

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136451.2 - phase: 0 /pseudo
(355 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
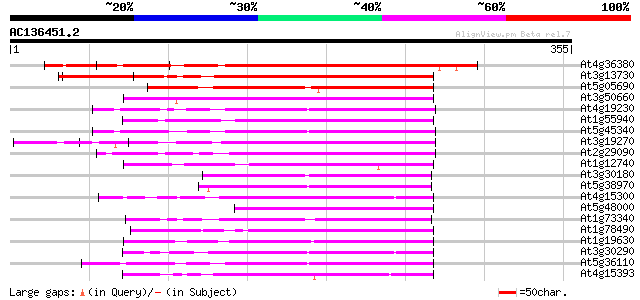
Score E
Sequences producing significant alignments: (bits) Value
At4g36380 cytochrome P450 (ROTUNDIFOLIA3) 248 3e-66
At3g13730 CYP90D 201 4e-52
At5g05690 cytochrome P450 90A1 (sp|Q42569) 141 6e-34
At3g50660 steroid 22-alpha-hydroxylase (DWF4) 130 1e-30
At4g19230 cytochrome P450 protein like 124 8e-29
At1g55940 cytochrome P450, putative 117 1e-26
At5g45340 cytochrome P450 115 4e-26
At3g19270 cytochrome P450, putative 109 2e-24
At2g29090 putative cytochrome P450 107 8e-24
At1g12740 hypothetical protein 107 1e-23
At3g30180 cytochrome P450 homolog, putative 107 1e-23
At5g38970 cytochrome P450 - like protein 106 2e-23
At4g15300 cytochrome P450 like protein 106 2e-23
At5g48000 cytochrome P450-like protein 103 1e-22
At1g73340 102 2e-22
At1g78490 unknown protein 102 3e-22
At1g19630 cytochrome P450, putative 98 8e-21
At3g30290 cytochrome P450, putative 94 9e-20
At5g36110 cytochrome P450-like 92 4e-19
At4g15393 putative protein 87 1e-17
>At4g36380 cytochrome P450 (ROTUNDIFOLIA3)
Length = 524
Score = 248 bits (634), Expect = 3e-66
Identities = 134/247 (54%), Positives = 170/247 (68%), Gaps = 21/247 (8%)
Query: 56 GYSSCPVTFMEKRKSILYKSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVNDVV 115
G P+ F R LYKSLKAKER++KMV ++VEER + + NDVV
Sbjct: 244 GLICIPIKFPGTR---LYKSLKAKERLIKMVKKVVEERQVAMTTTSP--------ANDVV 292
Query: 116 DALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKL 175
D LLRD G+ + S S + +S I+E MIPGEET+PTAMTLA+KFL+D P+AL+KL
Sbjct: 293 DVLLRDGGDSEKQSQPS----DFVSGKIVEMMIPGEETMPTAMTLAVKFLSDNPVALAKL 348
Query: 176 MEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYL 235
+EENME+K++K + Y WTDYMSL FTQNVI+ETLRMANI+NG+WRKA+KDVEIKGYL
Sbjct: 349 VEENMEMKRRKLELGEEYKWTDYMSLSFTQNVINETLRMANIINGVWRKALKDVEIKGYL 408
Query: 236 IPKDWGVMASLTSVHMDSKNYENPYKFDPWRWEII--YLHEAIIYTMF----YACRKLEL 289
IPK W V+AS SVHMD Y+NPY+FDPWRW+ I + +I +T F C LEL
Sbjct: 409 IPKGWCVLASFISVHMDEDIYDNPYQFDPWRWDRINGSANSSICFTPFGGGQRLCPGLEL 468
Query: 290 CLVTIAL 296
+ I++
Sbjct: 469 SKLEISI 475
Score = 65.1 bits (157), Expect = 6e-11
Identities = 34/79 (43%), Positives = 52/79 (65%), Gaps = 1/79 (1%)
Query: 23 KEKNNIKLRNGIKIPKGNSGWPLLGETLDFIASGYSSCPVTFMEKRKSILYKSLKAKERM 82
+E++N + + G+ IP G+ GWP++GETL+FIA GYSS PVTFM+KRKS+ K K
Sbjct: 57 EEEDNEEKKKGM-IPNGSLGWPVIGETLNFIACGYSSRPVTFMDKRKSLYGKVFKTNIIG 115
Query: 83 MKMVGRIVEERMKLIMDNN 101
++ E K+++ N+
Sbjct: 116 TPIIISTDAEVNKVVLQNH 134
>At3g13730 CYP90D
Length = 491
Score = 201 bits (512), Expect = 4e-52
Identities = 112/235 (47%), Positives = 153/235 (64%), Gaps = 16/235 (6%)
Query: 34 IKIPKGNSGWPLLGETLDFIASGYSSCPVTFMEKRKSILYKSLKAKERMMKMVGRIVEER 93
I + KG L E +FI SG S P+ F + L++SL+AK+ M+K V RI+E +
Sbjct: 205 ISVEKGEDLEELKREFENFI-SGLMSLPINFPGTQ---LHRSLQAKKNMVKQVERIIEGK 260
Query: 94 MKLIMDNNAEDDEKVGVVNDVVDALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEET 153
++ N EDD + DVVD LL+D SS +L +I+ N+I+ MIPG ++
Sbjct: 261 IRKTK-NKEEDDV---IAKDVVDVLLKD--------SSEHLTHNLIANNMIDMMIPGHDS 308
Query: 154 LPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLR 213
+P +TLA+KFL+D P AL+ L EENM+LK K + W DY+SLPFTQ VI+ETLR
Sbjct: 309 VPVLITLAVKFLSDSPAALNLLTEENMKLKSLKELTGEPLYWNDYLSLPFTQKVITETLR 368
Query: 214 MANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRWE 268
M N++ G+ RKA+KDVEIKGY+IPK W +A L SVH+D YE+PYKF+PWRW+
Sbjct: 369 MGNVIIGVMRKAMKDVEIKGYVIPKGWCFLAYLRSVHLDKLYYESPYKFNPWRWQ 423
Score = 51.2 bits (121), Expect = 8e-07
Identities = 20/47 (42%), Positives = 34/47 (71%)
Query: 32 NGIKIPKGNSGWPLLGETLDFIASGYSSCPVTFMEKRKSILYKSLKA 78
+G K P G+ GWP++GET++F++S YS P +FM+KR+ + + K+
Sbjct: 47 HGPKFPHGSLGWPVIGETIEFVSSAYSDRPESFMDKRRLMYGRVFKS 93
>At5g05690 cytochrome P450 90A1 (sp|Q42569)
Length = 472
Score = 141 bits (355), Expect = 6e-34
Identities = 72/183 (39%), Positives = 115/183 (62%), Gaps = 14/183 (7%)
Query: 88 RIVEERMKLIMDNNAEDDEKVGVVNDVVDALLRDKGELSQSSSSSNLMVEMISQNIIEFM 147
++ E ++M E++E D++ ALL ++ E I ++ +
Sbjct: 226 KVAEALTVVVMKRREEEEEGAERKKDMLAALL---------AADDGFSDEEIVDFLVALL 276
Query: 148 IPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGYT--WTDYMSLPFTQ 205
+ G ET T MTLA+KFLT+ PLAL++L EE+ +++ K+ D Y+ W+DY S+PFTQ
Sbjct: 277 VAGYETTSTIMTLAVKFLTETPLALAQLKEEHEKIRAMKS---DSYSLEWSDYKSMPFTQ 333
Query: 206 NVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPW 265
V++ETLR+ANI+ G++R+A+ DVEIKGY IPK W V +S +VH+D ++++ F+PW
Sbjct: 334 CVVNETLRVANIIGGVFRRAMTDVEIKGYKIPKGWKVFSSFRAVHLDPNHFKDARTFNPW 393
Query: 266 RWE 268
RW+
Sbjct: 394 RWQ 396
Score = 28.9 bits (63), Expect = 4.5
Identities = 12/33 (36%), Positives = 20/33 (60%)
Query: 36 IPKGNSGWPLLGETLDFIASGYSSCPVTFMEKR 68
+P G+ G PL+GET I + + P F+++R
Sbjct: 31 LPPGSLGLPLIGETFQLIGAYKTENPEPFIDER 63
>At3g50660 steroid 22-alpha-hydroxylase (DWF4)
Length = 513
Score = 130 bits (327), Expect = 1e-30
Identities = 73/203 (35%), Positives = 112/203 (54%), Gaps = 7/203 (3%)
Query: 73 YKSLKAKERMMKMVGRIVEERMKLIMDNNAED------DEKVGVVNDVVDALLRDKGELS 126
+K+L+++ ++K + R +EER I + + E+ DE +D V D L
Sbjct: 229 HKALQSRATILKFIERKMEERKLDIKEEDQEEEEVKTEDEAEMSKSDHVRKQRTDDDLLG 288
Query: 127 QSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQK 186
SNL E I I+ + G ET A+ LA+ FL P A+ +L EE++E+ + K
Sbjct: 289 WVLKHSNLSTEQILDLILSLLFAGHETSSVAIALAIFFLQACPKAVEELREEHLEIARAK 348
Query: 187 TNCPDG-YTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMAS 245
+ W DY + FTQ VI+ETLR+ N+V + RKA+KDV KGY IP W V+
Sbjct: 349 KELGESELNWDDYKKMDFTQCVINETLRLGNVVRFLHRKALKDVRYKGYDIPSGWKVLPV 408
Query: 246 LTSVHMDSKNYENPYKFDPWRWE 268
+++VH+D+ Y+ P F+PWRW+
Sbjct: 409 ISAVHLDNSRYDQPNLFNPWRWQ 431
Score = 35.4 bits (80), Expect = 0.048
Identities = 18/58 (31%), Positives = 32/58 (55%), Gaps = 2/58 (3%)
Query: 13 VLLMSLWFVKKEKNNIKLRNGIKIPKGNSGWPLLGETLDFIASGYSSCPVTFMEKRKS 70
+L + L+ + ++ N K R +P G SGWP LGET+ ++ ++ FM++ S
Sbjct: 18 LLSLLLFLILLKRRNRKTR--FNLPPGKSGWPFLGETIGYLKPYTATTLGDFMQQHVS 73
>At4g19230 cytochrome P450 protein like
Length = 467
Score = 124 bits (311), Expect = 8e-29
Identities = 73/217 (33%), Positives = 120/217 (54%), Gaps = 24/217 (11%)
Query: 53 IASGYSSCPVTFMEKRKSILYKSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVN 112
+ GY+S PV ++ +KS+KA++ + +++ RI+ ER + N + N
Sbjct: 202 LEKGYNSMPVNLPG---TLFHKSMKARKELSQILARILSERRQ----NGSSH-------N 247
Query: 113 DVVDALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLAL 172
D++ + + DK EL+ E I+ NII + +T + M+ LK+L + P L
Sbjct: 248 DLLGSFMGDKEELTD---------EQIADNIIGVIFAARDTTASVMSWILKYLAENPNVL 298
Query: 173 SKLMEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIK 232
+ EE M ++K K + TW D +P T VI ETLR+A+I++ +R+AV+DVE +
Sbjct: 299 EAVTEEQMAIRKDKEE-GESLTWGDTKKMPLTSRVIQETLRVASILSFTFREAVEDVEYE 357
Query: 233 GYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRWEI 269
GYLIPK W V+ ++H + + NP KFDP R+E+
Sbjct: 358 GYLIPKGWKVLPLFRNIHHSADIFSNPGKFDPSRFEV 394
Score = 28.5 bits (62), Expect = 5.8
Identities = 12/36 (33%), Positives = 18/36 (49%), Gaps = 4/36 (11%)
Query: 34 IKIPKGNSGWPLLGETLDFIASGYSSCPVTFMEKRK 69
+ +P G GWP +GET YS P F + ++
Sbjct: 34 LPLPPGTMGWPYVGETFQL----YSQDPNVFFQSKQ 65
>At1g55940 cytochrome P450, putative
Length = 639
Score = 117 bits (292), Expect = 1e-26
Identities = 67/198 (33%), Positives = 104/198 (51%), Gaps = 13/198 (6%)
Query: 72 LYKSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVNDVVDALLRDKGELSQSSSS 131
LY L A+ +MKM+ R+ +ER + DD K G + + + +G+ S
Sbjct: 371 LYNGLMARSNVMKMLKRMFKERR----EEATSDDSKYGDFMETMIYEVEKEGDTINEERS 426
Query: 132 SNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPD 191
L I+ +I ET T L +KF+ + P L +L E+ + + + +
Sbjct: 427 VEL--------ILSLLIASYETTSTMTALTVKFIAENPKVLMELKREHETILQNRADKES 478
Query: 192 GYTWTDYMSLP-FTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVH 250
G TW +Y S+ FT VI+E+LR+ ++ ++RKAV DVEIKGY IP W V+ + +H
Sbjct: 479 GVTWKEYRSMMNFTHMVINESLRLGSLSPAMFRKAVNDVEIKGYTIPAGWIVLVVPSLLH 538
Query: 251 MDSKNYENPYKFDPWRWE 268
D + YE P +F+PWRWE
Sbjct: 539 YDPQIYEQPCEFNPWRWE 556
Score = 33.1 bits (74), Expect = 0.24
Identities = 14/44 (31%), Positives = 28/44 (62%), Gaps = 4/44 (9%)
Query: 9 VSIIVLLMSLWFVKKEKNNIKLRNGIKIPKGNSGWPLLGETLDF 52
++++V+ +SLW + N K+P G+ G+P++GET++F
Sbjct: 157 IALVVVKISLWLYRWANPNCSG----KLPPGSMGFPVIGETVEF 196
>At5g45340 cytochrome P450
Length = 463
Score = 115 bits (288), Expect = 4e-26
Identities = 66/217 (30%), Positives = 115/217 (52%), Gaps = 24/217 (11%)
Query: 53 IASGYSSCPVTFMEKRKSILYKSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVN 112
+ GY+S P+ ++ +K++KA++ + +++ I+ +R + +
Sbjct: 202 LEKGYNSMPINLPG---TLFHKAMKARKELAQILANILSKRRQNPSSHT----------- 247
Query: 113 DVVDALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLAL 172
D++ + + DK L+ E I+ NII + +T + +T LK+L D P L
Sbjct: 248 DLLGSFMEDKAGLTD---------EQIADNIIGVIFAARDTTASVLTWILKYLADNPTVL 298
Query: 173 SKLMEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIK 232
+ EE M ++K K + TW D +P T VI ETLR A I++ +R+AV+DVE +
Sbjct: 299 EAVTEEQMAIRKDKKE-GESLTWEDTKKMPLTYRVIQETLRAATILSFTFREAVEDVEYE 357
Query: 233 GYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRWEI 269
GYLIPK W V+ ++H ++ + +P KFDP R+E+
Sbjct: 358 GYLIPKGWKVLPLFRNIHHNADIFSDPGKFDPSRFEV 394
>At3g19270 cytochrome P450, putative
Length = 468
Score = 109 bits (273), Expect = 2e-24
Identities = 71/225 (31%), Positives = 113/225 (49%), Gaps = 19/225 (8%)
Query: 45 LLGETLDFIASGYSSCPVTFMEKRKSILYKSLKAKERMMKMVGRIVEERMKLIMDNNAED 104
+L + + GY+S P M + +K+L A++++ +V I+ ER
Sbjct: 193 ILKHNYNIVDKGYNSFP---MSLPGTSYHKALMARKQLKTIVSEIICERR---------- 239
Query: 105 DEKVGVVNDVVDALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKF 164
EK + D + LL K E + L E I+ NII + ++T + +T LK+
Sbjct: 240 -EKRALQTDFLGHLLNFKNEKGRV-----LTQEQIADNIIGVLFAAQDTTASCLTWILKY 293
Query: 165 LTDYPLALSKLMEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRK 224
L D L + E + ++ + TW ++P T VI E+LRMA+I++ +R+
Sbjct: 294 LHDDQKLLEAVKAEQKAIYEENSREKKPLTWRQTRNMPLTHKVIVESLRMASIISFTFRE 353
Query: 225 AVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRWEI 269
AV DVE KGYLIPK W VM ++H + K + NP FDP R+E+
Sbjct: 354 AVVDVEYKGYLIPKGWKVMPLFRNIHHNPKYFSNPEVFDPSRFEV 398
Score = 43.5 bits (101), Expect = 2e-04
Identities = 28/76 (36%), Positives = 40/76 (51%), Gaps = 11/76 (14%)
Query: 3 WFIGVYVSIIVLLMSLWFVKKEKNNIKLRNGIKIPKGNSGWPLLGETLDFIASGYSSCPV 62
WF+ V + I+ LL+ V K+K N + K+P G+ GWP LGETL YS P
Sbjct: 5 WFLVVPILILCLLLVRVIVSKKKKNSRG----KLPPGSMGWPYLGETLQL----YSQNPN 56
Query: 63 TFM---EKRKSILYKS 75
F +KR ++K+
Sbjct: 57 VFFTSKQKRYGEIFKT 72
>At2g29090 putative cytochrome P450
Length = 482
Score = 107 bits (268), Expect = 8e-24
Identities = 65/214 (30%), Positives = 115/214 (53%), Gaps = 19/214 (8%)
Query: 56 GYSSCPVTFMEKRKSILYKSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVNDVV 115
GY+S P ++ ++ +KS+KA+ + + + +++E+R +N E+ +GV+
Sbjct: 220 GYNSMP---LDLPGTLFHKSMKARIELSEELRKVIEKRR----ENGREEGGLLGVLLGAK 272
Query: 116 DALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKL 175
D + + LS S I+ NII + +T + +T LK+L D+P L ++
Sbjct: 273 D---QKRNGLSDSQ---------IADNIIGVIFAATDTTASVLTWLLKYLHDHPNLLQEV 320
Query: 176 MEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYL 235
E ++++ +W D +P T VI ETLR A++++ +R+AV+DVE GYL
Sbjct: 321 SREQFSIRQKIKKENRRISWEDTRKMPLTTRVIQETLRAASVLSFTFREAVQDVEYDGYL 380
Query: 236 IPKDWGVMASLTSVHMDSKNYENPYKFDPWRWEI 269
IPK W V+ +H S+ + +P KFDP R+E+
Sbjct: 381 IPKGWKVLPLFRRIHHSSEFFPDPEKFDPSRFEV 414
Score = 32.3 bits (72), Expect = 0.40
Identities = 19/64 (29%), Positives = 32/64 (49%), Gaps = 11/64 (17%)
Query: 7 VYVSIIVLLMSLWFVKKEKNNIKLRNGIKIPKGNSGWPLLGETLDFIASGYSSCPVTFME 66
+ V ++VLL W KE+ +++P G+ G P +GETL Y+ P +F
Sbjct: 27 IVVVVVVLLFKWWLHWKEQR-------LRLPPGSMGLPYIGETLRL----YTENPNSFFA 75
Query: 67 KRKS 70
R++
Sbjct: 76 TRQN 79
>At1g12740 hypothetical protein
Length = 478
Score = 107 bits (267), Expect = 1e-23
Identities = 63/202 (31%), Positives = 109/202 (53%), Gaps = 21/202 (10%)
Query: 73 YKSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVNDVVDALLRDKGELSQSSSSS 132
+K L+ + + MKM+ +++ER + N ++ D V ++ +G + +
Sbjct: 219 HKCLQGRAKAMKMLRNMLQERRENPRKNPSD-------FFDYVIEEIQKEGTILTEEIAL 271
Query: 133 NLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDG 192
+LM ++ + ET A+TLA+KFL+D P L +L EE+ + + + + G
Sbjct: 272 DLMFVLLFASF--------ETTSLALTLAIKFLSDDPEVLKRLTEEHETILRNREDADSG 323
Query: 193 YTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIK------GYLIPKDWGVMASL 246
TW +Y S+ +T I+ET R+ANIV I+RKA++D++ K Y IP W VM
Sbjct: 324 LTWEEYKSMTYTFQFINETARLANIVPAIFRKALRDIKFKEFVNDTDYTIPAGWAVMVCP 383
Query: 247 TSVHMDSKNYENPYKFDPWRWE 268
+VH++ + Y++P F+P RWE
Sbjct: 384 PAVHLNPEMYKDPLVFNPSRWE 405
Score = 36.6 bits (83), Expect = 0.021
Identities = 20/66 (30%), Positives = 38/66 (57%), Gaps = 4/66 (6%)
Query: 3 WFIGVYVSIIVLLMSLWFVKKEKNNIKLRNGIKIPKGNSGWPLLGETLDFIASGYSSCPV 62
W + ++VS++++ ++ W N K R K+P G+ G+PLLGE++ F +S
Sbjct: 2 WALLIWVSLLLISITHWVYSWR--NPKCRG--KLPPGSMGFPLLGESIQFFKPNKTSDIP 57
Query: 63 TFMEKR 68
F+++R
Sbjct: 58 PFIKER 63
>At3g30180 cytochrome P450 homolog, putative
Length = 465
Score = 107 bits (266), Expect = 1e-23
Identities = 56/147 (38%), Positives = 86/147 (58%), Gaps = 4/147 (2%)
Query: 123 GELSQSSSSSNLMVEM-ISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENME 181
G L + + L+ + I ++ + G ET+ T +ALK+L D+P AL +L E++
Sbjct: 250 GYLMKKEDNRYLLTDKEIRDQVVTILYSGYETVSTTSMMALKYLHDHPKALEELRREHLA 309
Query: 182 LKKQKTNCPDG-YTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDW 240
++++K PD T D S+ FT+ VI ET R+A IVNG+ RK D+E+ GYLIPK W
Sbjct: 310 IRERKR--PDEPLTLDDIKSMKFTRAVIFETSRLATIVNGVLRKTTHDLELNGYLIPKGW 367
Query: 241 GVMASLTSVHMDSKNYENPYKFDPWRW 267
+ ++ D+ YE+P F+PWRW
Sbjct: 368 RIYVYTREINYDTSLYEDPMIFNPWRW 394
Score = 31.6 bits (70), Expect = 0.69
Identities = 17/56 (30%), Positives = 28/56 (49%), Gaps = 5/56 (8%)
Query: 1 MEWFIGVYVSIIVLLMSLWFVKKEKNNIKLRNGIKIPKGNSGWPLLGETLDFIASG 56
M +G+ V I+ L +L + + + K +P G GWP+ GET +F+ G
Sbjct: 4 MMMILGLLVIIVCLCTALLRWNQMRYSKK-----GLPPGTMGWPIFGETTEFLKQG 54
>At5g38970 cytochrome P450 - like protein
Length = 465
Score = 106 bits (265), Expect = 2e-23
Identities = 57/157 (36%), Positives = 86/157 (54%), Gaps = 10/157 (6%)
Query: 120 RDKGE---------LSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPL 170
RD GE + + + L E I ++ + G ET+ T +ALK+L D+P
Sbjct: 239 RDSGETFTDMLGYLMKKEGNRYPLTDEEIRDQVVTILYSGYETVSTTSMMALKYLHDHPK 298
Query: 171 ALSKLMEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVE 230
AL +L E++ +++K + D S+ FT+ VI ET R+A IVNG+ RK +D+E
Sbjct: 299 ALQELRAEHLAFRERKRQ-DEPLGLEDVKSMKFTRAVIYETSRLATIVNGVLRKTTRDLE 357
Query: 231 IKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRW 267
I GYLIPK W + ++ D+ YE+P F+PWRW
Sbjct: 358 INGYLIPKGWRIYVYTREINYDANLYEDPLIFNPWRW 394
Score = 34.3 bits (77), Expect = 0.11
Identities = 16/51 (31%), Positives = 30/51 (58%), Gaps = 3/51 (5%)
Query: 7 VYVSIIVLLMSLWFVKKEKNNIKL-RNGIKIPKGNSGWPLLGETLDFIASG 56
V + ++++++SL N ++ +NG+ P G GWP+ GET +F+ G
Sbjct: 6 VMMGLLLIIVSLCSALLRWNQMRYTKNGL--PPGTMGWPIFGETTEFLKQG 54
>At4g15300 cytochrome P450 like protein
Length = 487
Score = 106 bits (264), Expect = 2e-23
Identities = 63/212 (29%), Positives = 107/212 (49%), Gaps = 19/212 (8%)
Query: 57 YSSCPVTFMEKRKSILYKSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVNDVVD 116
Y V F++ ++ + S +A++RMMK++ K ++ A +E +G +++
Sbjct: 219 YKMMKVLFVQYTEADI--SWQARKRMMKLL-------RKTVLTKRASGEE-LGEFFNIIF 268
Query: 117 ALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLM 176
+ +GE + VE + I F + ET P + +KF++D+P +L
Sbjct: 269 GEMEGEGE--------TMSVENAVEYIYTFFLVANETTPRILAATVKFISDHPKVKQELQ 320
Query: 177 EENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLI 236
E+ E+ + K G TW DY S+ FTQ VI+E+LR+ + + R D ++ Y I
Sbjct: 321 REHEEIVRGKAEKEGGLTWEDYKSMHFTQMVINESLRIISTAPTVLRVLEHDFQVGDYTI 380
Query: 237 PKDWGVMASLTSVHMDSKNYENPYKFDPWRWE 268
P W M +H +S+ YE+PY F+PWRWE
Sbjct: 381 PAGWTFM-GYPHIHFNSEKYEDPYAFNPWRWE 411
>At5g48000 cytochrome P450-like protein
Length = 477
Score = 103 bits (257), Expect = 1e-22
Identities = 52/127 (40%), Positives = 77/127 (59%), Gaps = 1/127 (0%)
Query: 143 IIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGYTWTDYM-SL 201
I ++ +E+ + +LA+KFL + AL++L E+ + + + G +W +Y +
Sbjct: 279 IFAILVVAKESTSSVTSLAIKFLAENHKALAELKREHAAILQNRNGKGAGVSWEEYRHQM 338
Query: 202 PFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYK 261
FT VI+ETLRMAN+ ++RKAV DVEIKGY IP W V +VH + YENP +
Sbjct: 339 TFTNMVINETLRMANMAPIMYRKAVNDVEIKGYTIPAGWIVAVIPPAVHFNDAIYENPLE 398
Query: 262 FDPWRWE 268
F+PWRWE
Sbjct: 399 FNPWRWE 405
Score = 30.4 bits (67), Expect = 1.5
Identities = 17/51 (33%), Positives = 28/51 (54%), Gaps = 5/51 (9%)
Query: 3 WFIGVYV-SIIVLLMSLWFVKKEKNNIKLRNGIKIPKGNSGWPLLGETLDF 52
W V+V ++ +++S W + NG K+P G+ G P++GET DF
Sbjct: 5 WSAAVWVIAVAAVVISKWLYRWSNPKC---NG-KLPPGSMGLPIIGETCDF 51
>At1g73340
Length = 512
Score = 102 bits (255), Expect = 2e-22
Identities = 63/194 (32%), Positives = 103/194 (52%), Gaps = 23/194 (11%)
Query: 74 KSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVNDVVDALLRDKGELSQSSSSSN 133
K++KA++ +++ + + +E+R++ N A D N V+ LL ++ +
Sbjct: 258 KAMKARKEIIRKINKTIEKRLQ----NKAASDT---AGNGVLGRLLEEE----------S 300
Query: 134 LMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGY 193
L E ++ II + G ET M A+ FLT P A+++L+EE+ L
Sbjct: 301 LPNESMADFIINLLFAGNETTSKTMLFAVYFLTHCPKAMTQLLEEHDRLAGGML------ 354
Query: 194 TWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDS 253
TW DY ++ FTQ VI ETLR+ I + R+A +DV + Y+IPK V+ L++VH+D
Sbjct: 355 TWQDYKTMDFTQCVIDETLRLGGIAIWLMREAKEDVSYQDYVIPKGCFVVPFLSAVHLDE 414
Query: 254 KNYENPYKFDPWRW 267
Y+ F+PWRW
Sbjct: 415 SYYKESLSFNPWRW 428
Score = 35.4 bits (80), Expect = 0.048
Identities = 15/40 (37%), Positives = 26/40 (64%)
Query: 29 KLRNGIKIPKGNSGWPLLGETLDFIASGYSSCPVTFMEKR 68
K R ++P G+ GWPL+G+T ++ + S P +F+EK+
Sbjct: 36 KRRAPHRLPPGSRGWPLIGDTFAWLNAVAGSHPSSFVEKQ 75
>At1g78490 unknown protein
Length = 479
Score = 102 bits (254), Expect = 3e-22
Identities = 60/193 (31%), Positives = 104/193 (53%), Gaps = 10/193 (5%)
Query: 77 KAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVNDVVDALLRDKGELSQSSSSSNLMV 136
KA + +K ++ ++M +++ +N +++ L +D G S+ NL+
Sbjct: 222 KAVTKALKSREEAIQVMKDVLMMRKETREKQEDFLNTLLEELEKD-GSFFDQGSAINLIF 280
Query: 137 EMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGYTWT 196
++ F + E + LA+KF++ P L++L E+ + + + G +W
Sbjct: 281 ------LLAFAL--REGTSSCTALAVKFISKDPKVLAELKREHKAIVDNRKDKEAGVSWE 332
Query: 197 DYM-SLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKN 255
+Y ++ FT V +E LR+AN ++RKAV+DVEIKGY IP W V + ++VH D
Sbjct: 333 EYRHNMTFTNMVSNEVLRLANTTPLLFRKAVQDVEIKGYTIPAGWIVAVAPSAVHFDPAI 392
Query: 256 YENPYKFDPWRWE 268
YENP++F+PWRWE
Sbjct: 393 YENPFEFNPWRWE 405
Score = 35.8 bits (81), Expect = 0.037
Identities = 23/61 (37%), Positives = 37/61 (59%), Gaps = 6/61 (9%)
Query: 9 VSIIVLLMSLWFVKKEKNNIKLRNGIKIPKGNSGWPLLGETLDFI-ASGYSSCPVTFMEK 67
V+++V+ +S W + +N K K+P G+ G+P++GETLDF G P TF++K
Sbjct: 12 VALVVVRISHWLYRW--SNPKCPG--KLPPGSMGFPIIGETLDFFKPCGVEGIP-TFVKK 66
Query: 68 R 68
R
Sbjct: 67 R 67
>At1g19630 cytochrome P450, putative
Length = 411
Score = 97.8 bits (242), Expect = 8e-21
Identities = 57/196 (29%), Positives = 102/196 (51%), Gaps = 14/196 (7%)
Query: 73 YKSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVNDVVDALLRDKGELSQSSSSS 132
+K + A+ R+M+M+ +I+ ER I +N + D + LL + Q + +
Sbjct: 218 HKGIMARGRVMEMLEKIIRERRNEINSHNNHHE-------DFLQQLLAVDNDTPQLTDAE 270
Query: 133 NLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDG 192
I NI+ +I G++T +A+T +K+L + L L+EE ++ K+ +N P
Sbjct: 271 ------IKDNILTMIIAGQDTTASALTWMVKYLGENQKVLDILIEEQSQITKKASNKPF- 323
Query: 193 YTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMD 252
D +P+ ++ E+LRMA++V R ++D E++GY I K W + S+H+D
Sbjct: 324 LELEDLSEMPYASKMVKESLRMASVVPWFPRLVLQDCEMEGYKIKKGWNINIDARSIHLD 383
Query: 253 SKNYENPYKFDPWRWE 268
Y P+KF+P R+E
Sbjct: 384 PTVYSEPHKFNPLRFE 399
Score = 28.9 bits (63), Expect = 4.5
Identities = 10/20 (50%), Positives = 16/20 (80%)
Query: 36 IPKGNSGWPLLGETLDFIAS 55
+P G+ G+P++GETL F+ S
Sbjct: 35 VPPGSDGFPVIGETLQFMLS 54
>At3g30290 cytochrome P450, putative
Length = 408
Score = 94.4 bits (233), Expect = 9e-20
Identities = 56/197 (28%), Positives = 96/197 (48%), Gaps = 18/197 (9%)
Query: 72 LYKSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVNDVVDALLRDKGELSQSSSS 131
+Y +KA++RM + + EE +K E E+ G + ++ +
Sbjct: 151 VYNMMKARKRMKTL---LKEEVLK-----KREAGEEFGEFSKIIFG--------EKEGEK 194
Query: 132 SNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPD 191
+ ++ + + I F + ET P + +KF+++ P + +L E+ + + K+
Sbjct: 195 ETMSMKNVIEYIYTFFVIANETTPRILAATVKFISENPKVMQELQREHAMIFENKSE-EA 253
Query: 192 GYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHM 251
G TW DY S+ FT VI+E+LR++ V I RK D ++ Y IP W M S H
Sbjct: 254 GLTWEDYKSMTFTNMVINESLRISTTVPVILRKPDHDTKVGDYTIPAGWNFM-GYPSAHF 312
Query: 252 DSKNYENPYKFDPWRWE 268
D YE+P +F+PWRW+
Sbjct: 313 DPTKYEDPLEFNPWRWK 329
>At5g36110 cytochrome P450-like
Length = 477
Score = 92.0 bits (227), Expect = 4e-19
Identities = 57/223 (25%), Positives = 108/223 (47%), Gaps = 19/223 (8%)
Query: 46 LGETLDFIASGYSSCPVTFMEKRKSILYKSLKAKERMMKMVGRIVEERMKLIMDNNAEDD 105
L E + +A G S P+ R + +++KA + K V IV +R + + A ++
Sbjct: 202 LEEQFNTVAVGIFSIPIDLPGTRFN---RAIKASRLLRKEVSAIVRQRKEELKAGKALEE 258
Query: 106 EKVGVVNDVVDALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFL 165
+D++ +L + GE E ++ II +I G +T T + +L
Sbjct: 259 ------HDILSHMLMNIGETKD---------EDLADKIIGLLIGGHDTASIVCTFVVNYL 303
Query: 166 TDYPLALSKLMEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRKA 225
++P ++++E E+ K+K +G W D + ++ NV E +R+ ++G +R+A
Sbjct: 304 AEFPHVYQRVLQEQKEILKEKKE-KEGLRWEDIEKMRYSWNVACEVMRIVPPLSGTFREA 362
Query: 226 VKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRWE 268
+ KG+ IPK W + S T+ HM+ + P +F+P R+E
Sbjct: 363 IDHFSFKGFYIPKGWKLYWSATATHMNPDYFPEPERFEPNRFE 405
Score = 36.2 bits (82), Expect = 0.028
Identities = 18/56 (32%), Positives = 31/56 (55%), Gaps = 3/56 (5%)
Query: 13 VLLMSLWFVKKEKNNIKLRNGIKIPKGNSGWPLLGETLDFIASGYSSCPVTFMEKR 68
+LL L ++K ++ N +P GN+G PL+GE+ F+++G P F+ R
Sbjct: 14 ILLSLLLLLRKHLSHFSYPN---LPPGNTGLPLIGESFSFLSAGRQGHPEKFITDR 66
>At4g15393 putative protein
Length = 453
Score = 87.0 bits (214), Expect = 1e-17
Identities = 51/199 (25%), Positives = 97/199 (48%), Gaps = 21/199 (10%)
Query: 72 LYKSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVNDVVDALLRDKGELSQSSSS 131
+Y+ +KA+ RMMK++ V ++ +K+G + + + D + S
Sbjct: 205 VYRIVKARNRMMKVIKETVVKKRA--------SGKKLG---EFFETIFGD-------TES 246
Query: 132 SNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPD 191
+ +E+ ++ I + ET P + +K ++D P + +L E+ + + K +
Sbjct: 247 VTMSIEIATEYIFTLFVLANETTPGVLAATIKLISDNPKVMQELRREHEGIVQDKIKKDE 306
Query: 192 --GYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSV 249
TW DY S+ FTQ VI+E+LR+ + V + R +++ Y IP W + V
Sbjct: 307 TADLTWEDYKSMTFTQMVINESLRITSTVPTVLRIIDHEIQFGDYTIPAGW-IFMGYPYV 365
Query: 250 HMDSKNYENPYKFDPWRWE 268
H + + Y++P F+PWRW+
Sbjct: 366 HFNPEKYDDPLAFNPWRWK 384
Score = 34.3 bits (77), Expect = 0.11
Identities = 20/52 (38%), Positives = 29/52 (55%), Gaps = 8/52 (15%)
Query: 4 FIGVYVSIIVLLMSLWFV--KKEKNNIKLRNGIKIPKGNSGWPLLGETLDFI 53
F V VS+IV+ + W K K N KL P G+ G+P++GET +F+
Sbjct: 7 FWAVIVSLIVVKLCHWIYQWKNPKGNGKL------PPGSMGYPIIGETFEFM 52
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.328 0.142 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,421,740
Number of Sequences: 26719
Number of extensions: 303052
Number of successful extensions: 1471
Number of sequences better than 10.0: 201
Number of HSP's better than 10.0 without gapping: 143
Number of HSP's successfully gapped in prelim test: 58
Number of HSP's that attempted gapping in prelim test: 1177
Number of HSP's gapped (non-prelim): 257
length of query: 355
length of database: 11,318,596
effective HSP length: 100
effective length of query: 255
effective length of database: 8,646,696
effective search space: 2204907480
effective search space used: 2204907480
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC136451.2


