

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136451.5 + phase: 0
(161 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
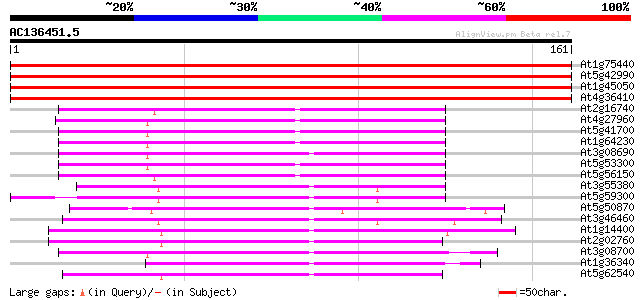
Score E
Sequences producing significant alignments: (bits) Value
At1g75440 ubiquitin-conjugating enzyme 16, putative 315 5e-87
At5g42990 ubiquitin-conjugating enzyme-like protein 309 4e-85
At1g45050 similar to ubiquitin-conjugating enzyme 15 gb|AAC39324... 306 4e-84
At4g36410 ubiquitin-conjugating enzyme 288 7e-79
At2g16740 putative ubiquitin-conjugating enzyme E2 86 9e-18
At4g27960 ubiquitin-protein ligase UBC9 84 3e-17
At5g41700 ubiquitin-conjugating enzyme E2-17 kD 8 (ubiquitin-pro... 84 4e-17
At1g64230 ubiquitin-conjugating enzyme like protein 84 4e-17
At3g08690 putative ubiquitin conjugating enzyme 83 6e-17
At5g53300 ubiquitin-conjugating enzyme E2-17 kD 10 (ubiquitin-pr... 82 1e-16
At5g56150 ubiquitin-conjugating enzyme-like protein 81 3e-16
At3g55380 ubiquitin-conjugating enzyme UBC3 79 1e-15
At5g59300 putative E2, ubiquitin-conjugating enzyme UBC7 79 1e-15
At5g50870 ubiquitin-conjugating enzyme-like protein 77 3e-15
At3g46460 ubiquitin conjugating enzyme E2 (UBC13) 77 3e-15
At1g14400 unknown protein 76 9e-15
At2g02760 putative ubiquitin-conjugating enzyme E2 75 2e-14
At3g08700 putative ubiquitin-conjugating enzyme 73 6e-14
At1g36340 putative ubiquitin conjugating enzyme 72 2e-13
At5g62540 ubiquitin-conjugating enzyme E2-17 kd 3 (ubiquitin-pro... 71 3e-13
>At1g75440 ubiquitin-conjugating enzyme 16, putative
Length = 161
Score = 315 bits (808), Expect = 5e-87
Identities = 146/161 (90%), Positives = 155/161 (95%)
Query: 1 MTSSSAASRKTLSKIACNRLQKELVEWQVNPPTGFKHRVTDNLQRWVIEVGGAPGTLYSN 60
M+SS A SRKTLSKIA NRLQKELVEWQ+NPPTGFKH+VTDNLQRW+IEV GAPGTLY+N
Sbjct: 1 MSSSGAPSRKTLSKIATNRLQKELVEWQMNPPTGFKHKVTDNLQRWIIEVIGAPGTLYAN 60
Query: 61 ETYQLQVDFPENYPMEAPQVIFLHPAPLHPHIYSNGHICLDILYDSWSPAMTVSSICISI 120
+TYQLQVDFPE+YPME+PQVIFLHPAPLHPHIYSNGHICLDILYDSWSPAMTVSSICISI
Sbjct: 61 DTYQLQVDFPEHYPMESPQVIFLHPAPLHPHIYSNGHICLDILYDSWSPAMTVSSICISI 120
Query: 121 LSMLSSSTAKQRPEDNDRYVKNCRNGRSPKETRWWFHDDKV 161
LSMLSSST KQRP DNDRYVKNC+NGRSPKETRWWFHDDKV
Sbjct: 121 LSMLSSSTEKQRPTDNDRYVKNCKNGRSPKETRWWFHDDKV 161
>At5g42990 ubiquitin-conjugating enzyme-like protein
Length = 161
Score = 309 bits (792), Expect = 4e-85
Identities = 143/161 (88%), Positives = 152/161 (93%)
Query: 1 MTSSSAASRKTLSKIACNRLQKELVEWQVNPPTGFKHRVTDNLQRWVIEVGGAPGTLYSN 60
MTSSS+ SRK LSKIACNRLQKEL EWQVNPPTGFKHRVTDNLQ+WVIEV GAPGTLY+N
Sbjct: 1 MTSSSSPSRKALSKIACNRLQKELSEWQVNPPTGFKHRVTDNLQKWVIEVTGAPGTLYAN 60
Query: 61 ETYQLQVDFPENYPMEAPQVIFLHPAPLHPHIYSNGHICLDILYDSWSPAMTVSSICISI 120
ETY LQV+FP++YPMEAPQVIF+ PAPLHPHIYSNGHICLDILYDSWSPAMTVSS+CISI
Sbjct: 61 ETYNLQVEFPQHYPMEAPQVIFVPPAPLHPHIYSNGHICLDILYDSWSPAMTVSSVCISI 120
Query: 121 LSMLSSSTAKQRPEDNDRYVKNCRNGRSPKETRWWFHDDKV 161
LSMLSSS KQRP DNDRYVKNC+NGRSPKETRWWFHDDKV
Sbjct: 121 LSMLSSSPEKQRPTDNDRYVKNCKNGRSPKETRWWFHDDKV 161
>At1g45050 similar to ubiquitin-conjugating enzyme 15 gb|AAC39324.1;
similar to ESTs emb|Z47633.1, gb|L47909.1, gb|T42291.1
Length = 161
Score = 306 bits (783), Expect = 4e-84
Identities = 139/161 (86%), Positives = 152/161 (94%)
Query: 1 MTSSSAASRKTLSKIACNRLQKELVEWQVNPPTGFKHRVTDNLQRWVIEVGGAPGTLYSN 60
MTSSSA SRK LSKIACNRLQKEL EWQ+NPPTGF+H+VTDNLQ+W I+V GAPGTLY+N
Sbjct: 1 MTSSSAPSRKALSKIACNRLQKELSEWQLNPPTGFRHKVTDNLQKWTIDVTGAPGTLYAN 60
Query: 61 ETYQLQVDFPENYPMEAPQVIFLHPAPLHPHIYSNGHICLDILYDSWSPAMTVSSICISI 120
ETYQLQV+FPE+YPMEAPQV+F+ PAP HPHIYSNGHICLDILYDSWSPAMTV+S+CISI
Sbjct: 61 ETYQLQVEFPEHYPMEAPQVVFVSPAPSHPHIYSNGHICLDILYDSWSPAMTVNSVCISI 120
Query: 121 LSMLSSSTAKQRPEDNDRYVKNCRNGRSPKETRWWFHDDKV 161
LSMLSSS AKQRP DNDRYVKNC+NGRSPKETRWWFHDDKV
Sbjct: 121 LSMLSSSPAKQRPADNDRYVKNCKNGRSPKETRWWFHDDKV 161
>At4g36410 ubiquitin-conjugating enzyme
Length = 161
Score = 288 bits (738), Expect = 7e-79
Identities = 130/161 (80%), Positives = 149/161 (91%)
Query: 1 MTSSSAASRKTLSKIACNRLQKELVEWQVNPPTGFKHRVTDNLQRWVIEVGGAPGTLYSN 60
MTSSS ++RK L+KIA NRLQKE +EWQ NPP+GFKHRV+DNLQRW+IEV G PGTLY+N
Sbjct: 1 MTSSSESTRKGLTKIATNRLQKEFMEWQTNPPSGFKHRVSDNLQRWIIEVHGVPGTLYAN 60
Query: 61 ETYQLQVDFPENYPMEAPQVIFLHPAPLHPHIYSNGHICLDILYDSWSPAMTVSSICISI 120
ETYQLQV+FPE+YPMEAPQVIF HPAPLHPHIYSNGHICLD+LYDSWSPAM +SSIC+SI
Sbjct: 61 ETYQLQVEFPEHYPMEAPQVIFQHPAPLHPHIYSNGHICLDVLYDSWSPAMRLSSICLSI 120
Query: 121 LSMLSSSTAKQRPEDNDRYVKNCRNGRSPKETRWWFHDDKV 161
LSMLSSS+ KQ+P+DND Y+KNC++GRSPKETRW FHDDKV
Sbjct: 121 LSMLSSSSVKQKPKDNDHYLKNCKHGRSPKETRWRFHDDKV 161
>At2g16740 putative ubiquitin-conjugating enzyme E2
Length = 148
Score = 85.9 bits (211), Expect = 9e-18
Identities = 44/112 (39%), Positives = 67/112 (59%), Gaps = 2/112 (1%)
Query: 15 IACNRLQKELVEWQVNPPTGFKHRVT-DNLQRWVIEVGGAPGTLYSNETYQLQVDFPENY 73
+A R+ KEL E Q +PP T +++ W + G + YS + + + FP +Y
Sbjct: 1 MATRRILKELKELQRDPPVSCSAGPTGEDMFHWQATIMGPNESPYSGGVFLVNIHFPPDY 60
Query: 74 PMEAPQVIFLHPAPLHPHIYSNGHICLDILYDSWSPAMTVSSICISILSMLS 125
P + P+V+F HP+I SNG+ICLDIL D WSPA+T+S + +SI S+L+
Sbjct: 61 PFKPPKVVF-RTKVFHPNINSNGNICLDILKDQWSPALTISKVLLSICSLLT 111
>At4g27960 ubiquitin-protein ligase UBC9
Length = 178
Score = 84.3 bits (207), Expect = 3e-17
Identities = 43/113 (38%), Positives = 67/113 (59%), Gaps = 2/113 (1%)
Query: 14 KIACNRLQKELVEWQVNPPTGFKHR-VTDNLQRWVIEVGGAPGTLYSNETYQLQVDFPEN 72
++A R+ KEL + Q +PPT V +++ W + G + YS + + + FP +
Sbjct: 30 EMASKRILKELKDLQKDPPTSCSAGPVAEDMFHWQATIMGPSDSPYSGGVFLVTIHFPPD 89
Query: 73 YPMEAPQVIFLHPAPLHPHIYSNGHICLDILYDSWSPAMTVSSICISILSMLS 125
YP + P+V F HP+I SNG ICLDIL + WSPA+T+S + +SI S+L+
Sbjct: 90 YPFKPPKVAF-RTKVFHPNINSNGSICLDILKEQWSPALTISKVLLSICSLLT 141
>At5g41700 ubiquitin-conjugating enzyme E2-17 kD 8
(ubiquitin-protein ligase 8) (ubiquitin carrier protein
8) (sp|P35131)
Length = 148
Score = 83.6 bits (205), Expect = 4e-17
Identities = 43/112 (38%), Positives = 66/112 (58%), Gaps = 2/112 (1%)
Query: 15 IACNRLQKELVEWQVNPPTGFKHR-VTDNLQRWVIEVGGAPGTLYSNETYQLQVDFPENY 73
+A R+ KEL + Q +PPT V +++ W + G + YS + + + FP +Y
Sbjct: 1 MASKRILKELKDLQKDPPTSCSAGPVAEDMFHWQATIMGPAESPYSGGVFLVTIHFPPDY 60
Query: 74 PMEAPQVIFLHPAPLHPHIYSNGHICLDILYDSWSPAMTVSSICISILSMLS 125
P + P+V F HP+I SNG ICLDIL + WSPA+T+S + +SI S+L+
Sbjct: 61 PFKPPKVAF-RTKVFHPNINSNGSICLDILKEQWSPALTISKVLLSICSLLT 111
>At1g64230 ubiquitin-conjugating enzyme like protein
Length = 148
Score = 83.6 bits (205), Expect = 4e-17
Identities = 42/112 (37%), Positives = 66/112 (58%), Gaps = 2/112 (1%)
Query: 15 IACNRLQKELVEWQVNPPTGFKHR-VTDNLQRWVIEVGGAPGTLYSNETYQLQVDFPENY 73
+A R+ KEL + Q +PPT V +++ W + G + YS + + + FP +Y
Sbjct: 1 MASKRILKELKDLQKDPPTSCSAGPVAEDMFHWQATIMGPSDSPYSGGVFLVTIHFPPDY 60
Query: 74 PMEAPQVIFLHPAPLHPHIYSNGHICLDILYDSWSPAMTVSSICISILSMLS 125
P + P+V F HP++ SNG ICLDIL + WSPA+T+S + +SI S+L+
Sbjct: 61 PFKPPKVAF-RTKVFHPNVNSNGSICLDILKEQWSPALTISKVLLSICSLLT 111
>At3g08690 putative ubiquitin conjugating enzyme
Length = 148
Score = 83.2 bits (204), Expect = 6e-17
Identities = 42/112 (37%), Positives = 67/112 (59%), Gaps = 2/112 (1%)
Query: 15 IACNRLQKELVEWQVNPPTGFKHR-VTDNLQRWVIEVGGAPGTLYSNETYQLQVDFPENY 73
+A R+ KEL + Q +PP+ V +++ W + G P + Y+ + + + FP +Y
Sbjct: 1 MASKRILKELKDLQKDPPSNCSAGPVAEDMFHWQATIMGPPESPYAGGVFLVSIHFPPDY 60
Query: 74 PMEAPQVIFLHPAPLHPHIYSNGHICLDILYDSWSPAMTVSSICISILSMLS 125
P + P+V F HP+I SNG ICLDIL + WSPA+T+S + +SI S+L+
Sbjct: 61 PFKPPKVSFKTKV-YHPNINSNGSICLDILKEQWSPALTISKVLLSICSLLT 111
>At5g53300 ubiquitin-conjugating enzyme E2-17 kD 10
(ubiquitin-protein ligase 10) (ubiquitin carrier protein
10) (sp|P35133)
Length = 148
Score = 82.4 bits (202), Expect = 1e-16
Identities = 42/112 (37%), Positives = 66/112 (58%), Gaps = 2/112 (1%)
Query: 15 IACNRLQKELVEWQVNPPTGFKHR-VTDNLQRWVIEVGGAPGTLYSNETYQLQVDFPENY 73
+A R+ KEL + Q +PPT V +++ W + G + Y+ + + + FP +Y
Sbjct: 1 MASKRILKELKDLQKDPPTSCSAGPVAEDMFHWQATIMGPSESPYAGGVFLVTIHFPPDY 60
Query: 74 PMEAPQVIFLHPAPLHPHIYSNGHICLDILYDSWSPAMTVSSICISILSMLS 125
P + P+V F HP+I SNG ICLDIL + WSPA+T+S + +SI S+L+
Sbjct: 61 PFKPPKVAF-RTKVFHPNINSNGSICLDILKEQWSPALTISKVLLSICSLLT 111
>At5g56150 ubiquitin-conjugating enzyme-like protein
Length = 148
Score = 80.9 bits (198), Expect = 3e-16
Identities = 42/112 (37%), Positives = 66/112 (58%), Gaps = 2/112 (1%)
Query: 15 IACNRLQKELVEWQVNPPTGFKHRVT-DNLQRWVIEVGGAPGTLYSNETYQLQVDFPENY 73
+A R+ KEL + Q +PP T D++ +W + G + ++ + + + FP +Y
Sbjct: 1 MASKRINKELRDLQRDPPVSCSAGPTGDDMFQWQATIMGPADSPFAGGVFLVTIHFPPDY 60
Query: 74 PMEAPQVIFLHPAPLHPHIYSNGHICLDILYDSWSPAMTVSSICISILSMLS 125
P + P+V F HP+I SNG ICLDIL + WSPA+TVS + +SI S+L+
Sbjct: 61 PFKPPKVAF-RTKVYHPNINSNGSICLDILKEQWSPALTVSKVLLSICSLLT 111
>At3g55380 ubiquitin-conjugating enzyme UBC3
Length = 167
Score = 79.0 bits (193), Expect = 1e-15
Identities = 44/121 (36%), Positives = 64/121 (52%), Gaps = 16/121 (13%)
Query: 20 LQKELVEWQVNPPTGFKHRVTD--NLQRWVIEVGGAPGTLYSNETYQLQVDFPENYPMEA 77
LQK+L + P GF + D N+ +W + + G P TLY + + FPENYP+
Sbjct: 10 LQKQLKDLCKKPVDGFSAGLVDEKNVFQWSVSIMGPPDTLYEGGFFNAIMSFPENYPVSP 69
Query: 78 PQVIFLHPAPLHPHIYSNGHICLDILY-------------DSWSPAMTVSSICISILSML 124
P V F HP++YS+G +C+ IL+ + W+P TV SI +SI+SML
Sbjct: 70 PTVTFTSEM-WHPNVYSDGKVCISILHPPGDDPHGYELASERWTPVHTVESIVLSIISML 128
Query: 125 S 125
S
Sbjct: 129 S 129
>At5g59300 putative E2, ubiquitin-conjugating enzyme UBC7
Length = 203
Score = 78.6 bits (192), Expect = 1e-15
Identities = 49/140 (35%), Positives = 73/140 (52%), Gaps = 22/140 (15%)
Query: 1 MTSSSAASRKTLSKIACNRLQKELVEWQVNPPTGFKHRVTD--NLQRWVIEVGGAPGTLY 58
++SSS AS+ +L LQK+L + +P GF + D N+ W + + G P TLY
Sbjct: 33 LSSSSMASQASLL------LQKQLKDLCKHPVDGFSAGLVDEKNIFEWSVTIIGPPDTLY 86
Query: 59 SNETYQLQVDFPENYPMEAPQVIFLHPAPLHPHIYSNGHICLDILY-------------D 105
+ + FP+NYP P V F HP++YS+G +C+ IL+ +
Sbjct: 87 EGGFFNAIMTFPQNYPNSPPTVRFTSDM-WHPNVYSDGRVCISILHPPGDDPSGYELASE 145
Query: 106 SWSPAMTVSSICISILSMLS 125
W+P TV SI +SI+SMLS
Sbjct: 146 RWTPVHTVESIMLSIISMLS 165
>At5g50870 ubiquitin-conjugating enzyme-like protein
Length = 192
Score = 77.4 bits (189), Expect = 3e-15
Identities = 51/132 (38%), Positives = 77/132 (57%), Gaps = 10/132 (7%)
Query: 18 NRLQKELVEWQVNPPTGFKHRV---TDNLQRWVIEVGGAPGTLYSNETYQLQVDFPENYP 74
+R+QKEL + + N + RV +DNL R + G GT Y T+Q+ + P+ YP
Sbjct: 5 SRIQKELQDCERNQDSS-GIRVCPKSDNLTRLTGTIPGPIGTPYEGGTFQIDITMPDGYP 63
Query: 75 MEAPQVIFLHPAPLHPHIYS-NGHICLDILYDSWSPAMTVSSICISILSMLSSSTAKQRP 133
E P++ F HP+I S +G ICLDIL D WSPA+T+ + +SI ++LS+ K P
Sbjct: 64 FEPPKMQFSTKV-WHPNISSQSGAICLDILKDQWSPALTLKTALVSIQALLSAPEPKD-P 121
Query: 134 ED---NDRYVKN 142
+D ++Y+KN
Sbjct: 122 QDAVVAEQYMKN 133
>At3g46460 ubiquitin conjugating enzyme E2 (UBC13)
Length = 166
Score = 77.4 bits (189), Expect = 3e-15
Identities = 49/150 (32%), Positives = 71/150 (46%), Gaps = 25/150 (16%)
Query: 16 ACNRLQKELVEWQVNPPTGFKHRVTD--NLQRWVIEVGGAPGTLYSNETYQLQVDFPENY 73
AC LQK+L + +P GF + D N+ W + + G P TLY + + FP+NY
Sbjct: 5 ACLLLQKQLKDLCKHPVDGFSAGLVDEKNIFEWSVTIIGPPDTLYEGGFFYAIMSFPQNY 64
Query: 74 PMEAPQVIFLHPAPLHPHIYSNGHICLDILY-------------DSWSPAMTVSSICISI 120
P P V F HP++Y +G +C+ IL+ + W+P TV SI +SI
Sbjct: 65 PNSPPTVRFTSDI-WHPNVYPDGRVCISILHPPGDDPSGYELASERWTPVHTVESIMLSI 123
Query: 121 LSMLSS---------STAKQRPEDNDRYVK 141
+SMLS AK+ E D + K
Sbjct: 124 ISMLSGPNDESPANVEAAKEWREKRDEFKK 153
>At1g14400 unknown protein
Length = 152
Score = 75.9 bits (185), Expect = 9e-15
Identities = 47/144 (32%), Positives = 71/144 (48%), Gaps = 11/144 (7%)
Query: 12 LSKIACNRLQKELVEWQVNPPTGFKHRVTDN-LQRWVIEVGGAPGTLYSNETYQLQVDFP 70
+S A RL ++ Q +PP G DN + W + G T + T++L + F
Sbjct: 1 MSTPARKRLMRDFKRLQQDPPAGISGAPQDNNIMLWNAVIFGPDDTPWDGGTFKLSLQFS 60
Query: 71 ENYPMEAPQVIFLHPAPLHPHIYSNGHICLDILYDSWSPAMTVSSICISILSML------ 124
E+YP + P V F+ HP+IY++G ICLDIL + WSP V++I SI S+L
Sbjct: 61 EDYPNKPPTVRFVSRM-FHPNIYADGSICLDILQNQWSPIYDVAAILTSIQSLLCDPNPN 119
Query: 125 ---SSSTAKQRPEDNDRYVKNCRN 145
+S A+ E Y + R+
Sbjct: 120 SPANSEAARMYSESKREYNRRVRD 143
>At2g02760 putative ubiquitin-conjugating enzyme E2
Length = 152
Score = 75.1 bits (183), Expect = 2e-14
Identities = 42/114 (36%), Positives = 62/114 (53%), Gaps = 2/114 (1%)
Query: 12 LSKIACNRLQKELVEWQVNPPTGFKHRVTDN-LQRWVIEVGGAPGTLYSNETYQLQVDFP 70
+S A RL ++ Q +PP G DN + W + G T + T++L + F
Sbjct: 1 MSTPARKRLMRDFKRLQQDPPAGISGAPQDNNIMLWNAVIFGPDDTPWDGGTFKLSLQFS 60
Query: 71 ENYPMEAPQVIFLHPAPLHPHIYSNGHICLDILYDSWSPAMTVSSICISILSML 124
E+YP + P V F+ HP+IY++G ICLDIL + WSP V++I SI S+L
Sbjct: 61 EDYPNKPPTVRFVSRM-FHPNIYADGSICLDILQNQWSPIYDVAAILTSIQSLL 113
>At3g08700 putative ubiquitin-conjugating enzyme
Length = 149
Score = 73.2 bits (178), Expect = 6e-14
Identities = 43/128 (33%), Positives = 65/128 (50%), Gaps = 9/128 (7%)
Query: 15 IACNRLQKELVEWQVNPPTGFKHR--VTDNLQRWVIEVGGAPGTLYSNETYQLQVDFPEN 72
+A R+ +EL + Q +PP +++ W + G + YS + + +DF +
Sbjct: 1 MASKRISRELRDMQRHPPANCSAGPVAEEDIFHWQATIMGPHDSPYSGGVFTVSIDFSSD 60
Query: 73 YPMEAPQVIFLHPAPLHPHIYSNGHICLDILYDSWSPAMTVSSICISILSMLSSSTAKQR 132
YP + P+V F HP+I S G ICLDIL + WSPA T S + +SI S+L+
Sbjct: 61 YPFKPPKVNFKTKV-YHPNIDSKGSICLDILKEQWSPAPTTSKVLLSICSLLTD------ 113
Query: 133 PEDNDRYV 140
P ND V
Sbjct: 114 PNPNDPLV 121
>At1g36340 putative ubiquitin conjugating enzyme
Length = 154
Score = 71.6 bits (174), Expect = 2e-13
Identities = 34/96 (35%), Positives = 52/96 (53%), Gaps = 5/96 (5%)
Query: 40 TDNLQRWVIEVGGAPGTLYSNETYQLQVDFPENYPMEAPQVIFLHPAPLHPHIYSNGHIC 99
++N+ W + G GT Y + L + FP +YP + P+ F P HP+I G IC
Sbjct: 33 SNNIYEWTAVIRGPDGTPYEGGMFNLSIKFPTDYPFKPPKFTFKTPI-YHPNINDEGSIC 91
Query: 100 LDILYDSWSPAMTVSSICISILSMLSSSTAKQRPED 135
++IL D W+PA+ V + +SIL +L K P+D
Sbjct: 92 MNILKDKWTPALMVEKVLLSILLLLE----KPNPDD 123
>At5g62540 ubiquitin-conjugating enzyme E2-17 kd 3
(ubiquitin-protein ligase 3) (ubiquitin carrier protein
3)-like protein
Length = 150
Score = 70.9 bits (172), Expect = 3e-13
Identities = 40/110 (36%), Positives = 59/110 (53%), Gaps = 2/110 (1%)
Query: 16 ACNRLQKELVEWQVNPPTGFKHRVTDN-LQRWVIEVGGAPGTLYSNETYQLQVDFPENYP 74
A RL + Q +PP G DN + W + G T + T++L + F E+YP
Sbjct: 5 AKKRLMWDFKRLQKDPPVGISGAPQDNNIMHWNALIFGPEDTPWDGGTFKLTLHFTEDYP 64
Query: 75 MEAPQVIFLHPAPLHPHIYSNGHICLDILYDSWSPAMTVSSICISILSML 124
+ P V F+ HP+IY++G ICLDIL + WSP V+++ SI S+L
Sbjct: 65 NKPPIVRFVSRM-FHPNIYADGSICLDILQNQWSPIYDVAAVLTSIQSLL 113
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.132 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,133,065
Number of Sequences: 26719
Number of extensions: 170991
Number of successful extensions: 468
Number of sequences better than 10.0: 61
Number of HSP's better than 10.0 without gapping: 49
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 374
Number of HSP's gapped (non-prelim): 63
length of query: 161
length of database: 11,318,596
effective HSP length: 91
effective length of query: 70
effective length of database: 8,887,167
effective search space: 622101690
effective search space used: 622101690
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC136451.5


