

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136973.16 + phase: 0 /partial
(373 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
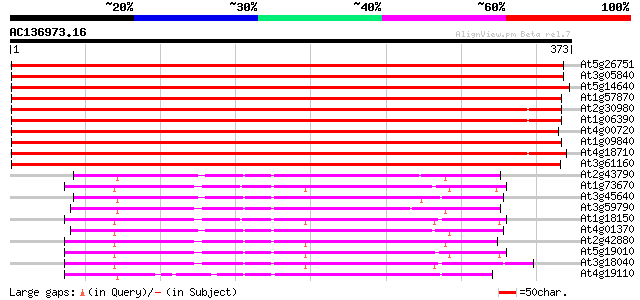
Score E
Sequences producing significant alignments: (bits) Value
At5g26751 shaggy-like kinase alpha 691 0.0
At3g05840 shaggy related protein kinase, ASK-GAMMA 687 0.0
At5g14640 protein kinase MSK-3 - like 681 0.0
At1g57870 protein kinase, putative 612 e-175
At2g30980 putative shaggy-like protein kinase dzeta 608 e-174
At1g06390 putative shaggy kinase iota protein 608 e-174
At4g00720 Shaggy related protein kinase tetha 607 e-174
At1g09840 shaggy-like protien kinase, kappa 605 e-173
At4g18710 shaggy-like protein kinase etha (EC 2.7.1.-) 601 e-172
At3g61160 shaggy-like kinase beta 552 e-158
At2g43790 MAP kinase (ATMPK6) 195 3e-50
At1g73670 MAP kinase like protein 185 4e-47
At3g45640 mitogen-activated protein kinase 3 184 9e-47
At3g59790 mitogen-activated protein kinase-like protein 183 1e-46
At1g18150 unknown protein 183 1e-46
At4g01370 MAP kinase 4 182 2e-46
At2g42880 MAP kinase 4 homolog 179 2e-45
At5g19010 MAP kinase -like protein 179 3e-45
At3g18040 MAP kinase like protein 178 5e-45
At4g19110 putative serine/threonine-protein kinase 177 8e-45
>At5g26751 shaggy-like kinase alpha
Length = 405
Score = 691 bits (1782), Expect = 0.0
Identities = 329/367 (89%), Positives = 348/367 (94%)
Query: 2 QEMEAHVVDGNSTEAGHVIVTTIGGKNGQPKQTISYMAERAVGQGSFGVVFQAKCLETGE 61
+EMEA VVDGN TE GH+IVTTIGG+NGQPKQTISYMAER VG GSFGVVFQAKCLETGE
Sbjct: 34 KEMEATVVDGNGTETGHIIVTTIGGRNGQPKQTISYMAERVVGHGSFGVVFQAKCLETGE 93
Query: 62 TVAIKKVLQDKRYKNRELQTMRLLDHPNVVTLKHCFFSTTEKDELYLNLVLEFVPETVHR 121
TVAIKKVLQD+RYKNRELQTMRLLDHPNVV+LKHCFFSTTEKDELYLNLVLE+VPETVHR
Sbjct: 94 TVAIKKVLQDRRYKNRELQTMRLLDHPNVVSLKHCFFSTTEKDELYLNLVLEYVPETVHR 153
Query: 122 VIRHYSKMNQRMPLIYVKLYSYQILRSLAYIHNCVGVSHRDIKPQNLLVNPHTHQLKLCD 181
VI+HY+K+NQRMPLIYVKLY+YQI R+L+YIH C+GV HRDIKPQNLLVNPHTHQ+KLCD
Sbjct: 154 VIKHYNKLNQRMPLIYVKLYTYQIFRALSYIHRCIGVCHRDIKPQNLLVNPHTHQVKLCD 213
Query: 182 FGSAKVLVKGEPNISYICSRYYRAPELIFGATEYTSAIDIWSAGCVLGELLLGQPLFPGA 241
FGSAKVLVKGEPNISYICSRYYRAPELIFGATEYT+AID+WSAGCVL ELLLGQPLFPG
Sbjct: 214 FGSAKVLVKGEPNISYICSRYYRAPELIFGATEYTTAIDVWSAGCVLAELLLGQPLFPGE 273
Query: 242 SGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIKAHPWHKIFRKRMPPEAVDLVSRL 301
SGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIKAHPWHKIF KRMPPEAVDLVSRL
Sbjct: 274 SGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIKAHPWHKIFHKRMPPEAVDLVSRL 333
Query: 302 LQYSPNLRSTALEALVHPFFDELRDPNTRLPNGRHLPPLFNFKANELKGVPAEMLVKLVP 361
LQYSPNLRS AL+ LVHPFFDELRDPN RLPNGR LPPLFNFK +ELKGVP EM+ KLVP
Sbjct: 334 LQYSPNLRSAALDTLVHPFFDELRDPNARLPNGRFLPPLFNFKPHELKGVPLEMVAKLVP 393
Query: 362 SHARKQC 368
HARKQC
Sbjct: 394 EHARKQC 400
>At3g05840 shaggy related protein kinase, ASK-GAMMA
Length = 409
Score = 687 bits (1773), Expect = 0.0
Identities = 326/367 (88%), Positives = 348/367 (93%)
Query: 2 QEMEAHVVDGNSTEAGHVIVTTIGGKNGQPKQTISYMAERAVGQGSFGVVFQAKCLETGE 61
+EMEA +V+GN TE GH+IVTTIGG+NGQPKQTISYMAER VG GSFGVVFQAKCLETGE
Sbjct: 38 KEMEATIVNGNVTETGHIIVTTIGGRNGQPKQTISYMAERVVGHGSFGVVFQAKCLETGE 97
Query: 62 TVAIKKVLQDKRYKNRELQTMRLLDHPNVVTLKHCFFSTTEKDELYLNLVLEFVPETVHR 121
TVAIKKVLQD+RYKNRELQTMRLLDHPNVV+LKHCFFSTTEKDELYLNLVLE+VPETVHR
Sbjct: 98 TVAIKKVLQDRRYKNRELQTMRLLDHPNVVSLKHCFFSTTEKDELYLNLVLEYVPETVHR 157
Query: 122 VIRHYSKMNQRMPLIYVKLYSYQILRSLAYIHNCVGVSHRDIKPQNLLVNPHTHQLKLCD 181
VI+HY+K+NQRMPL+YVKLY+YQI RSL+YIH C+GV HRDIKPQNLLVNPHTHQ+KLCD
Sbjct: 158 VIKHYNKLNQRMPLVYVKLYTYQIFRSLSYIHRCIGVCHRDIKPQNLLVNPHTHQVKLCD 217
Query: 182 FGSAKVLVKGEPNISYICSRYYRAPELIFGATEYTSAIDIWSAGCVLGELLLGQPLFPGA 241
FGSAKVLVKGEPNISYICSRYYRAPELIFGATEYT+AID+WSAGCVL ELLLGQPLFPG
Sbjct: 218 FGSAKVLVKGEPNISYICSRYYRAPELIFGATEYTTAIDVWSAGCVLAELLLGQPLFPGE 277
Query: 242 SGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIKAHPWHKIFRKRMPPEAVDLVSRL 301
SGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIKAHPWHKIF KRMPPEAVDLVSRL
Sbjct: 278 SGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIKAHPWHKIFHKRMPPEAVDLVSRL 337
Query: 302 LQYSPNLRSTALEALVHPFFDELRDPNTRLPNGRHLPPLFNFKANELKGVPAEMLVKLVP 361
LQYSPNLR AL++LVHPFFDELRDPN RLPNGR LPPLFNFK +ELKGVP EM+ KLVP
Sbjct: 338 LQYSPNLRCAALDSLVHPFFDELRDPNARLPNGRFLPPLFNFKPHELKGVPVEMVAKLVP 397
Query: 362 SHARKQC 368
HARKQC
Sbjct: 398 EHARKQC 404
>At5g14640 protein kinase MSK-3 - like
Length = 410
Score = 681 bits (1757), Expect = 0.0
Identities = 328/371 (88%), Positives = 346/371 (92%)
Query: 2 QEMEAHVVDGNSTEAGHVIVTTIGGKNGQPKQTISYMAERAVGQGSFGVVFQAKCLETGE 61
+EMEA VVDGN TE GH+IVTTIGGKNGQPKQTISYMAER VGQGSFG+VFQAKCLETGE
Sbjct: 39 KEMEAAVVDGNGTETGHIIVTTIGGKNGQPKQTISYMAERIVGQGSFGIVFQAKCLETGE 98
Query: 62 TVAIKKVLQDKRYKNRELQTMRLLDHPNVVTLKHCFFSTTEKDELYLNLVLEFVPETVHR 121
TVAIKKVLQDKRYKNRELQTMRLLDHPNVV+LKHCFFSTTEKDELYLNLVLE+VPETV+R
Sbjct: 99 TVAIKKVLQDKRYKNRELQTMRLLDHPNVVSLKHCFFSTTEKDELYLNLVLEYVPETVYR 158
Query: 122 VIRHYSKMNQRMPLIYVKLYSYQILRSLAYIHNCVGVSHRDIKPQNLLVNPHTHQLKLCD 181
V +HYS+ NQRMP+IYVKLY+YQI R+LAYIH VGV HRDIKPQNLLVNPHTHQ+KLCD
Sbjct: 159 VSKHYSRANQRMPIIYVKLYTYQICRALAYIHGGVGVCHRDIKPQNLLVNPHTHQVKLCD 218
Query: 182 FGSAKVLVKGEPNISYICSRYYRAPELIFGATEYTSAIDIWSAGCVLGELLLGQPLFPGA 241
FGSAKVLVKGEPNISYICSRYYRAPELIFGATEYT+ IDIWSAGCVL ELLLGQPLFPG
Sbjct: 219 FGSAKVLVKGEPNISYICSRYYRAPELIFGATEYTTTIDIWSAGCVLAELLLGQPLFPGE 278
Query: 242 SGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIKAHPWHKIFRKRMPPEAVDLVSRL 301
SGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIKAHPWHKIF KR PPEAVDLVSRL
Sbjct: 279 SGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIKAHPWHKIFHKRTPPEAVDLVSRL 338
Query: 302 LQYSPNLRSTALEALVHPFFDELRDPNTRLPNGRHLPPLFNFKANELKGVPAEMLVKLVP 361
LQYSPNLRSTA+EA+VHPFFDELRDPNTRLPNGR LPPLFNFK ELKG E+L KL+P
Sbjct: 339 LQYSPNLRSTAMEAIVHPFFDELRDPNTRLPNGRALPPLFNFKPQELKGASLELLSKLIP 398
Query: 362 SHARKQCTLFA 372
HARKQC+ A
Sbjct: 399 DHARKQCSFLA 409
>At1g57870 protein kinase, putative
Length = 420
Score = 612 bits (1577), Expect = e-175
Identities = 288/366 (78%), Positives = 320/366 (86%)
Query: 2 QEMEAHVVDGNSTEAGHVIVTTIGGKNGQPKQTISYMAERAVGQGSFGVVFQAKCLETGE 61
++ E ++DG E GHVI TT+ G+NGQ +QT+SY+AE VG GSFG+VFQAKC ETGE
Sbjct: 47 RDSEPDIIDGVGAEPGHVITTTLPGRNGQSRQTVSYIAEHVVGTGSFGMVFQAKCRETGE 106
Query: 62 TVAIKKVLQDKRYKNRELQTMRLLDHPNVVTLKHCFFSTTEKDELYLNLVLEFVPETVHR 121
VAIKKVLQDKRYKNRELQ M++LDHPNVV LKH F+S TE +E+YLNLVLEFVPETV+R
Sbjct: 107 VVAIKKVLQDKRYKNRELQIMQMLDHPNVVCLKHSFYSRTENEEVYLNLVLEFVPETVNR 166
Query: 122 VIRHYSKMNQRMPLIYVKLYSYQILRSLAYIHNCVGVSHRDIKPQNLLVNPHTHQLKLCD 181
R YS+MNQ MPLIYVKLY+YQI R LAY+HNC G+ HRDIKPQNLLVNPHTHQLK+CD
Sbjct: 167 TARSYSRMNQLMPLIYVKLYTYQICRGLAYLHNCCGLCHRDIKPQNLLVNPHTHQLKICD 226
Query: 182 FGSAKVLVKGEPNISYICSRYYRAPELIFGATEYTSAIDIWSAGCVLGELLLGQPLFPGA 241
FGSAKVLVKGEPNISYICSRYYRAPELIFGATEYT+AIDIWS GCV+ ELLLGQPLFPG
Sbjct: 227 FGSAKVLVKGEPNISYICSRYYRAPELIFGATEYTTAIDIWSTGCVMAELLLGQPLFPGE 286
Query: 242 SGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIKAHPWHKIFRKRMPPEAVDLVSRL 301
SGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIK HPWHK+F+KR+PPEAVDL+ R
Sbjct: 287 SGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIKPHPWHKVFQKRLPPEAVDLLCRF 346
Query: 302 LQYSPNLRSTALEALVHPFFDELRDPNTRLPNGRHLPPLFNFKANELKGVPAEMLVKLVP 361
QYSPNLR TA+EA +HPFFDELRDPN RLPNGR LPPLFNFK EL G+P E + +LVP
Sbjct: 347 FQYSPNLRCTAVEACIHPFFDELRDPNARLPNGRPLPPLFNFKPQELSGIPPETVDRLVP 406
Query: 362 SHARKQ 367
HARKQ
Sbjct: 407 EHARKQ 412
>At2g30980 putative shaggy-like protein kinase dzeta
Length = 412
Score = 608 bits (1567), Expect = e-174
Identities = 286/366 (78%), Positives = 324/366 (88%), Gaps = 1/366 (0%)
Query: 2 QEMEAHVVDGNSTEAGHVIVTTIGGKNGQPKQTISYMAERAVGQGSFGVVFQAKCLETGE 61
+EM A V++GN GH+I TTIGGKNG+PKQTISYMAER VG GSFG+VFQAKCLETGE
Sbjct: 37 KEMSAAVIEGNDAVTGHIISTTIGGKNGEPKQTISYMAERVVGTGSFGIVFQAKCLETGE 96
Query: 62 TVAIKKVLQDKRYKNRELQTMRLLDHPNVVTLKHCFFSTTEKDELYLNLVLEFVPETVHR 121
+VAIKKVLQD+RYKNRELQ MRL+DHPNVV+LKHCFFSTT +DEL+LNLV+E+VPET++R
Sbjct: 97 SVAIKKVLQDRRYKNRELQLMRLMDHPNVVSLKHCFFSTTTRDELFLNLVMEYVPETLYR 156
Query: 122 VIRHYSKMNQRMPLIYVKLYSYQILRSLAYIHNCVGVSHRDIKPQNLLVNPHTHQLKLCD 181
V++HY+ NQRMP+ YVKLY+YQI R LAYIH GV HRD+KPQNLLV+P THQ KLCD
Sbjct: 157 VLKHYTSSNQRMPIFYVKLYTYQIFRGLAYIHTAPGVCHRDVKPQNLLVDPLTHQCKLCD 216
Query: 182 FGSAKVLVKGEPNISYICSRYYRAPELIFGATEYTSAIDIWSAGCVLGELLLGQPLFPGA 241
FGSAKVLVKGE NISYICSRYYRAPELIFGATEYTS+IDIWSAGCVL ELLLGQPLFPG
Sbjct: 217 FGSAKVLVKGEANISYICSRYYRAPELIFGATEYTSSIDIWSAGCVLAELLLGQPLFPGE 276
Query: 242 SGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIKAHPWHKIFRKRMPPEAVDLVSRL 301
+ VDQLVEIIKVLGTPTREEI+CMNPNYT+F+FPQIKAHPWHK+F KRMPPEA+DL SRL
Sbjct: 277 NSVDQLVEIIKVLGTPTREEIRCMNPNYTDFRFPQIKAHPWHKVFHKRMPPEAIDLASRL 336
Query: 302 LQYSPNLRSTALEALVHPFFDELRDPNTRLPNGRHLPPLFNFKANELKGVPAEMLVKLVP 361
LQYSP+LR TALEA HPFF+ELR+PN RLPNGR LPPLFNFK EL G E++ +L+P
Sbjct: 337 LQYSPSLRCTALEACAHPFFNELREPNARLPNGRPLPPLFNFK-QELSGASPELINRLIP 395
Query: 362 SHARKQ 367
H R+Q
Sbjct: 396 EHVRRQ 401
>At1g06390 putative shaggy kinase iota protein
Length = 407
Score = 608 bits (1567), Expect = e-174
Identities = 285/366 (77%), Positives = 325/366 (87%), Gaps = 1/366 (0%)
Query: 2 QEMEAHVVDGNSTEAGHVIVTTIGGKNGQPKQTISYMAERAVGQGSFGVVFQAKCLETGE 61
+EM A V++GN GH+I TTIGGKNG+PKQTISYMAER VG GSFG+VFQAKCLETGE
Sbjct: 35 KEMSAAVIEGNDAVTGHIISTTIGGKNGEPKQTISYMAERVVGTGSFGIVFQAKCLETGE 94
Query: 62 TVAIKKVLQDKRYKNRELQTMRLLDHPNVVTLKHCFFSTTEKDELYLNLVLEFVPETVHR 121
+VAIKKVLQD+RYKNRELQ MR +DHPNV++LKHCFFSTT +DEL+LNLV+E+VPET++R
Sbjct: 95 SVAIKKVLQDRRYKNRELQLMRPMDHPNVISLKHCFFSTTSRDELFLNLVMEYVPETLYR 154
Query: 122 VIRHYSKMNQRMPLIYVKLYSYQILRSLAYIHNCVGVSHRDIKPQNLLVNPHTHQLKLCD 181
V+RHY+ NQRMP+ YVKLY+YQI R LAYIH GV HRD+KPQNLLV+P THQ+KLCD
Sbjct: 155 VLRHYTSSNQRMPIFYVKLYTYQIFRGLAYIHTVPGVCHRDVKPQNLLVDPLTHQVKLCD 214
Query: 182 FGSAKVLVKGEPNISYICSRYYRAPELIFGATEYTSAIDIWSAGCVLGELLLGQPLFPGA 241
FGSAKVLVKGEPNISYICSRYYRAPELIFGATEYT++IDIWSAGCVL ELLLGQPLFPG
Sbjct: 215 FGSAKVLVKGEPNISYICSRYYRAPELIFGATEYTASIDIWSAGCVLAELLLGQPLFPGE 274
Query: 242 SGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIKAHPWHKIFRKRMPPEAVDLVSRL 301
+ VDQLVEIIKVLGTPTREEI+CMNPNYT+F+FPQIKAHPWHK+F KRMPPEA+DL SRL
Sbjct: 275 NSVDQLVEIIKVLGTPTREEIRCMNPNYTDFRFPQIKAHPWHKVFHKRMPPEAIDLASRL 334
Query: 302 LQYSPNLRSTALEALVHPFFDELRDPNTRLPNGRHLPPLFNFKANELKGVPAEMLVKLVP 361
LQYSP+LR TALEA HPFF+ELR+PN RLPNGR LPPLFNFK EL G E++ +L+P
Sbjct: 335 LQYSPSLRCTALEACAHPFFNELREPNARLPNGRPLPPLFNFK-QELGGASMELINRLIP 393
Query: 362 SHARKQ 367
H R+Q
Sbjct: 394 EHVRRQ 399
>At4g00720 Shaggy related protein kinase tetha
Length = 472
Score = 607 bits (1566), Expect = e-174
Identities = 287/364 (78%), Positives = 322/364 (87%)
Query: 2 QEMEAHVVDGNSTEAGHVIVTTIGGKNGQPKQTISYMAERAVGQGSFGVVFQAKCLETGE 61
++ME VV+G+ TE G VI TT+GG++G+PKQTISYMA+R VG GSFGVVFQAKCLETGE
Sbjct: 103 KDMETTVVNGSGTETGQVITTTVGGRDGKPKQTISYMAQRVVGTGSFGVVFQAKCLETGE 162
Query: 62 TVAIKKVLQDKRYKNRELQTMRLLDHPNVVTLKHCFFSTTEKDELYLNLVLEFVPETVHR 121
VAIKKVLQDKRYKNRELQ MRL DHPNVV L+H FFSTT+KDELYLNLVLE+VPETV+R
Sbjct: 163 QVAIKKVLQDKRYKNRELQIMRLQDHPNVVRLRHSFFSTTDKDELYLNLVLEYVPETVYR 222
Query: 122 VIRHYSKMNQRMPLIYVKLYSYQILRSLAYIHNCVGVSHRDIKPQNLLVNPHTHQLKLCD 181
+HY+KMNQ MP+I+V+LY+YQI R+L Y+H VGV HRDIKPQNLLVNP THQLK+CD
Sbjct: 223 ASKHYTKMNQHMPIIFVQLYTYQICRALNYLHRVVGVCHRDIKPQNLLVNPQTHQLKICD 282
Query: 182 FGSAKVLVKGEPNISYICSRYYRAPELIFGATEYTSAIDIWSAGCVLGELLLGQPLFPGA 241
FGSAK+LV GEPNISYICSRYYRAPELIFGATEYT+AID+WS GCV+ ELLLGQPLFPG
Sbjct: 283 FGSAKMLVPGEPNISYICSRYYRAPELIFGATEYTNAIDMWSGGCVMAELLLGQPLFPGE 342
Query: 242 SGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIKAHPWHKIFRKRMPPEAVDLVSRL 301
SG+DQLVEIIK+LGTPTREEI+CMNPNYTEFKFPQIKAHPWHKIF KRMPPEAVDLVSRL
Sbjct: 343 SGIDQLVEIIKILGTPTREEIRCMNPNYTEFKFPQIKAHPWHKIFHKRMPPEAVDLVSRL 402
Query: 302 LQYSPNLRSTALEALVHPFFDELRDPNTRLPNGRHLPPLFNFKANELKGVPAEMLVKLVP 361
LQYSPNLR TALEA HPFFD+LRDPN LPNGR LPPLFNF A EL G E+ +L+P
Sbjct: 403 LQYSPNLRCTALEACAHPFFDDLRDPNVSLPNGRALPPLFNFTAQELAGASTELRQRLIP 462
Query: 362 SHAR 365
+H +
Sbjct: 463 AHCQ 466
>At1g09840 shaggy-like protien kinase, kappa
Length = 421
Score = 605 bits (1559), Expect = e-173
Identities = 285/366 (77%), Positives = 320/366 (86%)
Query: 2 QEMEAHVVDGNSTEAGHVIVTTIGGKNGQPKQTISYMAERAVGQGSFGVVFQAKCLETGE 61
++ E ++DG E GHVI TT+ G+NGQ +QT+SY++E VG GSFG+VFQAKC ETGE
Sbjct: 48 RDSEPDIIDGAGAEPGHVIRTTLRGRNGQSRQTVSYISEHVVGTGSFGMVFQAKCRETGE 107
Query: 62 TVAIKKVLQDKRYKNRELQTMRLLDHPNVVTLKHCFFSTTEKDELYLNLVLEFVPETVHR 121
VAIKKVLQDKRYKNRELQ M++LDHPN V LKH FFS T+ +E+YLNLVLEFVPETV+R
Sbjct: 108 VVAIKKVLQDKRYKNRELQIMQMLDHPNAVALKHSFFSRTDNEEVYLNLVLEFVPETVNR 167
Query: 122 VIRHYSKMNQRMPLIYVKLYSYQILRSLAYIHNCVGVSHRDIKPQNLLVNPHTHQLKLCD 181
V R YS+ NQ MPLIYVKLY+YQI R+LAYIHN G+ HRDIKPQNLLVNPHTHQLK+CD
Sbjct: 168 VARSYSRTNQLMPLIYVKLYTYQICRALAYIHNSFGLCHRDIKPQNLLVNPHTHQLKICD 227
Query: 182 FGSAKVLVKGEPNISYICSRYYRAPELIFGATEYTSAIDIWSAGCVLGELLLGQPLFPGA 241
FGSAKVLVKGEPN+SYICSRYYRAPELIFGA+EYT+AIDIWS GCV+ ELLLGQPLFPG
Sbjct: 228 FGSAKVLVKGEPNVSYICSRYYRAPELIFGASEYTTAIDIWSTGCVMAELLLGQPLFPGE 287
Query: 242 SGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIKAHPWHKIFRKRMPPEAVDLVSRL 301
SGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIK HPWHK+F+KR+PPEAVDL+ R
Sbjct: 288 SGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIKPHPWHKVFQKRLPPEAVDLLCRF 347
Query: 302 LQYSPNLRSTALEALVHPFFDELRDPNTRLPNGRHLPPLFNFKANELKGVPAEMLVKLVP 361
QYSPNLR TALEA +HP FDELRDPNTRLPNGR LPPLFNFK EL G+P E++ +LVP
Sbjct: 348 FQYSPNLRCTALEACIHPLFDELRDPNTRLPNGRPLPPLFNFKPQELSGIPPEIVNRLVP 407
Query: 362 SHARKQ 367
HARKQ
Sbjct: 408 EHARKQ 413
>At4g18710 shaggy-like protein kinase etha (EC 2.7.1.-)
Length = 380
Score = 601 bits (1549), Expect = e-172
Identities = 285/369 (77%), Positives = 323/369 (87%), Gaps = 1/369 (0%)
Query: 2 QEMEAHVVDGNSTEAGHVIVTTIGGKNGQPKQTISYMAERAVGQGSFGVVFQAKCLETGE 61
+EM A VVDG+ GH+I TTIGGKNG+PKQTISYMAER VG GSFG+VFQAKCLETGE
Sbjct: 5 KEMPAAVVDGHDQVTGHIISTTIGGKNGEPKQTISYMAERVVGTGSFGIVFQAKCLETGE 64
Query: 62 TVAIKKVLQDKRYKNRELQTMRLLDHPNVVTLKHCFFSTTEKDELYLNLVLEFVPETVHR 121
TVAIKKVLQD+RYKNRELQ MR++DHPNVV LKHCFFSTT KDEL+LNLV+E+VPE+++R
Sbjct: 65 TVAIKKVLQDRRYKNRELQLMRVMDHPNVVCLKHCFFSTTSKDELFLNLVMEYVPESLYR 124
Query: 122 VIRHYSKMNQRMPLIYVKLYSYQILRSLAYIHNCVGVSHRDIKPQNLLVNPHTHQLKLCD 181
V++HYS NQRMPL+YVKLY YQI R LAYIHN GV HRD+KPQNLLV+P THQ+K+CD
Sbjct: 125 VLKHYSSANQRMPLVYVKLYMYQIFRGLAYIHNVAGVCHRDLKPQNLLVDPLTHQVKICD 184
Query: 182 FGSAKVLVKGEPNISYICSRYYRAPELIFGATEYTSAIDIWSAGCVLGELLLGQPLFPGA 241
FGSAK LVKGE NISYICSR+YRAPELIFGATEYT++IDIWSAGCVL ELLLGQPLFPG
Sbjct: 185 FGSAKQLVKGEANISYICSRFYRAPELIFGATEYTTSIDIWSAGCVLAELLLGQPLFPGE 244
Query: 242 SGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIKAHPWHKIFRKRMPPEAVDLVSRL 301
+ VDQLVEIIKVLGTPTREEI+CMNP+YT+F+FPQIKAHPWHKIF KRMPPEA+D SRL
Sbjct: 245 NAVDQLVEIIKVLGTPTREEIRCMNPHYTDFRFPQIKAHPWHKIFHKRMPPEAIDFASRL 304
Query: 302 LQYSPNLRSTALEALVHPFFDELRDPNTRLPNGRHLPPLFNFKANELKGVPAEMLVKLVP 361
LQYSP+LR TALEA HPFFDELR+PN RLPNGR PPLFNFK E+ G E++ KL+P
Sbjct: 305 LQYSPSLRCTALEACAHPFFDELREPNARLPNGRPFPPLFNFK-QEVAGSSPELVNKLIP 363
Query: 362 SHARKQCTL 370
H ++Q L
Sbjct: 364 DHIKRQLGL 372
>At3g61160 shaggy-like kinase beta
Length = 438
Score = 552 bits (1423), Expect = e-158
Identities = 263/365 (72%), Positives = 302/365 (82%)
Query: 2 QEMEAHVVDGNSTEAGHVIVTTIGGKNGQPKQTISYMAERAVGQGSFGVVFQAKCLETGE 61
++M+ ++ GN TE+G +I T G N Q +TISY AE +G GSFGVVFQAKCLET E
Sbjct: 74 KDMDCGIIKGNGTESGRIITTKKKGLNDQKDKTISYRAEHVIGTGSFGVVFQAKCLETEE 133
Query: 62 TVAIKKVLQDKRYKNRELQTMRLLDHPNVVTLKHCFFSTTEKDELYLNLVLEFVPETVHR 121
VAIKKVLQDKRYKNRELQ MR+LDHPNVV LKH FFSTTEKDELYLNLVLE+VPET++R
Sbjct: 134 KVAIKKVLQDKRYKNRELQIMRMLDHPNVVELKHSFFSTTEKDELYLNLVLEYVPETIYR 193
Query: 122 VIRHYSKMNQRMPLIYVKLYSYQILRSLAYIHNCVGVSHRDIKPQNLLVNPHTHQLKLCD 181
R Y+KMNQ MPLIY++LY+YQI R++ Y+H VGV HRDIKPQNLLVN TH++K+CD
Sbjct: 194 ASRSYTKMNQHMPLIYIQLYTYQICRAMNYLHQVVGVCHRDIKPQNLLVNNVTHEVKICD 253
Query: 182 FGSAKVLVKGEPNISYICSRYYRAPELIFGATEYTSAIDIWSAGCVLGELLLGQPLFPGA 241
FGSAK+L+ GEPNISYICSRYYRAPELIFGATEYTSAID+WS GCV+ EL LG PLFPG
Sbjct: 254 FGSAKMLIPGEPNISYICSRYYRAPELIFGATEYTSAIDMWSVGCVMAELFLGHPLFPGE 313
Query: 242 SGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQIKAHPWHKIFRKRMPPEAVDLVSRL 301
+ VDQLVEIIK+LGTP REEIK MNP Y +FKFPQIKA PWHKIFR+++ PEA+DL SRL
Sbjct: 314 TSVDQLVEIIKILGTPAREEIKNMNPRYNDFKFPQIKAQPWHKIFRRQVSPEAMDLASRL 373
Query: 302 LQYSPNLRSTALEALVHPFFDELRDPNTRLPNGRHLPPLFNFKANELKGVPAEMLVKLVP 361
LQYSPNLR TALEA HPFFD+LRDP LPNGR LPPLF+F A EL G E+ +L+P
Sbjct: 374 LQYSPNLRCTALEACAHPFFDDLRDPRASLPNGRALPPLFDFTAQELAGASVELRHRLIP 433
Query: 362 SHARK 366
HARK
Sbjct: 434 EHARK 438
>At2g43790 MAP kinase (ATMPK6)
Length = 395
Score = 195 bits (496), Expect = 3e-50
Identities = 107/292 (36%), Positives = 173/292 (58%), Gaps = 15/292 (5%)
Query: 43 VGQGSFGVVFQAKCLETGETVAIKKVLQ------DKRYKNRELQTMRLLDHPNVVTLKHC 96
+G+G++G+V A ET E+VAIKK+ D + RE++ +R +DH N+V ++
Sbjct: 69 IGKGAYGIVCSAMNSETNESVAIKKIANAFDNKIDAKRTLREIKLLRHMDHENIVAIRDI 128
Query: 97 FFSTTEKDELYLNLVLEFVPETVHRVIRHYSKMNQRMPLIYVKLYSYQILRSLAYIHNCV 156
+ + E + +H++IR NQ + + + + YQILR L YIH+
Sbjct: 129 IPPPLRNAFNDVYIAYELMDTDLHQIIRS----NQALSEEHCQYFLYQILRGLKYIHSA- 183
Query: 157 GVSHRDIKPQNLLVNPHTHQLKLCDFGSAKVLVKGEPNISYICSRYYRAPELIFGATEYT 216
V HRD+KP NLL+N + LK+CDFG A+V + + Y+ +R+YRAPEL+ +++YT
Sbjct: 184 NVLHRDLKPSNLLLNANC-DLKICDFGLARVTSESDFMTEYVVTRWYRAPELLLNSSDYT 242
Query: 217 SAIDIWSAGCVLGELLLGQPLFPGASGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQ 276
+AID+WS GC+ EL+ +PLFPG V QL +++++GTP+ EE++ +N N + Q
Sbjct: 243 AAIDVWSVGCIFMELMDRKPLFPGRDHVHQLRLLMELIGTPSEEELEFLNENAKRY-IRQ 301
Query: 277 IKAHPWHKIFRK--RMPPEAVDLVSRLLQYSPNLRSTALEALVHPFFDELRD 326
+ +P I K + P A+DL+ ++L + P R T L+AL HP+ + L D
Sbjct: 302 LPPYPRQSITDKFPTVHPLAIDLIEKMLTFDPRRRITVLDALAHPYLNSLHD 353
>At1g73670 MAP kinase like protein
Length = 576
Score = 185 bits (469), Expect = 4e-47
Identities = 120/313 (38%), Positives = 174/313 (55%), Gaps = 27/313 (8%)
Query: 37 YMAERAVGQGSFGVVFQAKCLETGETVAIKKV------LQDKRYKNRELQTMRLLDHPNV 90
Y + VG+GS+GVV A TGE VAIKK+ + D RE++ +RLL HP+V
Sbjct: 90 YQIQEVVGKGSYGVVGSAIDTHTGERVAIKKINDVFDHISDATRILREIKLLRLLLHPDV 149
Query: 91 VTLKHCFFSTTEKDELYLNLVLEFVPETVHRVIRHYSKMNQRMPLIYVKLYSYQILRSLA 150
V +KH + ++ + +V E + +H+VI K N + + + + YQ+LR L
Sbjct: 150 VEIKHIMLPPSRREFRDVYVVFELMESDLHQVI----KANDDLTPEHHQFFLYQLLRGLK 205
Query: 151 YIHNCVGVSHRDIKPQNLLVNPHTHQLKLCDFGSAKVLVKGEPNI----SYICSRYYRAP 206
Y+H V HRD+KP+N+L N +LK+CDFG A+V P Y+ +R+YRAP
Sbjct: 206 YVH-AANVFHRDLKPKNILANADC-KLKICDFGLARVSFNDAPTAIFWTDYVATRWYRAP 263
Query: 207 ELIFGA-TEYTSAIDIWSAGCVLGELLLGQPLFPGASGVDQLVEIIKVLGTPTREEI-KC 264
EL ++YT AIDIWS GC+ E+LLG+PLFPG + V QL + LGTP E I K
Sbjct: 264 ELCGSFFSKYTPAIDIWSVGCIFAEMLLGKPLFPGKNVVHQLDIMTDFLGTPPPEAISKI 323
Query: 265 MNPNYTEFKFPQIKAHPWHKIFRKRMP---PEAVDLVSRLLQYSPNLRSTALEALVHPFF 321
N + K P F K+ P P A+ L+ RL+ + P R +A EAL P+F
Sbjct: 324 RNDKARRYLGNMRKKQP--VPFSKKFPKADPSALRLLERLIAFDPKDRPSAEEALADPYF 381
Query: 322 D----ELRDPNTR 330
+ ++R+P+T+
Sbjct: 382 NGLSSKVREPSTQ 394
>At3g45640 mitogen-activated protein kinase 3
Length = 370
Score = 184 bits (466), Expect = 9e-47
Identities = 103/295 (34%), Positives = 168/295 (56%), Gaps = 16/295 (5%)
Query: 43 VGQGSFGVVFQAKCLETGETVAIKKVLQ------DKRYKNRELQTMRLLDHPNVVTLKHC 96
+G+G++G+V ET E VA+KK+ D + RE++ +R LDH N++ ++
Sbjct: 44 IGRGAYGIVCSVLDTETNELVAMKKIANAFDNHMDAKRTLREIKLLRHLDHENIIAIRDV 103
Query: 97 FFSTTEKDELYLNLVLEFVPETVHRVIRHYSKMNQRMPLIYVKLYSYQILRSLAYIHNCV 156
+ + + E + +H++IR NQ + + + + YQ+LR L YIH+
Sbjct: 104 VPPPLRRQFSDVYISTELMDTDLHQIIRS----NQSLSEEHCQYFLYQLLRGLKYIHSA- 158
Query: 157 GVSHRDIKPQNLLVNPHTHQLKLCDFGSAKVLVKGEPNISYICSRYYRAPELIFGATEYT 216
+ HRD+KP NLL+N + LK+CDFG A+ + + Y+ +R+YRAPEL+ +++YT
Sbjct: 159 NIIHRDLKPSNLLLNANC-DLKICDFGLARPTSENDFMTEYVVTRWYRAPELLLNSSDYT 217
Query: 217 SAIDIWSAGCVLGELLLGQPLFPGASGVDQLVEIIKVLGTPTREEIK-CMNPNYTEF--K 273
+AID+WS GC+ EL+ +PLFPG V Q+ + ++LGTPT ++ N + + +
Sbjct: 218 AAIDVWSVGCIFMELMNRKPLFPGKDHVHQMRLLTELLGTPTESDLGFTHNEDAKRYIRQ 277
Query: 274 FPQIKAHPWHKIFRKRMPPEAVDLVSRLLQYSPNLRSTALEALVHPFFDELRDPN 328
P P K+F + P A+DLV R+L + PN R T +AL H + +L DPN
Sbjct: 278 LPNFPRQPLAKLF-SHVNPMAIDLVDRMLTFDPNRRITVEQALNHQYLAKLHDPN 331
>At3g59790 mitogen-activated protein kinase-like protein
Length = 393
Score = 183 bits (465), Expect = 1e-46
Identities = 103/294 (35%), Positives = 165/294 (56%), Gaps = 15/294 (5%)
Query: 41 RAVGQGSFGVVFQAKCLETGETVAIKKVLQ------DKRYKNRELQTMRLLDHPNVVTLK 94
R +G+G+ G+V A ET E VAIKK+ Q + + RE++ +R DH N+V ++
Sbjct: 64 RPIGRGACGIVCSAVDSETNEKVAIKKITQVFDNTIEAKRTLREIKLLRHFDHENIVAIR 123
Query: 95 HCFFSTTEKDELYLNLVLEFVPETVHRVIRHYSKMNQRMPLIYVKLYSYQILRSLAYIHN 154
+ +V E + ++R + K +Q + + + YQILR L YIH+
Sbjct: 124 DVILPPQRDSFEDVYIVNELMEFDLYRTL----KSDQELTKDHGMYFMYQILRGLKYIHS 179
Query: 155 CVGVSHRDIKPQNLLVNPHTHQLKLCDFGSAKVLVKGEPNISYICSRYYRAPELIFGATE 214
V HRD+KP NLL++ LK+CDFG A+ + Y+ +R+YRAPEL+ G+++
Sbjct: 180 A-NVLHRDLKPSNLLLSTQC-DLKICDFGLARATPESNLMTEYVVTRWYRAPELLLGSSD 237
Query: 215 YTSAIDIWSAGCVLGELLLGQPLFPGASGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKF 274
YT+AID+WS GC+ E++ +PLFPG V+QL +++++GTP+ EE+ ++ Y +
Sbjct: 238 YTAAIDVWSVGCIFMEIMNREPLFPGKDQVNQLRLLLELIGTPSEEELGSLS-EYAKRYI 296
Query: 275 PQIKAHPWHKIFRK--RMPPEAVDLVSRLLQYSPNLRSTALEALVHPFFDELRD 326
Q+ P K +PP A+DLV ++L + P R + EAL HP+ D
Sbjct: 297 RQLPTLPRQSFTEKFPNVPPLAIDLVEKMLTFDPKQRISVKEALAHPYLSSFHD 350
>At1g18150 unknown protein
Length = 589
Score = 183 bits (465), Expect = 1e-46
Identities = 119/313 (38%), Positives = 171/313 (54%), Gaps = 27/313 (8%)
Query: 37 YMAERAVGQGSFGVVFQAKCLETGETVAIKKV------LQDKRYKNRELQTMRLLDHPNV 90
Y + VG+GS+GVV A TGE VAIKK+ + D RE++ +RLL HP+V
Sbjct: 104 YQIQEVVGKGSYGVVASAVDSHTGERVAIKKINDVFEHVSDATRILREIKLLRLLRHPDV 163
Query: 91 VTLKHCFFSTTEKDELYLNLVLEFVPETVHRVIRHYSKMNQRMPLIYVKLYSYQILRSLA 150
V +KH + ++ + +V E + +H+VI K N + + + + YQ+LR L
Sbjct: 164 VEIKHIMLPPSRREFRDIYVVFELMESDLHQVI----KANDDLTPEHYQFFLYQLLRGLK 219
Query: 151 YIHNCVGVSHRDIKPQNLLVNPHTHQLKLCDFGSAKVLVKGEPNI----SYICSRYYRAP 206
Y+H V HRD+KP+N+L N +LK+CDFG A+V P Y+ +R+YRAP
Sbjct: 220 YVH-AANVFHRDLKPKNILANADC-KLKICDFGLARVSFNDAPTAIFWTDYVATRWYRAP 277
Query: 207 ELIFGA-TEYTSAIDIWSAGCVLGELLLGQPLFPGASGVDQLVEIIKVLGTPTREEI-KC 264
EL ++YT AIDIWS GC+ E+LLG+PLFPG + V QL + LGTP E I +
Sbjct: 278 ELCGSFFSKYTPAIDIWSVGCIFAEMLLGKPLFPGKNVVHQLDLMTDFLGTPPPESISRI 337
Query: 265 MNPNYTEFKFPQIKAHP---WHKIFRKRMPPEAVDLVSRLLQYSPNLRSTALEALVHPFF 321
N + K P HK + P A+ L+ RLL + P R++A +AL P+F
Sbjct: 338 RNEKARRYLSSMRKKQPVPFSHKF--PKADPLALRLLERLLAFDPKDRASAEDALADPYF 395
Query: 322 DEL----RDPNTR 330
L R+P T+
Sbjct: 396 SGLSNSEREPTTQ 408
>At4g01370 MAP kinase 4
Length = 376
Score = 182 bits (463), Expect = 2e-46
Identities = 108/297 (36%), Positives = 169/297 (56%), Gaps = 16/297 (5%)
Query: 41 RAVGQGSFGVVFQAKCLETGETVAIKKV------LQDKRYKNRELQTMRLLDHPNVVTLK 94
R +G+G++G+V A ETGE VAIKK+ + D + RE++ ++ +DH NV+ +K
Sbjct: 47 RPIGRGAYGIVCAATNSETGEEVAIKKIGNAFDNIIDAKRTLREIKLLKHMDHENVIAVK 106
Query: 95 HCFFSTTEKDELYLNLVLEFVPETVHRVIRHYSKMNQRMPLIYVKLYSYQILRSLAYIHN 154
++ + +V E + +H++IR NQ + + + + YQ+LR L Y+H+
Sbjct: 107 DIIKPPQRENFNDVYIVYELMDTDLHQIIRS----NQPLTDDHCRFFLYQLLRGLKYVHS 162
Query: 155 CVGVSHRDIKPQNLLVNPHTHQLKLCDFGSAKVLVKGEPNISYICSRYYRAPELIFGATE 214
V HRD+KP NLL+N + LKL DFG A+ + + Y+ +R+YRAPEL+ +E
Sbjct: 163 A-NVLHRDLKPSNLLLNANC-DLKLGDFGLARTKSETDFMTEYVVTRWYRAPELLLNCSE 220
Query: 215 YTSAIDIWSAGCVLGELLLGQPLFPGASGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKF 274
YT+AIDIWS GC+LGE + +PLFPG V QL I +++G+P + + +
Sbjct: 221 YTAAIDIWSVGCILGETMTREPLFPGKDYVHQLRLITELIGSPDDSSLGFLRSDNARRYV 280
Query: 275 PQIKAHPWHKIFRKRMP---PEAVDLVSRLLQYSPNLRSTALEALVHPFFDELRDPN 328
Q+ +P + F R P AVDL+ ++L + P+ R T EAL HP+ L D N
Sbjct: 281 RQLPQYP-RQNFAARFPNMSAGAVDLLEKMLVFDPSRRITVDEALCHPYLAPLHDIN 336
>At2g42880 MAP kinase 4 homolog
Length = 606
Score = 179 bits (455), Expect = 2e-45
Identities = 110/301 (36%), Positives = 167/301 (54%), Gaps = 19/301 (6%)
Query: 37 YMAERAVGQGSFGVVFQAKCLETGETVAIKKV------LQDKRYKNRELQTMRLLDHPNV 90
+ + +G+GS+GVV A TGE VAIKK+ + D RE++ +RLL HP++
Sbjct: 25 FKVQEVIGKGSYGVVCSAIDTLTGEKVAIKKIHDIFEHISDAARILREIKLLRLLRHPDI 84
Query: 91 VTLKHCFFSTTEKDELYLNLVLEFVPETVHRVIRHYSKMNQRMPLIYVKLYSYQILRSLA 150
V +KH + ++ + +V E + +H+VI K N + + + + YQ+LR+L
Sbjct: 85 VEIKHIMLPPSRREFKDIYVVFELMESDLHQVI----KANDDLTREHYQFFLYQLLRALK 140
Query: 151 YIHNCVGVSHRDIKPQNLLVNPHTHQLKLCDFGSAKVLVKGEPNI----SYICSRYYRAP 206
YIH V HRD+KP+N+L N + +LK+CDFG A+V P Y+ +R+YRAP
Sbjct: 141 YIHTA-NVYHRDLKPKNILANANC-KLKICDFGLARVAFNDTPTTIFWTDYVATRWYRAP 198
Query: 207 ELIFGA-TEYTSAIDIWSAGCVLGELLLGQPLFPGASGVDQLVEIIKVLGTPTREEIKCM 265
EL ++YT AIDIWS GC+ E+L+G+PLFPG + V QL + +LGTP+ + I +
Sbjct: 199 ELCGSFYSKYTPAIDIWSIGCIFAEVLMGKPLFPGKNVVHQLDLMTDLLGTPSLDTISRV 258
Query: 266 NPNYTEFKFPQIKAHPWHKIFRK--RMPPEAVDLVSRLLQYSPNLRSTALEALVHPFFDE 323
++ P +K P ++ L+ RLL + P R TA EAL P+F
Sbjct: 259 RNEKARRYLTSMRKKPPIPFAQKFPNADPLSLKLLERLLAFDPKDRPTAEEALADPYFKG 318
Query: 324 L 324
L
Sbjct: 319 L 319
>At5g19010 MAP kinase -like protein
Length = 567
Score = 179 bits (453), Expect = 3e-45
Identities = 117/313 (37%), Positives = 170/313 (53%), Gaps = 27/313 (8%)
Query: 37 YMAERAVGQGSFGVVFQAKCLETGETVAIKKV------LQDKRYKNRELQTMRLLDHPNV 90
Y E +G+GS+GVV A TGE VAIKK+ + D RE++ +RLL HP++
Sbjct: 25 YRIEEVIGKGSYGVVCSAYDTHTGEKVAIKKINDIFEHVSDATRILREIKLLRLLRHPDI 84
Query: 91 VTLKHCFFSTTEKDELYLNLVLEFVPETVHRVIRHYSKMNQRMPLIYVKLYSYQILRSLA 150
V +KH + ++ + +V E + +H+VI K N + + + + YQ+LR L
Sbjct: 85 VEIKHILLPPSRREFRDIYVVFELMESDLHQVI----KANDDLTPEHYQFFLYQLLRGLK 140
Query: 151 YIHNCVGVSHRDIKPQNLLVNPHTHQLKLCDFGSAKVLVKGEPNI----SYICSRYYRAP 206
YIH V HRD+KP+N+L N +LK+CDFG A+V P Y+ +R+YRAP
Sbjct: 141 YIHTA-NVFHRDLKPKNILANADC-KLKICDFGLARVAFNDTPTAIFWTDYVATRWYRAP 198
Query: 207 ELIFGA-TEYTSAIDIWSAGCVLGELLLGQPLFPGASGVDQLVEIIKVLGTPTREEI-KC 264
EL ++YT AIDIWS GC+ ELL G+PLFPG + V QL + +LGTP+ E I +
Sbjct: 199 ELCGSFFSKYTPAIDIWSIGCIFAELLTGKPLFPGKNVVHQLDLMTDMLGTPSAEAIGRV 258
Query: 265 MNPNYTEFKFPQIKAHPWHKIFRKRMP---PEAVDLVSRLLQYSPNLRSTALEALVHPFF 321
N + K P F + P P A+ L+ ++L + P R TA EAL +F
Sbjct: 259 RNEKARRYLSSMRKKKPIP--FSHKFPHTDPLALRLLEKMLSFEPKDRPTAEEALADVYF 316
Query: 322 DEL----RDPNTR 330
L R+P+ +
Sbjct: 317 KGLAKVEREPSAQ 329
>At3g18040 MAP kinase like protein
Length = 510
Score = 178 bits (451), Expect = 5e-45
Identities = 117/328 (35%), Positives = 179/328 (53%), Gaps = 25/328 (7%)
Query: 37 YMAERAVGQGSFGVVFQAKCLETGETVAIKKV------LQDKRYKNRELQTMRLLDHPNV 90
Y + +G+GS+GVV A +GE VAIKK+ + D RE++ +RLL HP++
Sbjct: 23 YQIQEVIGKGSYGVVASAIDTHSGEKVAIKKINDVFEHVSDATRILREIKLLRLLRHPDI 82
Query: 91 VTLKHCFFSTTEKDELYLNLVLEFVPETVHRVIRHYSKMNQRMPLIYVKLYSYQILRSLA 150
V +KH + ++ + +V E + +H+VI K N + + + + YQ+LR L
Sbjct: 83 VEIKHVMLPPSRREFRDIYVVFELMESDLHQVI----KANDDLTPEHYQFFLYQLLRGLK 138
Query: 151 YIHNCVGVSHRDIKPQNLLVNPHTHQLKLCDFGSAKVLVKGEPNI----SYICSRYYRAP 206
+IH V HRD+KP+N+L N +LK+CDFG A+V P+ Y+ +R+YRAP
Sbjct: 139 FIHTA-NVFHRDLKPKNILANSDC-KLKICDFGLARVSFNDAPSAIFWTDYVATRWYRAP 196
Query: 207 ELIFGA-TEYTSAIDIWSAGCVLGELLLGQPLFPGASGVDQLVEIIKVLGTPTREEIKCM 265
EL ++YT AIDIWS GC+ E+L G+PLFPG + V QL + +LGTP E I +
Sbjct: 197 ELCGSFFSKYTPAIDIWSIGCIFAEMLTGKPLFPGKNVVHQLDIMTDLLGTPPPEAIARI 256
Query: 266 NPNYTEFKFPQIKAHP----WHKIFRKRMPPEAVDLVSRLLQYSPNLRSTALEALVHPFF 321
++ P HK + P A+ L+ RLL + P R +A EAL P+F
Sbjct: 257 RNEKARRYLGNMRRKPPVPFTHKF--PHVDPLALRLLHRLLAFDPKDRPSAEEALADPYF 314
Query: 322 DELRDPNTRLPNGRHLPPL-FNFKANEL 348
L + + R P+ + +P L F F+ ++
Sbjct: 315 YGLANVD-REPSTQPIPKLEFEFERRKI 341
>At4g19110 putative serine/threonine-protein kinase
Length = 461
Score = 177 bits (449), Expect = 8e-45
Identities = 104/292 (35%), Positives = 172/292 (58%), Gaps = 19/292 (6%)
Query: 37 YMAERAVGQGSFGVVFQAKCLETGETVAIKKVLQ-----DKRYKNRELQTMRLLDHPNVV 91
Y + VG G+FG V++A +TGE VAIKK+ + D+ RE++++R ++HPN+V
Sbjct: 4 YKLIKEVGDGTFGSVWRAINKQTGEVVAIKKMKKKYYSWDECINLREVKSLRRMNHPNIV 63
Query: 92 TLKHCFFSTTEKDELYLNLVLEFVPETVHRVIRHYSKMNQRMPLIYVKLYSYQILRSLAY 151
LK E D LY V E++ ++++++ K+ +K + +Q+ + L+Y
Sbjct: 64 KLKEVI---RENDILYF--VFEYMECNLYQLMKDRQKLFAEAD---IKNWCFQVFQGLSY 115
Query: 152 IHNCVGVSHRDIKPQNLLVNPHTHQLKLCDFGSAKVLVKGEPNISYICSRYYRAPELIFG 211
+H G HRD+KP+NLLV+ +K+ DFG A+ + P Y+ +R+YRAPE++
Sbjct: 116 MHQR-GYFHRDLKPENLLVSKDI--IKIADFGLAREVNSSPPFTEYVSTRWYRAPEVLLQ 172
Query: 212 ATEYTSAIDIWSAGCVLGELLLGQPLFPGASGVDQLVEIIKVLGTPTREE-IKCMN-PNY 269
+ YTS +D+W+ G ++ ELL +P+FPGAS D++ +I V+GTPT E ++ +N N
Sbjct: 173 SYVYTSKVDMWAMGAIMAELLSLRPIFPGASEADEIYKICSVIGTPTEETWLEGLNLANT 232
Query: 270 TEFKFPQIKAHPWHKIFRKRMPPEAVDLVSRLLQYSPNLRSTALEALVHPFF 321
++FPQ+ P + +A++L+ RL + P+ R TA E L HPFF
Sbjct: 233 INYQFPQLPGVPLSSLM-PSASEDAINLIERLCSWDPSSRPTAAEVLQHPFF 283
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.138 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,604,001
Number of Sequences: 26719
Number of extensions: 376988
Number of successful extensions: 2893
Number of sequences better than 10.0: 872
Number of HSP's better than 10.0 without gapping: 279
Number of HSP's successfully gapped in prelim test: 593
Number of HSP's that attempted gapping in prelim test: 1371
Number of HSP's gapped (non-prelim): 1005
length of query: 373
length of database: 11,318,596
effective HSP length: 101
effective length of query: 272
effective length of database: 8,619,977
effective search space: 2344633744
effective search space used: 2344633744
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC136973.16


