

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
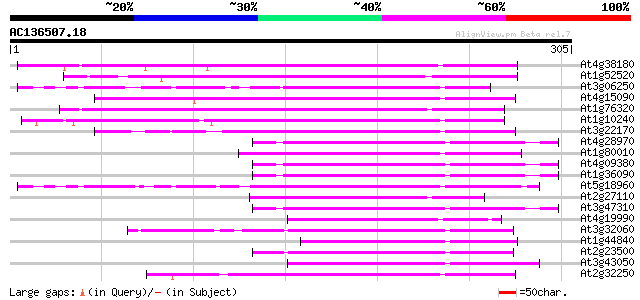
Query= AC136507.18 - phase: 1 /pseudo
(305 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At4g38180 unknown protein (At4g38180) 84 1e-16
At1g52520 F6D8.26 70 2e-12
At3g06250 unknown protein 68 7e-12
At4g15090 unknown protein 67 1e-11
At1g76320 putative phytochrome A signaling protein 66 3e-11
At1g10240 unknown protein 65 3e-11
At3g22170 far-red impaired response protein, putative 64 8e-11
At4g28970 putative protein 62 3e-10
At1g80010 hypothetical protein 62 5e-10
At4g09380 putative protein 61 7e-10
At1g36090 hypothetical protein 61 7e-10
At5g18960 FAR1 - like protein 61 9e-10
At2g27110 Mutator-like transposase 60 1e-09
At3g47310 putative protein 59 3e-09
At4g19990 putative protein 58 7e-09
At3g32060 hypothetical protein 57 1e-08
At1g44840 hypothetical protein 56 3e-08
At2g23500 Mutator-like transposase 55 5e-08
At3g43050 putative protein 55 6e-08
At2g32250 Mutator-like transposase 54 1e-07
>At4g38180 unknown protein (At4g38180)
Length = 788
Score = 84.0 bits (206), Expect = 1e-16
Identities = 69/283 (24%), Positives = 122/283 (42%), Gaps = 14/283 (4%)
Query: 5 RLVCERSGSHIVPKKKPKHAGTGS----RKCGCLFMISGYQSKQTKEWRLNILNGVHNHP 60
+ VC + G + +K+ K + GC +S + + + +W ++ HNH
Sbjct: 118 QFVCAKEGFRNMNEKRTKDREIKRPRTITRVGCKASLS-VKMQDSGKWLVSGFVKDHNHE 176
Query: 61 MEPALEGHILAG--RLKEDDKKIVRDLTKNKMLPRNILIHLKNQRPHC----MTNVKQVY 114
+ P + H L ++ K ++ L M PR I+ L + T V
Sbjct: 177 LVPPDQVHCLRSHRQISGPAKTLIDTLQAAGMGPRRIMSALIKEYGGISKVGFTEVDCRN 236
Query: 115 NERQQIWKANRGD-KKPLQYLISKLEEHKYTYYSRTQLESNTIEDIFWAHPTSIKLFNNF 173
R K+ G+ + L YL ++ +YS E ++ ++FWA P +I F +F
Sbjct: 237 YMRNNRQKSIEGEIQLLLDYLRQMNADNPNFFYSVQGSEDQSVGNVFWADPKAIMDFTHF 296
Query: 174 PTVLVMDSTYKTNIYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSS 233
+ D+TY++N YR+P GV G F+ +E E +FVW+ +S+
Sbjct: 297 GDTVTFDTTYRSNRYRLPFAPFTGVNHHGQPILFGCAFIINETEASFVWLFNTWLAAMSA 356
Query: 234 KMNMPKVIVTDRDMSLMKAVAHVFPESYAMNCYFHVQANVKQR 276
+ P I TD D + A+ HVFP + C +H+ +++
Sbjct: 357 --HPPVSITTDHDAVIRAAIMHVFPGARHRFCKWHILKKCQEK 397
>At1g52520 F6D8.26
Length = 703
Score = 69.7 bits (169), Expect = 2e-12
Identities = 63/252 (25%), Positives = 107/252 (42%), Gaps = 14/252 (5%)
Query: 30 KCGCLFMISGYQSKQTKEWRLNILNGVHNHPMEPALEGHILAGRLKEDDKKI---VRDLT 86
+ GC MI Q +K WR+ + HNH L G L +K K + V D
Sbjct: 152 RTGCPAMIRMRQV-DSKRWRVVEVTLDHNH-----LLGCKLYKSVKRKRKCVSSPVSDAK 205
Query: 87 KNKMLPRNILIHLKNQRPHCMTNVK-QVYNERQQIWKANRGDKKPLQYLISKLE-EHKYT 144
K+ ++ + N P+ N K Q + RGD + +++ +
Sbjct: 206 TIKLYRACVVDNGSNVNPNSTLNKKFQNSTGSPDLLNLKRGDSAAIYNYFCRMQLTNPNF 265
Query: 145 YYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMPMFEVVGVTSTDLT 204
+Y + + ++FWA S + F V+ +DS+Y + + +P+ GV T
Sbjct: 266 FYLMDVNDEGQLRNVFWADAFSKVSCSYFGDVIFIDSSYISGKFEIPLVTFTGVNHHGKT 325
Query: 205 YSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYAMN 264
+ GF+ E E++ W+LK+ LS P+ IVTDR L A++ VFP S+
Sbjct: 326 TLLSCGFLAGETMESYHWLLKV---WLSVMKRSPQTIVTDRCKPLEAAISQVFPRSHQRF 382
Query: 265 CYFHVQANVKQR 276
H+ + ++
Sbjct: 383 SLTHIMRKIPEK 394
>At3g06250 unknown protein
Length = 764
Score = 67.8 bits (164), Expect = 7e-12
Identities = 66/258 (25%), Positives = 108/258 (41%), Gaps = 35/258 (13%)
Query: 5 RLVCERSGSHIVPKKKPKHAGTGSRKCGCLFMISGYQSKQTKEWRLNILNGVHNHPMEPA 64
R VC + G + P G CG I + + + W ++ LN HNH +EP
Sbjct: 235 RFVCSKEGF-----QHPSRMG-----CGAYMRI---KRQDSGGWIVDRLNKDHNHDLEP- 280
Query: 65 LEGHILAGRLKEDDKKIVRDLTKNKMLPRNILIHLKNQRPHCMTNVKQVYNERQQIWKAN 124
G+ KKI D+T L LI L + H + + + W
Sbjct: 281 -------GKKNAGMKKITDDVTGG--LDSVDLIELNDLSNHISSTRENTIGKE---WYPV 328
Query: 125 RGDKKPLQYLISKLEEHKYTYYSRTQLESN-TIEDIFWAHPTSIKLFNNFPTVLVMDSTY 183
L Y SK E +Y+ +L+SN + IFWA S + F +V D++Y
Sbjct: 329 L-----LDYFQSKQAEDMGFFYA-IELDSNGSCMSIFWADSRSRFACSQFGDAVVFDTSY 382
Query: 184 KTNIYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVT 243
+ Y +P +G +G + E +E F W+ + + +S + P+ +V
Sbjct: 383 RKGDYSVPFATFIGFNHHRQPVLLGGALVADESKEAFSWLFQTWLRAMSGR--RPRSMVA 440
Query: 244 DRDMSLMKAVAHVFPESY 261
D+D+ + +AVA VFP ++
Sbjct: 441 DQDLPIQQAVAQVFPGTH 458
>At4g15090 unknown protein
Length = 768
Score = 67.0 bits (162), Expect = 1e-11
Identities = 54/237 (22%), Positives = 102/237 (42%), Gaps = 10/237 (4%)
Query: 47 EWRLNILNGVHNHPMEPALEGHILAGR-LKEDDKKIVRDLTKNKMLPRNILIHL------ 99
+W ++ HNH + PAL H R +K +K + L + + + +
Sbjct: 79 KWIIHEFVKDHNHELLPALAYHFRIQRNVKLAEKNNIDILHAVSERTKKMYVEMSRQSGG 138
Query: 100 -KNQRPHCMTNVKQVYNERQQIWKANRGDKKPLQYLISKLEEHKYTYYSRTQLESNTIED 158
KN T+V ++ + + + L+Y +E+ +Y+ E + +
Sbjct: 139 YKNIGSLLQTDVSSQVDKGRYLALEEGDSQVLLEYFKRIKKENPKFFYAIDLNEDQRLRN 198
Query: 159 IFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEE 218
+FWA S + +F V+ D+TY ++P+ +GV +G + E E
Sbjct: 199 LFWADAKSRDDYLSFNDVVSFDTTYVKFNDKLPLALFIGVNHHSQPMLLGCALVADESME 258
Query: 219 NFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYAMNCYFHVQANVKQ 275
FVW++K + + + PKVI+TD+D LM AV+ + P + +HV + +
Sbjct: 259 TFVWLIKTWLRAMGGR--APKVILTDQDKFLMSAVSELLPNTRHCFALWHVLEKIPE 313
>At1g76320 putative phytochrome A signaling protein
Length = 670
Score = 65.9 bits (159), Expect = 3e-11
Identities = 53/244 (21%), Positives = 110/244 (44%), Gaps = 5/244 (2%)
Query: 28 SRKCGCLFMISGYQSKQTKEWRLNILNGVHNHPMEPALEGHILAGRLKEDDKKIVRDLTK 87
S K GC + + + +W + HNH + P + + R E K L +
Sbjct: 63 SPKIGCKASMH-VKRRPDGKWYVYSFVKEHNHDLLPEQAHYFRSHRNTELVKSNDSRLRR 121
Query: 88 NKMLPRNILIHLKNQRP-HCMTNVKQVYNERQQIWKANRGDKKPL-QYLISKLEEHKYTY 145
K P HL + + +++ + + GD + L ++L+ EE+ +
Sbjct: 122 KKNTPLTDCKHLSAYHDLDFIDGYMRNQHDKGRRLVLDTGDAEILLEFLMRMQEENPKFF 181
Query: 146 YSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMPMFEVVGVTSTDLTY 205
++ E + + ++FW I+ + +F V+ +++Y + Y++P+ VGV
Sbjct: 182 FAVDFSEDHLLRNVFWVDAKGIEDYKSFSDVVSFETSYFVSKYKVPLVLFVGVNHHVQPV 241
Query: 206 SVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYAMNC 265
+G G + + +VW+++ L++ PKV++TD++ ++ A+A V PE+ C
Sbjct: 242 LLGCGLLADDTVYTYVWLMQSW--LVAMGGQKPKVMLTDQNNAIKAAIAAVLPETRHCYC 299
Query: 266 YFHV 269
+HV
Sbjct: 300 LWHV 303
>At1g10240 unknown protein
Length = 680
Score = 65.5 bits (158), Expect = 3e-11
Identities = 67/278 (24%), Positives = 124/278 (44%), Gaps = 20/278 (7%)
Query: 7 VCERSGS---HIVPKKKPKHAGTGSRKCGC--LFMISGYQSKQTKEWRLNILNGVHNHPM 61
VC R+G+ + + KP+ SR CGC IS + EWR+ HNH +
Sbjct: 96 VCHRAGNTPIKTLSEGKPQRNRRSSR-CGCQAYLRISKLTELGSTEWRVTGFANHHNHEL 154
Query: 62 -EPALEGHILAGR-LKEDDKKIVRDLTKNKMLPRNILIHLKNQRPHCMT------NVKQV 113
EP + A R + + DK + +K + + ++ L+ ++ C+ K V
Sbjct: 155 LEPNQVRFLPAYRSISDADKSRILMFSKTGISVQQMMRLLELEK--CVEPGFLPFTEKDV 212
Query: 114 YNERQQIWKANRGDKK-PLQYLISKLEEHKYTYYSRTQLESNT-IEDIFWAHPTSIKLFN 171
N Q K + D+ + ++E + L++N +E+I W++ +SI+ +
Sbjct: 213 RNLLQSFKKLDPEDENIDFLRMCQSIKEKDPNFKFEFTLDANDKLENIAWSYASSIQSYE 272
Query: 172 NFPTVLVMDSTYKTNIYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLL 231
F +V D+T++ + MP+ VGV + + G + E ++ W L+ +
Sbjct: 273 LFGDAVVFDTTHRLSAVEMPLGIWVGVNNYGVPCFFGCVLLRDENLRSWSWALQAFTGFM 332
Query: 232 SSKMNMPKVIVTDRDMSLMKAVAHVFPESYAMNCYFHV 269
+ K P+ I+TD +M L +A+A P + C + V
Sbjct: 333 NGK--APQTILTDHNMCLKEAIAGEMPATKHALCIWMV 368
>At3g22170 far-red impaired response protein, putative
Length = 814
Score = 64.3 bits (155), Expect = 8e-11
Identities = 49/230 (21%), Positives = 98/230 (42%), Gaps = 19/230 (8%)
Query: 47 EWRLNILNGVHNHPMEPALEGHILAGRLKEDDKKIVRDLTKNKMLPRNILIHLKNQRPHC 106
+W ++ HNH + PA + E +KI + K + +I LK+
Sbjct: 170 KWVIHSFVREHNHELLPAQA-------VSEQTRKIYAAMAK-QFAEYKTVISLKSDSKSS 221
Query: 107 MTNVKQVYNERQQIWKANRGDKKPLQYLISKLEEHKYTYYSRTQL-ESNTIEDIFWAHPT 165
E+ + GD K L +S+++ ++ L + ++++FW
Sbjct: 222 F--------EKGRTLSVETGDFKILLDFLSRMQSLNSNFFYAVDLGDDQRVKNVFWVDAK 273
Query: 166 SIKLFNNFPTVLVMDSTYKTNIYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLK 225
S + +F V+ +D+TY N Y+MP+ VGV +G ++ E + W+++
Sbjct: 274 SRHNYGSFCDVVSLDTTYVRNKYKMPLAIFVGVNQHYQYMVLGCALISDESAATYSWLME 333
Query: 226 MLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYAMNCYFHVQANVKQ 275
+ + + PKV++T+ D+ + V +FP + +HV V +
Sbjct: 334 TWLRAIGGQ--APKVLITELDVVMNSIVPEIFPNTRHCLFLWHVLMKVSE 381
>At4g28970 putative protein
Length = 914
Score = 62.4 bits (150), Expect = 3e-10
Identities = 44/167 (26%), Positives = 77/167 (45%), Gaps = 14/167 (8%)
Query: 133 YLISKLEEHKYTYYSRTQLESNT-IEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMP 191
Y++ K+ TY +LE + +F A I+ F V+V+D+T+ +Y
Sbjct: 369 YMLEKVNPGTVTY---VELEGEKKFKYLFIALGACIEGFRAMRKVIVVDATHLKTVYGGM 425
Query: 192 MFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLMK 251
+ Y + FG + EK+ +++W L+ L+ + S + V ++DR S+ K
Sbjct: 426 LVIATAQDPNHHHYPLAFGIIDSEKDVSWIWFLEKLKTVYSDVPGL--VFISDRHQSIKK 483
Query: 252 AVAHVFPESYAMNCYFHVQANVKQRCVLDCKYPLGFKKDGKEVSNRD 298
AV V+P + C +H+ N++ R +D KDG V RD
Sbjct: 484 AVKTVYPNALHAACIWHLCQNMRDRVTID--------KDGAAVKFRD 522
>At1g80010 hypothetical protein
Length = 696
Score = 61.6 bits (148), Expect = 5e-10
Identities = 35/155 (22%), Positives = 74/155 (47%), Gaps = 3/155 (1%)
Query: 125 RGDKKPLQYLISKLEEHKYTY-YSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTY 183
RG + LQ +++ + Y + ++ ++FW + +++F VL+ D+T
Sbjct: 240 RGGFRALQDFFFQIQLSSPNFLYLMDLADDGSLRNVFWIDARARAAYSHFGDVLLFDTTC 299
Query: 184 KTNIYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVT 243
+N Y +P+ VG+ T +G G + + E +VW+ + + + P++ +T
Sbjct: 300 LSNAYELPLVAFVGINHHGDTILLGCGLLADQSFETYVWLFRAWLTCMLGR--PPQIFIT 357
Query: 244 DRDMSLMKAVAHVFPESYAMNCYFHVQANVKQRCV 278
++ ++ AV+ VFP ++ HV N+ Q V
Sbjct: 358 EQCKAMRTAVSEVFPRAHHRLSLTHVLHNICQSVV 392
>At4g09380 putative protein
Length = 960
Score = 61.2 bits (147), Expect = 7e-10
Identities = 44/167 (26%), Positives = 77/167 (45%), Gaps = 14/167 (8%)
Query: 133 YLISKLEEHKYTYYSRTQLESNT-IEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMP 191
Y++ K+ TY +LE + +F A I+ F V+V+D+T+ +Y
Sbjct: 369 YMLEKVNPGTVTY---VELEGEKKFKYLFIALGACIEGFRAMRKVIVVDATHLKTVYGGM 425
Query: 192 MFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLMK 251
+ Y + FG + EK+ +++W L+ L+ + S + V ++DR S+ K
Sbjct: 426 LVIATAQDPNHHHYPLAFGIIDSEKDVSWIWFLENLKTVYSDVPGL--VFISDRHQSIKK 483
Query: 252 AVAHVFPESYAMNCYFHVQANVKQRCVLDCKYPLGFKKDGKEVSNRD 298
AV V+P + C +H+ N++ R +D KDG V RD
Sbjct: 484 AVKTVYPNALHAACIWHLCQNMRDRVKID--------KDGAAVKFRD 522
>At1g36090 hypothetical protein
Length = 645
Score = 61.2 bits (147), Expect = 7e-10
Identities = 44/167 (26%), Positives = 77/167 (45%), Gaps = 14/167 (8%)
Query: 133 YLISKLEEHKYTYYSRTQLESNT-IEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMP 191
Y++ K+ TY +LE + +F A I+ F V+V+D+T+ +Y
Sbjct: 329 YMLEKVNPGTVTY---VELEGEKKFKYLFIALGACIEGFRAMRKVIVVDATHLKTVYGGM 385
Query: 192 MFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLMK 251
+ Y + FG + EK+ +++W L+ L+ + S + V ++DR S+ K
Sbjct: 386 LVIATAHDPNHHHYPLAFGIIDSEKDVSWIWFLEKLKTVYSDVPRL--VFISDRHQSIKK 443
Query: 252 AVAHVFPESYAMNCYFHVQANVKQRCVLDCKYPLGFKKDGKEVSNRD 298
AV V+P + C +H+ N++ R +D KDG V RD
Sbjct: 444 AVKTVYPNALHAACIWHLCQNMRDRVKID--------KDGAAVKFRD 482
>At5g18960 FAR1 - like protein
Length = 788
Score = 60.8 bits (146), Expect = 9e-10
Identities = 65/285 (22%), Positives = 118/285 (40%), Gaps = 35/285 (12%)
Query: 5 RLVCERSGSHIVPKKKPKHAGTGSRKCGCLFMISGYQSKQTKEWRLNILNGVHNHPMEPA 64
R VC R G + P G CG I + + + W ++ LN HNH +EP
Sbjct: 256 RFVCSREGF-----QHPSRMG-----CGAYMRI---KRQDSGGWIVDRLNKDHNHDLEPG 302
Query: 65 LEGHILAGRLKEDDKKIVRDLTKNKMLPRNILIHLKNQRPHCMTNVKQVYNERQQIWKAN 124
+ AG KKI D T L LI L + + + ++ N + W
Sbjct: 303 KKND--AGM-----KKIPDDGTGG--LDSVDLIELNDFGNNHIKKTRE--NRIGKEWYPL 351
Query: 125 RGDKKPLQYLISKLEEHKYTYYS-RTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTY 183
L Y S+ E +Y+ + + + IFWA + + F +V D++Y
Sbjct: 352 L-----LDYFQSRQTEDMGFFYAVELDVNNGSCMSIFWADSRARFACSQFGDSVVFDTSY 406
Query: 184 KTNIYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVT 243
+ Y +P ++G +G + E +E F+W+ + + +S + P+ IV
Sbjct: 407 RKGSYSVPFATIIGFNHHRQPVLLGCAMVADESKEAFLWLFQTWLRAMSGR--RPRSIVA 464
Query: 244 DRDMSLMKAVAHVFPESYAMNCYFHVQANVKQRCVLDCKYPLGFK 288
D+D+ + +A+ VFP ++ + ++ ++ + +P FK
Sbjct: 465 DQDLPIQQALVQVFPGAHHRYSAWQIREKERENLI---PFPSEFK 506
>At2g27110 Mutator-like transposase
Length = 851
Score = 60.5 bits (145), Expect = 1e-09
Identities = 33/128 (25%), Positives = 59/128 (45%), Gaps = 2/128 (1%)
Query: 131 LQYLISKLEEHKYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRM 190
L+Y E+ +Y+ E N + ++FWA S + +F + +D+ Y+ N +R+
Sbjct: 201 LEYFKRMQAENPGFFYAVQLDEDNQMSNVFWADSRSRVAYTHFGDTVTLDTRYRCNQFRV 260
Query: 191 PMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLM 250
P GV G + E + +F+W+ K L + + P +VTD+D ++
Sbjct: 261 PFAPFTGVNHHGQAILFGCALILDESDTSFIWLFKTF--LTAMRDQPPVSLVTDQDRAIQ 318
Query: 251 KAVAHVFP 258
A VFP
Sbjct: 319 IAAGQVFP 326
>At3g47310 putative protein
Length = 735
Score = 58.9 bits (141), Expect = 3e-09
Identities = 42/167 (25%), Positives = 75/167 (44%), Gaps = 14/167 (8%)
Query: 133 YLISKLEEHKYTYYSRTQLESNT-IEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMP 191
Y++ K+ TY +LE + +F A I+ F V+V+D+T+ +Y
Sbjct: 318 YMLEKVNPGTVTY---VELEGEKKFKYLFIALGACIEGFRTMRKVIVVDATHLKTVYGGM 374
Query: 192 MFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLMK 251
+ Y + FG + E + +++W L+ L+ + S + V ++DR S+ K
Sbjct: 375 LVIATAQDPNHHHYPLAFGIIDSENDVSWIWFLEKLKTVYSDVPGL--VFISDRHQSIKK 432
Query: 252 AVAHVFPESYAMNCYFHVQANVKQRCVLDCKYPLGFKKDGKEVSNRD 298
V V+P + C +H+ N++ R +D KDG V RD
Sbjct: 433 VVKTVYPNALHAACIWHLCQNMRDRVKID--------KDGAAVKFRD 471
>At4g19990 putative protein
Length = 672
Score = 57.8 bits (138), Expect = 7e-09
Identities = 34/117 (29%), Positives = 58/117 (49%), Gaps = 5/117 (4%)
Query: 152 ESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMPMFEVVGVTSTDLTYSVGFG- 210
+ ++ +IFW + F V+ +D+T+ N Y++P+ GV +GFG
Sbjct: 182 DMQSLRNIFWVDAKGRFDYTCFSDVVSIDTTFIKNEYKLPLVAFTGVNHHGQFLLLGFGL 241
Query: 211 FMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYAMNCYF 267
+T E + FVW+ + K + P+VI+T D L +AV VFP S +C++
Sbjct: 242 LLTDESKSGFVWLFRAWLKAMHG--CRPRVILTKHDQMLKEAVLEVFPSS--RHCFY 294
>At3g32060 hypothetical protein
Length = 487
Score = 57.4 bits (137), Expect = 1e-08
Identities = 47/210 (22%), Positives = 98/210 (46%), Gaps = 12/210 (5%)
Query: 65 LEGHILAGRLKEDDKKIVRDLTKNKMLPRNILIHLKNQRPHCMTNVKQVYNERQQIWKAN 124
L H L G+++ +I+ +L + K+ + + + H + ++K R++ +K
Sbjct: 165 LHDHFL-GQVETPVPRIIMELVQTKLGVKVSYSTVLRGKYHAIYDLK---GSREESYK-- 218
Query: 125 RGDKKPLQYLISKLEEHKYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYK 184
D Y++ K+ + TY E++ + +F A SI+ F VL++D+T+
Sbjct: 219 --DINCYLYMLKKVNDGTVTYLKLD--ENDKFQYVFVALGASIEGFRVMRKVLIVDATHL 274
Query: 185 TNIYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTD 244
N Y + Y + F + E + ++ W + L+ ++ + V +TD
Sbjct: 275 KNGYGGVLVFASAQDPNRHHYIIAFAVLDGENDASWEWFFEKLKTVVPDTSEL--VFMTD 332
Query: 245 RDMSLMKAVAHVFPESYAMNCYFHVQANVK 274
R+ SL+KA+ +V+ ++ C +H+ NVK
Sbjct: 333 RNASLIKAIRNVYTAAHHGYCIWHLSQNVK 362
>At1g44840 hypothetical protein
Length = 926
Score = 55.8 bits (133), Expect = 3e-08
Identities = 30/118 (25%), Positives = 58/118 (48%), Gaps = 2/118 (1%)
Query: 159 IFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEE 218
+F A SI F + +LV+D T+ Y+ + G + Y +GF + E +E
Sbjct: 557 MFLAFGASIAGFRHLRRILVVDGTHLKGKYKGVLLTSSGQDANFQVYPLGFAVVDSENDE 616
Query: 219 NFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYAMNCYFHVQANVKQR 276
++ W L ++++ + I++DR S++ AV VFP++ C H+ N++ +
Sbjct: 617 SWTWFFTKLERIIADSKTL--TILSDRHSSILVAVKRVFPQANHGACIIHLCRNIQTK 672
>At2g23500 Mutator-like transposase
Length = 784
Score = 55.1 bits (131), Expect = 5e-08
Identities = 35/142 (24%), Positives = 69/142 (47%), Gaps = 4/142 (2%)
Query: 133 YLISKLEEHKYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMPM 192
Y++ K+ + TY E++ + +F A SI+ F VL++D+T+ N Y +
Sbjct: 355 YMLKKVNDGTVTYLKLD--ENDKFQYVFVALGASIEGFRVMRKVLIVDATHLKNGYGGVL 412
Query: 193 FEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLMKA 252
Y + F + E + ++ W + L+ ++ + V +TDR+ SL+KA
Sbjct: 413 VFASAQDPNRHHYIIAFAVLDGENDASWEWFFEKLKTVVPDTSEL--VFMTDRNASLIKA 470
Query: 253 VAHVFPESYAMNCYFHVQANVK 274
+ +V+ ++ C +H+ NVK
Sbjct: 471 IRNVYTAAHHGYCIWHLSQNVK 492
>At3g43050 putative protein
Length = 608
Score = 54.7 bits (130), Expect = 6e-08
Identities = 35/137 (25%), Positives = 64/137 (46%), Gaps = 2/137 (1%)
Query: 152 ESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMPMFEVVGVTSTDLTYSVGFGF 211
E + +F+A I+ FN V+++D+T+ + + Y + FG
Sbjct: 310 EKKNFQYLFFALGACIEGFNAMRKVIIVDATHLKIAQGGVLVVALAQDLEHHHYPIDFGV 369
Query: 212 MTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYAMNCYFHVQA 271
+ EK+ ++ W + L+ ++ + V V+DR+ SLMK+ ++P S C +H+
Sbjct: 370 LDGEKDVSWSWFFEKLKSVIPDSSEL--VFVSDRNQSLMKSQRELYPLSQHGCCIYHLCQ 427
Query: 272 NVKQRCVLDCKYPLGFK 288
NVK C K +G K
Sbjct: 428 NVKGACNYSKKKVVGKK 444
>At2g32250 Mutator-like transposase
Length = 684
Score = 53.5 bits (127), Expect = 1e-07
Identities = 40/205 (19%), Positives = 91/205 (43%), Gaps = 10/205 (4%)
Query: 75 KEDDKKIVRDLTK---NKMLPRNILIHLKNQ-RPHCMTNVKQVYNERQQIWKANRGDKKP 130
KED+K ++ + K +++ P + + ++ + +P +K+ Q+ K
Sbjct: 118 KEDEKWVIYNFVKEHNHEICPDDFYVSVRGKNKPAGALAIKKGL----QLALEEEDLKLL 173
Query: 131 LQYLISKLEEHKYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRM 190
L++ + ++ +Y+ + ++FW + + +F V++ D+ Y N YR+
Sbjct: 174 LEHFMEMQDKQPGFFYAVDFDSDKRVRNVFWLDAKAKHDYCSFSDVVLFDTFYVRNGYRI 233
Query: 191 PMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLM 250
P +GV+ +G + E + W+ + K + + P V++TD+D L
Sbjct: 234 PFAPFIGVSHHRQYVLLGCALIGEVSESTYSWLFRTWLKAVGGQ--APGVMITDQDKLLS 291
Query: 251 KAVAHVFPESYAMNCYFHVQANVKQ 275
V VFP+ + C + V + + +
Sbjct: 292 DIVVEVFPDVRHIFCLWSVLSKISE 316
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.136 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,000,615
Number of Sequences: 26719
Number of extensions: 288148
Number of successful extensions: 805
Number of sequences better than 10.0: 85
Number of HSP's better than 10.0 without gapping: 56
Number of HSP's successfully gapped in prelim test: 29
Number of HSP's that attempted gapping in prelim test: 709
Number of HSP's gapped (non-prelim): 92
length of query: 305
length of database: 11,318,596
effective HSP length: 99
effective length of query: 206
effective length of database: 8,673,415
effective search space: 1786723490
effective search space used: 1786723490
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC136507.18


