKLGKNGVYYNLV 321
+TRPRFPLMLWKPV PVYPRLIQQVPEGLT +EAT MR+KGR L PICKLGKNGVY +LV
Sbjct: 333 RTRPRFPLMLWKPVAPVYPRLIQQVPEGLTRQEATNMRRKGRELMPICKLGKNGVYCDLV 392
Query: 322 NNVREAFEECELVRVNCQGLNKSDYRKIGAKLR 354
NV+EAFE CELVR++CQG+ SD+RKIGAKL+
Sbjct: 393 KNVKEAFEVCELVRIDCQGMKGSDFRKIGAKLK 425
Score = 28.5 bits (62), Expect = 5.8
Identities = 18/90 (20%), Positives = 42/90 (46%)
Query: 170 EPLTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMD 229
E LT++E + R + LG++G +++ N+ ++ + +I C G+ D
Sbjct: 359 EGLTRQEATNMRRKGRELMPICKLGKNGVYCDLVKNVKEAFEVCELVRIDCQGMKGSDFR 418
Query: 230 NVCQQLEEKTGGKVIYRRGGVIYLFRGRNY 259
+ +L++ ++ I ++RGR +
Sbjct: 419 KIGAKLKDLVPCVLVSFENEQILIWRGREW 448
>At1g23400 CAF2-like protein
Length = 564
Score = 410 bits (1054), Expect = e-115
Identities = 203/351 (57%), Positives = 252/351 (70%), Gaps = 6/351 (1%)
Query: 10 PISASNADQSSRRPTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKT------PEN 63
PI S+R K N PQS+PALK + + KPV +
Sbjct: 35 PIPIPKYAPSNRNRNQKTNHQTDTNPKKPQSNPALKLPHHRTRYYKPVKEGVISSDGDRT 94
Query: 64 VKISEDGVSYVIEGAPFEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPS 123
+ I + GVSY + GAPFEF++SY+ETPK KPV +REP F+PF P TMPRPWTG+ PL S
Sbjct: 95 ILIGDSGVSYQLPGAPFEFQFSYSETPKVKPVGIREPAFMPFAPPTMPRPWTGKAPLKKS 154
Query: 124 KKKLKEFDSFVLPPPHKKGVKPVQSPGPFLPGTSPRYVMSREEVLGEPLTKEEINELVRS 183
KKK+ FDSF PP K GVK V+ PGP G P+ M+REEVLGEPL + E L++
Sbjct: 155 KKKIPLFDSFNPPPAGKSGVKYVEMPGPLPFGRYPKEGMNREEVLGEPLKRWEKGMLIKP 214
Query: 184 TLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMDNVCQQLEEKTGGKV 243
+ +RQ+NLGRDGF HNML+ IH+HWKRRRVCK++C GV TVDM+NVC+ LEEKTGG++
Sbjct: 215 HMHDNRQVNLGRDGFTHNMLELIHSHWKRRRVCKVRCKGVPTVDMNNVCRVLEEKTGGEI 274
Query: 244 IYRRGGVIYLFRGRNYNHKTRPRFPLMLWKPVPPVYPRLIQQVPEGLTLEEATEMRQKGR 303
I+R GGV+YLFRGRNYN++TRP++PLMLWKP PVYP+LIQ+VPEGLT EEA E R KG+
Sbjct: 275 IHRVGGVVYLFRGRNYNYRTRPQYPLMLWKPAAPVYPKLIQEVPEGLTKEEAHEFRVKGK 334
Query: 304 TLTPICKLGKNGVYYNLVNNVREAFEECELVRVNCQGLNKSDYRKIGAKLR 354
+L PICKL KNGVY +LV +VR+AFE LV+V+C GL SDY+KIGAKL+
Sbjct: 335 SLRPICKLSKNGVYVSLVKDVRDAFELSSLVKVDCPGLEPSDYKKIGAKLK 385
>At5g54890 putative protein
Length = 358
Score = 254 bits (648), Expect = 7e-68
Identities = 138/323 (42%), Positives = 190/323 (58%), Gaps = 35/323 (10%)
Query: 37 DPQSHPALKFSNIPKQKLKP-VNKTPENVKISEDGVSYVIEGAPFEFKYSYTET-PKSKP 94
DP P K + PK+K K K ++ ++ D VI PF+F+YSY+ET P+ +P
Sbjct: 35 DPPFSPLSKPTKPPKEKKKQKTKKQDQSSELVNDLKIPVISDLPFDFRYSYSETNPEIEP 94
Query: 95 VQMREPP-FVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKGVKPVQSPGPFL 153
+ REP F PFGP + R WTG L E D
Sbjct: 95 IGFREPKRFSPFGPGRLDRKWTGTTALASP-----EIDQ--------------------- 128
Query: 154 PGTSPRYVMSREEVLGEPLTKEEINELVRSTLKS--SRQLNLGRDGFIHNMLDNIHAHWK 211
++V R VLGE LT++E+ EL+ S +RQ+NLG+ G HNM+D+IH HWK
Sbjct: 129 ----SQWVEERARVLGETLTEDEVTELIERYRHSDCTRQINLGKGGVTHNMIDDIHNHWK 184
Query: 212 RRRVCKIKCIGVCTVDMDNVCQQLEEKTGGKVIYRRGGVIYLFRGRNYNHKTRPRFPLML 271
+ +IKC+GV T+DMDN+C LEEK+GGK++YR ++ L+RGRNY+ K+RP PLML
Sbjct: 185 KAEAVRIKCLGVPTLDMDNICFHLEEKSGGKIVYRNINILVLYRGRNYDPKSRPIIPLML 244
Query: 272 WKPVPPVYPRLIQQVPEGLTLEEATEMRQKGRTLTPICKLGKNGVYYNLVNNVREAFEEC 331
WKP PP+YPRL++ V +GL EE EMR +G + KL +NGVY N+V VRE FE
Sbjct: 245 WKPHPPIYPRLVKNVADGLEFEETKEMRNRGLHSPALMKLTRNGVYVNVVGRVREEFETE 304
Query: 332 ELVRVNCQGLNKSDYRKIGAKLR 354
E+VR++C + SD ++IG KL+
Sbjct: 305 EIVRLDCTHVGMSDCKRIGVKLK 327
>At4g31010 unknown protein
Length = 405
Score = 240 bits (613), Expect = 7e-64
Identities = 134/329 (40%), Positives = 190/329 (57%), Gaps = 26/329 (7%)
Query: 29 KNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAPFEFKYSYTE 88
++P +P N K+K KP + P ++ +GV V PF+F++SYTE
Sbjct: 38 QSPPPLSSSASENPDFNQKNNNKKKPKPQYRPPSSL----EGVKTVHSDLPFDFRFSYTE 93
Query: 89 TPKS-KPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKGVKPVQ 147
+ + +P+ +REP + PFGP + R WTG P K++ D P +K K
Sbjct: 94 SCSNVRPIGLREPKYSPFGPDRLDREWTG-VCAPAVNPKVESVDGVEDPKLEEKRRKV-- 150
Query: 148 SPGPFLPGTSPRYVMSREEVLGEPLTKEEINELVR--STLKSSRQLNLGRDGFIHNMLDN 205
RE++ G LT+ E LV K+ RQ+NLGRDG HNML++
Sbjct: 151 ----------------REKIQGASLTEAERKFLVELCQRNKTKRQVNLGRDGLTHNMLND 194
Query: 206 IHAHWKRRRVCKIKCIGVCTVDMDNVCQQLEEKTGGKVIYRRGGVIYLFRGRNYNHKTRP 265
++ HWK ++KC+GV T+DM NV LE+KT G+V+ + G + L+RGRNY+ K RP
Sbjct: 195 VYNHWKHAEAVRVKCLGVPTLDMKNVIFHLEDKTFGQVVSKHSGTLVLYRGRNYDPKKRP 254
Query: 266 RFPLMLWKPVPPVYPRLIQQVPEGLTLEEATEMRQKGRTLTPICKLGKNGVYYNLVNNVR 325
+ PLMLWKP PVYPRLI+ +GL+++E MR+KG + + KL KNG Y +LV VR
Sbjct: 255 KIPLMLWKPHEPVYPRLIKTTIDGLSIDETKAMRKKGLAVPALTKLAKNGYYGSLVPMVR 314
Query: 326 EAFEECELVRVNCQGLNKSDYRKIGAKLR 354
+AF ELVR++C GL + DY+KIGAKLR
Sbjct: 315 DAFLVSELVRIDCLGLERKDYKKIGAKLR 343
>At3g01370 unknown protein
Length = 1020
Score = 61.2 bits (147), Expect = 8e-10
Identities = 30/90 (33%), Positives = 48/90 (53%)
Query: 172 LTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMDNV 231
L E+ L ++ +++L +G+ G +++ IH W+ V KI C + ++M
Sbjct: 166 LPPAELRRLRTVGIRLTKKLKIGKAGITEGIVNGIHERWRTTEVVKIFCEDISRMNMKRT 225
Query: 232 CQQLEEKTGGKVIYRRGGVIYLFRGRNYNH 261
LE KTGG VI+R G I L+RG NY +
Sbjct: 226 HDVLETKTGGLVIWRSGSKILLYRGVNYQY 255
Score = 51.6 bits (122), Expect = 6e-07
Identities = 31/94 (32%), Positives = 47/94 (49%), Gaps = 4/94 (4%)
Query: 170 EPLTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMD 229
E +T +E L + LK L LGR G ++N+H HWK R + KI C
Sbjct: 586 EGITNDEKYMLRKIGLKMKPFLLLGRRGVFDGTIENMHLHWKYRELVKIICNEYSIEAAH 645
Query: 230 NVCQQLEEKTGGKVI----YRRGGVIYLFRGRNY 259
V + LE ++GG ++ +G I ++RG+NY
Sbjct: 646 KVAEILEAESGGILVAVEMVSKGYAIIVYRGKNY 679
Score = 37.7 bits (86), Expect = 0.010
Identities = 22/82 (26%), Positives = 43/82 (51%), Gaps = 4/82 (4%)
Query: 271 LWKPVPPVYPRLIQQVPE--GLTLE--EATEMRQKGRTLTPICKLGKNGVYYNLVNNVRE 326
+WK + + ++VP LTL E +R G LT K+GK G+ +VN + E
Sbjct: 143 VWKKETEMERKKEEKVPSLAELTLPPAELRRLRTVGIRLTKKLKIGKAGITEGIVNGIHE 202
Query: 327 AFEECELVRVNCQGLNKSDYRK 348
+ E+V++ C+ +++ + ++
Sbjct: 203 RWRTTEVVKIFCEDISRMNMKR 224
Score = 35.4 bits (80), Expect = 0.048
Identities = 18/59 (30%), Positives = 32/59 (53%)
Query: 280 PRLIQQVPEGLTLEEATEMRQKGRTLTPICKLGKNGVYYNLVNNVREAFEECELVRVNC 338
P+L EG+T +E +R+ G + P LG+ GV+ + N+ ++ ELV++ C
Sbjct: 578 PQLSDIDKEGITNDEKYMLRKIGLKMKPFLLLGRRGVFDGTIENMHLHWKYRELVKIIC 636
Score = 33.1 bits (74), Expect = 0.24
Identities = 24/86 (27%), Positives = 44/86 (50%), Gaps = 5/86 (5%)
Query: 275 VPPVYPRLIQQVPEG----LTLEEATEMRQKGRTLTPICKLGKNGVYYNLVNNVREAFEE 330
V P Y R + +P G LT +E T +R+ GR L LG+N L + + +E+
Sbjct: 359 VVPDYRRPFRLLPYGVSPKLTDDEMTTIRRLGRPLPCHFALGRNRNLQGLAVAIVKLWEK 418
Query: 331 CELVRVNC-QGLNKSDYRKIGAKLRI 355
CEL ++ +G+ ++ + +L++
Sbjct: 419 CELAKIAVKRGVQNTNSELMAEELKV 444
>At5g16180 putative protein
Length = 718
Score = 52.8 bits (125), Expect = 3e-07
Identities = 35/105 (33%), Positives = 51/105 (48%), Gaps = 7/105 (6%)
Query: 170 EPLTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMD 229
E LT EE L R LK + L LGR G +++ +H HWK R V K+ + +
Sbjct: 568 EILTNEERECLRRIGLKMNSSLVLGRRGVFFGVMEGLHQHWKHREVAKVITMQKLFSRVV 627
Query: 230 NVCQQLEEKTGGKVI----YRRGGVIYLFRGRNYNHKTRPRFPLM 270
+ LE ++ G +I + G I ++RG+NY RP LM
Sbjct: 628 YTAKALETESNGVLISIEKLKEGHAILIYRGKNYK---RPSSKLM 669
Score = 32.3 bits (72), Expect = 0.40
Identities = 23/89 (25%), Positives = 40/89 (44%), Gaps = 1/89 (1%)
Query: 172 LTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCI-GVCTVDMDN 230
LT EE+ L LGR+ + + I W++ + KI G + +
Sbjct: 359 LTDEELTYLRNIAQPLPFHFVLGRNYGLQGLASAIVKLWEKCIIAKIAIKWGALNTNNEE 418
Query: 231 VCQQLEEKTGGKVIYRRGGVIYLFRGRNY 259
+ +L TGG +I R +I L+RG+++
Sbjct: 419 MADELRYLTGGVLILRNKYLIVLYRGKDF 447
Score = 32.0 bits (71), Expect = 0.53
Identities = 45/220 (20%), Positives = 86/220 (38%), Gaps = 44/220 (20%)
Query: 159 RYVMSREEVLGEPLTKEEI------NELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKR 212
R+++ R + P T E I N L R K + +N+ + G +++ I + WK
Sbjct: 188 RFILRRMKKESVPTTAELILDEGLLNRLRREASKMRKWVNVRKAGVTELVVNKIKSMWKL 247
Query: 213 RRVCKIKCIGVCTVDMDNVCQQLEEKTGGKVIYRRGGVIYLFRGR-NYNHKTRPRFPLML 271
+ ++ +M+ + +E TGG V+ + + ++RG +Y+ + + L
Sbjct: 248 NELAMVRFDVPLCRNMERAQEIIE--TGGLVVLSKKEFLVVYRGGPSYSSEGQDEISSSL 305
Query: 272 -------------------WKPVP-PVYPRLIQQVPEG---------------LTLEEAT 296
W P PV L+ +V G LT EE T
Sbjct: 306 YEREADRLLDGLGPRYMDWWMRRPFPVDADLLPEVVNGYMTPSRRCPPNTRAKLTDEELT 365
Query: 297 EMRQKGRTLTPICKLGKNGVYYNLVNNVREAFEECELVRV 336
+R + L LG+N L + + + +E+C + ++
Sbjct: 366 YLRNIAQPLPFHFVLGRNYGLQGLASAIVKLWEKCIIAKI 405
>At4g29750 putative protein
Length = 818
Score = 52.0 bits (123), Expect = 5e-07
Identities = 27/89 (30%), Positives = 47/89 (52%)
Query: 174 KEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMDNVCQ 233
+ E+ L L+ ++ +G G +++ IH W+ V K+K +++M +
Sbjct: 246 EHELKRLRNVALRMVERVKVGSAGITQALVEAIHEKWEVDEVVKLKFSEPYSLNMKRTHE 305
Query: 234 QLEEKTGGKVIYRRGGVIYLFRGRNYNHK 262
LE+KTGG VI+R G + L+RG +Y K
Sbjct: 306 VLEKKTGGLVIWRSGSSVVLYRGISYKLK 334
Score = 46.6 bits (109), Expect = 2e-05
Identities = 30/101 (29%), Positives = 48/101 (46%), Gaps = 4/101 (3%)
Query: 170 EPLTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMD 229
E +T+EE + L L LGR ++N+H HWK R + K+ G +
Sbjct: 645 EIITEEERLLYRKIGLSMDPFLLLGRREVYDGTIENMHLHWKHRELVKVIVRGKSLPQVK 704
Query: 230 NVCQQLEEKTGGKVI----YRRGGVIYLFRGRNYNHKTRPR 266
++ LE ++GG ++ +G I L+RG+NY R R
Sbjct: 705 HIAISLEAESGGVLVSVDKTMKGYAIILYRGKNYQMPFRLR 745
Score = 30.8 bits (68), Expect = 1.2
Identities = 19/66 (28%), Positives = 32/66 (47%)
Query: 288 EGLTLEEATEMRQKGRTLTPICKLGKNGVYYNLVNNVREAFEECELVRVNCQGLNKSDYR 347
E +T EE R+ G ++ P LG+ VY + N+ ++ ELV+V +G + +
Sbjct: 645 EIITEEERLLYRKIGLSMDPFLLLGRREVYDGTIENMHLHWKHRELVKVIVRGKSLPQVK 704
Query: 348 KIGAKL 353
I L
Sbjct: 705 HIAISL 710
>At3g23070 hypothetical protein
Length = 850
Score = 52.0 bits (123), Expect = 5e-07
Identities = 27/88 (30%), Positives = 47/88 (52%)
Query: 172 LTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMDNV 231
L + E+ L T +++ ++ + G +D I WK + ++K G ++M +
Sbjct: 190 LPESELRRLRNLTFRTASKMRIRGGGVTQVAVDAIKEKWKSAEIVRLKIEGASALNMRKM 249
Query: 232 CQQLEEKTGGKVIYRRGGVIYLFRGRNY 259
+ LE+KTGG VI+R G I L+RG +Y
Sbjct: 250 HEILEKKTGGLVIWRSGTSISLYRGVSY 277
Score = 47.4 bits (111), Expect = 1e-05
Identities = 43/141 (30%), Positives = 65/141 (45%), Gaps = 8/141 (5%)
Query: 170 EPLTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMD 229
E +T EE + LK L LGR G ++N+H HWK R + KI +
Sbjct: 605 ESITDEERFMFRKLGLKMKAFLLLGRRGVFDGTVENMHLHWKYRELVKIIVKAKTFDGVK 664
Query: 230 NVCQQLEEKTGGKVI----YRRGGVIYLFRGRNYNHKTRPRFPLMLWKPVPPVYPRLIQ- 284
V LE ++GG ++ +G I ++RG++Y T R +L K R I+
Sbjct: 665 KVALALEAESGGILVSIDKVTKGYAIIVYRGQDYKRPTMLRPKNLLTK--RKALARSIEL 722
Query: 285 QVPEGLTLEEATEMRQKGRTL 305
Q EGL L+ + M+ K + L
Sbjct: 723 QRREGL-LKHISTMQAKAKQL 742
Score = 33.1 bits (74), Expect = 0.24
Identities = 27/122 (22%), Positives = 49/122 (40%), Gaps = 4/122 (3%)
Query: 142 GVKPVQSPGPFLPGTSPRYVMSREEV---LGEPLTKEEINELVRSTLKSSRQLNLGRDGF 198
G P+ LPG P Y + + L +E L R LGR
Sbjct: 359 GDNPLPVDADLLPGAIPDYEPPFRVLPYGVRSSLGPKEATALRRLARSIPPHFALGRSRQ 418
Query: 199 IHNMLDNIHAHWKRRRVCKIKCI-GVCTVDMDNVCQQLEEKTGGKVIYRRGGVIYLFRGR 257
+ + + W++ + KI GV + + + + L++ TGG ++ R + +RG+
Sbjct: 419 LQGLATAMVRLWEKSMLAKIAIKRGVQSTTSERMAEDLKKLTGGIMLSRNKDFLVFYRGK 478
Query: 258 NY 259
N+
Sbjct: 479 NF 480
Score = 30.8 bits (68), Expect = 1.2
Identities = 18/67 (26%), Positives = 31/67 (45%)
Query: 287 PEGLTLEEATEMRQKGRTLTPICKLGKNGVYYNLVNNVREAFEECELVRVNCQGLNKSDY 346
PE +T EE R+ G + LG+ GV+ V N+ ++ ELV++ +
Sbjct: 604 PESITDEERFMFRKLGLKMKAFLLLGRRGVFDGTVENMHLHWKYRELVKIIVKAKTFDGV 663
Query: 347 RKIGAKL 353
+K+ L
Sbjct: 664 KKVALAL 670
>At4g14510 hypothetical protein
Length = 918
Score = 49.3 bits (116), Expect = 3e-06
Identities = 32/108 (29%), Positives = 50/108 (45%), Gaps = 4/108 (3%)
Query: 170 EPLTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMD 229
E +T+EE + LK L LGR G ++N+H HWK R + KI
Sbjct: 674 EGITEEERFMFQKLGLKMKAFLLLGRRGVFDGTVENMHLHWKYRELIKILVKAKTLEGAQ 733
Query: 230 NVCQQLEEKTGGKVI----YRRGGVIYLFRGRNYNHKTRPRFPLMLWK 273
V LE ++GG ++ +G + ++RG++Y T R +L K
Sbjct: 734 KVAMALEAESGGILVSVDKISKGYAVIVYRGKDYKRPTTLRPKNLLTK 781
Score = 45.4 bits (106), Expect = 5e-05
Identities = 29/100 (29%), Positives = 49/100 (49%), Gaps = 5/100 (5%)
Query: 163 SREEVLGEPLTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIG 222
SR + L++ E+N L ++ ++ + G ++D I WK + ++K G
Sbjct: 199 SRYSLAEMTLSEFELNRLRNVMFRTKSKMRVTGAGVTQAVVDAIQEKWKGSEIVRLKIEG 258
Query: 223 VCTVDMDNVCQQLEEKTGGKVIYRRGGVIYLFRGRNYNHK 262
++M + + LE KTGG VI+R G I L YN+K
Sbjct: 259 SSALNMRRMHEILERKTGGLVIWRSGTSIAL-----YNYK 293
Score = 28.5 bits (62), Expect = 5.8
Identities = 16/66 (24%), Positives = 31/66 (46%)
Query: 288 EGLTLEEATEMRQKGRTLTPICKLGKNGVYYNLVNNVREAFEECELVRVNCQGLNKSDYR 347
EG+T EE ++ G + LG+ GV+ V N+ ++ EL+++ + +
Sbjct: 674 EGITEEERFMFQKLGLKMKAFLLLGRRGVFDGTVENMHLHWKYRELIKILVKAKTLEGAQ 733
Query: 348 KIGAKL 353
K+ L
Sbjct: 734 KVAMAL 739
>At3g18390 unknown protein
Length = 848
Score = 49.3 bits (116), Expect = 3e-06
Identities = 23/84 (27%), Positives = 46/84 (54%)
Query: 176 EINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMDNVCQQL 235
E+ L R + ++N+ + G +++ I+ W++ + ++K V DM + +
Sbjct: 247 ELRRLRRDGMYLRVRINIPKAGLTQAVMEKIYDTWRKEELVRLKFHEVLARDMKTAHEIV 306
Query: 236 EEKTGGKVIYRRGGVIYLFRGRNY 259
E +TGG VI+R G V+ ++RG +Y
Sbjct: 307 ERRTGGMVIWRAGSVMVVYRGLDY 330
Score = 43.5 bits (101), Expect = 2e-04
Identities = 27/103 (26%), Positives = 49/103 (47%), Gaps = 6/103 (5%)
Query: 170 EPLTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMD 229
E +++EE + LK L +G G +++N+H HWK R + K+ ++
Sbjct: 651 EVISEEERAMFRKVGLKMKAYLPIGIRGVFDGVIENMHLHWKHRELVKLISKQKNQAFVE 710
Query: 230 NVCQQLEEKTGGKVI----YRRGGVIYLFRGRNYNH--KTRPR 266
+ LE ++GG ++ +G + +RG+NY RPR
Sbjct: 711 ETARLLEYESGGVLVAIEKVPKGFALIYYRGKNYRRPISLRPR 753
Score = 27.7 bits (60), Expect = 10.0
Identities = 19/70 (27%), Positives = 33/70 (47%), Gaps = 4/70 (5%)
Query: 271 LWKPVPPVYPRLIQQVPEG----LTLEEATEMRQKGRTLTPICKLGKNGVYYNLVNNVRE 326
L P P Y + +P G LT E T +R+ G+TL LG+N + L + +
Sbjct: 419 LLPPTIPGYKTPFRLLPTGMRSNLTNAEMTNLRKIGKTLPCHFALGRNRNHQGLAAAILQ 478
Query: 327 AFEECELVRV 336
+E+ + ++
Sbjct: 479 IWEKSLIAKI 488
>At3g25440 unknown protein
Length = 380
Score = 48.5 bits (114), Expect = 5e-06
Identities = 26/91 (28%), Positives = 45/91 (48%), Gaps = 1/91 (1%)
Query: 170 EPLTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMD 229
E LT EE ++ LK + +GR G ++ N+H HWK+ + ++ ++
Sbjct: 111 EILTPEEHFYYLKMGLKCKNYVPVGRRGIYQGVILNMHLHWKKHQTLQVVIKTFTPDEVK 170
Query: 230 NVCQQLEEKTGGKVI-YRRGGVIYLFRGRNY 259
+ +L TGG V+ G I ++RG+NY
Sbjct: 171 EIAVELARLTGGIVLDVHEGNTIIMYRGKNY 201
>At1g28290 proline-rich protein, putative
Length = 359
Score = 44.7 bits (104), Expect = 8e-05
Identities = 46/146 (31%), Positives = 59/146 (39%), Gaps = 25/146 (17%)
Query: 20 SRRPTGKPNKNPSKPKVDPQSHPALKFSNIP--KQKLKPVNKTPENVKISEDGVSYVIEG 77
++ P P K P KP V P + P +K P K +KP K P + Y
Sbjct: 96 TKAPVKPPTKPPVKPPVSPPAKPPVKPPVYPPTKAPVKPPTKPPVKPPV------YPPTK 149
Query: 78 APFEFKYSYTETPKSKPVQMREPPFVP--FGPVTMPRPWTGRPPL-PPSKKKLKEFDSFV 134
AP + P PV +PP P PV P +PP+ PP+K +K
Sbjct: 150 AP-------VKPPTKPPV---KPPVYPPTKAPVKPPTKPPVKPPVSPPAKPPVKP----P 195
Query: 135 LPPPHKKGVKPVQSPGPFLPGTSPRY 160
+ PP K VKP SP P T P Y
Sbjct: 196 VYPPTKAPVKPPVSPPTKPPVTPPVY 221
Score = 40.8 bits (94), Expect = 0.001
Identities = 45/151 (29%), Positives = 58/151 (37%), Gaps = 19/151 (12%)
Query: 23 PTGKPNKNPSKPKVDPQSHPALKFSNIPKQK--LKPVNKTPENVKISEDGVSYVIEGAPF 80
P P K P K V P + P +K P K +KP K P +S V
Sbjct: 67 PAKSPVKPPVKAPVSPPAKPPVKPPVYPPTKAPVKPPTKPPVKPPVSPPAKPPVKPPV-- 124
Query: 81 EFKYSYTETPKSKPVQMR-EPPFVP--FGPVTMPRPWTGRPPL-PPSKKKLKEFDSFVLP 136
Y T+ P P + +PP P PV P +PP+ PP+K +K
Sbjct: 125 ---YPPTKAPVKPPTKPPVKPPVYPPTKAPVKPPTKPPVKPPVYPPTKAPVK-------- 173
Query: 137 PPHKKGVKPVQSPGPFLPGTSPRYVMSREEV 167
PP K VKP SP P P Y ++ V
Sbjct: 174 PPTKPPVKPPVSPPAKPPVKPPVYPPTKAPV 204
Score = 30.0 bits (66), Expect = 2.0
Identities = 31/105 (29%), Positives = 44/105 (41%), Gaps = 16/105 (15%)
Query: 84 YSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPL-PPSKKKLKEFDSFVLPPPHKKG 142
+ + P PV+ PP PV+ P +PP+ PP+K +K PP K
Sbjct: 61 HPHPHPPAKSPVK---PPVK--APVSPPAKPPVKPPVYPPTKAPVK--------PPTKPP 107
Query: 143 VKPVQSPGPFLPGTSPRYVMSREEVLGEPLTKEEINELVRSTLKS 187
VKP SP P P Y ++ V +P TK + V K+
Sbjct: 108 VKPPVSPPAKPPVKPPVYPPTKAPV--KPPTKPPVKPPVYPPTKA 150
>At4g13070 putative protein
Length = 332
Score = 43.9 bits (102), Expect = 1e-04
Identities = 29/93 (31%), Positives = 46/93 (49%), Gaps = 4/93 (4%)
Query: 170 EPLTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTV--D 227
E LT+EE + L R+ K + +GR G ++ N+H HWK+ K+ C C
Sbjct: 169 ESLTEEEQHYLKRTGEKRKNFVLVGRRGVFGGVVLNLHLHWKKHETVKVIC-KPCNKPGQ 227
Query: 228 MDNVCQQLEEKTGGKVI-YRRGGVIYLFRGRNY 259
+ ++L + G VI + I L+RG+NY
Sbjct: 228 VHEYAEELARLSKGIVIDVKPNNTIVLYRGKNY 260
Score = 33.9 bits (76), Expect = 0.14
Identities = 18/57 (31%), Positives = 31/57 (53%)
Query: 287 PEGLTLEEATEMRQKGRTLTPICKLGKNGVYYNLVNNVREAFEECELVRVNCQGLNK 343
PE LT EE +++ G +G+ GV+ +V N+ +++ E V+V C+ NK
Sbjct: 168 PESLTEEEQHYLKRTGEKRKNFVLVGRRGVFGGVVLNLHLHWKKHETVKVICKPCNK 224
>At3g22120 unknown protein
Length = 351
Score = 43.1 bits (100), Expect = 2e-04
Identities = 40/133 (30%), Positives = 50/133 (37%), Gaps = 5/133 (3%)
Query: 22 RPTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAPFE 81
+P P K P P V P PA+K +KP K P VK + P
Sbjct: 55 KPPKHPVKPPKPPAVKPPKPPAVKPPTPKPPTVKPHPKPP-TVKPHPKPPTVKPHPKPPT 113
Query: 82 FKYSYTETPKSKPVQMREPPFVP---FGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPP 138
K + + P +KP +PP V P + P+P T PP P K S PP
Sbjct: 114 VKPPHPKPP-TKPHPHPKPPIVKPPTKPPPSTPKPPTKPPPSTPKPPTTKPPPSTPKPPH 172
Query: 139 HKKGVKPVQSPGP 151
HK P P P
Sbjct: 173 HKPPPTPCPPPTP 185
Score = 33.5 bits (75), Expect = 0.18
Identities = 22/69 (31%), Positives = 26/69 (36%), Gaps = 4/69 (5%)
Query: 90 PKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKGVKPVQSP 149
PK P + P PV P+P +PP PP+ K V P P VKP P
Sbjct: 47 PKPSPAPHKPPKH----PVKPPKPPAVKPPKPPAVKPPTPKPPTVKPHPKPPTVKPHPKP 102
Query: 150 GPFLPGTSP 158
P P
Sbjct: 103 PTVKPHPKP 111
>At2g28480 hypothetical protein
Length = 372
Score = 42.7 bits (99), Expect = 3e-04
Identities = 26/95 (27%), Positives = 43/95 (44%), Gaps = 1/95 (1%)
Query: 166 EVLGEPLTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCT 225
EV +T EE L + K S + +GR G ++ N+H HWK+ K+ C
Sbjct: 158 EVRPHEITGEERFYLKKMGQKRSNYVPIGRRGVFGGVILNMHLHWKKHETVKVICNNSKP 217
Query: 226 VDMDNVCQQLEEKTGGKVIYRRG-GVIYLFRGRNY 259
+ ++L + +GG + G I +RG+ Y
Sbjct: 218 GQVQQYAEELAKLSGGVPVNIIGDDTIIFYRGKGY 252
>At5g14920 unknown protein
Length = 275
Score = 42.4 bits (98), Expect = 4e-04
Identities = 37/136 (27%), Positives = 53/136 (38%), Gaps = 16/136 (11%)
Query: 23 PTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAPFEF 82
P+ KP P+ P P + P +K IP +KP TP I+ P +
Sbjct: 57 PSYKPPTLPTTPIKPPTTKPPVKPPTIPVTPVKPPVSTPP------------IKLPPVQP 104
Query: 83 KYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKG 142
TP KP ++ P + P P +P T P PP+ ++ V PP +K
Sbjct: 105 PTYKPPTPTVKPPSVQPPTYKP--PTPTVKPPTTSPVKPPTTPPVQ--SPPVQPPTYKPP 160
Query: 143 VKPVQSPGPFLPGTSP 158
PV+ P P P
Sbjct: 161 TSPVKPPTTTPPVKPP 176
Score = 30.0 bits (66), Expect = 2.0
Identities = 37/145 (25%), Positives = 47/145 (31%), Gaps = 19/145 (13%)
Query: 22 RPTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKIS----EDGVSYVIEG 77
+P KP P P P S P +K + KP T + + + V
Sbjct: 75 KPPVKPPTIPVTPVKPPVSTPPIKLPPVQPPTYKPPTPTVKPPSVQPPTYKPPTPTVKPP 134
Query: 78 APFEFKYSYTETPKSKPVQ--MREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVL 135
K T +S PVQ +PP P P T P PP V
Sbjct: 135 TTSPVKPPTTPPVQSPPVQPPTYKPPTSPVKPPTTTPPVKPPTTTPP-----------VQ 183
Query: 136 PPPHKKGVKPVQSP--GPFLPGTSP 158
PP + PV+ P P P T P
Sbjct: 184 PPTYNPPTTPVKPPTAPPVKPPTPP 208
>At3g27550 hypothetical protein
Length = 491
Score = 42.0 bits (97), Expect = 5e-04
Identities = 23/91 (25%), Positives = 39/91 (42%), Gaps = 1/91 (1%)
Query: 170 EPLTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMD 229
E T E++ + K+ + +G G ++ N+H HWK ++ C +
Sbjct: 90 ELFTSEQVQAFKKIGFKNKNYVPVGVRGVFGGVVQNMHMHWKFHETVQVCCDNFPKEKIK 149
Query: 230 NVCQQLEEKTGGKVI-YRRGGVIYLFRGRNY 259
+ + +GG VI I +FRGRNY
Sbjct: 150 EMASMIARLSGGVVINIHNVKTIIMFRGRNY 180
>At4g07680 hypothetical protein
Length = 684
Score = 40.4 bits (93), Expect = 0.001
Identities = 39/159 (24%), Positives = 62/159 (38%), Gaps = 30/159 (18%)
Query: 26 KPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENV--KISEDGVSYVIEGA----- 78
+P PK Q P N+PK+ + TP K ++ GV +EG
Sbjct: 202 QPQTQEDNPKAQTQKTP-----NVPKKTTNNQSATPSPPPSKQADVGVCRNLEGEFDKVS 256
Query: 79 -PFEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPP 137
+F + T +P + ++ R+PP P P +KK K + + P
Sbjct: 257 IVSDFDFVETVSPLNCRLRSRKPPV----------------PNVPKQKKEKTLNELIQKP 300
Query: 138 PHKKGVKPVQSPGPFLPGTSPRYVMSREEVLGEPLTKEE 176
P ++G KP Q P +P T P+ + + E K E
Sbjct: 301 PARRGRKPSQQPKK-VPPTVPKITIKNPKPSEEAKEKAE 338
>At4g33970 extensin-like protein
Length = 699
Score = 38.5 bits (88), Expect = 0.006
Identities = 44/158 (27%), Positives = 55/158 (33%), Gaps = 35/158 (22%)
Query: 10 PISASNADQSSRRPTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKP---VNKTPENVKI 66
P+ A+ D+ S P+ +P + P PK PQ S + K++ P VN P V
Sbjct: 474 PVLATPVDKPSPVPS-RPVQKPQPPKESPQPDDPYDQSPVTKRRSPPPAPVNSPPPPV-- 530
Query: 67 SEDGVSYVIEGAPFEFKYSYTETPKSKPVQMREPPF--VPFGPVTMPRPWTGRPPLPPSK 124
Y+ P PV PP P PV P P PP P
Sbjct: 531 -------------------YSPPPPPPPVHSPPPPVHSPPPPPVYSPPP----PPPPVHS 567
Query: 125 KKLKEFDS----FVLPPPHKKGVKPVQSPGPFLPGTSP 158
F + PPP PV SP P P SP
Sbjct: 568 PPPPVFSPPPPVYSPPPPVHSPPPPVHSPPPPAPVHSP 605
Score = 30.0 bits (66), Expect = 2.0
Identities = 34/129 (26%), Positives = 38/129 (29%), Gaps = 5/129 (3%)
Query: 23 PTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAPFEF 82
P P +P P S P FS P PV P V V AP
Sbjct: 550 PPPPPVYSPPPPPPPVHSPPPPVFSPPP-----PVYSPPPPVHSPPPPVHSPPPPAPVHS 604
Query: 83 KYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKG 142
+P P PP V P + P PP P K S P KK
Sbjct: 605 PPPPVHSPPPPPPVYSPPPPVFSPPPSQSPPVVYSPPPRPPKINSPPVQSPPPAPVEKKE 664
Query: 143 VKPVQSPGP 151
P +P P
Sbjct: 665 TPPAHAPAP 673
Score = 28.1 bits (61), Expect = 7.6
Identities = 26/80 (32%), Positives = 31/80 (38%), Gaps = 8/80 (10%)
Query: 87 TETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKGVKPV 146
+ TP SKP + +P VP PV P P P PS + + P PV
Sbjct: 416 SSTP-SKPSPVHKPTPVPTTPVHKPTPVPTTPVQKPS-----PVPTTPVQKPSPVPTTPV 469
Query: 147 QSPGPFL--PGTSPRYVMSR 164
P P L P P V SR
Sbjct: 470 HEPSPVLATPVDKPSPVPSR 489
>At5g62640 unknown protein
Length = 520
Score = 38.1 bits (87), Expect = 0.007
Identities = 41/146 (28%), Positives = 57/146 (38%), Gaps = 19/146 (13%)
Query: 21 RRPTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAPF 80
RR TG+ + KP+ HP L + P P+ + + IS DG S +
Sbjct: 112 RRLTGEEDL---KPEDSVYYHPTLNPTGAPPPGKPPMYNSSIGLAISSDGAS-----SSS 163
Query: 81 EFKYSYTETPKSKPVQMREPPFVPF----GPVTMPRPWTGRPPLPPSKKKLKEFDSFVLP 136
S TE+ S V + PP P ++ P PPLPP+ F P
Sbjct: 164 AALSSITESEDS--VLVNPPPLPPLPDGDNALSASLPLPPLPPLPPTTGLTLPHSPFPPP 221
Query: 137 PP-----HKKGVKPVQSPGPFLPGTS 157
PP + V+P P P LP +S
Sbjct: 222 PPGPPPKEQDFVRPPLPPPPQLPQSS 247
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.137 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,119,579
Number of Sequences: 26719
Number of extensions: 465858
Number of successful extensions: 2028
Number of sequences better than 10.0: 136
Number of HSP's better than 10.0 without gapping: 37
Number of HSP's successfully gapped in prelim test: 101
Number of HSP's that attempted gapping in prelim test: 1563
Number of HSP's gapped (non-prelim): 358
length of query: 355
length of database: 11,318,596
effective HSP length: 100
effective length of query: 255
effective length of database: 8,646,696
effective search space: 2204907480
effective search space used: 2204907480
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC137546.15


