

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136472.16 - phase: 0
(212 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
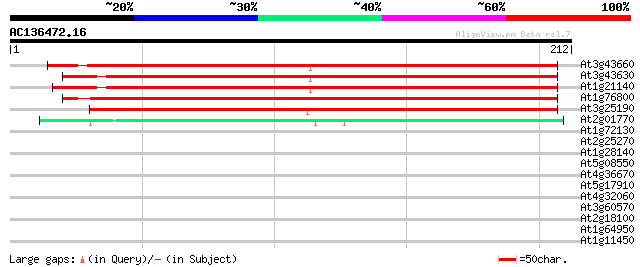
Score E
Sequences producing significant alignments: (bits) Value
At3g43660 nodulin - like protein 258 2e-69
At3g43630 nodulin -like protein 257 4e-69
At1g21140 putative tonoplast intrinsic protein, alpha (alpha-TIP) 253 4e-68
At1g76800 nodulin-like protein 244 2e-65
At3g25190 unknown protein 223 8e-59
At2g01770 putative membrane protein 57 5e-09
At1g72130 peptide transporter PTR2-B -like protein 29 1.6
At2g25270 unknown protein 28 2.7
At1g28140 unknown protein 28 2.7
At5g08550 putative protein 28 4.6
At4g36670 sugar transporter like protein 28 4.6
At5g17910 putative protein 27 6.1
At4g32060 unknown protein (At4g32060) 27 7.9
At3g60570 putative protein 27 7.9
At2g18100 unknown protein 27 7.9
At1g64950 unknown protein 27 7.9
At1g11450 similar to MtN21 emb|CAA75575; similar to EST gb|H77065 27 7.9
>At3g43660 nodulin - like protein
Length = 198
Score = 258 bits (658), Expect = 2e-69
Identities = 134/194 (69%), Positives = 162/194 (83%), Gaps = 4/194 (2%)
Query: 15 SESKVYNMDVEKKGEEENDEKDYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVKT 74
S + N+D+EK ++ DY++RAQWLRAAVLGANDGL+STASLMMG+GAV +DV+
Sbjct: 3 SNNNNLNLDMEK---DQETTFDYSKRAQWLRAAVLGANDGLVSTASLMMGIGAVKQDVRI 59
Query: 75 MILTGIAGLVAGACSMAIGEFVSVYSQYDIEFAQMKRQ-GNISQKDKLPNPYYAAFASAI 133
M+LTG AGLVAGACSMAIGEF+SVYSQYDIE AQMKR+ G ++K+KLP+P AA ASA+
Sbjct: 60 MLLTGFAGLVAGACSMAIGEFISVYSQYDIEVAQMKRESGGETKKEKLPSPTQAAIASAL 119
Query: 134 AFAVGAFVPLLGAAFVKDYKVRLGVVVGVVSLALFGFGLLSAVLGKAPLVKSSLRVLIGG 193
AF +GA VPLL AAFVK+YKVR+GV+V V+LAL FG L AVLGKAP+VKS +RVLIGG
Sbjct: 120 AFTLGAIVPLLAAAFVKEYKVRIGVIVAAVTLALVMFGWLGAVLGKAPVVKSLVRVLIGG 179
Query: 194 WLAMSLTFGLTKLV 207
WLAM++TFG TKLV
Sbjct: 180 WLAMAITFGFTKLV 193
>At3g43630 nodulin -like protein
Length = 200
Score = 257 bits (656), Expect = 4e-69
Identities = 135/188 (71%), Positives = 160/188 (84%), Gaps = 4/188 (2%)
Query: 21 NMDVEKKGEEENDEKDYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVKTMILTGI 80
N+D+EK E+ D Y++RAQWLRAAVLGANDGL+STASLMMGVGAV ++VK MILTG
Sbjct: 8 NLDMEKDQEKAFD---YSKRAQWLRAAVLGANDGLVSTASLMMGVGAVKQNVKIMILTGF 64
Query: 81 AGLVAGACSMAIGEFVSVYSQYDIEFAQMKRQ-GNISQKDKLPNPYYAAFASAIAFAVGA 139
AGLVAGACSMAIGEFVSVYSQYDIE AQMKR+ G +K+KLP+P AA ASA+AF++GA
Sbjct: 65 AGLVAGACSMAIGEFVSVYSQYDIEVAQMKRETGGEIEKEKLPSPTQAAAASALAFSLGA 124
Query: 140 FVPLLGAAFVKDYKVRLGVVVGVVSLALFGFGLLSAVLGKAPLVKSSLRVLIGGWLAMSL 199
VPLL AAFVK+YKVR+G +V V+LAL FG L AVLGKAP+VKSSLRVL+GGWLAM++
Sbjct: 125 MVPLLAAAFVKEYKVRIGAIVAAVTLALVMFGWLGAVLGKAPVVKSSLRVLVGGWLAMAI 184
Query: 200 TFGLTKLV 207
T+G TKL+
Sbjct: 185 TYGFTKLI 192
>At1g21140 putative tonoplast intrinsic protein, alpha (alpha-TIP)
Length = 200
Score = 253 bits (647), Expect = 4e-68
Identities = 135/192 (70%), Positives = 158/192 (81%), Gaps = 4/192 (2%)
Query: 17 SKVYNMDVEKKGEEENDEKDYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVKTMI 76
S N+D+E E+ D Y++RAQWLRAAVLGANDGL+STASLMMGVGAV +DVK MI
Sbjct: 7 SNSLNLDMEMDQEKAFD---YSKRAQWLRAAVLGANDGLVSTASLMMGVGAVKQDVKVMI 63
Query: 77 LTGIAGLVAGACSMAIGEFVSVYSQYDIEFAQMKRQ-GNISQKDKLPNPYYAAFASAIAF 135
L+G AGLVAGACSMAIGEFVSVYSQYDIE AQMKR+ G +K+KLP+P AA ASA+AF
Sbjct: 64 LSGFAGLVAGACSMAIGEFVSVYSQYDIEVAQMKRENGGQVEKEKLPSPMQAAAASALAF 123
Query: 136 AVGAFVPLLGAAFVKDYKVRLGVVVGVVSLALFGFGLLSAVLGKAPLVKSSLRVLIGGWL 195
++GA VPL+ AAFVKDY VR+G +V V+LAL FG L AVLGKAP+ KSS RVLIGGWL
Sbjct: 124 SLGAIVPLMAAAFVKDYHVRIGAIVAAVTLALVMFGWLGAVLGKAPVFKSSARVLIGGWL 183
Query: 196 AMSLTFGLTKLV 207
AM++TFGLTKL+
Sbjct: 184 AMAVTFGLTKLI 195
>At1g76800 nodulin-like protein
Length = 196
Score = 244 bits (624), Expect = 2e-65
Identities = 129/187 (68%), Positives = 152/187 (80%), Gaps = 4/187 (2%)
Query: 21 NMDVEKKGEEENDEKDYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVKTMILTGI 80
NMD+EK E+ DY++R+QWLRAAVLGANDGL+STASLMMGVGAV DVK MIL+G
Sbjct: 9 NMDIEK----ESTTFDYSKRSQWLRAAVLGANDGLVSTASLMMGVGAVKHDVKAMILSGF 64
Query: 81 AGLVAGACSMAIGEFVSVYSQYDIEFAQMKRQGNISQKDKLPNPYYAAFASAIAFAVGAF 140
AG+VAGACSMAIGEFVSVYSQYDIE AQM+R +K+KLP+P AA ASA+AF+ GA
Sbjct: 65 AGMVAGACSMAIGEFVSVYSQYDIEVAQMERDSVEIEKEKLPSPMQAAAASALAFSAGAI 124
Query: 141 VPLLGAAFVKDYKVRLGVVVGVVSLALFGFGLLSAVLGKAPLVKSSLRVLIGGWLAMSLT 200
VPLL AAFVK+YK+R+ VV V++AL FG L A LGKAP V+SS RVL GGWLAM++T
Sbjct: 125 VPLLAAAFVKEYKMRIISVVVAVTVALMVFGWLGAALGKAPAVRSSARVLFGGWLAMAVT 184
Query: 201 FGLTKLV 207
FGLTKL+
Sbjct: 185 FGLTKLI 191
>At3g25190 unknown protein
Length = 219
Score = 223 bits (567), Expect = 8e-59
Identities = 117/190 (61%), Positives = 146/190 (76%), Gaps = 13/190 (6%)
Query: 31 ENDEKDYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVKTMILTGIAGLVAGACSM 90
E +E DY QRAQWLRAA+LGANDGL++ ASLMMGVG++ +DVK M+L G AGLVAGACSM
Sbjct: 23 EKEEVDYMQRAQWLRAALLGANDGLVTVASLMMGVGSIKEDVKAMLLVGFAGLVAGACSM 82
Query: 91 AIGEFVSVYSQYDIEFAQMKR-------------QGNISQKDKLPNPYYAAFASAIAFAV 137
AIGEFVSV +Q DIE AQMKR Q +K++LPNP AA ASA+AF+V
Sbjct: 83 AIGEFVSVCTQRDIETAQMKRAIEHKTSLSAIDEQEEEEKKERLPNPGQAAIASALAFSV 142
Query: 138 GAFVPLLGAAFVKDYKVRLGVVVGVVSLALFGFGLLSAVLGKAPLVKSSLRVLIGGWLAM 197
GA +PLLGA F++++KVR+ VV V ++AL FG+ AVLGK +VKSS+RV+IGGW+AM
Sbjct: 143 GAAMPLLGAVFIENHKVRMVVVAVVATIALVVFGVTGAVLGKTSVVKSSVRVVIGGWMAM 202
Query: 198 SLTFGLTKLV 207
+LTFGLTK +
Sbjct: 203 ALTFGLTKFI 212
>At2g01770 putative membrane protein
Length = 250
Score = 57.4 bits (137), Expect = 5e-09
Identities = 54/251 (21%), Positives = 98/251 (38%), Gaps = 54/251 (21%)
Query: 12 LARSESKVYNMDVEKKGE---EENDEKDYTQRAQWLRAAVLGANDGLLSTASLMMGVGAV 68
++ E K+ + +E + + + + EK +T + +R ++G +DGL +L G+
Sbjct: 1 MSSEEDKITRISIEPEKQTLLDHHTEKHFTA-GEIVRDIIIGVSDGLTVPFALAAGLSGA 59
Query: 69 TKDVKTMILTGIAGLVAGACSMAIGEFVSVYSQYDIEFAQMKRQGN-------------- 114
++ GIA + AGA SM +G +++ S+ D +MKR+
Sbjct: 60 NASSSIVLTAGIAEVAAGAISMGLGGYLAAKSEEDHYAREMKREQEEIVAVPETEAAEVA 119
Query: 115 --ISQKDKLPNPY----------------------------------YAAFASAIAFAVG 138
++Q P+ Y +AF AIA+ +G
Sbjct: 120 EILAQYGIEPHEYSPVVNALRKNPQAWLDFMMRFELGLEKPDPKRALQSAFTIAIAYVLG 179
Query: 139 AFVPLLGAAFVKDYKVRLGVVVGVVSLALFGFGLLSAVLGKAPLVKSSLRVLIGGWLAMS 198
F+PLL + + V + ALF FG + ++S+ G +A +
Sbjct: 180 GFIPLLPYMLIPHAMDAVVASVVITLFALFIFGYAKGHFTGSKPLRSAFETAFIGAIASA 239
Query: 199 LTFGLTKLVNH 209
F L K+V H
Sbjct: 240 AAFCLAKVVQH 250
>At1g72130 peptide transporter PTR2-B -like protein
Length = 538
Score = 29.3 bits (64), Expect = 1.6
Identities = 15/36 (41%), Positives = 19/36 (52%)
Query: 139 AFVPLLGAAFVKDYKVRLGVVVGVVSLALFGFGLLS 174
AF+PLLG Y R ++ SL + G GLLS
Sbjct: 80 AFLPLLGGFLADSYLGRFRTIIISSSLYILGLGLLS 115
>At2g25270 unknown protein
Length = 545
Score = 28.5 bits (62), Expect = 2.7
Identities = 22/64 (34%), Positives = 37/64 (57%), Gaps = 6/64 (9%)
Query: 148 FVKDYKVRLGVVVGVVSLALFGFGLLSAVLGKAPLVKSSLRVLIGGWLAMSLTFGL--TK 205
F+ +V L +VV +V L + GL+S++ G +V + ++I GW+ ++ TF L T
Sbjct: 254 FLDSVRVAL-IVVSIVMLVVTFLGLVSSIFGMQVIVYT---LVILGWILVTGTFILSGTF 309
Query: 206 LVNH 209
LV H
Sbjct: 310 LVLH 313
>At1g28140 unknown protein
Length = 280
Score = 28.5 bits (62), Expect = 2.7
Identities = 24/88 (27%), Positives = 38/88 (42%), Gaps = 8/88 (9%)
Query: 125 YYAAFASAIAFAVGAFVPLLGAAFVKDYKVRLGVVVGVVSLALFGFGLLSAVLGK-APLV 183
Y SA+ F F L G F + G+ G + L F + S +LG + L+
Sbjct: 166 YVVDHPSAVWFVGPLFASLTGLVFKE------GLCYGKLEAGLLTFIIPSVLLGHLSGLM 219
Query: 184 KSSLR-VLIGGWLAMSLTFGLTKLVNHV 210
++ VL+G W+A+ L F K +
Sbjct: 220 NDEVKLVLLGTWMALFLVFAGRKFTQPI 247
>At5g08550 putative protein
Length = 908
Score = 27.7 bits (60), Expect = 4.6
Identities = 13/42 (30%), Positives = 23/42 (53%)
Query: 5 QHHHAATLARSESKVYNMDVEKKGEEENDEKDYTQRAQWLRA 46
Q H +L +V ++D +++GEEE E + +A +RA
Sbjct: 153 QDHEQQSLKDVVKQVSDLDFDEEGEEEQHEDAFADQAAIIRA 194
>At4g36670 sugar transporter like protein
Length = 493
Score = 27.7 bits (60), Expect = 4.6
Identities = 34/164 (20%), Positives = 64/164 (38%), Gaps = 31/164 (18%)
Query: 48 VLGANDGLLSTASLMMGVGAVTKDVKTMILTGIAGLVAGACSMAIGEFVSVYSQYDIEFA 107
+ G + G++S A + + T DV+ +LTGI L A S+ G + +
Sbjct: 30 IFGYDTGVMSGAMVFIEEDLKTNDVQIEVLTGILNLCALVGSLLAGRTSDIIGR------ 83
Query: 108 QMKRQGNISQKDKLPNPYYAAFASAIAFAVGAFVPLLGAAFVKDYKVRLGVVVGVVSLAL 167
Y ++I F +G+ + G + R +GV
Sbjct: 84 -----------------RYTIVLASILFMLGSILMGWGPNYPVLLSGRCTAGLGV----- 121
Query: 168 FGFGLLSAVLGKAPLVKSSLRVLIGGWLAMSLTFGLTKLVNHVV 211
GF L+ A + A + +S R L+ + ++ G+ L+ ++V
Sbjct: 122 -GFALMVAPVYSAEIATASHRGLLASLPHLCISIGI--LLGYIV 162
>At5g17910 putative protein
Length = 1342
Score = 27.3 bits (59), Expect = 6.1
Identities = 11/24 (45%), Positives = 17/24 (70%)
Query: 11 TLARSESKVYNMDVEKKGEEENDE 34
T R E K + +DVEKKG+ E+++
Sbjct: 171 THVRFEEKAFILDVEKKGDREDEK 194
>At4g32060 unknown protein (At4g32060)
Length = 498
Score = 26.9 bits (58), Expect = 7.9
Identities = 25/85 (29%), Positives = 40/85 (46%), Gaps = 11/85 (12%)
Query: 104 IEFAQM--KRQGNISQKDKLPNPYYAAFASAIAFAVGAFVPLLGAAFVKDYKV------- 154
+EFA KR+G+IS KD + AA AS ++ + L ++D ++
Sbjct: 336 LEFAHYDYKRRGSISAKDFALSMVAAADASHLSKLLDRVESLSEHPHLRDMRISLKEFKQ 395
Query: 155 --RLGVVVGVVSLALFGFGLLSAVL 177
L +G SLALF +G + +L
Sbjct: 396 FDELRSKLGPFSLALFAYGKANGLL 420
>At3g60570 putative protein
Length = 252
Score = 26.9 bits (58), Expect = 7.9
Identities = 24/69 (34%), Positives = 30/69 (42%), Gaps = 7/69 (10%)
Query: 123 NPYYAAFASAIAFAVGAF----VPLLGAAFVKDYKVRLGV---VVGVVSLALFGFGLLSA 175
NPYY +FA A G F V G ++K ++R V GV F L SA
Sbjct: 158 NPYYMSFAVKFANGDGNFACIEVQPAGGQYMKMEEMRSAVWRLSPGVPLKGPFNIRLTSA 217
Query: 176 VLGKAPLVK 184
V GK + K
Sbjct: 218 VSGKKIIAK 226
>At2g18100 unknown protein
Length = 656
Score = 26.9 bits (58), Expect = 7.9
Identities = 20/57 (35%), Positives = 27/57 (47%), Gaps = 1/57 (1%)
Query: 121 LPNPYYAAFASAIAFAVGAFVPLLGAAFVKDYKVRLGVVVGVVSLALFGFGLLSAVL 177
L P AA A+A +G VP++GA+ G V G V++A FG A L
Sbjct: 321 LAAPAIAAGFGALAPTLGTIVPVIGASGFAAAAEAAGTVAGSVAVAA-SFGAAGAGL 376
>At1g64950 unknown protein
Length = 510
Score = 26.9 bits (58), Expect = 7.9
Identities = 13/39 (33%), Positives = 17/39 (43%)
Query: 1 MAEIQHHHAATLARSESKVYNMDVEKKGEEENDEKDYTQ 39
M QH L R+ K+ + EEE D K+Y Q
Sbjct: 241 MRREQHDVLLPLIRARRKIVEERKNRSSEEEEDNKEYVQ 279
>At1g11450 similar to MtN21 emb|CAA75575; similar to EST gb|H77065
Length = 352
Score = 26.9 bits (58), Expect = 7.9
Identities = 14/43 (32%), Positives = 23/43 (52%)
Query: 63 MGVGAVTKDVKTMILTGIAGLVAGACSMAIGEFVSVYSQYDIE 105
+ +G+V VK + G+ ++ GA MAI F+ V Y +E
Sbjct: 25 VAMGSVNALVKKALDVGVNHMIIGAYRMAISSFILVPIAYFLE 67
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.135 0.380
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,087,904
Number of Sequences: 26719
Number of extensions: 152405
Number of successful extensions: 590
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 12
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 572
Number of HSP's gapped (non-prelim): 19
length of query: 212
length of database: 11,318,596
effective HSP length: 95
effective length of query: 117
effective length of database: 8,780,291
effective search space: 1027294047
effective search space used: 1027294047
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC136472.16


