

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136451.10 - phase: 0 /pseudo
(820 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
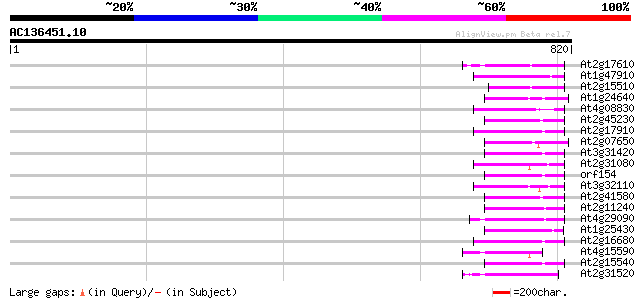
Score E
Sequences producing significant alignments: (bits) Value
At2g17610 putative non-LTR retroelement reverse transcriptase 78 2e-14
At1g47910 reverse transcriptase, putative 76 7e-14
At2g15510 putative non-LTR retroelement reverse transcriptase 74 4e-13
At1g24640 hypothetical protein 73 7e-13
At4g08830 putative protein 72 1e-12
At2g45230 putative non-LTR retroelement reverse transcriptase 72 1e-12
At2g17910 putative non-LTR retroelement reverse transcriptase 72 1e-12
At2g07650 putative non-LTR retrolelement reverse transcriptase 69 8e-12
At3g31420 hypothetical protein 67 4e-11
At2g31080 putative non-LTR retroelement reverse transcriptase 67 5e-11
orf154 -mitochondrial genome- 66 9e-11
At3g32110 non-LTR reverse transcriptase, putative 66 9e-11
At2g41580 putative non-LTR retroelement reverse transcriptase 64 3e-10
At2g11240 pseudogene 64 3e-10
At4g29090 putative protein 64 4e-10
At1g25430 hypothetical protein 61 2e-09
At2g16680 putative non-LTR retroelement reverse transcriptase 60 4e-09
At4g15590 reverse transcriptase like protein 60 5e-09
At2g15540 putative non-LTR retroelement reverse transcriptase 59 1e-08
At2g31520 putative non-LTR retroelement reverse transcriptase 59 1e-08
>At2g17610 putative non-LTR retroelement reverse transcriptase
Length = 773
Score = 77.8 bits (190), Expect = 2e-14
Identities = 52/149 (34%), Positives = 75/149 (49%), Gaps = 13/149 (8%)
Query: 663 SMSCHLCLFTLFPSSRLPQVSSPLLNLF*IIFFLGGNDDHK-KITWVDWNYVCLSKEVGG 721
SMSC L P + Q++S + +F N K KI WV W+ + K++GG
Sbjct: 259 SMSCFL-----LPMRLIHQITSAMR------WFWWSNTKVKHKIPWVAWSKLNDPKKMGG 307
Query: 722 LGVRRIREFNSALLGKWSWMVLVEKESLWYRVLVARYGEEGGRLLDGGRTASAWWRAIAT 781
L +R +++FN ALL K SW +L + SL RV A+Y + RLLD T+ + + +
Sbjct: 308 LAIRDLKDFNIALLAKQSWRILQQPFSLMARVFKAKYFPK-ERLLDAKATSQSSYAWKSI 366
Query: 782 LHSEAWFGTNVKRSVGNGENIYFWSDVWV 810
LH +K GNG NI W D W+
Sbjct: 367 LHGTKLISRGLKYIAGNGNNIQLWKDNWL 395
>At1g47910 reverse transcriptase, putative
Length = 1142
Score = 76.3 bits (186), Expect = 7e-14
Identities = 45/134 (33%), Positives = 71/134 (52%), Gaps = 3/134 (2%)
Query: 678 RLPQVSSPLLNLF*IIFFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGK 737
RLP+ + L F+ N D + + W+ W+ +C SK GGLG R + +FNSALL K
Sbjct: 593 RLPKAITSKLTSAVAKFWWSSNGDSRGMHWMAWDKLCSSKSDGGLGFRNVDDFNSALLAK 652
Query: 738 WSWMVLVEKESLWYRVLVARYGEEGGRLLD-GGRTASAWWRAIATLHSEAWFGTNVKRSV 796
W ++ +SL+ +V RY + L + S WR++ + S + G +KR V
Sbjct: 653 QLWRLITAPDSLFAKVFKGRYFRKSNPLDSIKSYSPSYGWRSMISARSLVYKGL-IKR-V 710
Query: 797 GNGENIYFWSDVWV 810
G+G +I W+D W+
Sbjct: 711 GSGASISVWNDPWI 724
>At2g15510 putative non-LTR retroelement reverse transcriptase
Length = 1138
Score = 73.6 bits (179), Expect = 4e-13
Identities = 38/111 (34%), Positives = 57/111 (51%), Gaps = 2/111 (1%)
Query: 701 DHKKIT-WVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRVLVARYG 759
+HK+ T WV W +CL+KE GGLG R I FN ALL K SW +L L+ R +RY
Sbjct: 607 EHKRKTHWVSWEKLCLAKESGGLGFRDIESFNQALLAKQSWRILQYPSFLFARFFKSRYF 666
Query: 760 EEGGRLLDGGRTASAWWRAIATLHSEAWFGTNVKRSVGNGENIYFWSDVWV 810
++ L+ + + L +++ VGNG+++ W D W+
Sbjct: 667 DD-EEFLEADLGVRPSYACCSILFGRELLAKGLRKEVGNGKSLNVWMDPWI 716
>At1g24640 hypothetical protein
Length = 1270
Score = 72.8 bits (177), Expect = 7e-13
Identities = 42/125 (33%), Positives = 64/125 (50%), Gaps = 5/125 (4%)
Query: 694 FFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRV 753
F+ D +KI W W +CL K+ GGLG R I+ FN ALL K +W +L + L R+
Sbjct: 788 FWWDSCDHSRKIHWQSWERLCLPKDSGGLGFRDIQSFNQALLAKQAWRLLHFPDCLLSRL 847
Query: 754 LVARYGEEGGRLLDG--GRTASAWWRAIATLHSEAWFGTNVKRSVGNGENIYFWSDVWVG 811
L +RY + LD + S WR+I L +++ VG+G +++ W D W+
Sbjct: 848 LKSRY-FDATDFLDAALSQRPSFGWRSI--LFGRELLSKGLQKRVGDGASLFVWIDPWID 904
Query: 812 GVSFK 816
F+
Sbjct: 905 DNGFR 909
>At4g08830 putative protein
Length = 947
Score = 72.4 bits (176), Expect = 1e-12
Identities = 44/132 (33%), Positives = 65/132 (48%), Gaps = 22/132 (16%)
Query: 679 LPQVSSPLLNLF*IIFFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKW 738
LPQ + L+ +F LG + + KK+ V W+ VCL K GGLG+R + N AL+ K
Sbjct: 487 LPQSTLEGLDKLARVFLLGSSAEKKKLHLVAWDRVCLPKSEGGLGIRTSKCMNKALVSKV 546
Query: 739 SWMVLVEKESLWYRVLVARYGEEGGRLLDGGRTASAWWRAIATLHSEAWFGTNVKRSVGN 798
W ++ ++ SLW R+L ++Y ++ G S W VGN
Sbjct: 547 GWRLINDRYSLWARILRSKYRVGLREVVSRG---SRW-------------------VVGN 584
Query: 799 GENIYFWSDVWV 810
G +I FWSD W+
Sbjct: 585 GRDILFWSDNWL 596
>At2g45230 putative non-LTR retroelement reverse transcriptase
Length = 1374
Score = 72.4 bits (176), Expect = 1e-12
Identities = 39/120 (32%), Positives = 61/120 (50%), Gaps = 5/120 (4%)
Query: 694 FFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRV 753
F+ + + + W W ++ K VGGLG + I FN ALLGK W ++ EK+SL +V
Sbjct: 825 FWWKNKKEGRGLHWKAWCHLSRPKAVGGLGFKEIEAFNIALLGKQLWRMITEKDSLMAKV 884
Query: 754 LVARYGEEGGRLLD--GGRTASAWWRAIATLHSEAWFGTNVKRSVGNGENIYFWSDVWVG 811
+RY + L G R + AW + ++ ++ +GNGE I W+D W+G
Sbjct: 885 FKSRYFSKSDPLNAPLGSRPSFAW---KSIYEAQVLIKQGIRAVIGNGETINVWTDPWIG 941
>At2g17910 putative non-LTR retroelement reverse transcriptase
Length = 1344
Score = 72.0 bits (175), Expect = 1e-12
Identities = 43/134 (32%), Positives = 65/134 (48%), Gaps = 3/134 (2%)
Query: 678 RLPQVSSPLLNLF*IIFFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGK 737
+LP+ L + F+ +KI W+ W + L K+ GG G + ++ FN ALL K
Sbjct: 787 KLPKNLCQKLTTVMMDFWWNSMQQKRKIHWLSWQRLTLPKDQGGFGFKDLQCFNQALLAK 846
Query: 738 WSWMVLVEKESLWYRVLVARYGEEGGRL-LDGGRTASAWWRAIATLHSEAWFGTNVKRSV 796
+W VL EK SL+ RV +RY L G S WR+I L ++ +
Sbjct: 847 QAWRVLQEKGSLFSRVFQSRYFSNSDFLSATRGSRPSYAWRSI--LFGRELLMQGLRTVI 904
Query: 797 GNGENIYFWSDVWV 810
GNG+ + W+D W+
Sbjct: 905 GNGQKTFVWTDKWL 918
>At2g07650 putative non-LTR retrolelement reverse transcriptase
Length = 732
Score = 69.3 bits (168), Expect = 8e-12
Identities = 39/129 (30%), Positives = 61/129 (47%), Gaps = 8/129 (6%)
Query: 694 FFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRV 753
F G + +K + W VC + GGLG+R+ ++ N ALL K W ++ + SLW R+
Sbjct: 350 FLWGSSVTQRKQHLISWKRVCKPRSEGGLGIRKAQDMNKALLSKVGWRLIQDYHSLWARI 409
Query: 754 LVARYGEEGGRLLDGGRT-----ASAWWRAIATLHSEAWFGTNVKRSVGNGENIYFWSDV 808
+ Y + R DG T S+ WR++A E + +G+G I FW D
Sbjct: 410 MRCNYRVQDVR--DGAWTKVRSVCSSTWRSVALGMREVVI-PGLSWVIGDGREILFWMDK 466
Query: 809 WVGGVSFKE 817
W+ + E
Sbjct: 467 WLTNIPLAE 475
>At3g31420 hypothetical protein
Length = 1491
Score = 67.0 bits (162), Expect = 4e-11
Identities = 40/118 (33%), Positives = 58/118 (48%), Gaps = 3/118 (2%)
Query: 694 FFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRV 753
F+ ++ K + W W + + K+ GGLG + + FN ALLGK +W +L L RV
Sbjct: 983 FWWSSGNETKGMHWFTWKRMSIPKKEGGLGFKELENFNLALLGKQTWHLLQHPNCLMARV 1042
Query: 754 LVARYGEEGGRL-LDGGRTASAWWRAIATLHSEAWFGTNVKRSVGNGENIYFWSDVWV 810
L RY E + GR AS W++I LH ++ VG+G I W D W+
Sbjct: 1043 LRGRYFPETNVMNAVQGRRASFVWKSI--LHGRNLLKKGLRCCVGDGSLINAWLDPWL 1098
>At2g31080 putative non-LTR retroelement reverse transcriptase
Length = 1231
Score = 66.6 bits (161), Expect = 5e-11
Identities = 44/135 (32%), Positives = 64/135 (46%), Gaps = 4/135 (2%)
Query: 679 LPQVSSPLLNLF*IIFFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKW 738
LP + L+ + F G + KK + W +C K GG+G+R R+ N AL+ K
Sbjct: 673 LPVSTLDTLDRYSRTFLWGSTMEKKKQHLLSWRKICKPKAEGGIGLRSARDMNKALVAKV 732
Query: 739 SWMVLVEKESLWYRVLVARY---GEEGGRLLDGGRTASAWWRAIATLHSEAWFGTNVKRS 795
W +L +KESLW RV+ +Y G + L S+ WR++A E V
Sbjct: 733 GWRLLQDKESLWARVVRKKYKVGGVQDTSWLKPQPRWSSTWRSVAVGLREV-VVKGVGWV 791
Query: 796 VGNGENIYFWSDVWV 810
G+G I FW D W+
Sbjct: 792 PGDGCTIRFWLDRWL 806
>orf154 -mitochondrial genome-
Length = 154
Score = 65.9 bits (159), Expect = 9e-11
Identities = 39/119 (32%), Positives = 62/119 (51%), Gaps = 4/119 (3%)
Query: 694 FFLGGNDDHKKITWVDWNYVCLSKEV-GGLGVRRIREFNSALLGKWSWMVLVEKESLWYR 752
F+ ++ +KI+WV W +C SKE GGLG R + FN ALL K S+ ++ + +L R
Sbjct: 28 FWWSSCENKRKISWVAWQKLCKSKEDDGGLGFRDLGWFNQALLAKQSFRIIHQPHTLLSR 87
Query: 753 VLVARYGEEGGRL-LDGGRTASAWWRAIATLHSEAWFGTNVKRSVGNGENIYFWSDVWV 810
+L +RY + G S WR+I +H + R++G+G + W D W+
Sbjct: 88 LLRSRYFPHSSMMECSVGTRPSYAWRSI--IHGRELLSRGLLRTIGDGIHTKVWLDRWI 144
>At3g32110 non-LTR reverse transcriptase, putative
Length = 1911
Score = 65.9 bits (159), Expect = 9e-11
Identities = 45/139 (32%), Positives = 66/139 (47%), Gaps = 10/139 (7%)
Query: 678 RLPQVSSPLLNLF*IIFFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGK 737
+LPQ + L+ F G + KK V W VCL + GGLG+R N AL+ K
Sbjct: 1348 KLPQSTLDGLDKVSRAFLWGSTLEKKKQHLVAWTRVCLPRREGGLGIRSATAMNKALIAK 1407
Query: 738 WSWMVLVEKESLWYRVLVARYGEEGGRLLDGGRTA-----SAWWRAIAT-LHSEAWFGTN 791
W VL + SLW +V+ ++Y + G + D T S+ WR++ L W
Sbjct: 1408 VGWRVLNDGSSLWAQVVRSKY--KVGDVHDRNWTVAKSNWSSTWRSVGVGLRDVIW--RE 1463
Query: 792 VKRSVGNGENIYFWSDVWV 810
+G+G I FW+D W+
Sbjct: 1464 QHWVIGDGRQIRFWTDRWL 1482
>At2g41580 putative non-LTR retroelement reverse transcriptase
Length = 1094
Score = 64.3 bits (155), Expect = 3e-10
Identities = 38/119 (31%), Positives = 60/119 (49%), Gaps = 5/119 (4%)
Query: 694 FFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRV 753
F+ D KKI W+ + L K +GG G + ++ FN ALL K + + + +SL ++
Sbjct: 554 FWWNSVQDKKKIHWIGAQKLMLPKFLGGFGFKDLQCFNQALLAKQASRLHTDSDSLLSQI 613
Query: 754 LVARYGEEGGRL--LDGGRTASAWWRAIATLHSEAWFGTNVKRSVGNGENIYFWSDVWV 810
L +RY L G R + AW + L+ + +K+ +GNGEN Y W D W+
Sbjct: 614 LKSRYYMNSDFLSATKGTRPSYAWQ---SILYGRELLVSGLKKIIGNGENTYVWMDNWI 669
>At2g11240 pseudogene
Length = 1044
Score = 64.3 bits (155), Expect = 3e-10
Identities = 36/119 (30%), Positives = 61/119 (51%), Gaps = 5/119 (4%)
Query: 694 FFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRV 753
F+ + D KK+ W+ W+ + +K+ GGLG + I FN ALL K SW ++ + R+
Sbjct: 651 FWWDSSADKKKMCWIAWSKMAKNKKEGGLGFKDITNFNDALLAKLSWRIVQSPSCVLVRI 710
Query: 754 LVARYGEEGGRLLDGGRTA--SAWWRAIATLHSEAWFGTNVKRSVGNGENIYFWSDVWV 810
L+ +Y LD TA S WR I T + + + + +G+G + W++ W+
Sbjct: 711 LLGKYCRTSS-FLDCSVTAASSHGWRGICT--GKDLIKSQLGKVIGSGLDTLVWNEPWL 766
>At4g29090 putative protein
Length = 575
Score = 63.5 bits (153), Expect = 4e-10
Identities = 43/139 (30%), Positives = 64/139 (45%), Gaps = 8/139 (5%)
Query: 673 LFPSSRLPQVSSPLLNLF*IIFFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNS 732
L P + Q+ S L + F+ + K + W W+++ K GG+G + I FN
Sbjct: 12 LLPKTVCKQIISVLAD-----FWWRNKQEAKGMHWKAWDHLSCYKAEGGIGFKDIEAFNL 66
Query: 733 ALLGKWSWMVLVEKESLWYRVLVARYGEEGGRL-LDGGRTASAWWRAIATLHSEAWFGTN 791
ALLGK W +L ESL +V +RY + L G S W++I S+
Sbjct: 67 ALLGKQMWRMLSRPESLMAKVFKSRYFHKSDPLNAPLGSRPSFVWKSIHA--SQEILRQG 124
Query: 792 VKRSVGNGENIYFWSDVWV 810
+ VGNGE+I W W+
Sbjct: 125 ARAVVGNGEDIIIWRHKWL 143
>At1g25430 hypothetical protein
Length = 1213
Score = 61.2 bits (147), Expect = 2e-09
Identities = 39/117 (33%), Positives = 56/117 (47%), Gaps = 3/117 (2%)
Query: 694 FFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRV 753
F GN + K V W +CL K GGLG+RR+ E+N L + W + V K+SLW
Sbjct: 839 FLWSGNIEQAKGIKVSWAALCLPKSEGGLGLRRLLEWNKTLSMRLIWRLFVAKDSLWADW 898
Query: 754 LVARYGEEGG-RLLDGGRTASAWWRAIATLHSEAWFGTNVKRSVGNGENIYFWSDVW 809
+ G ++GG++ S W+ + +L A K VGNG +W D W
Sbjct: 899 QHLHHLSRGSFWAVEGGQSDSWTWKRLLSLRPLAHQFLVCK--VGNGLKADYWYDNW 953
>At2g16680 putative non-LTR retroelement reverse transcriptase
Length = 1319
Score = 60.5 bits (145), Expect = 4e-09
Identities = 39/134 (29%), Positives = 62/134 (46%), Gaps = 3/134 (2%)
Query: 678 RLPQVSSPLLNLF*IIFFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGK 737
RLP+ L + F+ + KI W+ + L K +GG G + ++ FN ALL K
Sbjct: 761 RLPKGLCTKLTSVMMDFWWNSMEFSNKIHWIGGKKLTLPKSLGGFGFKDLQCFNQALLAK 820
Query: 738 WSWMVLVEKESLWYRVLVARYGEEGGRL-LDGGRTASAWWRAIATLHSEAWFGTNVKRSV 796
+W + + +S+ ++ +RY L G S WR+I L+ +KR +
Sbjct: 821 QAWRLFSDSKSIVSQIFKSRYFMNTDFLNARQGTRPSYTWRSI--LYGRELLNGGLKRLI 878
Query: 797 GNGENIYFWSDVWV 810
GNGE W D W+
Sbjct: 879 GNGEQTNVWIDKWL 892
>At4g15590 reverse transcriptase like protein
Length = 929
Score = 60.1 bits (144), Expect = 5e-09
Identities = 41/120 (34%), Positives = 55/120 (45%), Gaps = 8/120 (6%)
Query: 663 SMSCHLCLFTLFPSSRLPQVSSPLLNLF*IIFFLGGNDDHKKITWVDWNYVCLSKEVGGL 722
S+ H L P+S L Q+ N F G + +K + W VC K GGL
Sbjct: 597 SIPIHTMSSILLPASLLEQLDKVSRN-----FLWGSTVEKRKQHLLSWKKVCRPKAAGGL 651
Query: 723 GVRRIREFNSALLGKWSWMVLVEKESLWYRVLVARY---GEEGGRLLDGGRTASAWWRAI 779
G+R ++ N ALL K W +L +K SLW RVL +Y L T S+ WR+I
Sbjct: 652 GLRASKDMNRALLAKVGWRLLNDKVSLWARVLRRKYKVTDVHDSSWLVPKATWSSTWRSI 711
>At2g15540 putative non-LTR retroelement reverse transcriptase
Length = 1225
Score = 58.9 bits (141), Expect = 1e-08
Identities = 35/118 (29%), Positives = 56/118 (46%), Gaps = 3/118 (2%)
Query: 694 FFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRV 753
F+ ++ I W+ W+ +C GGLG R + EFN LL K W ++ SL RV
Sbjct: 705 FWWKTREESNGIHWIAWDKLCTPFSDGGLGFRTLEEFNLVLLAKQLWRLIRFPNSLLSRV 764
Query: 754 LVARYGEEGGRLLDG-GRTASAWWRAIATLHSEAWFGTNVKRSVGNGENIYFWSDVWV 810
L RY + G S WR+I + ++ + ++R++G+G W D W+
Sbjct: 765 LRGRYFRYSDPIQIGKANRPSFGWRSI--MAAKPLLLSGLRRTIGSGMLTRVWEDPWI 820
>At2g31520 putative non-LTR retroelement reverse transcriptase
Length = 1524
Score = 58.5 bits (140), Expect = 1e-08
Identities = 43/140 (30%), Positives = 69/140 (48%), Gaps = 11/140 (7%)
Query: 663 SMSCHLCLFTLFPSSRLPQVSSPLLNLF*IIFFLGGNDDHKKITWVDWNYVCLSKEVGGL 722
+MSC F L P + ++ S L+N F+ + + I WV W + SK+ GGL
Sbjct: 960 AMSC----FKL-PKGIVSEIESLLMN-----FWWEKASNQRGIPWVAWKRLQYSKKEGGL 1009
Query: 723 GVRRIREFNSALLGKWSWMVLVEKESLWYRVLVARYGEEGGRLLDGGRTASAWWRAIATL 782
G R + +FN ALL K +W ++ SL+ RV+ ARY ++ +LD + + L
Sbjct: 1010 GFRDLAKFNDALLAKQAWRLIQYPNSLFARVMKARYFKDVS-ILDAKVRKQQSYGWASLL 1068
Query: 783 HSEAWFGTNVKRSVGNGENI 802
A + +G+G+NI
Sbjct: 1069 DGIALLKKGTRHLIGDGQNI 1088
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.361 0.164 0.647
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,836,640
Number of Sequences: 26719
Number of extensions: 684765
Number of successful extensions: 2278
Number of sequences better than 10.0: 57
Number of HSP's better than 10.0 without gapping: 52
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 2161
Number of HSP's gapped (non-prelim): 72
length of query: 820
length of database: 11,318,596
effective HSP length: 108
effective length of query: 712
effective length of database: 8,432,944
effective search space: 6004256128
effective search space used: 6004256128
T: 11
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.9 bits)
S2: 64 (29.3 bits)
Medicago: description of AC136451.10


