

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136973.11 - phase: 2 /pseudo
(192 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
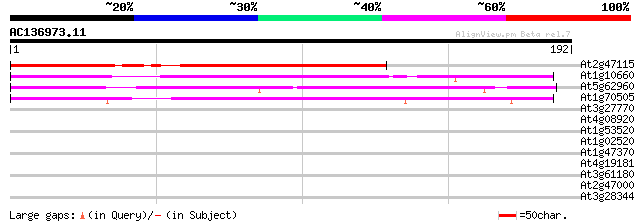
Score E
Sequences producing significant alignments: (bits) Value
At2g47115 putative protein 113 5e-26
At1g10660 unknown protein 100 4e-22
At5g62960 unknown protein 85 3e-17
At1g70505 putative protein 68 3e-12
At3g27770 unknown protein 33 0.071
At4g08920 flavin-type blue-light photoreceptor (HY4) 30 1.0
At1g53520 chalcone isomerase, putative 29 1.3
At1g02520 P-glycoprotein, putative 29 1.8
At1g47370 disease resistance protein, putative 28 2.3
At4g19181 putative protein 28 3.9
At3g61180 unknown protein 28 3.9
At2g47000 putative ABC transporter 27 6.7
At3g28344 P-glycoprotein, 5' partial 27 8.7
>At2g47115 putative protein
Length = 289
Score = 113 bits (283), Expect = 5e-26
Identities = 61/129 (47%), Positives = 81/129 (62%), Gaps = 10/129 (7%)
Query: 1 WYDFVCFAIVGVSILGALWVLWTNEGSSTSQSDSNIFVESLLVANSPSSDNRVAIGHVST 60
WYDF+CFAIV +I+ +LW L + D++ +SLL S+NR +
Sbjct: 7 WYDFICFAIVAAAIVTSLWFLSRRDRGCVVIDDTSH--DSLLPLYR--SNNR------GS 56
Query: 61 SQLWTSCWRGVHPLVLLTTRLFSFVSLAMLLYLDIHEYDASIFYYYTEWTFTLVMIYFAL 120
++LW SCW +HP LL TR SF+S+A LL D+ ++DASIF YYTEWTF LV+IYFA+
Sbjct: 57 ARLWASCWTRLHPGWLLFTRSTSFLSMAALLAWDVIKWDASIFVYYTEWTFMLVIIYFAM 116
Query: 121 GTTVSAYGC 129
G S YGC
Sbjct: 117 GIVASVYGC 125
>At1g10660 unknown protein
Length = 320
Score = 100 bits (250), Expect = 4e-22
Identities = 63/198 (31%), Positives = 95/198 (47%), Gaps = 32/198 (16%)
Query: 1 WYDFVCFAIVGVSILGALWVLWTNEGSSTSQSDSNIFVESLLVANSPSSDNRVAIGHVST 60
W +C I+ I+ A ++W EG Q +S R G +
Sbjct: 14 WRVLLCALILLAPIVLAAVLIWKYEGKRRRQRESQ----------------RELPGTLFQ 57
Query: 61 SQLWTSCWRGVHPLVLLTTRLFSFVSLAMLLYLDIHEYDASIFYYYTEWTFTLVMIYFAL 120
+ WT+C++ +HPL LL R+FSFV++ LL ++ A IFY+YT+WTFTLV +YF
Sbjct: 58 DEAWTTCFKRIHPLWLLAFRVFSFVAMLTLLISNVVRDGAGIFYFYTQWTFTLVTLYFGY 117
Query: 121 GTTVSAYGCWKVLNKPPPLQNGEMTEFLRRD------------LETKGSIFTFQSRYAEE 168
+ +S YGC + NK +G M + L+ +G+ +R +E
Sbjct: 118 ASVLSVYGCC-IYNKE---ASGNMESYTSIGDTEQGTYRPPIALDGEGNTSKASNRPSEA 173
Query: 169 EFEQTAGFWGYLMQITFQ 186
+TAGFW Y+ QI FQ
Sbjct: 174 PARKTAGFWVYIFQILFQ 191
>At5g62960 unknown protein
Length = 347
Score = 84.7 bits (208), Expect = 3e-17
Identities = 53/189 (28%), Positives = 92/189 (48%), Gaps = 17/189 (8%)
Query: 1 WYDFVCFAIVGVSILGALWVLWTNEGSSTSQSDSNIFVESLLVANSPSSDNRVAIGHVST 60
W +C + ++ + ++++ EG +SD + G+V
Sbjct: 38 WRVMICCIWMAIATVITAFLIFKYEGFRRKRSD----------VGEVDGGEKEWSGNVYE 87
Query: 61 SQLWTSCWRGVHPLVLLTTRLFSF-VSLAMLLYLDIHEYDASIFYYYTEWTFTLVMIYFA 119
+ W C R +HP LL R+ +F V L ML+ + + + +IF+YYT+WTF L+ +YF
Sbjct: 88 DETWRPCLRNIHPAWLLAFRVVAFFVLLVMLIVIGLVD-GPTIFFYYTQWTFGLITLYFG 146
Query: 120 LGTTVSAYGCWKVLNKPPPLQNGEMTEFLRRDLETKGSIFTF-QSRYAEEEFEQTAGFWG 178
LG+ +S +GC++ + + + +KG+ T QS+Y+ AGFWG
Sbjct: 147 LGSLLSLHGCYQYNKRAAGDRVDSIEAIDSERARSKGADNTIQQSQYS----SNPAGFWG 202
Query: 179 YLMQITFQV 187
Y+ QI FQ+
Sbjct: 203 YVFQIIFQM 211
>At1g70505 putative protein
Length = 338
Score = 67.8 bits (164), Expect = 3e-12
Identities = 56/193 (29%), Positives = 82/193 (42%), Gaps = 20/193 (10%)
Query: 1 WYDFVCFAIVGVSILGALWVLWTNEGSSTSQS---DSNIFVESLLVANSPSSDNRVAIGH 57
W FVC V S+ + +++W EG + D ++ +E L G
Sbjct: 40 WRFFVCAIFVLTSLFLSSYLIWRYEGPIKRKKRGDDQSLELEQLT-------------GV 86
Query: 58 VSTSQLWTSCWRGVHPLVLLTTRLFSFVSLAMLLYLDIHEYDASIFYYYTEWTFTLVMIY 117
V + W + + +HP LL R+F FV L L+ + IF +YT+WTFTLV IY
Sbjct: 87 VYDDESWNTSVKEIHPNWLLGFRVFGFVVLLGLISGNAIADGTGIFIFYTQWTFTLVTIY 146
Query: 118 FALGTTVSAYGCWKVLN--KPPPLQNGEMTEFLRRDLETKGSIFTFQSRYAEEEF--EQT 173
F LG+ VS Y N + + E + ++F S + E Q
Sbjct: 147 FGLGSLVSIYRFRSPDNGENRVSIVDEEQGTYRPPGNAENSNVFKSSSGHDRENMSTRQV 206
Query: 174 AGFWGYLMQITFQ 186
A GY+ QI FQ
Sbjct: 207 ATTLGYIHQILFQ 219
>At3g27770 unknown protein
Length = 108
Score = 33.5 bits (75), Expect = 0.071
Identities = 28/108 (25%), Positives = 40/108 (36%), Gaps = 17/108 (15%)
Query: 1 WYDFVCFAIVGVSILGALWVLWTNEGSSTSQSDSNIFVESLLVANSPSSDNRVAIGHVST 60
W +C V V ++ +L VLW E SS V PS + +
Sbjct: 18 WRVLLCAIWVIVPMIVSLLVLWKYEDSS--------------VQTQPSLNGNDVL---CI 60
Query: 61 SQLWTSCWRGVHPLVLLTTRLFSFVSLAMLLYLDIHEYDASIFYYYTE 108
+W C+ +HP LL R+ F L I+YYYT+
Sbjct: 61 DDVWRPCFERIHPGWLLGFRVLGFCFLLANNIARFANRGWRIYYYYTQ 108
>At4g08920 flavin-type blue-light photoreceptor (HY4)
Length = 681
Score = 29.6 bits (65), Expect = 1.0
Identities = 14/47 (29%), Positives = 23/47 (48%), Gaps = 8/47 (17%)
Query: 20 VLWTNEGSSTSQSDSNIFVESL--------LVANSPSSDNRVAIGHV 58
V W NEG+ + N+F++S+ + N P S R +GH+
Sbjct: 273 VAWANEGNEAGEESVNLFLKSIGLREYSRYISFNHPYSHERPLLGHL 319
>At1g53520 chalcone isomerase, putative
Length = 287
Score = 29.3 bits (64), Expect = 1.3
Identities = 20/79 (25%), Positives = 33/79 (41%), Gaps = 11/79 (13%)
Query: 7 FAIVGVSILGALWVL----------WTNEGSSTSQSDSNIFVESLLVANSPSSDNRVAIG 56
FAI+GV + A + + WT + Q DS++FV S+ A + S V +
Sbjct: 118 FAIIGVKVYAAGYYVNESILSGLSAWTGRSADEIQRDSSLFV-SIFQAQAEKSLQIVLVR 176
Query: 57 HVSTSQLWTSCWRGVHPLV 75
V W + + P +
Sbjct: 177 DVDGKTFWDALDEAISPRI 195
>At1g02520 P-glycoprotein, putative
Length = 1278
Score = 28.9 bits (63), Expect = 1.8
Identities = 18/58 (31%), Positives = 32/58 (55%), Gaps = 5/58 (8%)
Query: 118 FALGTTVSAYGCWKVLNKPPPLQ----NGEMTEFLRRDLETKGSIFTFQSRYAEEEFE 171
FA G +AY ++ + + P + NG++ E +R D+E K F++ +R EE F+
Sbjct: 344 FAAGQA-AAYKMFETIKRKPLIDAYDVNGKVLEDIRGDIELKDVHFSYPARPDEEIFD 400
>At1g47370 disease resistance protein, putative
Length = 363
Score = 28.5 bits (62), Expect = 2.3
Identities = 16/51 (31%), Positives = 26/51 (50%), Gaps = 1/51 (1%)
Query: 120 LGTTVSAYGCWKVLNKPPPLQNGEMTEFLRRDLETKGSIFTFQSRYAEEEF 170
L + YG +++K N ++T L R E+K ++ F SR+AE F
Sbjct: 33 LADALERYGIMFIIDKDEQRGN-DLTSLLLRIKESKVALVIFSSRFAESRF 82
>At4g19181 putative protein
Length = 740
Score = 27.7 bits (60), Expect = 3.9
Identities = 15/41 (36%), Positives = 23/41 (55%)
Query: 26 GSSTSQSDSNIFVESLLVANSPSSDNRVAIGHVSTSQLWTS 66
GSS +S+++ S V +S SSD+ I + ST W+S
Sbjct: 669 GSSIQLMESSLYSSSSCVMHSCSSDSLGDIQYDSTGSFWSS 709
>At3g61180 unknown protein
Length = 379
Score = 27.7 bits (60), Expect = 3.9
Identities = 24/91 (26%), Positives = 40/91 (43%), Gaps = 8/91 (8%)
Query: 53 VAIGHVSTSQLWT--SCWRGVHPLVLLTTR---LFSFVSLAMLLYLDIHEYDASIFYYYT 107
V + V+ QL S W P+++L LF VS+A+L + + D + +
Sbjct: 61 VRVREVAAEQLEERQSQWAYSKPIIVLDILWNFLFVIVSIAILGFSSDEDPDVPLRLWII 120
Query: 108 EWTFTLVMIYFALGTTVSAYGCWKVLNKPPP 138
+ V F +G ++ Y +V N PPP
Sbjct: 121 GYN---VQCLFHVGCVIAEYKRRRVANSPPP 148
>At2g47000 putative ABC transporter
Length = 1286
Score = 26.9 bits (58), Expect = 6.7
Identities = 16/57 (28%), Positives = 31/57 (54%), Gaps = 5/57 (8%)
Query: 118 FALGTTVSAYGCWKVLNKPPPLQ----NGEMTEFLRRDLETKGSIFTFQSRYAEEEF 170
FA G +AY ++ + + P + NG++ + ++ D+E K FT+ +R E+ F
Sbjct: 347 FAAGQA-AAYKMFETIERRPNIDSYSTNGKVLDDIKGDIELKDVYFTYPARPDEQIF 402
>At3g28344 P-glycoprotein, 5' partial
Length = 626
Score = 26.6 bits (57), Expect = 8.7
Identities = 17/49 (34%), Positives = 24/49 (48%), Gaps = 2/49 (4%)
Query: 142 GEMTEFLRRDLETKGSIFTFQSRYAEEEFEQTAGFWGYLMQITFQVKFL 190
G MT L + + GS+F RY + E G+ +IT QV+FL
Sbjct: 345 GSMTTDLAKGSDAVGSVFAVLDRYTSIDPEDPDGY--ETERITGQVEFL 391
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.137 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,225,460
Number of Sequences: 26719
Number of extensions: 159100
Number of successful extensions: 515
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 503
Number of HSP's gapped (non-prelim): 14
length of query: 192
length of database: 11,318,596
effective HSP length: 94
effective length of query: 98
effective length of database: 8,807,010
effective search space: 863086980
effective search space used: 863086980
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC136973.11


