

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
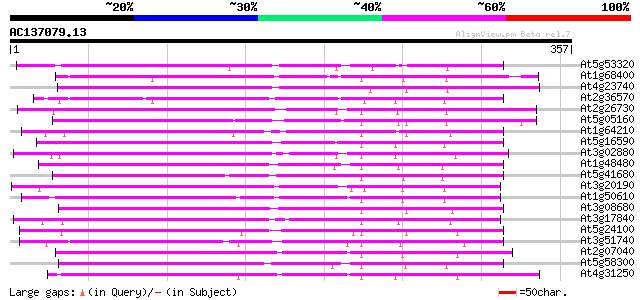
Query= AC137079.13 + phase: 0
(357 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At5g53320 receptor protein kinase-like protein 194 5e-50
At1g68400 receptor kinase like protein 193 1e-49
At4g23740 putative receptor kinase 192 2e-49
At2g36570 putative receptor-like protein kinase 189 3e-48
At2g26730 putative receptor-like protein kinase 185 4e-47
At5g05160 receptor-like protein kinase 181 5e-46
At1g64210 hypothetical protein 180 1e-45
At5g16590 receptor-like protein kinase 179 2e-45
At3g02880 putative protein kinase 177 6e-45
At1g48480 protein kinase like protein 177 8e-45
At5g41680 Pto kinase interactor 1-like protein 175 4e-44
At3g20190 receptor kinase like protein 170 1e-42
At1g50610 receptor-like protein kinase, putative 170 1e-42
At3g08680 putative protein kinase 169 2e-42
At3g17840 receptor kinase, putative 168 5e-42
At5g24100 receptor-like protein kinase 167 1e-41
At3g51740 unknown protein 165 3e-41
At2g07040 putative leucine-rich repeat transmembrane protein kinase 164 9e-41
At5g58300 receptor-like protein kinase 163 1e-40
At4g31250 receptor kinase - like protein 161 6e-40
>At5g53320 receptor protein kinase-like protein
Length = 601
Score = 194 bits (494), Expect = 5e-50
Identities = 120/321 (37%), Positives = 187/321 (57%), Gaps = 26/321 (8%)
Query: 5 EVKKNNLDSPMKKAT-SEGRLKGGDNNNSELVFFVEDHERFKLEDLLRATADLRSENFWS 63
E ++++ D P K+ S+ + GDN ++VFF + F LEDLLRA+A++ + +
Sbjct: 264 EQRRSSKDKPSKRRKDSDPNVGEGDN---KIVFFEGKNLVFDLEDLLRASAEVLGKGPFG 320
Query: 64 SLFKVKFENNVEYAVKRLKNLQVSCDEFREILKQISKVKHQNILSLVGYRSTKEEKLIIY 123
+ +KV E++ VKR+K + V EF + ++ I +KH+N+ +L GY +K+EKL++Y
Sbjct: 321 TTYKVDLEDSATIVVKRIKEVSVPQREFEQQIENIGSIKHENVATLRGYFYSKDEKLVVY 380
Query: 124 KYQSNGSVLNLLNDY--IARRKDFPWKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNL 181
Y +GS+ LL+ + RK W+ RLN+ G ARG+A I+ + + HGN+
Sbjct: 381 DYYEHGSLSTLLHGQKGLRDRKRLEWETRLNMVYGTARGVAHIHSQ----SGGKLVHGNI 436
Query: 182 KLSNILLDDKNEALISEHGLSKFFE--PDRGTFFSSHGYTAPEKSLTEKG----DVYSFG 235
K SNI L+ K IS G++ P GY APE + T KG DVYSFG
Sbjct: 437 KSSNIFLNGKGYGCISGTGMATLMHSLPRHAV-----GYRAPEITDTRKGTQPSDVYSFG 491
Query: 236 VILLELLTGQSIEVSRIDLVRWVRSMVREEWTGEVFDKEVRE--NDHQGAFSLLNIALMC 293
+++ E+LTG+S EV+ +LVRWV S+VREEWTGEVFD+E+ + +L + ++C
Sbjct: 492 ILIFEVLTGKS-EVA--NLVRWVNSVVREEWTGEVFDEELLRCTQVEEEMVEMLQVGMVC 548
Query: 294 VSRSQENRPNFGEILETIEGV 314
+R E RPN E++ +E +
Sbjct: 549 TARLPEKRPNMIEVVRMVEEI 569
>At1g68400 receptor kinase like protein
Length = 670
Score = 193 bits (490), Expect = 1e-49
Identities = 122/323 (37%), Positives = 186/323 (56%), Gaps = 29/323 (8%)
Query: 30 NNSELVFFVEDHERFKLEDLLRATADLRSENFWSSLFKVKFENNVEYAVKRLKNLQVSCD 89
+ ++VFF E RF+LEDLLRA+A++ + + + +K E+ E AVKRLK+
Sbjct: 342 DKGKMVFF-EGTRRFELEDLLRASAEMLGKGGFGTAYKAVLEDGNEVAVKRLKDAVTVAG 400
Query: 90 --EFREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLN-DYIARRKDFP 146
EF + ++ + +++H N++SL Y +EEKL++Y Y NGS+ LL+ + R
Sbjct: 401 KKEFEQQMEVLGRLRHTNLVSLKAYYFAREEKLLVYDYMPNGSLFWLLHGNRGPGRTPLD 460
Query: 147 WKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFE 206
W RL IA G ARGLAFI+ + + HG++K +N+LLD A +S+ GLS F
Sbjct: 461 WTTRLKIAAGAARGLAFIHGSC---KTLKLTHGDIKSTNVLLDRSGNARVSDFGLS-IFA 516
Query: 207 PDRGTFFSSHGYTAPE----KSLTEKGDVYSFGVILLELLTGQSIEV-------SRIDLV 255
P + T S+GY APE + T+K DVYSFGV+LLE+LTG+ + +DL
Sbjct: 517 PSQ-TVAKSNGYRAPELIDGRKHTQKSDVYSFGVLLLEILTGKCPNMVETGHSGGAVDLP 575
Query: 256 RWVRSMVREEWTGEVFDKEVR--ENDHQGAFSLLNIALMCVSRSQENRPNFGEILETIEG 313
RWV+S+VREEWT EVFD E+ ++ + LL IA+ C + + ++RP G +++ IE
Sbjct: 576 RWVQSVVREEWTAEVFDLELMRYKDIEEEMVGLLQIAMACTAVAADHRPKMGHVVKLIED 635
Query: 314 VMNAHDQQQMELSASKCCSNGSN 336
+ S + C++G N
Sbjct: 636 IRGGG-------SEASPCNDGIN 651
>At4g23740 putative receptor kinase
Length = 638
Score = 192 bits (489), Expect = 2e-49
Identities = 119/319 (37%), Positives = 179/319 (55%), Gaps = 16/319 (5%)
Query: 31 NSELVFFVEDHERFKLEDLLRATADLRSENFWSSLFKVKFENNVEYAVKRLKNLQVSCDE 90
N+ L FF + F LEDLLRA+A++ + + + +K E+ AVKRLK++ +
Sbjct: 317 NNRLSFFEGCNYSFDLEDLLRASAEVLGKGTFGTTYKAVLEDATSVAVKRLKDVAAGKRD 376
Query: 91 FREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLN-DYIARRKDFPWKL 149
F + ++ I +KH+N++ L Y +K+EKL++Y Y S GSV +LL+ + R W+
Sbjct: 377 FEQQMEIIGGIKHENVVELKAYYYSKDEKLMVYDYFSRGSVASLLHGNRGENRIPLDWET 436
Query: 150 RLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEPDR 209
R+ IA G A+G+A I+K+ + HGN+K SNI L+ ++ +S+ GL+ P
Sbjct: 437 RMKIAIGAAKGIARIHKE----NNGKLVHGNIKSSNIFLNSESNGCVSDLGLTAVMSPLA 492
Query: 210 GTFFSSHGYTAPEKSLTEK----GDVYSFGVILLELLTGQS-IEVSR----IDLVRWVRS 260
GY APE + T K DVYSFGV+LLELLTG+S I + I LVRWV S
Sbjct: 493 PPISRQAGYRAPEVTDTRKSSQLSDVYSFGVVLLELLTGKSPIHTTAGDEIIHLVRWVHS 552
Query: 261 MVREEWTGEVFDKEVRE--NDHQGAFSLLNIALMCVSRSQENRPNFGEILETIEGVMNAH 318
+VREEWT EVFD E+ N + +L IA+ CV ++ + RP +++ IE V N
Sbjct: 553 VVREEWTAEVFDIELLRYTNIEEEMVEMLQIAMSCVVKAADQRPKMSDLVRLIENVGNRR 612
Query: 319 DQQQMELSASKCCSNGSNQ 337
+ E NG+++
Sbjct: 613 TSIEPEPELKPKSENGASE 631
>At2g36570 putative receptor-like protein kinase
Length = 672
Score = 189 bits (479), Expect = 3e-48
Identities = 125/335 (37%), Positives = 194/335 (57%), Gaps = 44/335 (13%)
Query: 16 KKATSEGRLKGGDNN------NSELVFFVEDHERFKLEDLLRATADLRSENFWSSLFKVK 69
K+ +S G +GG+++ S LVFF E ++F+L+DLL+A+A++ + +++K
Sbjct: 319 KRRSSYG--EGGESDATSATDRSRLVFF-ERRKQFELDDLLKASAEMLGKGSLGTVYKAV 375
Query: 70 FEN-NVEYAVKRLKNLQVSCD--EFREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQ 126
++ + AVKRLK+ C EF + ++ I ++KHQN++ L Y KEEKL++Y+Y
Sbjct: 376 LDDGSTTVAVKRLKDAN-PCPRKEFEQYMEIIGRLKHQNVVKLRAYYYAKEEKLLVYEYL 434
Query: 127 SNGSVLNLLN-DYIARRKDFPWKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSN 185
NGS+ +LL+ + R W R+++ G ARGLA I+ +E ++ IPHGN+K SN
Sbjct: 435 PNGSLHSLLHGNRGPGRIPLDWTTRISLMLGAARGLAKIH---DEYSISKIPHGNIKSSN 491
Query: 186 ILLDDKNEALISEHGLSKFFEPDRGTFFSSHGYTAPEKS----LTEKGDVYSFGVILLEL 241
+LLD ALI++ GLS P GY APE+S L++K DVYSFGV+LLE+
Sbjct: 492 VLLDRNGVALIADFGLSLLLNPVH-AIARLGGYRAPEQSEIKRLSQKADVYSFGVLLLEV 550
Query: 242 LTG--------------------QSIEVSRIDLVRWVRSMVREEWTGEVFDKEV--REND 279
LTG + E + +DL +WVRS+V+EEWT EVFD E+ +N
Sbjct: 551 LTGKAPSIFPSPSRPRSAASVAVEEEEEAVVDLPKWVRSVVKEEWTAEVFDPELLRYKNI 610
Query: 280 HQGAFSLLNIALMCVSRSQENRPNFGEILETIEGV 314
+ ++L+I L CV E RP E+++ +E +
Sbjct: 611 EEEMVAMLHIGLACVVPQPEKRPTMAEVVKMVEEI 645
>At2g26730 putative receptor-like protein kinase
Length = 658
Score = 185 bits (469), Expect = 4e-47
Identities = 127/352 (36%), Positives = 184/352 (52%), Gaps = 33/352 (9%)
Query: 6 VKKNNLDSPMKKATSEGRLKG------GDNNNSELVFFVEDHERFKLEDLLRATADLRSE 59
V N+D P ++S+ + G G+ ++LVF F LEDLLRA+A++ +
Sbjct: 300 VATRNVDLPPGASSSKEEVTGTSSGMGGETERNKLVFTEGGVYSFDLEDLLRASAEVLGK 359
Query: 60 NFWSSLFKVKFENNVEYAVKRLKNLQVSCDEFREILKQISKVKHQNILSLVGYRSTKEEK 119
+ +K E VKRLK++ S EF ++ + K+KH N++ L Y +K+EK
Sbjct: 360 GSVGTSYKAVLEEGTTVVVKRLKDVMASKKEFETQMEVVGKIKHPNVIPLRAYYYSKDEK 419
Query: 120 LIIYKYQSNGSVLNLLN-DYIARRKDFPWKLRLNIACGIARGLAFIYKKLEEGEVNSIPH 178
L+++ + GS+ LL+ + R W R+ IA ARGLA ++ + + H
Sbjct: 420 LLVFDFMPTGSLSALLHGSRGSGRTPLDWDNRMRIAITAARGLAHLHVSAK------LVH 473
Query: 179 GNLKLSNILLDDKNEALISEHGLSKFFE----PDRGTFFSSHGYTAPE----KSLTEKGD 230
GN+K SNILL + +S++GL++ F P+R GY APE + +T K D
Sbjct: 474 GNIKASNILLHPNQDTCVSDYGLNQLFSNSSPPNR-----LAGYHAPEVLETRKVTFKSD 528
Query: 231 VYSFGVILLELLTGQ-----SIEVSRIDLVRWVRSMVREEWTGEVFDKEVR--ENDHQGA 283
VYSFGV+LLELLTG+ S+ IDL RWV S+VREEWT EVFD E+ N +
Sbjct: 529 VYSFGVLLLELLTGKSPNQASLGEEGIDLPRWVLSVVREEWTAEVFDVELMRYHNIEEEM 588
Query: 284 FSLLNIALMCVSRSQENRPNFGEILETIEGVMNAHDQQQMELSASKCCSNGS 335
LL IA+ CVS + RP E+L IE V + +S S GS
Sbjct: 589 VQLLQIAMACVSTVPDQRPVMQEVLRMIEDVNRSETTDDGLRQSSDDPSKGS 640
>At5g05160 receptor-like protein kinase
Length = 640
Score = 181 bits (459), Expect = 5e-46
Identities = 112/325 (34%), Positives = 184/325 (56%), Gaps = 26/325 (8%)
Query: 28 DNNNSELVFFVEDHERFKLEDLLRATADLRSENFWSSLFKVKFENNVEYAVKRLKNLQVS 87
D ++L FF + F LEDLL+A+A++ + + + +K E+ VKRL+ + S
Sbjct: 324 DPEKNKLFFFERCNHNFDLEDLLKASAEVLGKGSFGTAYKAVLEDTTAVVVKRLREVVAS 383
Query: 88 CDEFREILKQISKV-KHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFP 146
EF + ++ + K+ +H N + L+ Y +K+EKL++YKY + GS+ +++ R
Sbjct: 384 KKEFEQQMEIVGKINQHSNFVPLLAYYYSKDEKLLVYKYMTKGSLFGIMHGNRGDR-GVD 442
Query: 147 WKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFE 206
W+ R+ IA G ++ +++++ HG++K SNILL + E +S+ L F
Sbjct: 443 WETRMKIATGTSKAISYLHSL-------KFVHGDIKSSNILLTEDLEPCLSDTSLVTLFN 495
Query: 207 PDRGTFFSSHGYTAPE----KSLTEKGDVYSFGVILLELLTGQS------IEVSR--IDL 254
T + GY APE + ++++ DVYSFGV++LE+LTG++ +E R IDL
Sbjct: 496 LPTHT-PRTIGYNAPEVIETRRVSQRSDVYSFGVVILEMLTGKTPLTQPGLEDERVVIDL 554
Query: 255 VRWVRSMVREEWTGEVFDKEVR--ENDHQGAFSLLNIALMCVSRSQENRPNFGEILETIE 312
RWVRS+VREEWT EVFD E+ +N + +L +AL CV+R+ E+RP E+ IE
Sbjct: 555 PRWVRSVVREEWTAEVFDVELLKFQNIEEEMVQMLQLALACVARNPESRPKMEEVARMIE 614
Query: 313 GVMNAHDQQQME--LSASKCCSNGS 335
V QQ++ ++S+ SN S
Sbjct: 615 DVRRLDQSQQLQQNRTSSEATSNVS 639
>At1g64210 hypothetical protein
Length = 587
Score = 180 bits (456), Expect = 1e-45
Identities = 117/332 (35%), Positives = 177/332 (53%), Gaps = 33/332 (9%)
Query: 8 KNNLDSPMKKATSE---GRLKGGDNNNSE---LVFFVEDHERFKLEDLLRATADLRSENF 61
K + ++K S G D+N E ++FF + F L+DLL ++A++ +
Sbjct: 258 KTRISGKLRKRDSSSPPGNWTSRDDNTEEGGKIIFFGGRNHLFDLDDLLSSSAEVLGKGA 317
Query: 62 WSSLFKVKFENNVEYAVKRLKNLQVSCDEFREILKQISKVKHQNILSLVGYRSTKEEKLI 121
+ + +KV E+ VKRLK + V EF + ++ I ++H+N+ L Y +K++KL
Sbjct: 318 FGTTYKVTMEDMSTVVVKRLKEVVVGRREFEQQMEIIGMIRHENVAELKAYYYSKDDKLA 377
Query: 122 IYKYQSNGSVLNLLNDYIAR--RKDFPWKLRLNIACGIARGLAFIYKKLEEGEVNSIPHG 179
+Y Y ++GS+ +L+ R R W RL IA G ARGLA K+ EG+ HG
Sbjct: 378 VYSYYNHGSLFEILHGNRGRYHRVPLDWDARLRIATGAARGLA----KIHEGK---FIHG 430
Query: 180 NLKLSNILLDDKNEALISEHGLSKFFEPDRGTFFSSHGYTAPE----KSLTEKGDVYSFG 235
N+K SNI LD + I + GL+ T + GY APE + T+ DVYSFG
Sbjct: 431 NIKSSNIFLDSQCYGCIGDVGLTTIMRSLPQTTCLTSGYHAPEITDTRRSTQFSDVYSFG 490
Query: 236 VILLELLTGQSIEVSR----------IDLVRWVRSMVREEWTGEVFDKEVREND---HQG 282
V+LLELLTG+S VS+ +DL W+RS+V +EWTGEVFD E+ +
Sbjct: 491 VVLLELLTGKS-PVSQAELVPTGGENMDLASWIRSVVAKEWTGEVFDMEILSQSGGFEEE 549
Query: 283 AFSLLNIALMCVSRSQENRPNFGEILETIEGV 314
+L I L CV+ Q+ RP+ ++L+ IE +
Sbjct: 550 MVEMLQIGLACVALKQQERPHIAQVLKLIEDI 581
>At5g16590 receptor-like protein kinase
Length = 625
Score = 179 bits (455), Expect = 2e-45
Identities = 105/310 (33%), Positives = 178/310 (56%), Gaps = 19/310 (6%)
Query: 18 ATSEGRLKGGDNNNSELVFFVEDHERFKLEDLLRATADLRSENFWSSLFKVKFENNVEYA 77
A+ G K + +L FFV+ F L+ LL+A+A++ + + S +K F++ + A
Sbjct: 310 ASENGVSKNPAAVSKDLTFFVKSFGEFDLDGLLKASAEVLGKGTFGSSYKASFDHGLVVA 369
Query: 78 VKRLKNLQVSCDEFREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLN- 136
VKRL+++ V EFRE L+ + + H N+++L+ Y +++EKL++++Y S GS+ LL+
Sbjct: 370 VKRLRDVVVPEKEFREKLQVLGSISHANLVTLIAYYFSRDEKLVVFEYMSRGSLSALLHG 429
Query: 137 DYIARRKDFPWKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALI 196
+ + R W+ R NIA G AR +++++ + + HGN+K SNILL + EA +
Sbjct: 430 NKGSGRSPLNWETRANIALGAARAISYLHSR-----DATTSHGNIKSSNILLSESFEAKV 484
Query: 197 SEHGLSKFFEPDRGTFFSSHGYTAPE----KSLTEKGDVYSFGVILLELLTG-----QSI 247
S++ L+ P T GY APE + +++K DVYSFGV++LELLTG Q +
Sbjct: 485 SDYCLAPMISP-TSTPNRIDGYRAPEVTDARKISQKADVYSFGVLILELLTGKSPTHQQL 543
Query: 248 EVSRIDLVRWVRSMVREEWTGEVFDKEV---RENDHQGAFSLLNIALMCVSRSQENRPNF 304
+DL RWV S+ ++ +VFD E+ + + ++ LLNI + C ++ ++RP
Sbjct: 544 HEEGVDLPRWVSSITEQQSPSDVFDPELTRYQSDSNENMIRLLNIGISCTTQYPDSRPTM 603
Query: 305 GEILETIEGV 314
E+ IE V
Sbjct: 604 PEVTRLIEEV 613
>At3g02880 putative protein kinase
Length = 627
Score = 177 bits (450), Expect = 6e-45
Identities = 112/347 (32%), Positives = 188/347 (53%), Gaps = 42/347 (12%)
Query: 3 EIEVKKNNLDSPMKKATSEGRLK-------------GGDNN--NSELVFFVEDHERFKLE 47
E V N+++P+ ATS + G ++ N +L FFV+ F L+
Sbjct: 282 EENVPSRNVEAPVAAATSSAAIPKETVVVVPPAKATGSESGAVNKDLTFFVKSFGEFDLD 341
Query: 48 DLLRATADLRSENFWSSLFKVKFENNVEYAVKRLKNLQVSCDEFREILKQISKVKHQNIL 107
LL+A+A++ + S +K FE+ + AVKRL+++ V EFRE L + + H N++
Sbjct: 342 GLLKASAEVLGKGTVGSSYKASFEHGLVVAVKRLRDVVVPEKEFRERLHVLGSMSHANLV 401
Query: 108 SLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIAR-RKDFPWKLRLNIACGIARGLAFIYK 166
+L+ Y +++EKL++++Y S GS+ +L+ R W+ R IA G AR +++++
Sbjct: 402 TLIAYYFSRDEKLLVFEYMSKGSLSAILHGNKGNGRTPLNWETRAGIALGAARAISYLHS 461
Query: 167 KLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFE----PDRGTFFSSHGYTAPE 222
+ +G + HGN+K SNILL D EA +S++GL+ P+R GY APE
Sbjct: 462 R--DGTTS---HGNIKSSNILLSDSYEAKVSDYGLAPIISSTSAPNR-----IDGYRAPE 511
Query: 223 ----KSLTEKGDVYSFGVILLELLTG-----QSIEVSRIDLVRWVRSMVREEWTGEVFDK 273
+ +++K DVYSFGV++LELLTG Q + +DL RWV+S+ ++ +V D
Sbjct: 512 ITDARKISQKADVYSFGVLILELLTGKSPTHQQLNEEGVDLPRWVQSVTEQQTPSDVLDP 571
Query: 274 EVRENDHQG---AFSLLNIALMCVSRSQENRPNFGEILETIEGVMNA 317
E+ +G LL I + C ++ ++RP+ E+ IE V ++
Sbjct: 572 ELTRYQPEGNENIIRLLKIGMSCTAQFPDSRPSMAEVTRLIEEVSHS 618
>At1g48480 protein kinase like protein
Length = 655
Score = 177 bits (449), Expect = 8e-45
Identities = 108/315 (34%), Positives = 179/315 (56%), Gaps = 29/315 (9%)
Query: 19 TSEGRLKGGDNN-NSELVFFVEDHERFKLEDLLRATADLRSENFWSSLFKVKFENNVEYA 77
T G+ G+ +LVFF + F LEDLLRA+A++ + + + +K + A
Sbjct: 338 TGNGKASEGNGPATKKLVFFGNATKVFDLEDLLRASAEVLGKGTFGTAYKAVLDAVTVVA 397
Query: 78 VKRLKNLQVSCDEFREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLN- 136
VKRLK++ ++ EF+E ++ + + H+N++ L Y +++EKL++Y + GS+ LL+
Sbjct: 398 VKRLKDVMMADKEFKEKIELVGAMDHENLVPLRAYYFSRDEKLLVYDFMPMGSLSALLHG 457
Query: 137 DYIARRKDFPWKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALI 196
+ A R W +R IA G ARGL +++ + S HGN+K SNILL ++A +
Sbjct: 458 NRGAGRSPLNWDVRSRIAIGAARGLDYLH-----SQGTSTSHGNIKSSNILLTKSHDAKV 512
Query: 197 SEHGLSKFF-----EPDRGTFFSSHGYTAPE----KSLTEKGDVYSFGVILLELLTGQS- 246
S+ GL++ P+R T GY APE K +++KGDVYSFGV+LLEL+TG++
Sbjct: 513 SDFGLAQLVGSSATNPNRAT-----GYRAPEVTDPKRVSQKGDVYSFGVVLLELITGKAP 567
Query: 247 ----IEVSRIDLVRWVRSMVREEWTGEVFDKE---VRENDHQGAFSLLNIALMCVSRSQE 299
+ +DL RWV+S+ R+EW EVFD E + ++ + ++ + L C S+ +
Sbjct: 568 SNSVMNEEGVDLPRWVKSVARDEWRREVFDSELLSLATDEEEMMAEMVQLGLECTSQHPD 627
Query: 300 NRPNFGEILETIEGV 314
RP E++ +E +
Sbjct: 628 QRPEMSEVVRKMENL 642
>At5g41680 Pto kinase interactor 1-like protein
Length = 333
Score = 175 bits (443), Expect = 4e-44
Identities = 105/298 (35%), Positives = 163/298 (54%), Gaps = 17/298 (5%)
Query: 28 DNNNSELVFFVEDHERFKLEDLLRATADLRSENFWSSLFKVKFENNVEYAVKRLKNLQVS 87
DN+ ++VFF + F L+DLL A+A++ + + +KV E+ VKRL+ + V
Sbjct: 36 DNDEGKIVFFGGSNYTFDLDDLLAASAEILGKGAHVTTYKVAVEDTATVVVKRLEEVVVG 95
Query: 88 CDEFREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPW 147
EF + ++ + +++H N+ L Y +K +KL +Y Y S G++ +L+ + W
Sbjct: 96 RREFEQQMEIVGRIRHDNVAELKAYYYSKIDKLAVYSYYSQGNLFEMLHG--ESQVPLDW 153
Query: 148 KLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEP 207
+ RL IA G ARGLA I+ E + HGN+K SNI + K I + GL+ +
Sbjct: 154 ESRLRIAIGAARGLAIIH----EADDGKFVHGNIKSSNIFTNSKCYGCICDLGLTHITKS 209
Query: 208 DRGTFFSSHGYTAPE----KSLTEKGDVYSFGVILLELLTGQSIEV-----SRIDLVRWV 258
T S GY APE + T+ DVYSFGV+LLELLTG+S +DL W+
Sbjct: 210 LPQTTLRSSGYHAPEITDTRKSTQFSDVYSFGVVLLELLTGKSPASPLSLDENMDLASWI 269
Query: 259 RSMVREEWTGEVFDKE--VRENDHQGAFSLLNIALMCVSRSQENRPNFGEILETIEGV 314
RS+V +EWTGEVFD E ++ + +L I L CV+ ++RP+ I++ I+ +
Sbjct: 270 RSVVSKEWTGEVFDNELMMQMGIEEELVEMLQIGLACVALKPQDRPHITHIVKLIQDI 327
>At3g20190 receptor kinase like protein
Length = 679
Score = 170 bits (430), Expect = 1e-42
Identities = 112/336 (33%), Positives = 184/336 (54%), Gaps = 33/336 (9%)
Query: 2 GEIEVKKNNLDSPMKK------ATSEGRLKGGDNNNSELVFFVEDHERFKLEDLLRATAD 55
G+ +K N D K TS +G + ++L+F +D +RF L+DLLRA+A+
Sbjct: 317 GQDRTEKYNYDQSTDKDKAADSVTSYTSRRGAVPDQNKLLFLQDDIQRFDLQDLLRASAE 376
Query: 56 LRSENFWSSLFKVKFENNVEYAVKRLKNLQ-VSCDEFREILKQISKVKHQNILSLVGYRS 114
+ + S +K + VKR K++ V DEF E ++++ ++KH N+L +V Y
Sbjct: 377 VLGSGSFGSSYKTGINSGQMLVVKRYKHMNNVGRDEFHEHMRRLGRLKHPNLLPIVAYYY 436
Query: 115 TKEEKLIIYKYQSNGSVLNLLN-DYIARRKDFPWKLRLNIACGIARGLAFIYKKLEEGEV 173
+EEKL+I ++ N S+ + L+ ++ + W RL I G+A+GL +++ +L
Sbjct: 437 RREEKLLIAEFMPNRSLASHLHANHSVDQPGLDWPTRLKIIQGVAKGLGYLFNEL---TT 493
Query: 174 NSIPHGNLKLSNILLDDKNEALISEHGLSKFFEPDRGTFFSSH----GYTAPEKS----L 225
+IPHG+LK SN++LD+ E L++++ L ++ SH Y +PE S L
Sbjct: 494 LTIPHGHLKSSNVVLDESFEPLLTDYALRPVMNSEQ-----SHNLMISYKSPEYSLKGHL 548
Query: 226 TEKGDVYSFGVILLELLTGQSIE-------VSRIDLVRWVRSMVREEWTGEVFDKEV--R 276
T+K DV+ GV++LELLTG+ E + + LV WV +MV+E+ TG+VFDKE+ +
Sbjct: 549 TKKTDVWCLGVLILELLTGRFPENYLSQGYDANMSLVTWVSNMVKEKKTGDVFDKEMTGK 608
Query: 277 ENDHQGAFSLLNIALMCVSRSQENRPNFGEILETIE 312
+N +LL I L C +E R + +E IE
Sbjct: 609 KNCKAEMLNLLKIGLSCCEEDEERRMEMRDAVEKIE 644
>At1g50610 receptor-like protein kinase, putative
Length = 686
Score = 170 bits (430), Expect = 1e-42
Identities = 108/321 (33%), Positives = 178/321 (54%), Gaps = 25/321 (7%)
Query: 8 KNNLDSPMKKATSEGRLKGGDNNNSELVFFVEDHERFKLEDLLRATADLRSENFWSSLFK 67
KNN + T G + + L+F +D +RF L+DLLRA+A++ + + +K
Sbjct: 335 KNNKPAESVNHTRRGSMP---DPGGRLLFVRDDIQRFDLQDLLRASAEVLGSGTFGASYK 391
Query: 68 VKFENNVEYAVKRLKNLQ-VSCDEFREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQ 126
+ VKR K++ V DEF E ++++ ++ H NIL LV Y +EEKL++ ++
Sbjct: 392 AAISSGQTLVVKRYKHMNNVGRDEFHEHMRRLGRLNHPNILPLVAYYYRREEKLLVTEFM 451
Query: 127 SNGSVLNLLNDYIARRKDFPWKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNI 186
N S+ + L+ + D W RL I G+A+GL++++ +L +IPHG++K SNI
Sbjct: 452 PNSSLASHLHANNSAGLD--WITRLKIIKGVAKGLSYLFDEL---PTLTIPHGHMKSSNI 506
Query: 187 LLDDKNEALISEHGLSKFFEPDRGTFFSSHGYTAPE------KSLTEKGDVYSFGVILLE 240
+LDD E L++++ L + F + Y +PE + +T+K DV+ FGV++LE
Sbjct: 507 VLDDSFEPLLTDYALRPMMSSEHAHNFMT-AYKSPEYRPSKGQIITKKTDVWCFGVLILE 565
Query: 241 LLTGQSIE-------VSRIDLVRWVRSMVREEWTGEVFDKEV--RENDHQGAFSLLNIAL 291
+LTG+ E S + LV WV MV+E+ TG+VFDKE+ ++N +LL I L
Sbjct: 566 VLTGRFPENYLTQGYDSNMSLVTWVNDMVKEKKTGDVFDKEMKGKKNCKAEMINLLKIGL 625
Query: 292 MCVSRSQENRPNFGEILETIE 312
C +E R + E++E +E
Sbjct: 626 RCCEEEEERRMDMREVVEMVE 646
>At3g08680 putative protein kinase
Length = 640
Score = 169 bits (428), Expect = 2e-42
Identities = 102/297 (34%), Positives = 162/297 (54%), Gaps = 18/297 (6%)
Query: 32 SELVFFVEDHERFKLEDLLRATADLRSENFWSSLFKVKFENNVEYAVKRLKNLQVSCDEF 91
++LVFF F LEDLLRA+A++ + + + +K E VKRLK + EF
Sbjct: 322 NKLVFFEGSSYNFDLEDLLRASAEVLGKGSYGTTYKAILEEGTTVVVKRLKEVAAGKREF 381
Query: 92 REILKQISKVK-HQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLN-DYIARRKDFPWKL 149
+ ++ + ++ H N+ L Y +K+EKL++Y Y G+ LL+ + R W+
Sbjct: 382 EQQMEAVGRISPHVNVAPLRAYYFSKDEKLLVYDYYQGGNFSMLLHGNNEGGRAALDWET 441
Query: 150 RLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEPDR 209
RL I ARG++ I+ + HGN+K N+LL + +S+ G++
Sbjct: 442 RLRICLEAARGISHIHS----ASGAKLLHGNIKSPNVLLTQELHVCVSDFGIAPLMSHHT 497
Query: 210 GTFFSSHGYTAPE----KSLTEKGDVYSFGVILLELLTGQSIEVSR-----IDLVRWVRS 260
S GY APE + T+K DVYSFGV+LLE+LTG++ + +DL +WV+S
Sbjct: 498 LIPSRSLGYRAPEAIETRKHTQKSDVYSFGVLLLEMLTGKAAGKTTGHEEVVDLPKWVQS 557
Query: 261 MVREEWTGEVFDKEVRENDH---QGAFSLLNIALMCVSRSQENRPNFGEILETIEGV 314
+VREEWTGEVFD E+ + H + +L IA+ CVS+ ++RP+ E++ +E +
Sbjct: 558 VVREEWTGEVFDVELIKQQHNVEEEMVQMLQIAMACVSKHPDSRPSMEEVVNMMEEI 614
>At3g17840 receptor kinase, putative
Length = 647
Score = 168 bits (425), Expect = 5e-42
Identities = 110/332 (33%), Positives = 187/332 (56%), Gaps = 27/332 (8%)
Query: 3 EIEVKKNNLDSPMKKAT----SEGRLKGGDNNNS---ELVFFVEDHERFKLEDLLRATAD 55
EI +K +++P ++ S +K + N+S +LVFF + F LEDLLRA+A+
Sbjct: 310 EIPGEKAAVEAPENRSYVNEYSPSAVKAVEVNSSGMKKLVFFGNATKVFDLEDLLRASAE 369
Query: 56 LRSENFWSSLFKVKFENNVEYAVKRLKNLQVSCDEFREILKQISKVKHQNILSLVGYRST 115
+ + + + +K + AVKRLK++ ++ EF+E ++ + + H+N++ L Y +
Sbjct: 370 VLGKGTFGTAYKAVLDAVTLVAVKRLKDVTMADREFKEKIEVVGAMDHENLVPLRAYYYS 429
Query: 116 KEEKLIIYKYQSNGSVLNLLN-DYIARRKDFPWKLRLNIACGIARGLAFIYKKLEEGEVN 174
+EKL++Y + GS+ LL+ + A R W++R IA G ARGL +++ + ++
Sbjct: 430 GDEKLLVYDFMPMGSLSALLHGNKGAGRPPLNWEVRSGIALGAARGLDYLH---SQDPLS 486
Query: 175 SIPHGNLKLSNILLDDKNEALISEHGLSKFFEPDRGTFFSSHGYTAPE----KSLTEKGD 230
S HGN+K SNILL + ++A +S+ GL++ T + GY APE + +++K D
Sbjct: 487 S--HGNVKSSNILLTNSHDARVSDFGLAQLVSASSTTPNRATGYRAPEVTDPRRVSQKAD 544
Query: 231 VYSFGVILLELLTGQS-----IEVSRIDLVRWVRSMVREEWTGEVFDKEVREND-----H 280
VYSFGV+LLELLTG++ + +DL RWV S+ REEW EVFD E+ +
Sbjct: 545 VYSFGVVLLELLTGKAPSNSVMNEEGMDLARWVHSVAREEWRNEVFDSELMSIETVVSVE 604
Query: 281 QGAFSLLNIALMCVSRSQENRPNFGEILETIE 312
+ +L + + C + + RP E++ I+
Sbjct: 605 EEMAEMLQLGIDCTEQHPDKRPVMVEVVRRIQ 636
>At5g24100 receptor-like protein kinase
Length = 614
Score = 167 bits (422), Expect = 1e-41
Identities = 105/324 (32%), Positives = 174/324 (53%), Gaps = 23/324 (7%)
Query: 7 KKNNLDSPMKKATSEGRLKGGDNNN--SELVFFVEDHERFKLEDLLRATADLRSENFWSS 64
KK + + K E ++ ++ + ++++FF + F LEDLL A+A+ + +
Sbjct: 295 KKMPSEKEVSKLGKEKNIEDMEDKSEINKVMFFEGSNLAFNLEDLLIASAEFLGKGVFGM 354
Query: 65 LFKVKFENNVEYAVKRLKNLQVSCDEFREILKQISKVKHQNILSLVGYRSTKEEKLIIYK 124
+K E++ AVKRLK++ VS +F+ ++ + +KH+N+ L Y +KEEKL++Y
Sbjct: 355 TYKAVLEDSKVIAVKRLKDIVVSRKDFKHQMEIVGNIKHENVAPLRAYVCSKEEKLMVYD 414
Query: 125 YQSNGSVLNLLNDYIARRKDFP--WKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLK 182
Y SNGS+ L+ A P W+ RL G+A+GL I+ ++ HGN+K
Sbjct: 415 YDSNGSLSLRLHGKNADEGHVPLNWETRLRFMIGVAKGLGHIH-------TQNLAHGNIK 467
Query: 183 LSNILLDDKNEALISEHGLSKFFEPDRGTFFSSHG---YTAPE----KSLTEKGDVYSFG 235
SN+ ++ + ISE GL P S+ Y APE + T + D+YSFG
Sbjct: 468 SSNVFMNSEGYGCISEAGLPLLTNPVVRADSSARSVLRYRAPEVTDTRRSTPESDIYSFG 527
Query: 236 VILLELLTGQSIEVSR---IDLVRWVRSMVREEWTGEVFDKEVRENDHQGA--FSLLNIA 290
+++LE LTG+SI R IDLV WV ++ ++WTGEVFD E+ + + A +L +
Sbjct: 528 ILMLETLTGRSIMDDRKEGIDLVVWVNDVISKQWTGEVFDLELVKTPNVEAKLLQMLQLG 587
Query: 291 LMCVSRSQENRPNFGEILETIEGV 314
C + RP+ +++ET+E +
Sbjct: 588 TSCTAMVPAKRPDMVKVVETLEEI 611
>At3g51740 unknown protein
Length = 836
Score = 165 bits (418), Expect = 3e-41
Identities = 117/328 (35%), Positives = 175/328 (52%), Gaps = 30/328 (9%)
Query: 7 KKNNLDSPMKKATSEG---RLKGGDNNNSELVFFVEDHERFKLEDLLRATADLRSENFWS 63
+K+ D +K S G G +LV F + F +DLL ATA++ ++ +
Sbjct: 491 QKDGKDKTSEKTVSAGVAGTASAGGEMGGKLVHF-DGPFVFTADDLLCATAEIMGKSTYG 549
Query: 64 SLFKVKFENNVEYAVKRLKNLQVS-CDEFREILKQISKVKHQNILSLVGYR-STKEEKLI 121
+ +K E+ E AVKRL+ EF + + K++HQN+L+L Y K EKL+
Sbjct: 550 TAYKATLEDGNEVAVKRLREKTTKGVKEFEGEVTALGKIRHQNLLALRAYYLGPKGEKLL 609
Query: 122 IYKYQSNGSVLNLLNDYIARRKD--FPWKLRLNIACGIARGLAFIYKKLEEGEVNSIPHG 179
++ Y S GS+ L+ AR + PW+ R+ IA GI+RGLA ++ ++ H
Sbjct: 610 VFDYMSKGSLSAFLH---ARGPETLIPWETRMKIAKGISRGLAHLHSN------ENMIHE 660
Query: 180 NLKLSNILLDDKNEALISEHGLSKFFEPDRGTFF----SSHGYTAPE----KSLTEKGDV 231
NL SNILLD++ A I+++GLS+ T + GY APE K+ + K DV
Sbjct: 661 NLTASNILLDEQTNAHIADYGLSRLMTAAAATNVIATAGTLGYRAPEFSKIKNASAKTDV 720
Query: 232 YSFGVILLELLTGQSI--EVSRIDLVRWVRSMVREEWTGEVFDKEVRENDHQGAFSLLN- 288
YS G+I+LELLTG+S + +DL +WV S+V+EEWT EVFD E+ LLN
Sbjct: 721 YSLGIIILELLTGKSPGEPTNGMDLPQWVASIVKEEWTNEVFDLELMRETQSVGDELLNT 780
Query: 289 --IALMCVSRSQENRPNFGEILETIEGV 314
+AL CV S RP +++E +E +
Sbjct: 781 LKLALHCVDPSPAARPEANQVVEQLEEI 808
>At2g07040 putative leucine-rich repeat transmembrane protein kinase
Length = 647
Score = 164 bits (414), Expect = 9e-41
Identities = 100/306 (32%), Positives = 166/306 (53%), Gaps = 19/306 (6%)
Query: 30 NNSELVFFVEDHERFKLEDLLRATADLRSENFWSSLFKVKFENNVEYAVKRLKNLQ-VSC 88
+ ++L F +D +F+L+DLL+A+A++ + + +K N VKR K++
Sbjct: 317 HTTKLSFLRDDKGKFELQDLLKASAEILGSGCFGASYKTLLSNGSVMVVKRFKHMNSAGI 376
Query: 89 DEFREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIA-RRKDFPW 147
DEF+E +K++ ++ H+N+L +V Y KEEKL + + +NGS+ L+ + + + W
Sbjct: 377 DEFQEHMKRLGRLNHENLLPIVAYYYKKEEKLFVSDFVANGSLAAHLHGHKSLGQPSLDW 436
Query: 148 KLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEP 207
R NI G+ RGL +++K L PHG+LK SN+LL +K E L+ ++GL
Sbjct: 437 PTRFNIVKGVGRGLLYLHKNLPS---LMAPHGHLKSSNVLLSEKFEPLLMDYGLIPMINE 493
Query: 208 DRGTFFSSHGYTAPE----KSLTEKGDVYSFGVILLELLTGQSIE-------VSRIDLVR 256
+ Y +PE +T+K DV+ GV++LE+LTG+ +E S DL
Sbjct: 494 ESAQELMV-AYKSPEYVKQSRVTKKTDVWGLGVLILEILTGKLLESFSQVDKESEEDLAS 552
Query: 257 WVRSMVREEWTGEVFDKEVRENDHQGA--FSLLNIALMCVSRSQENRPNFGEILETIEGV 314
WVRS + EWT E+FD+E+ + + A +L+ I L C E R + E +E +E +
Sbjct: 553 WVRSSFKGEWTQELFDQEMGKTSNCEAHILNLMRIGLSCCEVDVEKRLDIREAVEKMEDL 612
Query: 315 MNAHDQ 320
M +Q
Sbjct: 613 MKEREQ 618
>At5g58300 receptor-like protein kinase
Length = 654
Score = 163 bits (413), Expect = 1e-40
Identities = 107/300 (35%), Positives = 164/300 (54%), Gaps = 26/300 (8%)
Query: 32 SELVFFVEDHERFKLEDLLRATADLRSENFWSSLFKVKFENNVEYAVKRLKNLQVSCDEF 91
++LVFF F LEDLLRA+A++ + + + +K E + VKRLK + EF
Sbjct: 339 NKLVFFNGCSYNFDLEDLLRASAEVLGKGSYGTAYKAVLEESTTVVVKRLKEVAAGKREF 398
Query: 92 REILKQISKV-KHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRK-DFPWKL 149
+ ++ IS+V H +++ L Y +K+EKL++ Y G++ +LL+ K W
Sbjct: 399 EQQMEIISRVGNHPSVVPLRAYYYSKDEKLMVCDYYPAGNLSSLLHGNRGSEKTPLDWDS 458
Query: 150 RLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKF----F 205
R+ I A+G+A L HGN+K SN+++ +++A IS+ GL+
Sbjct: 459 RVKITLSAAKGIA----HLHAAGGPKFSHGNIKSSNVIMKQESDACISDFGLTPLMAVPI 514
Query: 206 EPDRGTFFSSHGYTAPE----KSLTEKGDVYSFGVILLELLTGQSIEVS-----RIDLVR 256
P RG GY APE + T K DVYSFGV++LE+LTG+S S +DL R
Sbjct: 515 APMRGA-----GYRAPEVMETRKHTHKSDVYSFGVLILEMLTGKSPVQSPSRDDMVDLPR 569
Query: 257 WVRSMVREEWTGEVFDKEVR--ENDHQGAFSLLNIALMCVSRSQENRPNFGEILETIEGV 314
WV+S+VREEWT EVFD E+ +N + +L IA+ CV++ E RP +++ IE +
Sbjct: 570 WVQSVVREEWTSEVFDIELMRFQNIEEEMVQMLQIAMACVAQVPEVRPTMDDVVRMIEEI 629
>At4g31250 receptor kinase - like protein
Length = 951
Score = 161 bits (407), Expect = 6e-40
Identities = 110/337 (32%), Positives = 173/337 (50%), Gaps = 30/337 (8%)
Query: 25 KGGDNNNSELVFFVEDHERFKLEDLLRATADLRSENFWSSLFKVKFENNVEYAVKRLKNL 84
K GD +L F D ERF L+D+LRA+A++ + S +K + VKR + +
Sbjct: 342 KRGDQR--KLHFVRNDQERFTLQDMLRASAEVLGSGGFGSSYKAALSSGRAVVVKRFRFM 399
Query: 85 Q-VSCDEFREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRK 143
+ +EF + +K+I ++ H N+L L+ + KEEKL++ Y SNGS+ NLL+ I
Sbjct: 400 SNIGREEFYDHMKKIGRLSHPNLLPLIAFYYRKEEKLLVTNYISNGSLANLLHGNIMELS 459
Query: 144 D---------FPWKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEA 194
W +RL I G+ RGLA++Y+ + ++PHG+LK SN+LLD E
Sbjct: 460 KSNRTPGQVVLDWPIRLKIVRGVTRGLAYLYRVFPD---LNLPHGHLKSSNVLLDPNFEP 516
Query: 195 LISEHGLSKFFEPDRGTFFSSHGYTAPE----KSLTEKGDVYSFGVILLELLTG------ 244
L++++ L D+ F Y APE + + DV+S G+++LE+LTG
Sbjct: 517 LLTDYALVPVVNRDQSQQFMV-AYKAPEFTQQDRTSRRSDVWSLGILILEILTGKFPANY 575
Query: 245 -QSIEVSRIDLVRWVRSMVREEWTGEVFDKEVRENDHQGA--FSLLNIALMCVSRSQENR 301
+ + + +L WV S+ R EWT +VFDKE++ A LL I L C E R
Sbjct: 576 LRQGKGADDELAAWVESVARTEWTADVFDKEMKAGKEHEAQMLKLLKIGLRCCDWDIEKR 635
Query: 302 PNFGEILETIEGV-MNAHDQQQMELSASKCCSNGSNQ 337
E ++ IE V +A Q+ S+ S+G ++
Sbjct: 636 IELHEAVDRIEEVDRDAGGGQESVRSSYVTASDGDHR 672
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.135 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,223,026
Number of Sequences: 26719
Number of extensions: 352791
Number of successful extensions: 4015
Number of sequences better than 10.0: 882
Number of HSP's better than 10.0 without gapping: 672
Number of HSP's successfully gapped in prelim test: 210
Number of HSP's that attempted gapping in prelim test: 1171
Number of HSP's gapped (non-prelim): 945
length of query: 357
length of database: 11,318,596
effective HSP length: 100
effective length of query: 257
effective length of database: 8,646,696
effective search space: 2222200872
effective search space used: 2222200872
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC137079.13


