

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137546.13 - phase: 0
(363 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
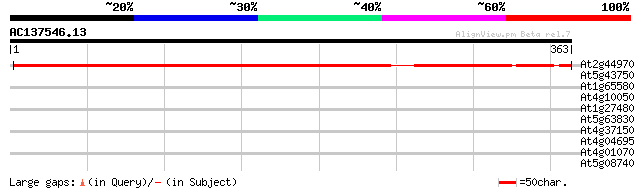
Score E
Sequences producing significant alignments: (bits) Value
At2g44970 unknown protein 440 e-124
At5g43750 unknown protein 33 0.24
At1g65580 hypothetical protein 32 0.41
At4g10050 lipase like protein 31 0.92
At1g27480 unknown protein 31 1.2
At5g63830 unknown protein 30 1.6
At4g37150 hydroxynitrile lyase like protein 29 3.5
At4g04695 Calcium-dependent Protein Kinase, isoform 31 (ppc:4.2.1) 28 6.0
At4g01070 putative flavonol glucosyltransferase 28 6.0
At5g08740 NADH dehydrogenase-like protein 28 7.8
>At2g44970 unknown protein
Length = 503
Score = 440 bits (1131), Expect = e-124
Identities = 223/362 (61%), Positives = 280/362 (76%), Gaps = 20/362 (5%)
Query: 3 FLRITSRVLKTLRGSSDDIGWLQHAPGMPPVHDGSSRFLELLSDIRNGKDSIPSSFVYLL 62
F + R +T+RGS+DDIGWLQ AP MPPV DG+ RF ++L DI +G +P++ VYLL
Sbjct: 156 FQGLIERARRTVRGSADDIGWLQRAPEMPPVEDGTDRFNKILEDIGHGVHRLPNTVVYLL 215
Query: 63 IPGLFSNHGPLYFVATKRFFSKMGLACHIAKVHSEASVEHNAMEIKQYIEEIYWGSGKPV 122
+PGLFSNHGPLYFV TK FSKMGLACHIAK+HSE+SVE NA EIK+YIEE+ WGS K V
Sbjct: 216 VPGLFSNHGPLYFVDTKTKFSKMGLACHIAKIHSESSVEKNAREIKEYIEELCWGSNKRV 275
Query: 123 MLLGHSKGGIDAAAALSLYWSDLKGKVAGLALVQSPYGGTPIASDILREGQIGD-KETRR 181
+LLGHSKGGIDAAAALSLYW +LK KVAGL L QSPYGG+PIA+DILREGQ+GD R+
Sbjct: 276 LLLGHSKGGIDAAAALSLYWPELKDKVAGLVLAQSPYGGSPIATDILREGQLGDYVNLRK 335
Query: 182 ILELIICKIIKGDIRALEDLTYEKRKDFIMKHKLPLDIPLISFRSEASITPSVLATMTQI 241
++E++I K+IKGDI+ALEDLTYE+RK+F+ H LP ++P +SFR+EASI+P+VL+T++ +
Sbjct: 336 MMEILISKVIKGDIQALEDLTYERRKEFLKNHPLPRELPTVSFRTEASISPAVLSTLSHV 395
Query: 242 AHAELPRLILPKFGSKVSDQFVESGRQVPVMVPVSAAMAAFALHLQLRYGEKSDGVVTCR 301
AHAELP ++PV++P+ AAMAA A LQ+RYGEKSDG+VTC
Sbjct: 396 AHAELP--------------LTNQAAKLPVVMPLGAAMAACAQLLQVRYGEKSDGLVTCC 441
Query: 302 DAEVPGSVVVRPNMKLDHAWMVYSSNSKKKKSSEPDAREMCQAIFTLLVELGKTEREVEQ 361
DAEVPGSVVVRP KLDHAWMVYS S + E DA ++C+A+ TLLV++ E+E +Q
Sbjct: 442 DAEVPGSVVVRPKRKLDHAWMVYS--SLNEVPLEADAAQVCEALLTLLVQV---EQERQQ 496
Query: 362 VL 363
L
Sbjct: 497 KL 498
>At5g43750 unknown protein
Length = 212
Score = 33.1 bits (74), Expect = 0.24
Identities = 19/50 (38%), Positives = 32/50 (64%), Gaps = 6/50 (12%)
Query: 235 LATMTQIAHAELPRLILPKFGSKVSDQFV------ESGRQVPVMVPVSAA 278
+AT+T ++ +P++ KFG++VSDQ V +SGR++ + VSAA
Sbjct: 1 MATVTILSPKSIPKVTDSKFGARVSDQIVNVVKCGKSGRRLKLAKLVSAA 50
>At1g65580 hypothetical protein
Length = 993
Score = 32.3 bits (72), Expect = 0.41
Identities = 29/105 (27%), Positives = 47/105 (44%), Gaps = 14/105 (13%)
Query: 111 IEEIYWGSGKPVMLLGHSKGGIDAAAALSLYWSDLKGKV---AGLAL-VQSPYGGTPIAS 166
++E+ G+G M G++ + L +W D+ GK AG + V P GG +
Sbjct: 570 LQEVEMGAGVLAMSAAKETVGLEGSP-LGQWWLDMIGKTLDEAGASFGVTMPRGGNALGV 628
Query: 167 DILREGQIGDKETRRILELIICKIIKGDIR-ALEDLTYEKRKDFI 210
+ + E R L I GD L+D+TY++ +DFI
Sbjct: 629 NTI--------EARPELSEADMVIFLGDFNYRLDDITYDETRDFI 665
>At4g10050 lipase like protein
Length = 350
Score = 31.2 bits (69), Expect = 0.92
Identities = 22/70 (31%), Positives = 38/70 (53%), Gaps = 4/70 (5%)
Query: 97 EASVEHNAMEIKQYIEEIYWGSGKPVMLLGHSKGGIDAAAALSLYWSDLKGKVAGLALVQ 156
E S+E + ++ I+E+Y S ++L+GHS GG + A+ + + +AGL +V
Sbjct: 125 ELSLETMSNDVLAVIKELYGDSPPAIVLVGHSMGG---SVAVQVAANKTLPSLAGLVVV- 180
Query: 157 SPYGGTPIAS 166
GT I+S
Sbjct: 181 DVVEGTAISS 190
>At1g27480 unknown protein
Length = 432
Score = 30.8 bits (68), Expect = 1.2
Identities = 24/81 (29%), Positives = 37/81 (45%), Gaps = 5/81 (6%)
Query: 84 KMGLACHIAKVHSEASVEHNAMEIKQYIEEIYW-GSGKPVMLLGHSKGGIDAAAALSLYW 142
+ GLA A H ++KQ +E+ GKPV+LL HS GG+ L+
Sbjct: 167 RYGLA---ASGHPSRVASQFLQDLKQLVEKTSSENEGKPVILLSHSLGGLFVLHFLNRTT 223
Query: 143 SDLKGK-VAGLALVQSPYGGT 162
+ K + + +P+GGT
Sbjct: 224 PSWRRKYIKHFVALAAPWGGT 244
>At5g63830 unknown protein
Length = 406
Score = 30.4 bits (67), Expect = 1.6
Identities = 17/36 (47%), Positives = 23/36 (63%)
Query: 174 IGDKETRRILELIICKIIKGDIRALEDLTYEKRKDF 209
I D E + E II KI+ GD +L+DL+ E+RK F
Sbjct: 106 ITDDEGLSLPEEIIQKIMNGDEVSLDDLSLEERKGF 141
>At4g37150 hydroxynitrile lyase like protein
Length = 256
Score = 29.3 bits (64), Expect = 3.5
Identities = 17/56 (30%), Positives = 32/56 (56%), Gaps = 2/56 (3%)
Query: 90 HIAKVHSEASVEHNAMEIKQYIEEIYWGSGKPVMLLGHSKGGIDAAAALSLYWSDL 145
++++V ++E A + + +E +GS V+L+ HS GGI AA A ++ S +
Sbjct: 42 NMSRVEDIQTLEDFAKPLLEVLES--FGSDDKVVLVAHSLGGIPAALAADMFPSKI 95
>At4g04695 Calcium-dependent Protein Kinase, isoform 31 (ppc:4.2.1)
Length = 484
Score = 28.5 bits (62), Expect = 6.0
Identities = 35/119 (29%), Positives = 55/119 (45%), Gaps = 14/119 (11%)
Query: 154 LVQSPY---GGTPIASDILREGQIGDKETRRILE-----LIICK-IIKGDIRALEDLTYE 204
+++ P+ G I D L +GQ G TR+ +E CK I+K ++++ ED
Sbjct: 16 ILEKPFVDIGKVYILGDELGQGQFGI--TRKCVEKTSGKTYACKTILKTNLKSREDEEAV 73
Query: 205 KRKDFIMKHKLPLDIPLISFRSEASITPSVLATMTQIAHAELPRLI--LPKFGSKVSDQ 261
KR+ IMKH L + ++ F+ SV M EL + I L K G S++
Sbjct: 74 KREIRIMKH-LSGEPNIVEFKKAYEDRDSVHIVMEYCGGGELFKKIEALSKDGKSYSEK 131
>At4g01070 putative flavonol glucosyltransferase
Length = 480
Score = 28.5 bits (62), Expect = 6.0
Identities = 29/102 (28%), Positives = 44/102 (42%), Gaps = 22/102 (21%)
Query: 249 LILPKFGSKVSDQFVESGRQVPVMVPVSAAMAAFALHLQLRYGEKSDGVVTCRDAE---- 304
L++ FG+ D VE + P +A + +F LHL K D V+C E
Sbjct: 114 LVVDLFGTDAFDVAVEFHVPPYIFYPTTANVLSFFLHL-----PKLDETVSCEFRELTEP 168
Query: 305 --VPGSVVVRPNMKLDHA---------WMVYSSNSKKKKSSE 335
+PG V V LD A W+++ N+K+ K +E
Sbjct: 169 LMLPGCVPVAGKDFLDPAQDRKDDAYKWLLH--NTKRYKEAE 208
>At5g08740 NADH dehydrogenase-like protein
Length = 519
Score = 28.1 bits (61), Expect = 7.8
Identities = 11/42 (26%), Positives = 23/42 (54%)
Query: 307 GSVVVRPNMKLDHAWMVYSSNSKKKKSSEPDAREMCQAIFTL 348
G+V++ K+++ W+V + ++ K P A E+ +TL
Sbjct: 181 GTVLLESGFKIEYDWLVLALGAESKLDVVPGAMELAFPFYTL 222
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.136 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,887,670
Number of Sequences: 26719
Number of extensions: 331514
Number of successful extensions: 806
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 798
Number of HSP's gapped (non-prelim): 10
length of query: 363
length of database: 11,318,596
effective HSP length: 101
effective length of query: 262
effective length of database: 8,619,977
effective search space: 2258433974
effective search space used: 2258433974
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC137546.13


