

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137077.10 + phase: 0
(247 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
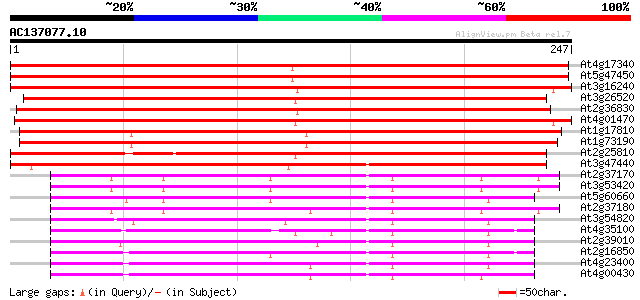
Score E
Sequences producing significant alignments: (bits) Value
At4g17340 membrane channel like protein 405 e-113
At5g47450 membrane channel protein-like; aquaporin (tonoplast in... 390 e-109
At3g16240 delta tonoplast integral protein (delta-TIP) 375 e-105
At3g26520 salt-stress induced tonoplast intrinsic protein 305 2e-83
At2g36830 putative aquaporin (tonoplast intrinsic protein gamma) 302 1e-82
At4g01470 putative water channel protein 278 1e-75
At1g17810 putative protein 260 4e-70
At1g73190 tonoplast intrinsic protein, alpha (alpha-TIP) 258 3e-69
At2g25810 putative aquaporin (tonoplast intrinsic protein) 256 6e-69
At3g47440 aquaporin-like protein 209 8e-55
At2g37170 aquaporin (plasma membrane intrinsic protein 2B) 144 4e-35
At3g53420 plasma membrane intrinsic protein 2a 143 7e-35
At5g60660 mipC protein - like (aquaporin) 143 1e-34
At2g37180 aquaporin (plasma membrane intrinsic protein 2C) 142 2e-34
At3g54820 aquaporin/MIP - like protein 142 2e-34
At4g35100 plasma membrane intrinsic protein PIP3 141 3e-34
At2g39010 putative aquaporin (water channel protein) 137 4e-33
At2g16850 putative aquaporin (plasma membrane intrinsic protein) 137 7e-33
At4g23400 water channel - like protein 127 4e-30
At4g00430 transmembrane protein 127 5e-30
>At4g17340 membrane channel like protein
Length = 250
Score = 405 bits (1041), Expect = e-113
Identities = 204/247 (82%), Positives = 223/247 (89%), Gaps = 1/247 (0%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
MVKI G+ DSFS ASLKAYLSEFIATL+FVFAGVGSA+A+ LTSDAALDPAGLVAVA
Sbjct: 1 MVKIEIGSVGDSFSVASLKAYLSEFIATLLFVFAGVGSALAFAKLTSDAALDPAGLVAVA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
VAHAFALFVGV+IAANISGGHLNPAVT GLA+GGNIT++TG FYWIAQ LGSIVA LLL
Sbjct: 61 VAHAFALFVGVSIAANISGGHLNPAVTLGLAVGGNITVITGFFYWIAQCLGSIVACLLLV 120
Query: 121 YVT-AKSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
+VT +SVPTHGVAAGL I G+V EI++TF LVYTVYATAADPKKGS+GTIAPIAIGF+
Sbjct: 121 FVTNGESVPTHGVAAGLGAIEGVVMEIVVTFALVYTVYATAADPKKGSLGTIAPIAIGFI 180
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGSY 239
VGANILAAGPFSGGSMNPARSFGPAVVSG+F+ WIYWVGPL+GG LAGLIYGDVFIGSY
Sbjct: 181 VGANILAAGPFSGGSMNPARSFGPAVVSGDFSQIWIYWVGPLVGGALAGLIYGDVFIGSY 240
Query: 240 TPAPASE 246
PAP +E
Sbjct: 241 APAPTTE 247
>At5g47450 membrane channel protein-like; aquaporin (tonoplast
intrinsic protein)-like
Length = 250
Score = 390 bits (1003), Expect = e-109
Identities = 196/247 (79%), Positives = 217/247 (87%), Gaps = 1/247 (0%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
MVKI G+ DSFS +SLKAYLSEFIATL+FVFAGVGSA+A+ LTSD ALDPAGLVA+A
Sbjct: 1 MVKIEVGSVGDSFSVSSLKAYLSEFIATLLFVFAGVGSAVAFAKLTSDGALDPAGLVAIA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
+AHAFALFVGV+IAANISGGHLNPAVT GLAIGGNIT++TG FYWIAQ LGSIVA LLL
Sbjct: 61 IAHAFALFVGVSIAANISGGHLNPAVTLGLAIGGNITLITGFFYWIAQCLGSIVACLLLV 120
Query: 121 YVT-AKSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
+VT KSVPTHGV+AGL + G+V EI++TF LVYTVYATAADPKKGS+GTIAPIAIGF+
Sbjct: 121 FVTNGKSVPTHGVSAGLGAVEGVVMEIVVTFALVYTVYATAADPKKGSLGTIAPIAIGFI 180
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGSY 239
VGANILAAGPFSGGSMNPARSFGPAVVSG+ + WIYWVGPL+GG LAGLIYGDVFIGSY
Sbjct: 181 VGANILAAGPFSGGSMNPARSFGPAVVSGDLSQIWIYWVGPLVGGALAGLIYGDVFIGSY 240
Query: 240 TPAPASE 246
E
Sbjct: 241 EAVETRE 247
>At3g16240 delta tonoplast integral protein (delta-TIP)
Length = 250
Score = 375 bits (964), Expect = e-105
Identities = 187/250 (74%), Positives = 215/250 (85%), Gaps = 3/250 (1%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
M + FG+FDDSFS ASL+AYL+EFI+TL+FVFAGVGSAIAY LTSDAALD GLVA+A
Sbjct: 1 MAGVAFGSFDDSFSLASLRAYLAEFISTLLFVFAGVGSAIAYAKLTSDAALDTPGLVAIA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
V H FALFV VAI ANISGGH+NPAVTFGLA+GG IT++TG+FYWIAQLLGS A LL
Sbjct: 61 VCHGFALFVAVAIGANISGGHVNPAVTFGLAVGGQITVITGVFYWIAQLLGSTAACFLLK 120
Query: 121 YVTAK-SVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
YVT +VPTH VAAGL I G+V EIIITF LVYTVYATAADPKKGS+GTIAP+AIG +
Sbjct: 121 YVTGGLAVPTHSVAAGLGSIEGVVMEIIITFALVYTVYATAADPKKGSLGTIAPLAIGLI 180
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGS- 238
VGANILAAGPFSGGSMNPARSFGPAV +G+F+ +W+YWVGPLIGGGLAGLIYG+VF+GS
Sbjct: 181 VGANILAAGPFSGGSMNPARSFGPAVAAGDFSGHWVYWVGPLIGGGLAGLIYGNVFMGSS 240
Query: 239 -YTPAPASEF 247
+ P +++F
Sbjct: 241 EHVPLASADF 250
>At3g26520 salt-stress induced tonoplast intrinsic protein
Length = 253
Score = 305 bits (780), Expect = 2e-83
Identities = 147/231 (63%), Positives = 185/231 (79%), Gaps = 1/231 (0%)
Query: 7 GTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVAVAHAFA 66
G ++ + +L+A L+EFI+TLIFVFAG GS IA+N +T + A P+GLVA A+AHAF
Sbjct: 10 GVQEEVYHPNALRAALAEFISTLIFVFAGSGSGIAFNKITDNGATTPSGLVAAALAHAFG 69
Query: 67 LFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLNYVTA-K 125
LFV V++ ANISGGH+NPAVTFG+ +GGNIT+L G+ YWIAQLLGS+ A LL++ T +
Sbjct: 70 LFVAVSVGANISGGHVNPAVTFGVLLGGNITLLRGILYWIAQLLGSVAACFLLSFATGGE 129
Query: 126 SVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFVVGANIL 185
+P G++AG+ + LVFEI++TFGLVYTVYATA DPK GS+GTIAPIAIGF+VGANIL
Sbjct: 130 PIPAFGLSAGVGSLNALVFEIVMTFGLVYTVYATAVDPKNGSLGTIAPIAIGFIVGANIL 189
Query: 186 AAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFI 236
A G FSG SMNPA +FGPAVVS + ++W+YW GPLIGGGLAG+IY VFI
Sbjct: 190 AGGAFSGASMNPAVAFGPAVVSWTWTNHWVYWAGPLIGGGLAGIIYDFVFI 240
>At2g36830 putative aquaporin (tonoplast intrinsic protein gamma)
Length = 251
Score = 302 bits (774), Expect = 1e-82
Identities = 150/236 (63%), Positives = 184/236 (77%), Gaps = 1/236 (0%)
Query: 4 IGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVAVAH 63
I G D++ +LKA L+EFI+TLIFV AG GS +A+N LT + A P+GLVA AVAH
Sbjct: 6 IAIGRPDEATRPDALKAALAEFISTLIFVVAGSGSGMAFNKLTENGATTPSGLVAAAVAH 65
Query: 64 AFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLNYVT 123
AF LFV V++ ANISGGH+NPAVTFG IGGNIT+L G+ YWIAQLLGS+VA L+L + T
Sbjct: 66 AFGLFVAVSVGANISGGHVNPAVTFGAFIGGNITLLRGILYWIAQLLGSVVACLILKFAT 125
Query: 124 AK-SVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFVVGA 182
+VP G++AG+ + VFEI++TFGLVYTVYATA DPK GS+GTIAPIAIGF+VGA
Sbjct: 126 GGLAVPAFGLSAGVGVLNAFVFEIVMTFGLVYTVYATAIDPKNGSLGTIAPIAIGFIVGA 185
Query: 183 NILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGS 238
NILA G FSG SMNPA +FGPAVVS + ++W+YW GPL+GGG+AGLIY FI +
Sbjct: 186 NILAGGAFSGASMNPAVAFGPAVVSWTWTNHWVYWAGPLVGGGIAGLIYEVFFINT 241
Score = 35.4 bits (80), Expect = 0.028
Identities = 26/65 (40%), Positives = 33/65 (50%), Gaps = 9/65 (13%)
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFA--DNWIYWVGPLIGGGLAGLIYGDVFIG 237
VGANI SGG +NPA +FG A + GN +YW+ L+G +A LI G
Sbjct: 75 VGANI------SGGHVNPAVTFG-AFIGGNITLLRGILYWIAQLLGSVVACLILKFATGG 127
Query: 238 SYTPA 242
PA
Sbjct: 128 LAVPA 132
>At4g01470 putative water channel protein
Length = 252
Score = 278 bits (712), Expect = 1e-75
Identities = 137/248 (55%), Positives = 178/248 (71%), Gaps = 3/248 (1%)
Query: 3 KIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVAVA 62
+I GT ++ +++A +EF + +IFVFAG GS +AY LT D PAGLVA +++
Sbjct: 5 RIAIGTPGEASRPDAIRAAFAEFFSMVIFVFAGQGSGMAYGKLTGDGPATPAGLVAASLS 64
Query: 63 HAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLNYV 122
HAFALFV V++ AN+SGGH+NPAVTFG IGGNIT+L + YWIAQLLG++VA LLL
Sbjct: 65 HAFALFVAVSVGANVSGGHVNPAVTFGAFIGGNITLLRAILYWIAQLLGAVVACLLLKVS 124
Query: 123 TA-KSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFVVG 181
T ++ G+ P +VFEI++TFGLVYTVYATA DPKKG IG IAP+AIG +VG
Sbjct: 125 TGGMETAAFSLSYGVTPWNAVVFEIVMTFGLVYTVYATAVDPKKGDIGIIAPLAIGLIVG 184
Query: 182 ANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGS--Y 239
ANIL G F G SMNPA SFGPAVVS + ++W+YWVGP IG +A ++Y +FIGS +
Sbjct: 185 ANILVGGAFDGASMNPAVSFGPAVVSWIWTNHWVYWVGPFIGAAIAAIVYDTIFIGSNGH 244
Query: 240 TPAPASEF 247
P P+++F
Sbjct: 245 EPLPSNDF 252
>At1g17810 putative protein
Length = 267
Score = 260 bits (665), Expect = 4e-70
Identities = 132/245 (53%), Positives = 172/245 (69%), Gaps = 6/245 (2%)
Query: 5 GFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALD-----PAGLVAV 59
GFG D++ S++A L+EF++T +FVFAG GS +A + L D A P GLV V
Sbjct: 10 GFGRADEATHPDSIRATLAEFLSTFVFVFAGEGSILALDKLYWDTAAHTGTNTPGGLVLV 69
Query: 60 AVAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLL 119
A+AHA ALF V+ A N+SGGH+NPAVTF IGG I+++ ++YW+AQL+G+I+A LLL
Sbjct: 70 ALAHALALFAAVSAAINVSGGHVNPAVTFAALIGGRISVIRAIYYWVAQLIGAILACLLL 129
Query: 120 NYVTAKSVPT-HGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGF 178
T P VA+G++ + GL+ EII+TF LVY VY+TA DPK+GSIG IAP+AIG
Sbjct: 130 RLATNGLRPVGFHVASGVSELHGLLMEIILTFALVYVVYSTAIDPKRGSIGIIAPLAIGL 189
Query: 179 VVGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGS 238
+VGANIL GPF G SMNPAR+FGPA+V ++++WIYWVGP IGG LA LIY + I S
Sbjct: 190 IVGANILVGGPFDGASMNPARAFGPALVGWRWSNHWIYWVGPFIGGALAALIYEYMIIPS 249
Query: 239 YTPAP 243
P
Sbjct: 250 VNEPP 254
>At1g73190 tonoplast intrinsic protein, alpha (alpha-TIP)
Length = 268
Score = 258 bits (658), Expect = 3e-69
Identities = 130/243 (53%), Positives = 170/243 (69%), Gaps = 6/243 (2%)
Query: 5 GFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALD-----PAGLVAV 59
GFG D++ S++A L+EF++T +FVFA GS ++ + L + A P GL+ V
Sbjct: 10 GFGRADEATHPDSIRATLAEFLSTFVFVFAAEGSILSLDKLYWEHAAHAGTNTPGGLILV 69
Query: 60 AVAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLL 119
A+AHAFALF V+ A N+SGGH+NPAVTFG +GG +T + ++YWIAQLLG+I+A LLL
Sbjct: 70 ALAHAFALFAAVSAAINVSGGHVNPAVTFGALVGGRVTAIRAIYYWIAQLLGAILACLLL 129
Query: 120 NYVTAKSVPT-HGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGF 178
T P +A+G+ + GLV EII+TFGLVY VY+T DPK+GS+G IAP+AIG
Sbjct: 130 RLTTNGMRPVGFRLASGVGAVNGLVLEIILTFGLVYVVYSTLIDPKRGSLGIIAPLAIGL 189
Query: 179 VVGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGS 238
+VGANIL GPFSG SMNPAR+FGPA+V + D+WIYWVGP IG LA LIY + I +
Sbjct: 190 IVGANILVGGPFSGASMNPARAFGPALVGWRWHDHWIYWVGPFIGSALAALIYEYMVIPT 249
Query: 239 YTP 241
P
Sbjct: 250 EPP 252
Score = 39.3 bits (90), Expect = 0.002
Identities = 29/95 (30%), Positives = 43/95 (44%), Gaps = 8/95 (8%)
Query: 51 LDPA-GLVAVAVAHAFALFVG--VAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIA 107
+DP G + + A L VG + + SG +NPA FG A+ G YW+
Sbjct: 172 IDPKRGSLGIIAPLAIGLIVGANILVGGPFSGASMNPARAFGPALVG-WRWHDHWIYWVG 230
Query: 108 QLLGSIVASLLLNYVTAKSVP----THGVAAGLNP 138
+GS +A+L+ Y+ + P HGV L P
Sbjct: 231 PFIGSALAALIYEYMVIPTEPPTHHAHGVHQPLAP 265
>At2g25810 putative aquaporin (tonoplast intrinsic protein)
Length = 249
Score = 256 bits (655), Expect = 6e-69
Identities = 132/237 (55%), Positives = 168/237 (70%), Gaps = 5/237 (2%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
M KI G ++ +KA + EFI T +FVFAGVGSA+A + L + + GL AVA
Sbjct: 1 MKKIELGHHSEAAKPDCIKALIVEFITTFLFVFAGVGSAMATDSLVGNTLV---GLFAVA 57
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
VAHAF + V ++ A +ISGGHLNPAVT GL +GG+I++ YWI QLL S A LL+
Sbjct: 58 VAHAFVVAVMIS-AGHISGGHLNPAVTLGLLLGGHISVFRAFLYWIDQLLASSAACFLLS 116
Query: 121 YVTA-KSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
Y+T P H +A+G++ G+++EII+TF L++TVYAT DPKKGS+ P+ GFV
Sbjct: 117 YLTGGMGTPVHTLASGVSYTQGIIWEIILTFSLLFTVYATIVDPKKGSLDGFGPLLTGFV 176
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFI 236
VGANILA G FSG SMNPARSFGPA+VSGN+ D+W+YWVGPLIGGGLAG IY +V I
Sbjct: 177 VGANILAGGAFSGASMNPARSFGPALVSGNWTDHWVYWVGPLIGGGLAGFIYENVLI 233
>At3g47440 aquaporin-like protein
Length = 256
Score = 209 bits (533), Expect = 8e-55
Identities = 108/238 (45%), Positives = 156/238 (65%), Gaps = 3/238 (1%)
Query: 1 MVKIGFGT-FDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAV 59
M+ F + F S +L+ Y+SEFI+T FV A VGS ++ L + P G++
Sbjct: 4 MIPTSFSSKFQGVLSMNALRCYVSEFISTFFFVLAAVGSVMSSRKLMAGDVSGPFGVLIP 63
Query: 60 AVAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLL 119
A+A+A AL V I+ N+SGGH+NPAVTF +A+ G I++ T +FYW +Q++ S++A L+L
Sbjct: 64 AIANALALSSSVYISWNVSGGHVNPAVTFAMAVAGRISVPTAMFYWTSQMIASVMACLVL 123
Query: 120 NY-VTAKSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGF 178
V + VP + +A + V E ++ F LVYTV+ TA+DP++G + PI IGF
Sbjct: 124 KVTVMEQHVPIYKIAGEMTGFGASVLEGVLAFVLVYTVF-TASDPRRGLPLAVGPIFIGF 182
Query: 179 VVGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFI 236
V GAN+LAAGPFSGGSMNPA +FG A+V G+F + +YWVGPL+GG A L+Y +V +
Sbjct: 183 VAGANVLAAGPFSGGSMNPACAFGSAMVYGSFKNQAVYWVGPLLGGATAALVYDNVVV 240
Score = 38.9 bits (89), Expect = 0.003
Identities = 29/89 (32%), Positives = 39/89 (43%), Gaps = 4/89 (4%)
Query: 50 ALDPAGLVAVAVAHAFALFVG---VAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWI 106
A DP + +AV F FV V A SGG +NPA FG A+ + YW+
Sbjct: 164 ASDPRRGLPLAVGPIFIGFVAGANVLAAGPFSGGSMNPACAFGSAMVYG-SFKNQAVYWV 222
Query: 107 AQLLGSIVASLLLNYVTAKSVPTHGVAAG 135
LLG A+L+ + V G + G
Sbjct: 223 GPLLGGATAALVYDNVVVPVEDDRGSSTG 251
>At2g37170 aquaporin (plasma membrane intrinsic protein 2B)
Length = 285
Score = 144 bits (363), Expect = 4e-35
Identities = 93/247 (37%), Positives = 137/247 (54%), Gaps = 24/247 (9%)
Query: 19 KAYLSEFIATLIFVFAGVGSAIAYN--DLTSDAALDPAGLVAVAVAHAFA--LFVGVAIA 74
+A ++EF+ATL+F++ V + I Y T +D G+ + +A AF +F+ V
Sbjct: 37 RAVIAEFVATLLFLYITVLTVIGYKIQSDTKAGGVDCGGVGILGIAWAFGGMIFILVYCT 96
Query: 75 ANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSI-----VASLLLNYVTAKSVPT 129
A ISGGH+NPAVTFGL + ++++ + Y +AQ LG+I V + +Y
Sbjct: 97 AGISGGHINPAVTFGLFLARKVSLIRAVLYMVAQCLGAICGVGFVKAFQSSYYDRYGGGA 156
Query: 130 HGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGS----IGTIAPIAIGFVVGANIL 185
+ +A G N GL EII TF LVYTV+ +A DPK+ + + +AP+ IGF V L
Sbjct: 157 NSLADGYNTGTGLAAEIIGTFVLVYTVF-SATDPKRNARDSHVPVLAPLPIGFAVFMVHL 215
Query: 186 AAGPFSGGSMNPARSFGPAVV---SGNFADNWIYWVGPLIGGGLAGLIY-------GDVF 235
A P +G +NPARSFG AV+ S + D+WI+WVGP IG +A + G
Sbjct: 216 ATIPITGTGINPARSFGAAVIYNKSKPWDDHWIFWVGPFIGAAIAAFYHQFVLRASGSKS 275
Query: 236 IGSYTPA 242
+GS+ A
Sbjct: 276 LGSFRSA 282
>At3g53420 plasma membrane intrinsic protein 2a
Length = 287
Score = 143 bits (361), Expect = 7e-35
Identities = 95/247 (38%), Positives = 137/247 (55%), Gaps = 24/247 (9%)
Query: 19 KAYLSEFIATLIFVFAGVGSAIAYN--DLTSDAALDPAGLVAVAVAHAFA--LFVGVAIA 74
+A ++EF+ATL+F++ V + I Y T +D G+ + +A AF +F+ V
Sbjct: 39 RAVIAEFVATLLFLYITVLTVIGYKIQSDTDAGGVDCGGVGILGIAWAFGGMIFILVYCT 98
Query: 75 ANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSI-----VASLLLNYVTAKSVPT 129
A ISGGH+NPAVTFGL + +++ L Y IAQ LG+I V + +Y T
Sbjct: 99 AGISGGHINPAVTFGLFLARKVSLPRALLYIIAQCLGAICGVGFVKAFQSSYYTRYGGGA 158
Query: 130 HGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGS----IGTIAPIAIGFVVGANIL 185
+ +A G + GL EII TF LVYTV+ +A DPK+ + + +AP+ IGF V L
Sbjct: 159 NSLADGYSTGTGLAAEIIGTFVLVYTVF-SATDPKRSARDSHVPVLAPLPIGFAVFMVHL 217
Query: 186 AAGPFSGGSMNPARSFGPAVV---SGNFADNWIYWVGPLIGGGLAGLIY-------GDVF 235
A P +G +NPARSFG AV+ S + D+WI+WVGP IG +A + G
Sbjct: 218 ATIPITGTGINPARSFGAAVIYNKSKPWDDHWIFWVGPFIGAAIAAFYHQFVLRASGSKS 277
Query: 236 IGSYTPA 242
+GS+ A
Sbjct: 278 LGSFRSA 284
>At5g60660 mipC protein - like (aquaporin)
Length = 291
Score = 143 bits (360), Expect = 1e-34
Identities = 90/229 (39%), Positives = 131/229 (56%), Gaps = 17/229 (7%)
Query: 19 KAYLSEFIATLIFVFAGVGSAIAYNDLTSDAA--LDPAGLVAVAVAHAFA--LFVGVAIA 74
+A ++EF+ATL+F++ + + I Y T A +D G+ + +A AF +FV V
Sbjct: 39 RAVIAEFVATLLFLYVSILTVIGYKAQTDATAGGVDCGGVGILGIAWAFGGMIFVLVYCT 98
Query: 75 ANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSI-----VASLLLNYVTAKSVPT 129
A ISGGH+NPAVT GL + ++++ + Y +AQ LG+I V + +Y T
Sbjct: 99 AGISGGHINPAVTVGLFLARKVSLVRTVLYIVAQCLGAICGCGFVKAFQSSYYTRYGGGA 158
Query: 130 HGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGS----IGTIAPIAIGFVVGANIL 185
+ +A G N GL EII TF LVYTV+ +A DPK+ + + +AP+ IGF V L
Sbjct: 159 NELADGYNKGTGLGAEIIGTFVLVYTVF-SATDPKRNARDSHVPVLAPLPIGFAVFMVHL 217
Query: 186 AAGPFSGGSMNPARSFGPAVVSGN---FADNWIYWVGPLIGGGLAGLIY 231
A P +G +NPARSFG AV+ N + D WI+WVGP+IG A +
Sbjct: 218 ATIPITGTGINPARSFGAAVIYNNEKAWDDQWIFWVGPMIGAAAAAFYH 266
>At2g37180 aquaporin (plasma membrane intrinsic protein 2C)
Length = 285
Score = 142 bits (358), Expect = 2e-34
Identities = 92/247 (37%), Positives = 136/247 (54%), Gaps = 24/247 (9%)
Query: 19 KAYLSEFIATLIFVFAGVGSAIAYN--DLTSDAALDPAGLVAVAVAHAFA--LFVGVAIA 74
+A ++EF+ATL+F++ V + I Y T +D G+ + +A AF +F+ V
Sbjct: 37 RAVIAEFVATLLFLYVTVLTVIGYKIQSDTKAGGVDCGGVGILGIAWAFGGMIFILVYCT 96
Query: 75 ANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLNYVTAKSVPTHG--- 131
A ISGGH+NPAVTFGL + ++++ + Y +AQ LG+I + + +G
Sbjct: 97 AGISGGHINPAVTFGLFLARKVSLIRAVLYMVAQCLGAICGVGFVKAFQSSHYVNYGGGA 156
Query: 132 --VAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGS----IGTIAPIAIGFVVGANIL 185
+A G N GL EII TF LVYTV+ +A DPK+ + + +AP+ IGF V L
Sbjct: 157 NFLADGYNTGTGLAAEIIGTFVLVYTVF-SATDPKRNARDSHVPVLAPLPIGFAVFMVHL 215
Query: 186 AAGPFSGGSMNPARSFGPAVV---SGNFADNWIYWVGPLIGGGLAGLIY-------GDVF 235
A P +G +NPARSFG AV+ S + D+WI+WVGP IG +A + G
Sbjct: 216 ATIPITGTGINPARSFGAAVIFNKSKPWDDHWIFWVGPFIGATIAAFYHQFVLRASGSKS 275
Query: 236 IGSYTPA 242
+GS+ A
Sbjct: 276 LGSFRSA 282
>At3g54820 aquaporin/MIP - like protein
Length = 286
Score = 142 bits (357), Expect = 2e-34
Identities = 86/230 (37%), Positives = 130/230 (56%), Gaps = 19/230 (8%)
Query: 19 KAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDP-----AGLVAVAVAHAFALFVGVAI 73
+A ++EFIATL+F++ + + I Y T D AL+P G++ +A A +F+ V
Sbjct: 38 RALIAEFIATLLFLYVTIMTVIGYKSQT-DPALNPDQCTGVGVLGIAWAFGGMIFILVYC 96
Query: 74 AANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN-----YVTAKSVP 128
A ISGGH+NPAVTFGL + +T++ + Y +AQ LG+I L+ Y T
Sbjct: 97 TAGISGGHINPAVTFGLLLARKVTLVRAVMYMVAQCLGAICGVALVKAFQSAYFTRYGGG 156
Query: 129 THGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGS----IGTIAPIAIGFVVGANI 184
+G++ G + G+ EII TF LVYTV+ +A DPK+ + + +AP+ IGF V
Sbjct: 157 ANGLSDGYSIGTGVAAEIIGTFVLVYTVF-SATDPKRSARDSHVPVLAPLPIGFAVFIVH 215
Query: 185 LAAGPFSGGSMNPARSFGPAVVSGN---FADNWIYWVGPLIGGGLAGLIY 231
LA P +G +NPARS G A++ + +WI+WVGP G +A +
Sbjct: 216 LATIPITGTGINPARSLGAAIIYNKDKAWDHHWIFWVGPFAGAAIAAFYH 265
>At4g35100 plasma membrane intrinsic protein PIP3
Length = 280
Score = 141 bits (356), Expect = 3e-34
Identities = 94/228 (41%), Positives = 129/228 (56%), Gaps = 22/228 (9%)
Query: 19 KAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVAVAHAFALFVGVAIAANIS 78
+A ++EFIATL+F++ V + I + T D GL+ +A A +FV V A IS
Sbjct: 38 RALIAEFIATLLFLYVTVATVIGHKKQTGPC--DGVGLLGIAWAFGGMIFVLVYCTAGIS 95
Query: 79 GGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLNYVTA-KSVPTHGVAAGLN 137
GGH+NPAVTFGL + ++++ L Y IAQ LG+I + +V A P + + G N
Sbjct: 96 GGHINPAVTFGLFLARKVSLVRALGYMIAQCLGAICG---VGFVKAFMKTPYNTLGGGAN 152
Query: 138 PIA-------GLVFEIIITFGLVYTVYATAADPKKGS----IGTIAPIAIGFVVGANILA 186
+A L EII TF LVYTV+ +A DPK+ + I +AP+ IGF V LA
Sbjct: 153 TVADGYSKGTALGAEIIGTFVLVYTVF-SATDPKRSARDSHIPVLAPLPIGFAVFMVHLA 211
Query: 187 AGPFSGGSMNPARSFGPAVVSGN---FADNWIYWVGPLIGGGLAGLIY 231
P +G +NPARSFG AV+ N + D WI+WVGP + G LA Y
Sbjct: 212 TIPITGTGINPARSFGAAVIYNNEKAWDDQWIFWVGPFL-GALAAAAY 258
>At2g39010 putative aquaporin (water channel protein)
Length = 289
Score = 137 bits (346), Expect = 4e-33
Identities = 86/229 (37%), Positives = 126/229 (54%), Gaps = 17/229 (7%)
Query: 19 KAYLSEFIATLIFVFAGVGSAIAYNDLTS----DAALDPAGLVAVAVAHAFALFVGVAIA 74
+A ++EFIATL+F++ V + I + T A GL+ ++ A +F+ V
Sbjct: 38 RAVIAEFIATLLFLYVTVLTVIGFKSQTDINAGGGACASVGLLGISWAFGGMIFILVYCT 97
Query: 75 ANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLNYVTAKSVPTHGVAA 134
A ISGGH+NPAVTFGL + ++++ + Y +AQ LG+ L+ + +G A
Sbjct: 98 AGISGGHINPAVTFGLFLASKVSLVRAVSYMVAQCLGATCGVGLVKVFQSTYYNRYGGGA 157
Query: 135 -----GLNPIAGLVFEIIITFGLVYTVYATAADPKKGS----IGTIAPIAIGFVVGANIL 185
G N G+ EII TF LVYTV+ +A DPK+ + I +AP+ IGF V L
Sbjct: 158 NMLSDGYNVGVGVGAEIIGTFVLVYTVF-SATDPKRNARDSHIPVLAPLPIGFSVFMVHL 216
Query: 186 AAGPFSGGSMNPARSFGPAVVSGN---FADNWIYWVGPLIGGGLAGLIY 231
A P +G +NPARSFG AV+ N + D WI+WVGP +G +A +
Sbjct: 217 ATIPITGTGINPARSFGAAVIYNNQKAWDDQWIFWVGPFVGAAIAAFYH 265
>At2g16850 putative aquaporin (plasma membrane intrinsic protein)
Length = 278
Score = 137 bits (344), Expect = 7e-33
Identities = 88/225 (39%), Positives = 127/225 (56%), Gaps = 16/225 (7%)
Query: 19 KAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVAVAHAFALFVGVAIAANIS 78
+A ++EFIATL+F++ V + I + + T GL+ +A A +FV V A IS
Sbjct: 36 RAIIAEFIATLLFLYVTVATVIGHKNQTGPCG--GVGLLGIAWAFGGMIFVLVYCTAGIS 93
Query: 79 GGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSI-----VASLLLNYVTAKSVPTHGVA 133
GGH+NPAVTFGL + +++ + Y +AQ LG+I V + ++ + VA
Sbjct: 94 GGHINPAVTFGLFLARKVSLPRAVAYMVAQCLGAICGVGLVKAFMMTPYKRLGGGANTVA 153
Query: 134 AGLNPIAGLVFEIIITFGLVYTVYATAADPKKGS----IGTIAPIAIGFVVGANILAAGP 189
G + L EII TF LVYTV+ +A DPK+ + + +AP+ IGF V LA P
Sbjct: 154 DGYSTGTALGAEIIGTFVLVYTVF-SATDPKRSARDSHVPVLAPLPIGFAVFMVHLATIP 212
Query: 190 FSGGSMNPARSFGPAVVSGN---FADNWIYWVGPLIGGGLAGLIY 231
+G +NPARSFG AV+ N + D+WI+WVGP + G LA Y
Sbjct: 213 ITGTGINPARSFGAAVIYNNEKAWDDHWIFWVGPFV-GALAAAAY 256
>At4g23400 water channel - like protein
Length = 287
Score = 127 bits (320), Expect = 4e-30
Identities = 83/225 (36%), Positives = 123/225 (53%), Gaps = 15/225 (6%)
Query: 19 KAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVAVAHAFALFVGVAIAANIS 78
+A ++EFIAT +F++ V + + + A G+ +A A +F V A IS
Sbjct: 53 RAGIAEFIATFLFLYVTVLTVMGVKRAPNMCA--SVGIQGIAWAFGGMIFALVYCTAGIS 110
Query: 79 GGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLNYVTAKSVPTHG-----VA 133
GGH+NPAVTFGL + +++ LFY + Q LG+I + ++ T+G VA
Sbjct: 111 GGHINPAVTFGLFLARKLSLTRALFYIVMQCLGAICGAGVVKGFQPGLYQTNGGGANVVA 170
Query: 134 AGLNPIAGLVFEIIITFGLVYTVYATAADPKKGS----IGTIAPIAIGFVVGANILAAGP 189
G +GL EI+ TF LVYTV+ +A D K+ + + +AP+ IGF V LA P
Sbjct: 171 HGYTKGSGLGAEIVGTFVLVYTVF-SATDAKRSARDSHVPILAPLPIGFAVFLVHLATIP 229
Query: 190 FSGGSMNPARSFGPAVVSGN---FADNWIYWVGPLIGGGLAGLIY 231
+G +NPARS G A++ + D+WI+WVGP IG LA L +
Sbjct: 230 ITGTGINPARSLGAAIIYNKDHAWDDHWIFWVGPFIGAALAALYH 274
>At4g00430 transmembrane protein
Length = 287
Score = 127 bits (319), Expect = 5e-30
Identities = 84/225 (37%), Positives = 123/225 (54%), Gaps = 15/225 (6%)
Query: 19 KAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVAVAHAFALFVGVAIAANIS 78
+A ++EFIAT +F++ V + + + A G+ +A A +F V A IS
Sbjct: 53 RAGIAEFIATFLFLYITVLTVMGVKRAPNMCA--SVGIQGIAWAFGGMIFALVYCTAGIS 110
Query: 79 GGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLNYVTAKSVPTHG-----VA 133
GGH+NPAVTFGL + +++ +FY I Q LG+I + ++ T G VA
Sbjct: 111 GGHINPAVTFGLFLARKLSLTRAVFYMIMQCLGAICGAGVVKGFQPTPYQTLGGGANTVA 170
Query: 134 AGLNPIAGLVFEIIITFGLVYTVYATAADPKKGS----IGTIAPIAIGFVVGANILAAGP 189
G +GL EII TF LVYTV+ +A D K+ + + +AP+ IGF V LA P
Sbjct: 171 HGYTKGSGLGAEIIGTFVLVYTVF-SATDAKRSARDSHVPILAPLPIGFAVFLVHLATIP 229
Query: 190 FSGGSMNPARSFGPAVV---SGNFADNWIYWVGPLIGGGLAGLIY 231
+G +NPARS G A++ ++ D+WI+WVGP IG LA L +
Sbjct: 230 ITGTGINPARSLGAAIIYNKDHSWDDHWIFWVGPFIGAALAALYH 274
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.323 0.141 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,281,858
Number of Sequences: 26719
Number of extensions: 217809
Number of successful extensions: 708
Number of sequences better than 10.0: 53
Number of HSP's better than 10.0 without gapping: 36
Number of HSP's successfully gapped in prelim test: 17
Number of HSP's that attempted gapping in prelim test: 515
Number of HSP's gapped (non-prelim): 65
length of query: 247
length of database: 11,318,596
effective HSP length: 97
effective length of query: 150
effective length of database: 8,726,853
effective search space: 1309027950
effective search space used: 1309027950
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC137077.10


