

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136507.1 - phase: 0
(236 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
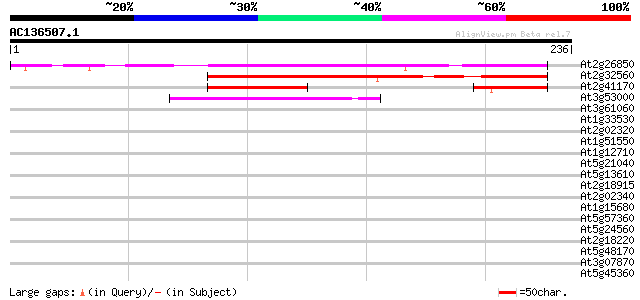
Score E
Sequences producing significant alignments: (bits) Value
At2g26850 unknown protein 152 1e-37
At2g32560 unknown protein 141 3e-34
At2g41170 unknown protein 54 7e-08
At3g53000 unknown protein 45 3e-05
At3g61060 unknown protein 37 0.007
At1g33530 hypothetical protein 35 0.027
At2g02320 unknown protein 35 0.035
At1g51550 unknown protein 35 0.035
At1g12710 unknown protein 33 0.10
At5g21040 unknown protein 33 0.13
At5g13610 unknown protein 31 0.50
At2g18915 Adagio 2 (ADO2)/ Lov kelch protein 2 (LKP2) 31 0.50
At2g02340 putative phloem-specific lectin 31 0.50
At1g15680 hypothetical protein 31 0.65
At5g57360 FKF1-like protein 2 (gb|AAF32300.1) 30 0.86
At5g24560 phloem-specific lectin-like protein 30 0.86
At2g18220 unknown protein 30 0.86
At5g48170 putative protein 30 1.1
At3g07870 unknown protein 30 1.1
At5g45360 putative F-box family protein 29 1.9
>At2g26850 unknown protein
Length = 371
Score = 152 bits (384), Expect = 1e-37
Identities = 95/229 (41%), Positives = 128/229 (55%), Gaps = 34/229 (14%)
Query: 1 MLLYF-ITCVSFFLFLKPLLLLKPLPSWTNEVR-LISLLFHKDFCLKSFCKLFRKTLVSR 58
MLLY ITC+SFF F K L P W ++ + L+S F K LF +L
Sbjct: 1 MLLYLLITCLSFFFFSKSL----SFPQWASKTKNLLSFYFSK--------YLFTNSLHQI 48
Query: 59 MSLVKKTLVSKVENVEEEVDSLQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRE 118
+ ++ K +S+LDLP+L L+CILE LPPS LC MA VC SLRE
Sbjct: 49 TPDLASPVLGK--------------MSILDLPDLPLDCILELLPPSELCTMARVCSSLRE 94
Query: 119 RCVSDHLWERHMKQKWGRVIGSVAYREWKWHVATKKNIGSLRHGKPR-SLIKFMSLYWPF 177
RCVSDHLWE+H+K KWG+++G A++EW+ ++++ ++ S H L K +SL
Sbjct: 95 RCVSDHLWEKHLKTKWGKILGPSAHKEWQCYLSSPYHLDSPHHKTSHLGLAKIISLMRSL 154
Query: 178 SWMRMKVDAAYSNSANKQNSYLPPDSFMTWYLALETGNFSFPAQVYNRE 226
S + D + S +P DS M +YL+LETG F FPAQVYNRE
Sbjct: 155 SSIFRDDD-----HRRRYPSSIPLDSTMNFYLSLETGRFWFPAQVYNRE 198
>At2g32560 unknown protein
Length = 228
Score = 141 bits (356), Expect = 3e-34
Identities = 76/151 (50%), Positives = 98/151 (64%), Gaps = 19/151 (12%)
Query: 84 LSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWGRVIGSVAY 143
+SVLDLPEL L+CIL+ LPPS LC MA VC SLRERCVSDHLWE+H+K KWG+++G A+
Sbjct: 1 MSVLDLPELALDCILDLLPPSGLCSMARVCSSLRERCVSDHLWEKHLKTKWGKILGPAAH 60
Query: 144 REWKWHVATK--------KNIGSLRHGKPRSLIKFMSLYWPFSWMRMKVDAAYSNSANKQ 195
REW+ ++++ G+L K SLI+ +S + K Y++S
Sbjct: 61 REWQCYISSSTYHLDSPHHQTGNLGFAKIISLIRSLSSV----FREDKQRRGYASS---- 112
Query: 196 NSYLPPDSFMTWYLALETGNFSFPAQVYNRE 226
LP DS M+ YL+LETG F FPAQVYNRE
Sbjct: 113 ---LPLDSSMSCYLSLETGRFWFPAQVYNRE 140
>At2g41170 unknown protein
Length = 371
Score = 53.9 bits (128), Expect = 7e-08
Identities = 24/42 (57%), Positives = 31/42 (73%)
Query: 84 LSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHL 125
+S+LDLP+L L+CILEKL PS LC M VC LR++ S +L
Sbjct: 1 MSLLDLPDLTLDCILEKLSPSELCAMTSVCSELRDKGSSSYL 42
Score = 44.3 bits (103), Expect = 6e-05
Identities = 22/32 (68%), Positives = 23/32 (71%), Gaps = 1/32 (3%)
Query: 196 NSYLPP-DSFMTWYLALETGNFSFPAQVYNRE 226
+SYL P DS M WY LE G F FPAQVYNRE
Sbjct: 39 SSYLAPIDSVMYWYSNLENGKFWFPAQVYNRE 70
>At3g53000 unknown protein
Length = 300
Score = 45.4 bits (106), Expect = 3e-05
Identities = 22/89 (24%), Positives = 45/89 (49%), Gaps = 2/89 (2%)
Query: 68 SKVENVEEEVDSLQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWE 127
S + N+ + + L + D+PE + C+ L P +C +AG+ S R SD +WE
Sbjct: 3 SSLSNLNDGTNGLAMGPGLGDIPESCVACVFMYLTPPEICNLAGLNRSFRGAASSDSVWE 62
Query: 128 RHMKQKWGRVIGSVAYREWKWHVATKKNI 156
+ + + + ++ + ++H +KK+I
Sbjct: 63 KKLPENYQDLLDLLPPE--RYHSLSKKDI 89
>At3g61060 unknown protein
Length = 290
Score = 37.4 bits (85), Expect = 0.007
Identities = 21/79 (26%), Positives = 38/79 (47%), Gaps = 1/79 (1%)
Query: 78 DSLQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWGRV 137
D + L ++DLPE + I+ +L P +C++A + R +D +WE + + RV
Sbjct: 17 DVYSRKLRLVDLPENCVALIMTRLDPPEICRLARLNRMFRRASSADFIWESKLPANY-RV 75
Query: 138 IGSVAYREWKWHVATKKNI 156
I + E KK++
Sbjct: 76 IAHKVFDEITLTKLIKKDL 94
>At1g33530 hypothetical protein
Length = 441
Score = 35.4 bits (80), Expect = 0.027
Identities = 15/53 (28%), Positives = 31/53 (58%)
Query: 79 SLQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMK 131
S + ++LP++++E IL++LP L ++ + + SDHL E+H++
Sbjct: 87 SCSETTLAVELPDVLVEEILQRLPVKYLVRLKSISKGWKSLIESDHLAEKHLR 139
>At2g02320 unknown protein
Length = 307
Score = 35.0 bits (79), Expect = 0.035
Identities = 27/106 (25%), Positives = 48/106 (44%), Gaps = 12/106 (11%)
Query: 51 FRKTLVSRMSLVKKTLVSKVENVEEEVDSLQQNLSVLDLPELVLECILEKLPPSSLCQMA 110
FRK L VKKTL ++ +++ + + LS+ DLPE + I+ P C A
Sbjct: 11 FRKIL----QRVKKTL--RLSASDQQSQGVTEPLSLGDLPEECISLIISFTSPRDACVFA 64
Query: 111 GVCHSLRERCVSDHLWERHMKQKWGRVIGSVAYREWKWHVATKKNI 156
V + SD +WE+ + ++ ++ H ++KK +
Sbjct: 65 LVSKTFESAVQSDIVWEKFIPPEYESLLSR------SQHFSSKKEL 104
>At1g51550 unknown protein
Length = 478
Score = 35.0 bits (79), Expect = 0.035
Identities = 17/55 (30%), Positives = 28/55 (50%)
Query: 81 QQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWG 135
Q + +++LP+ L IL LP S+ + C + SD LWE +++WG
Sbjct: 16 QSSSPIINLPDDHLLTILLLLPVDSILSFSMTCKRYKSLACSDSLWEALCEREWG 70
>At1g12710 unknown protein
Length = 291
Score = 33.5 bits (75), Expect = 0.10
Identities = 15/54 (27%), Positives = 28/54 (51%)
Query: 88 DLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWGRVIGSV 141
DLPE + I+E L P +C+ + + + R +D +WE + Q + V+ +
Sbjct: 30 DLPEACVAIIVENLDPVEICRFSKLNRAFRGASWADCVWESKLPQNYRDVLEKI 83
>At5g21040 unknown protein
Length = 539
Score = 33.1 bits (74), Expect = 0.13
Identities = 14/51 (27%), Positives = 26/51 (50%)
Query: 85 SVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWG 135
+++DLP+ ++ IL L P L ++ V L H W+ +++WG
Sbjct: 67 TIIDLPQALISEILNCLDPKELGLVSCVSTYLHRLASEHHAWKEFYRERWG 117
>At5g13610 unknown protein
Length = 402
Score = 31.2 bits (69), Expect = 0.50
Identities = 16/38 (42%), Positives = 22/38 (57%), Gaps = 1/38 (2%)
Query: 193 NKQNSYLPPDSFMTWYLALETGNFSFPAQVYNREVTNS 230
NKQN ++PP S MT Y+ L+ GN S P ++ S
Sbjct: 141 NKQN-FIPPTSRMTNYVVLKFGNHSDPTDTTRGRISGS 177
>At2g18915 Adagio 2 (ADO2)/ Lov kelch protein 2 (LKP2)
Length = 601
Score = 31.2 bits (69), Expect = 0.50
Identities = 17/55 (30%), Positives = 26/55 (46%), Gaps = 4/55 (7%)
Query: 91 ELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWG----RVIGSV 141
E++ IL +L P + + VC L E +D +W + WG RV+ SV
Sbjct: 194 EVIAIKILSQLTPGDIASVGCVCRRLNELTKNDDVWRMVCQNTWGTEATRVLESV 248
>At2g02340 putative phloem-specific lectin
Length = 305
Score = 31.2 bits (69), Expect = 0.50
Identities = 26/88 (29%), Positives = 38/88 (42%), Gaps = 10/88 (11%)
Query: 51 FRKTLVSRMSLVKKTLVSKVENVEEEVDSLQQNLSVLDLPELVLECILEKLPPSSLCQMA 110
FRK +V R VKKTL + V L DLPE + I+ P C +A
Sbjct: 11 FRK-IVQR---VKKTLRLSASDKSHGVAELD------DLPEECVSIIVSFTSPQDACVLA 60
Query: 111 GVCHSLRERCVSDHLWERHMKQKWGRVI 138
V + SD +WE+ + ++ +I
Sbjct: 61 SVSKTFASAVKSDIVWEKFIPPEYESLI 88
>At1g15680 hypothetical protein
Length = 410
Score = 30.8 bits (68), Expect = 0.65
Identities = 27/88 (30%), Positives = 39/88 (43%), Gaps = 12/88 (13%)
Query: 87 LDLPELVLECILEKLPPSSLCQMAGVC---HSLRERCVSDHLW---ERHMKQKWGRVIGS 140
++LPE +L I+ +LP S+ + VC SL E HL+ R+ W V G+
Sbjct: 17 IELPEELLAEIVARLPFISITRFKSVCKGWRSLIESTYFRHLFVFAHRNSSSSWSLVCGT 76
Query: 141 VAYREWKWHVATKKNI-GSLRHGKPRSL 167
+ W V G R+G PR L
Sbjct: 77 -----FGWSVEEMAGFYGCKRYGLPRRL 99
>At5g57360 FKF1-like protein 2 (gb|AAF32300.1)
Length = 609
Score = 30.4 bits (67), Expect = 0.86
Identities = 14/45 (31%), Positives = 22/45 (48%)
Query: 91 ELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWG 135
E+V IL +L P + ++ VC L ++ LW R + WG
Sbjct: 203 EVVSMKILSRLTPRDVASVSSVCRRLYVLTKNEDLWRRVCQNAWG 247
>At5g24560 phloem-specific lectin-like protein
Length = 251
Score = 30.4 bits (67), Expect = 0.86
Identities = 14/47 (29%), Positives = 24/47 (50%)
Query: 84 LSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHM 130
++ LDLPE + ++ P C+++ V LR S+ WER +
Sbjct: 1 MNFLDLPEECIATMISFTSPFDACRISAVSKLLRSAADSNTTWERFL 47
>At2g18220 unknown protein
Length = 779
Score = 30.4 bits (67), Expect = 0.86
Identities = 26/100 (26%), Positives = 41/100 (41%), Gaps = 4/100 (4%)
Query: 20 LLKPLPSWTNEVRLISLLFHKDFCLKSFCKLFRKTLVSRMSLVKKTLVSKVENVEEEVDS 79
L++ L W+ V L F L+SFCK T R K L+S++E E V+
Sbjct: 527 LVEHLSQWSCSVAFFELSFIPTIRLRSFCK---STKAERFRKEMKQLISQIEANSEFVNK 583
Query: 80 LQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRER 119
+ + L +L E LE + + +R+R
Sbjct: 584 KRALIKFLP-NDLAAESFLEDEKKAGKTPLLQYAEIIRQR 622
>At5g48170 putative protein
Length = 157
Score = 30.0 bits (66), Expect = 1.1
Identities = 20/62 (32%), Positives = 32/62 (51%), Gaps = 2/62 (3%)
Query: 66 LVSKVENVEEEVDSLQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHL 125
+V K N + V+ + S+ D ++++E IL +L SSLC A VC +D +
Sbjct: 9 MVEKSNNKRQRVNQVPV-FSINDHHDVLVE-ILRRLDGSSLCSAACVCRLWSAVARNDSI 66
Query: 126 WE 127
WE
Sbjct: 67 WE 68
>At3g07870 unknown protein
Length = 417
Score = 30.0 bits (66), Expect = 1.1
Identities = 15/49 (30%), Positives = 26/49 (52%)
Query: 69 KVENVEEEVDSLQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLR 117
K + ++VD + + LPE ++ I +LP SS+ ++ VC S R
Sbjct: 8 KKRKITDDVDGVGVGGGLESLPEDIIADIFSRLPISSIARLMFVCRSWR 56
>At5g45360 putative F-box family protein
Length = 316
Score = 29.3 bits (64), Expect = 1.9
Identities = 17/61 (27%), Positives = 25/61 (40%)
Query: 88 DLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWGRVIGSVAYREWK 147
++P + IL+ L L + VC L + LW R +WG + S RE
Sbjct: 73 NVPTELFRHILKFLSSEDLVSCSLVCKFLNFAAADESLWRRLYCIRWGLTLPSRKLRESA 132
Query: 148 W 148
W
Sbjct: 133 W 133
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.325 0.137 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,566,221
Number of Sequences: 26719
Number of extensions: 228548
Number of successful extensions: 932
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 31
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 895
Number of HSP's gapped (non-prelim): 43
length of query: 236
length of database: 11,318,596
effective HSP length: 96
effective length of query: 140
effective length of database: 8,753,572
effective search space: 1225500080
effective search space used: 1225500080
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 58 (26.9 bits)
Medicago: description of AC136507.1


