

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
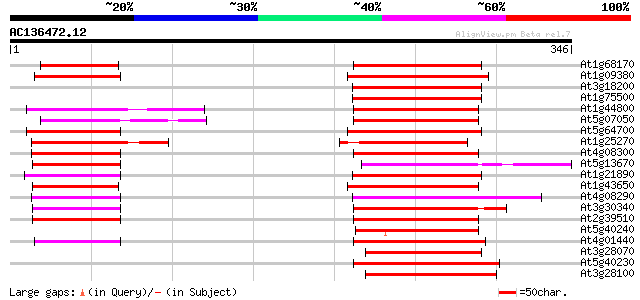
Query= AC136472.12 - phase: 0 /pseudo
(346 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At1g68170 MtN21-like protein 101 5e-22
At1g09380 nodulin N21 like protein 89 5e-18
At3g18200 integral membrane protein, putative 87 2e-17
At1g75500 nodulin-like protein 84 9e-17
At1g44800 nodulin like protein 84 9e-17
At5g07050 MtN21 nodulin protein-like 84 1e-16
At5g64700 nodulin-like protein 83 3e-16
At1g25270 83 3e-16
At4g08300 nodulin-like protein 82 6e-16
At5g13670 unknown protein 79 4e-15
At1g21890 nodulin-like protein 79 4e-15
At1g43650 nodulin-like protein 79 5e-15
At4g08290 nodulin-like protein 78 6e-15
At3g30340 unknown protein 78 8e-15
At2g39510 nodulin-like protein 74 2e-13
At5g40240 unknown protein 73 3e-13
At4g01440 predicted protein of unknown function 73 3e-13
At3g28070 unknown protein 72 3e-13
At5g40230 putative protein 72 4e-13
At3g28100 unknown protein 72 4e-13
>At1g68170 MtN21-like protein
Length = 329
Score = 101 bits (252), Expect = 5e-22
Identities = 47/79 (59%), Positives = 63/79 (79%)
Query: 213 SVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGA 272
++AYA I++SG +VAV +WC+ RGPLF SVF+P+ L++VAL G +L+E LHLGSIIG
Sbjct: 220 TIAYAAILISGMVVAVNAWCIESRGPLFVSVFSPVGLVIVALVGSFLLDETLHLGSIIGT 279
Query: 273 VLIVCGLYAVVWGKSKEMK 291
V+IV LY V+W K+KEMK
Sbjct: 280 VIIVGALYIVLWAKNKEMK 298
Score = 60.5 bits (145), Expect = 1e-09
Identities = 27/48 (56%), Positives = 38/48 (78%)
Query: 20 MVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILE 67
MV+VQIA AG+N+ +KLA+ DGM+ ++VAYR +FAT F+ PI FI +
Sbjct: 1 MVVVQIATAGLNIFFKLAMEDGMNPSVLVAYRLLFATLFMIPICFIFQ 48
>At1g09380 nodulin N21 like protein
Length = 374
Score = 88.6 bits (218), Expect = 5e-18
Identities = 37/87 (42%), Positives = 60/87 (68%)
Query: 209 LAYYSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGS 268
L + S YAG+V S ++SW + +GPL+ SVF+PL+L++VA+ +L EKL+ G+
Sbjct: 249 LRFISALYAGVVASALAFCLMSWAMQRKGPLYVSVFSPLLLVVVAIFSWALLEEKLYTGT 308
Query: 269 IIGAVLIVCGLYAVVWGKSKEMKKKNQ 295
+G+ L+V GLY V+WGK +E+ +K +
Sbjct: 309 FMGSALVVIGLYGVLWGKDREVSEKEE 335
Score = 57.4 bits (137), Expect = 1e-08
Identities = 29/53 (54%), Positives = 37/53 (69%)
Query: 16 PTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILER 68
P L MV+VQI +AG+N+ K+A+ GM I+VAYR IFAT P+AF LER
Sbjct: 8 PFLAMVLVQIGYAGMNITSKMAMEAGMKPLILVAYRQIFATIATFPVAFFLER 60
>At3g18200 integral membrane protein, putative
Length = 360
Score = 86.7 bits (213), Expect = 2e-17
Identities = 37/80 (46%), Positives = 57/80 (71%)
Query: 212 YSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIG 271
+++ YAGI+ SG +V + +WC++ GP+F +VF PL L+VA IL ++L+ G I+G
Sbjct: 250 FTILYAGIIASGLVVYLQTWCIYKSGPVFVAVFQPLQTLLVAAMAFLILGDQLYSGGIVG 309
Query: 272 AVLIVCGLYAVVWGKSKEMK 291
AV I+ GLY V+WGK++E K
Sbjct: 310 AVFIMLGLYLVLWGKNEERK 329
Score = 40.0 bits (92), Expect = 0.002
Identities = 22/85 (25%), Positives = 45/85 (52%), Gaps = 6/85 (7%)
Query: 10 VVQGLKPTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILERL 69
V + +K + ++ +Q FAG +++ ++A+N G+S + YR + A I P A+ E+
Sbjct: 6 VSEKVKLVVALITLQFCFAGFHIVSRVALNIGVSKVVYPVYRNLLALLLIGPFAYFFEK- 64
Query: 70 QEEEGKDDLDNTVSVISMRFIWGII 94
K+ T+S+++ F +I
Sbjct: 65 -----KERPPLTISLLAQFFFLALI 84
>At1g75500 nodulin-like protein
Length = 389
Score = 84.3 bits (207), Expect = 9e-17
Identities = 39/80 (48%), Positives = 55/80 (68%)
Query: 212 YSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIG 271
+++ YAGIV SG AV WC+ GP+F +V+ P+ L+VA+ L E+ +LG IIG
Sbjct: 267 FTILYAGIVASGIAFAVQIWCIDRGGPVFVAVYQPVQTLVVAIMASIALGEEFYLGGIIG 326
Query: 272 AVLIVCGLYAVVWGKSKEMK 291
AVLI+ GLY V++GKS+E K
Sbjct: 327 AVLIIAGLYFVLYGKSEERK 346
Score = 40.0 bits (92), Expect = 0.002
Identities = 20/65 (30%), Positives = 37/65 (56%)
Query: 4 KNNLCNVVQGLKPTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIA 63
+ +L V + L+ + M+ +Q +AG +V+ + A+N G+S + YR I A + P A
Sbjct: 8 RRSLWGVPEKLQLHIAMLTLQFGYAGFHVVSRAALNMGISKLVFPVYRNIIALLLLLPFA 67
Query: 64 FILER 68
+ LE+
Sbjct: 68 YFLEK 72
>At1g44800 nodulin like protein
Length = 370
Score = 84.3 bits (207), Expect = 9e-17
Identities = 38/77 (49%), Positives = 54/77 (69%)
Query: 213 SVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGA 272
+ Y+G+V SG + S + RGP+F + F+P+ +++ A G +L EK+HLGSIIGA
Sbjct: 252 AAVYSGVVCSGIAYYIQSIVIKQRGPVFTTSFSPMCMIITAFLGALVLAEKIHLGSIIGA 311
Query: 273 VLIVCGLYAVVWGKSKE 289
V IV GLY+VVWGKSK+
Sbjct: 312 VFIVLGLYSVVWGKSKD 328
Score = 42.7 bits (99), Expect = 3e-04
Identities = 26/111 (23%), Positives = 53/111 (47%), Gaps = 12/111 (10%)
Query: 11 VQGLKPTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILERLQ 70
++ +KP L ++ +Q +AG+ ++ ++ GM ++ YR + AT +AP A + ER
Sbjct: 6 MEKIKPILAIISLQFGYAGMYIITMVSFKHGMDHWVLATYRHVVATVVMAPFALMFERKI 65
Query: 71 EEEGKDDLDNTVSVISMRFIWGIISSKLLLTSPDFNFSNICIRN-GQPYTS 120
+ +++ W +++ +L D N I ++N YTS
Sbjct: 66 RPK-----------MTLAIFWRLLALGILEPLMDQNLYYIGLKNTSASYTS 105
>At5g07050 MtN21 nodulin protein-like
Length = 381
Score = 84.0 bits (206), Expect = 1e-16
Identities = 41/77 (53%), Positives = 55/77 (71%)
Query: 213 SVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGA 272
+ AY+GIV S V + RGP+FA+ F+PLM+++VA+ G +L EK+ LG +IGA
Sbjct: 246 AAAYSGIVASSISYYVQGIVMKKRGPVFATAFSPLMMVIVAVMGSFVLAEKIFLGGVIGA 305
Query: 273 VLIVCGLYAVVWGKSKE 289
VLIV GLYAV+WGK KE
Sbjct: 306 VLIVIGLYAVLWGKQKE 322
Score = 57.8 bits (138), Expect = 9e-09
Identities = 35/102 (34%), Positives = 53/102 (51%), Gaps = 11/102 (10%)
Query: 20 MVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILERLQEEEGKDDLD 79
M+ +Q +AG+N++ K+++N GMS ++V YR ATA IAP AF ER K
Sbjct: 1 MISLQFGYAGMNIITKISLNTGMSHYVLVVYRHAIATAVIAPFAFFFER------KAQPK 54
Query: 80 NTVSVISMRFIWGIISSKLLLTSPDFNFSNICIRNGQPYTSC 121
T S+ FI G++ + D NF + ++ P SC
Sbjct: 55 ITFSIFMQLFILGLLGPVI-----DQNFYYMGLKYTSPTFSC 91
>At5g64700 nodulin-like protein
Length = 359
Score = 82.8 bits (203), Expect = 3e-16
Identities = 38/83 (45%), Positives = 54/83 (64%)
Query: 209 LAYYSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGS 268
L +V Y G +V+G + SW + RGP+F S+FTPL LL L+ +L E + LGS
Sbjct: 252 LRLVAVIYCGFIVTGVAYYLQSWVIEKRGPVFLSMFTPLSLLFTLLSSAILLCEIISLGS 311
Query: 269 IIGAVLIVCGLYAVVWGKSKEMK 291
I+G +L++ GLY V+WGKS+E K
Sbjct: 312 IVGGLLLIIGLYCVLWGKSREEK 334
Score = 47.0 bits (110), Expect = 2e-05
Identities = 21/58 (36%), Positives = 36/58 (61%)
Query: 11 VQGLKPTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILER 68
++ KP L++ ++Q+ + + ++ K N GM+ + V YR FAT F+AP+AF ER
Sbjct: 3 MESKKPYLMVTIIQVIYTIMFLISKAVFNGGMNTFVFVFYRQAFATIFLAPLAFFFER 60
>At1g25270
Length = 356
Score = 82.8 bits (203), Expect = 3e-16
Identities = 41/79 (51%), Positives = 58/79 (72%), Gaps = 6/79 (7%)
Query: 204 HLLCTLAYYSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEK 263
+LL TL Y+GIVVSG +V +++WC+ +GPLF +VF+P+ L++VAL G L E
Sbjct: 233 NLLATL------YSGIVVSGMVVPLVAWCIATKGPLFVTVFSPIRLVIVALIGSFALEEP 286
Query: 264 LHLGSIIGAVLIVCGLYAV 282
LHLGSIIGA+++V G+Y V
Sbjct: 287 LHLGSIIGAMIMVGGVYLV 305
Score = 60.8 bits (146), Expect = 1e-09
Identities = 30/85 (35%), Positives = 53/85 (62%), Gaps = 6/85 (7%)
Query: 14 LKPTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILERLQEEE 73
+K + MV VQ FAG+ +L+K+ V+DG +L+++VAYR FAT F+ P+A I +R + E
Sbjct: 1 MKSVVAMVAVQFIFAGMFILFKITVDDGTNLKVLVAYRLSFATIFMLPLALIFQRKKRPE 60
Query: 74 GKDDLDNTVSVISMRFIWGIISSKL 98
T ++ + F+ G++ + +
Sbjct: 61 ------FTWRLLLLAFVSGLLGAAI 79
>At4g08300 nodulin-like protein
Length = 373
Score = 81.6 bits (200), Expect = 6e-16
Identities = 36/77 (46%), Positives = 54/77 (69%)
Query: 213 SVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGA 272
+ Y+G+V SG + S + RGP+F + F+P+ +++ A G +L EK+HLGSIIGA
Sbjct: 255 AAVYSGVVCSGMAYYIQSIVIRERGPVFTTSFSPMCMIITAFLGVLVLAEKIHLGSIIGA 314
Query: 273 VLIVCGLYAVVWGKSKE 289
+ IV GLY+VVWGK+K+
Sbjct: 315 IFIVFGLYSVVWGKAKD 331
Score = 45.8 bits (107), Expect = 3e-05
Identities = 21/55 (38%), Positives = 34/55 (61%)
Query: 14 LKPTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILER 68
LKP + ++ +Q +AG+ ++ ++ GM+ I+ YR + AT IAP A ILER
Sbjct: 9 LKPIIAIISLQFGYAGMYIITMVSFKHGMNHWILATYRHVVATIVIAPFALILER 63
>At5g13670 unknown protein
Length = 377
Score = 79.0 bits (193), Expect = 4e-15
Identities = 47/129 (36%), Positives = 71/129 (54%), Gaps = 8/129 (6%)
Query: 218 GIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGAVLIVC 277
G +VSG VI W RGP+F S F PL +++VA+ + EK+++G +IG+V+IV
Sbjct: 257 GGLVSGLAYYVIGWASKERGPVFVSAFNPLSMVLVAILSTFVFLEKVYVGRVIGSVVIVI 316
Query: 278 GLYAVVWGKSKEMKKKNQLVPSKNPHEFDPVEIVVRSIEKDKSNLNNNNQMVKDNEDSQT 337
G+Y V+WGKSK+ K L P+ E V I++ K +NNQ+V +
Sbjct: 317 GIYLVLWGKSKD--KGGMLQPNAGCAE------TVVKIDQQKVPTPDNNQVVSISYHLMI 368
Query: 338 DEHEQQSQE 346
+ +SQE
Sbjct: 369 PKAAARSQE 377
Score = 51.2 bits (121), Expect = 8e-07
Identities = 24/54 (44%), Positives = 37/54 (68%)
Query: 15 KPTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILER 68
+P + +V +Q +A ++++ KLA+N GMS ++VAYR A+A I P A ILER
Sbjct: 7 RPFIAIVFIQCLYALMSIVAKLALNKGMSPHVLVAYRMAVASALITPFALILER 60
>At1g21890 nodulin-like protein
Length = 389
Score = 79.0 bits (193), Expect = 4e-15
Identities = 33/80 (41%), Positives = 52/80 (64%)
Query: 212 YSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIG 271
++ AY+G++ SG V + RGP+F + F PL +++ A G +L+E +HLGS+IG
Sbjct: 260 FAAAYSGVICSGVAYYVQGVVMRERGPVFVATFNPLCVVITAALGVVVLSESIHLGSVIG 319
Query: 272 AVLIVCGLYAVVWGKSKEMK 291
+ I+ GLY VVWGK K+ +
Sbjct: 320 TLFIIVGLYTVVWGKGKDKR 339
Score = 45.8 bits (107), Expect = 3e-05
Identities = 21/59 (35%), Positives = 35/59 (58%)
Query: 10 VVQGLKPTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILER 68
++ LKP L M+ +Q +AG+ ++ +++ GM+ ++ YR ATA IAP A ER
Sbjct: 5 LMNSLKPYLAMISMQFGYAGMYIITMVSLKHGMNHYVLAVYRHAIATAVIAPFALFHER 63
>At1g43650 nodulin-like protein
Length = 341
Score = 78.6 bits (192), Expect = 5e-15
Identities = 34/81 (41%), Positives = 53/81 (64%)
Query: 209 LAYYSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGS 268
L S+AY GI+V+G + W + +GP+F +++TPL L++ + + E +LGS
Sbjct: 239 LPLLSMAYCGIMVTGLTYWLQVWAIEKKGPVFTALYTPLALILTCIVSSFLFKETFYLGS 298
Query: 269 IIGAVLIVCGLYAVVWGKSKE 289
+ GAVL+VCGLY +WGK+KE
Sbjct: 299 VGGAVLLVCGLYLGLWGKTKE 319
Score = 44.7 bits (104), Expect = 8e-05
Identities = 22/53 (41%), Positives = 33/53 (61%)
Query: 15 KPTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILE 67
K + MV VQI +AG+ +L K+A++ G + + V YR FA ++P AF LE
Sbjct: 4 KANMAMVFVQIVYAGMPLLSKVAISQGTNPFVFVFYRQAFAALALSPFAFFLE 56
>At4g08290 nodulin-like protein
Length = 384
Score = 78.2 bits (191), Expect = 6e-15
Identities = 40/117 (34%), Positives = 64/117 (54%)
Query: 212 YSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIG 271
++ Y GIV SG V + RGP+F + F PL +++VAL IL+E++H G +IG
Sbjct: 255 FAPLYTGIVSSGITYYVQGMVMKTRGPVFVTAFNPLCMILVALIASFILHEQIHFGCVIG 314
Query: 272 AVLIVCGLYAVVWGKSKEMKKKNQLVPSKNPHEFDPVEIVVRSIEKDKSNLNNNNQM 328
+I GLY VVWGK K+ + + KN + P+ K S++++N+ +
Sbjct: 315 GAVIAAGLYMVVWGKGKDYEVSGLDILEKNSLQELPITTKSEDDNKLVSSISDNSNV 371
Score = 44.3 bits (103), Expect = 1e-04
Identities = 20/55 (36%), Positives = 31/55 (56%)
Query: 14 LKPTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILER 68
L+P LLM+ +Q AG ++ +N G + +V+ YR + A +AP A I ER
Sbjct: 11 LRPYLLMIFLQFGAAGTYIVIMATLNQGQNRYVVIVYRNLVAALVLAPFALIFER 65
>At3g30340 unknown protein
Length = 364
Score = 77.8 bits (190), Expect = 8e-15
Identities = 36/94 (38%), Positives = 61/94 (64%), Gaps = 3/94 (3%)
Query: 213 SVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGA 272
++ Y+GIV SG +SWC+ RG +F S F PL+ + A+ + L+E+++ GS+IG+
Sbjct: 253 ALLYSGIVGSGLCYVGMSWCLRQRGAVFTSSFIPLIQVFAAIFSFSFLHEQIYCGSVIGS 312
Query: 273 VLIVCGLYAVVWGKSKEMKKKNQLVPSKNPHEFD 306
++I+ GLY ++WGKSK+ K+ V + P + D
Sbjct: 313 MVIIVGLYILLWGKSKD---KSASVTKQEPLDLD 343
Score = 45.1 bits (105), Expect = 6e-05
Identities = 19/54 (35%), Positives = 31/54 (57%)
Query: 15 KPTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILER 68
K L+M M+ I + VNV++K +++G++ + YR T F+ P A LER
Sbjct: 10 KAVLMMSMINIGLSVVNVMFKKMIDEGLNRMVATTYRLAVGTLFLIPFAIFLER 63
>At2g39510 nodulin-like protein
Length = 374
Score = 73.6 bits (179), Expect = 2e-13
Identities = 34/77 (44%), Positives = 51/77 (66%)
Query: 213 SVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGA 272
+ Y G++ SG V + RGP+F + F PL +++VA+ G IL E + LG I+GA
Sbjct: 250 AAVYGGVICSGIGYYVQGVIMKTRGPVFVTAFNPLSMVIVAILGSIILAEVMFLGRILGA 309
Query: 273 VLIVCGLYAVVWGKSKE 289
++IV GLY+V+WGKSK+
Sbjct: 310 IVIVLGLYSVLWGKSKD 326
Score = 55.8 bits (133), Expect = 3e-08
Identities = 24/54 (44%), Positives = 38/54 (69%)
Query: 15 KPTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILER 68
KP + +V +Q +AG++++ K A+N GMS ++ +YR I AT FIAP A+ L+R
Sbjct: 7 KPFITVVSLQFGYAGLSIIAKFALNQGMSPHVLASYRHIVATIFIAPFAYFLDR 60
>At5g40240 unknown protein
Length = 368
Score = 72.8 bits (177), Expect = 3e-13
Identities = 30/81 (37%), Positives = 52/81 (64%), Gaps = 5/81 (6%)
Query: 214 VAYAGIVVSGAMVAVIS-----WCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGS 268
++ A I+ SG V++ S W +H++GP++ S+F PL + + G L + LHLGS
Sbjct: 260 ISLAAIIYSGVFVSLFSALTHTWGLHLKGPVYISLFRPLSIAIAVAMGAIFLGDALHLGS 319
Query: 269 IIGAVLIVCGLYAVVWGKSKE 289
+IG++++ G Y V+WGK++E
Sbjct: 320 VIGSMILCIGFYTVIWGKARE 340
Score = 35.8 bits (81), Expect = 0.035
Identities = 18/53 (33%), Positives = 27/53 (49%)
Query: 16 PTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILER 68
P M V+ A G N L+K A G+S + V Y +I +T + P++ I R
Sbjct: 20 PFAAMFAVECATVGSNTLFKAATLRGLSFYVFVFYSYIVSTLLLLPLSVIFGR 72
>At4g01440 predicted protein of unknown function
Length = 365
Score = 72.8 bits (177), Expect = 3e-13
Identities = 31/81 (38%), Positives = 51/81 (62%)
Query: 213 SVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGA 272
++ YAG V G SWC+ RGP+F S+FTP+ L+ L IL+ ++ LGS++G+
Sbjct: 251 TIVYAGAVAQGICTVGTSWCIRKRGPIFTSIFTPVGLIFATLFDFLILHRQIFLGSVVGS 310
Query: 273 VLIVCGLYAVVWGKSKEMKKK 293
+++ GLY + GK + MK++
Sbjct: 311 GVVIFGLYIFLLGKVRLMKEE 331
Score = 45.4 bits (106), Expect = 4e-05
Identities = 20/53 (37%), Positives = 31/53 (57%)
Query: 16 PTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILER 68
P ++MVM+ A N L K ++ G++ ++ YR +T F+APIAF ER
Sbjct: 10 PVIIMVMINSALGLANALVKKVLDGGVNHMVIATYRLAISTLFLAPIAFFWER 62
>At3g28070 unknown protein
Length = 268
Score = 72.4 bits (176), Expect = 3e-13
Identities = 30/72 (41%), Positives = 46/72 (63%)
Query: 220 VVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGAVLIVCGL 279
+V+ + SW V +GPL+ ++F PL +L+ + G LN+ L+LG +IG +LI G
Sbjct: 175 IVTSVYYVIHSWTVRHKGPLYLAIFKPLSILIAVVMGAIFLNDSLYLGCLIGGILITLGF 234
Query: 280 YAVVWGKSKEMK 291
YAV+WGK+ E K
Sbjct: 235 YAVMWGKANEEK 246
>At5g40230 putative protein
Length = 370
Score = 72.0 bits (175), Expect = 4e-13
Identities = 31/90 (34%), Positives = 56/90 (61%)
Query: 213 SVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGA 272
SV Y+G +VS + +W +H++GP++ S+F PL +++ G L + L+LGS+IG+
Sbjct: 265 SVMYSGGLVSSFGSVIHTWGLHLKGPVYISLFKPLSIVIAVAMGVMFLGDALYLGSVIGS 324
Query: 273 VLIVCGLYAVVWGKSKEMKKKNQLVPSKNP 302
+++ G Y V+WGK++E K ++P
Sbjct: 325 LILSLGFYTVIWGKAREDSIKTVAGTEQSP 354
Score = 37.0 bits (84), Expect = 0.016
Identities = 18/53 (33%), Positives = 27/53 (49%)
Query: 16 PTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILER 68
P MV V+ G N L+K A G+S + V Y ++ AT + P++ I R
Sbjct: 21 PFTAMVAVECVTVGSNTLFKAATLRGLSFYVFVFYTYVVATLVLLPLSLIFGR 73
>At3g28100 unknown protein
Length = 227
Score = 72.0 bits (175), Expect = 4e-13
Identities = 31/81 (38%), Positives = 49/81 (60%)
Query: 220 VVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGAVLIVCGL 279
+++ + SW V +GPL+ ++F PL +L+ + LN+ L+LG +IG +LI G
Sbjct: 138 IITSVYYVIHSWTVRHKGPLYLAIFKPLSILIAVVMSAVFLNDSLYLGCLIGGLLITLGF 197
Query: 280 YAVVWGKSKEMKKKNQLVPSK 300
YAV+WGK+ E K + LV K
Sbjct: 198 YAVMWGKANEEKDQLLLVSGK 218
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.334 0.145 0.470
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,461,553
Number of Sequences: 26719
Number of extensions: 315016
Number of successful extensions: 1651
Number of sequences better than 10.0: 60
Number of HSP's better than 10.0 without gapping: 51
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 1553
Number of HSP's gapped (non-prelim): 105
length of query: 346
length of database: 11,318,596
effective HSP length: 100
effective length of query: 246
effective length of database: 8,646,696
effective search space: 2127087216
effective search space used: 2127087216
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC136472.12


