

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136449.11 - phase: 0 /pseudo
(982 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
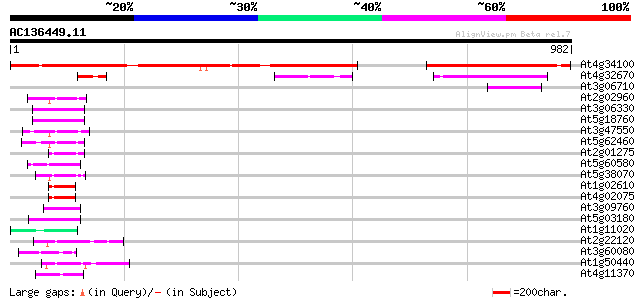
Score E
Sequences producing significant alignments: (bits) Value
At4g34100 unknown protein 795 0.0
At4g32670 putative protein 151 2e-36
At3g06710 hypothetical protein 74 3e-13
At2g02960 unknown protein 66 1e-10
At3g06330 unknown protein 65 1e-10
At5g18760 unknown protein 64 4e-10
At3g47550 unknown protein 63 7e-10
At5g62460 unknown protein 63 9e-10
At2g01275 unknown protein 57 5e-08
At5g60580 unknown protein 56 9e-08
At5g38070 putative protein 56 9e-08
At1g02610 hypothetical protein 53 7e-07
At4g02075 PIT1 (Pit1) 53 1e-06
At3g09760 unknown protein 51 3e-06
At5g03180 unknown protein 48 3e-05
At1g11020 unknown protein 47 4e-05
At2g22120 unknown protein 46 9e-05
At3g60080 unknown protein 46 1e-04
At1g50440 unknown protein 45 2e-04
At4g11370 RING-H2 finger protein RHA1a -like protein 44 3e-04
>At4g34100 unknown protein
Length = 1051
Score = 795 bits (2053), Expect = 0.0
Identities = 428/665 (64%), Positives = 481/665 (71%), Gaps = 98/665 (14%)
Query: 1 MEIANEPPPSLDGTPIA--ATTPSSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSAKY 58
MEI+ S+ G + + PS SS SSSSS + S + + + +T ++Y
Sbjct: 1 MEISPADSLSISGAAASEVVSEPSVSSSSSSSSPNQASPNPFSNMDPAVSTATG---SRY 57
Query: 59 DDDDEDEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHP 118
DDDEDEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHP
Sbjct: 58 VDDDEDEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHP 117
Query: 119 FSFSPVYAENAPARLPFQEFVVGMAMKACHVLQFFVRLSFVLSVWLLIIPFITFWIWRLA 178
FSFSPVYA+NAP+RLPFQEFVVG+AMKACHVLQFF+RLSFVLSVWLL IPFITFWIWRLA
Sbjct: 118 FSFSPVYADNAPSRLPFQEFVVGIAMKACHVLQFFLRLSFVLSVWLLTIPFITFWIWRLA 177
Query: 179 FVRSFGEAQRLFFYHLFTAVILTDCLHGFLLSASIVFIFFGGTSLRDYFRHLREIGGQDA 238
FVR+FGEAQRLF H+ T VILTDCLHG DYFRHLRE+GGQ+
Sbjct: 178 FVRTFGEAQRLFLSHISTTVILTDCLHG------------------DYFRHLRELGGQE- 218
Query: 239 EREDEVDRNGARVARRPAGQANRNVNGDANGEDAVAAQGVAGAGQVIRRNAENVAARWEM 298
ER+D+VDRNGAR ARRPAGQANRN+ G+ NGEDA QG A GQ+ RRN ENV AR ++
Sbjct: 219 ERDDDVDRNGARAARRPAGQANRNLAGEGNGEDA-GDQG-AAVGQIARRNPENVLARLDI 276
Query: 299 QAARLEAHVEQMFDGLDDADGAEDVPFDELVGMQ-------------------------- 332
QAARLEA VEQMFDGLDDADGAEDVPFDELVGMQ
Sbjct: 277 QAARLEAQVEQMFDGLDDADGAEDVPFDELVGMQGPVFHLVENAFTVLASNMIFLGVVIF 336
Query: 333 ---------------------VPPTDASLSLA-------NITLKNALTAVQNLSTATQES 364
P ASL L NITLK+ALTAV NL++ Q +
Sbjct: 337 VPFTLGRIILYHVSWLFAAARGPAVAASLHLTDTGLSLENITLKSALTAVSNLTSEGQGN 396
Query: 365 GSIGQIAEMLKVNASELSEMSNNITASVSDDLLKGGSIGTSRISDVTTLAXGYIFLSTLI 424
G +GQ+ EM+KVN SEL+ +N T SV+ DLLKG ++G S++SD+TTLA GY+F+ L+
Sbjct: 397 GLLGQLTEMMKVNGSELNGANN--TLSVATDLLKGSTVGASKLSDITTLAVGYMFIVFLV 454
Query: 425 FCYFGVVALIRYTKGEPLTAGRFYGIASIAETIPSLFRQFLAAMRHLMTMVKVAFLLVIE 484
F Y G++ALIRY K E +PSL RQFLAAMRHLMTM+KVAFLLVIE
Sbjct: 455 FLYLGIIALIRYAK----------------EAVPSLLRQFLAAMRHLMTMIKVAFLLVIE 498
Query: 485 LGVFPLMCGWWLDVCTIQMFGKTMVHRAQFFSASPLASSLAHWVVGIVYMLQISIFVSLL 544
LGVFPLMCGWWLDVCT++MFGKTM HR QF S SPLASSL HWVVGI+YMLQISIFVSLL
Sbjct: 499 LGVFPLMCGWWLDVCTVRMFGKTMSHRVQFLSISPLASSLVHWVVGIMYMLQISIFVSLL 558
Query: 545 RGVLRNGVLYFLRDPADPNYNPSRDLIDNPVHKHARRVLLSVAVYGSLMVMLVILPR*LG 604
RGVLR GVLYFLRDPADPNYNP RDLID+PVHKHARRVLLSVAVYGSL+VMLV LP L
Sbjct: 559 RGVLRPGVLYFLRDPADPNYNPFRDLIDDPVHKHARRVLLSVAVYGSLIVMLVFLPVKLA 618
Query: 605 IADGP 609
I P
Sbjct: 619 IRMAP 623
Score = 403 bits (1035), Expect = e-112
Identities = 194/253 (76%), Positives = 217/253 (85%), Gaps = 6/253 (2%)
Query: 730 GVRYSIEQIRKRRTSVLLNQIWKWCSIVVKSSALLSIWIFVIPVLIGLLFELLVIVPMRV 789
G++Y+IE ++ +RTSVLLNQIWKWC IV KSS LL+IW+F+IPVLIGLLFELLVIVPMRV
Sbjct: 802 GIKYAIEHVKSKRTSVLLNQIWKWCGIVFKSSVLLAIWVFIIPVLIGLLFELLVIVPMRV 861
Query: 790 PVDESPVFLLYQDWALGLIFLKIWTRLVMLDHMMPLMDESWRVKFERVRDDGFSRLQGLW 849
PVDESPVFLLYQDWALGLIFLKIWTRLVMLDHM+P++D+SWR KFERVR+DGFSRLQGLW
Sbjct: 862 PVDESPVFLLYQDWALGLIFLKIWTRLVMLDHMLPIVDDSWRAKFERVREDGFSRLQGLW 921
Query: 850 VLREIVLPIIMKLLTALCVPYVLARGVFPALGYPLVVNSAVYRFAWLGCLSFSFVCFCAK 909
VLREIV PI+MKLLTALCVPYVLARGVFP LGYPLVVNSAVYRFAW+GCLS S CFCAK
Sbjct: 922 VLREIVFPIVMKLLTALCVPYVLARGVFPMLGYPLVVNSAVYRFAWIGCLSVSLFCFCAK 981
Query: 910 RFHVWFTNLHNSIRDDRYLIGRRLHNFGE-HVEKANEAATSTGVQDAILLGPNINQQDRD 968
R HVWF NLHNSIRDDRYLIGRRLHNFGE + N+ +S D +L+G ++ D
Sbjct: 982 RCHVWFRNLHNSIRDDRYLIGRRLHNFGEAALANQNQNQSSEDAGDGVLIG-----REGD 1036
Query: 969 ADVGLRLRHINQQ 981
D GLRLR QQ
Sbjct: 1037 VDNGLRLRRAIQQ 1049
>At4g32670 putative protein
Length = 808
Score = 151 bits (382), Expect = 2e-36
Identities = 78/200 (39%), Positives = 116/200 (58%), Gaps = 4/200 (2%)
Query: 742 RTSVLLNQIWKWCSIVVKSSALLSIWIFVIPVLIGLLFELLVIVPMRVPVDESPVFLLYQ 801
RT +LLN + + +++ L SIWI VIP ++GLL +L++I+P +VP+ ESPV+ L
Sbjct: 613 RTDLLLNHVLMF----IRNVLLFSIWISVIPGVLGLLIDLMIIIPSQVPLGESPVYNLLH 668
Query: 802 DWALGLIFLKIWTRLVMLDHMMPLMDESWRVKFERVRDDGFSRLQGLWVLREIVLPIIMK 861
DW +G++ L IW L ML + +WR K +R+R +RL W++R+++ II+
Sbjct: 669 DWLIGVVVLHIWIFLTMLTRINCFATVAWREKLQRIRSVTINRLPFTWLIRDVIGSIIVS 728
Query: 862 LLTALCVPYVLARGVFPALGYPLVVNSAVYRFAWLGCLSFSFVCFCAKRFHVWFTNLHNS 921
LL LCVPYV+ +FP LG+ VN V RF W L+ + F K LH
Sbjct: 729 LLFTLCVPYVVVNSLFPILGFSSAVNLTVQRFIWPAILALIPIWFSVKLIRDLILYLHKL 788
Query: 922 IRDDRYLIGRRLHNFGEHVE 941
D+RY +G RL +F E +E
Sbjct: 789 EFDNRYKVGERLVDFTEDLE 808
Score = 83.6 bits (205), Expect = 5e-16
Identities = 42/137 (30%), Positives = 78/137 (56%), Gaps = 9/137 (6%)
Query: 464 FLAAMRHLMTMVKVAFLLVIELGVFPLMCGWWLDVCTIQMFGKTMVHRAQFFSASPLASS 523
+L ++R + VK F+L +LGV P + G WL CT + GKT H + S PL +
Sbjct: 259 YLTSVRTFLPSVKDTFILSFKLGVLPWLLGCWLHFCTFPILGKTASHTVEVLSDYPLMAD 318
Query: 524 LAHWVVGIVYMLQISIFVSLLRGVLRNGVLYFLRDPADPNYNPSRDLIDNPVHKHARRVL 583
HW++G +Y++ + L++ +++ L++L D A+PNY ++ H +L
Sbjct: 319 -KHWLMGTLYLVSALSCMELIQKIVQKRALWYLLDVAEPNYKVTK--------LHLGPIL 369
Query: 584 LSVAVYGSLMVMLVILP 600
L+ A++G+++V+++ LP
Sbjct: 370 LAFALHGTMVVIVLHLP 386
Score = 46.2 bits (108), Expect = 9e-05
Identities = 21/51 (41%), Positives = 34/51 (66%), Gaps = 7/51 (13%)
Query: 119 FSFSPVYAENAPARLPFQEFVVGMAMKACHVLQFFVRLSFVLSVWLLIIPF 169
+S PVY+ENAP RLP+ EF++G+ M+A +R ++ W+L++PF
Sbjct: 5 YSIVPVYSENAPERLPWHEFLMGLLMRA-------LRFMNLILPWILMMPF 48
>At3g06710 hypothetical protein
Length = 302
Score = 74.3 bits (181), Expect = 3e-13
Identities = 33/95 (34%), Positives = 56/95 (58%)
Query: 836 RVRDDGFSRLQGLWVLREIVLPIIMKLLTALCVPYVLARGVFPALGYPLVVNSAVYRFAW 895
R+R+ G ++L +W+L++++ II LL LC+PY+L + + P LG+ N A+ F W
Sbjct: 208 RIRNVGINKLPSMWLLQDVIGSIITTLLATLCIPYLLLKSLLPLLGFSRSANPAIEPFIW 267
Query: 896 LGCLSFSFVCFCAKRFHVWFTNLHNSIRDDRYLIG 930
L+ V F AK+ + + H + DDRY++G
Sbjct: 268 PALLALIAVWFMAKQTYDLVSYFHRLVFDDRYVVG 302
>At2g02960 unknown protein
Length = 271
Score = 65.9 bits (159), Expect = 1e-10
Identities = 36/107 (33%), Positives = 54/107 (49%), Gaps = 7/107 (6%)
Query: 32 SPRGSKGKEIDAEAVATASTAPPSAKYDDDDEDEEDV----CRICRNPGDADNPLRYPCA 87
SP ++ +E+ +E+ A + D DDE+E + CRIC + +N L PCA
Sbjct: 2 SPVNAEVEEMRSESPVVNDKALDISDDDHDDENEPLIVSGECRICSDESPVEN-LESPCA 60
Query: 88 CSGSIKFVHQDCLLQWLNHSNARQCEVCKHPFSFSPVYAENAPARLP 134
CSGS+K+ H+ C+ +W N CE+C P + P Y P P
Sbjct: 61 CSGSLKYAHRKCVQRWCNEKGNIICEICHQP--YQPGYTAPPPPLQP 105
>At3g06330 unknown protein
Length = 426
Score = 65.5 bits (158), Expect = 1e-10
Identities = 27/90 (30%), Positives = 47/90 (52%)
Query: 41 IDAEAVATASTAPPSAKYDDDDEDEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCL 100
+ +E A ++ P + D++ +EE VCRIC + + N L+ C+C G ++ VH+ C
Sbjct: 190 VSSETSADQVSSVPPEETDEEIPEEEAVCRICLDVCEEGNTLKMECSCKGDLRLVHEACA 249
Query: 101 LQWLNHSNARQCEVCKHPFSFSPVYAENAP 130
++W + R C+VC+ PV P
Sbjct: 250 MKWFSTKGTRTCDVCRQVVQNLPVTLVRVP 279
>At5g18760 unknown protein
Length = 411
Score = 63.9 bits (154), Expect = 4e-10
Identities = 28/90 (31%), Positives = 46/90 (51%)
Query: 41 IDAEAVATASTAPPSAKYDDDDEDEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCL 100
I EA T P+ + +++ +EE VCRIC + + N L+ C+C G ++ VH+ C
Sbjct: 175 ISNEANGDQITPVPAEETEEEIPEEEAVCRICLDVCEEGNTLKMECSCKGDLRLVHEHCA 234
Query: 101 LQWLNHSNARQCEVCKHPFSFSPVYAENAP 130
++W + R C+VC+ PV P
Sbjct: 235 IKWFSTKGTRICDVCRQEVRNLPVILLRVP 264
>At3g47550 unknown protein
Length = 288
Score = 63.2 bits (152), Expect = 7e-10
Identities = 38/122 (31%), Positives = 56/122 (45%), Gaps = 14/122 (11%)
Query: 23 SSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSAKYDDDDEDEEDV----CRICRNPGDA 78
S S SS P+G+ D + + T P Y DE+E + CRIC+
Sbjct: 24 SDSSGESSYRPQGT-----DLASSSVNETEVPREYYAVADEEEPLLQSVECRICQEEDST 78
Query: 79 DNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHPFSFSPVYAENAPARLPFQEF 138
N L PCAC+GS+K+ H+ C+ +W N CE+C P+ + AP P E
Sbjct: 79 KN-LEAPCACNGSLKYAHRKCVQRWCNEKGDITCEICHQPYQ----HGYTAPPPPPPDET 133
Query: 139 VV 140
++
Sbjct: 134 II 135
>At5g62460 unknown protein
Length = 307
Score = 62.8 bits (151), Expect = 9e-10
Identities = 36/113 (31%), Positives = 55/113 (47%), Gaps = 15/113 (13%)
Query: 22 SSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSAKYDDDDEDEEDV----CRICRNPGD 77
S + S+S S+ + GK+ + E + + D DDE+E + CRIC+
Sbjct: 35 SRAQGSTSLSTTKSMDGKKTEEEET--------TEQRDVDDEEEPLIQSVECRICQEEDS 86
Query: 78 ADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHPFSFSPVYAENAP 130
N L PC+CSGS+K+ H+ C+ +W N CE+C S+ P Y P
Sbjct: 87 VKN-LESPCSCSGSLKYAHRKCVQRWCNEKGDTTCEICHK--SYQPGYTAPPP 136
>At2g01275 unknown protein
Length = 259
Score = 57.0 bits (136), Expect = 5e-08
Identities = 24/62 (38%), Positives = 34/62 (54%), Gaps = 3/62 (4%)
Query: 69 CRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHPFSFSPVYAEN 128
CRIC + D D+ + PC+CSGS+K+ H+ C+ +W N CE+C F P Y
Sbjct: 59 CRICHDE-DLDSNMETPCSCSGSVKYAHRRCVQRWCNEKGDTTCEICHQ--EFKPDYTAP 115
Query: 129 AP 130
P
Sbjct: 116 PP 117
>At5g60580 unknown protein
Length = 487
Score = 56.2 bits (134), Expect = 9e-08
Identities = 32/95 (33%), Positives = 46/95 (47%), Gaps = 3/95 (3%)
Query: 31 SSPRGSKGKEIDAEAVATASTAPPSAKYDDDDEDEEDVCRICR-NPGDADNPLRYPCACS 89
S+PR +G ++ + A +T A +D EDE VCRIC + L+ C+C
Sbjct: 221 STPRVKEG-DVFSNASEAGNTETGDADGEDIPEDEA-VCRICLVELCEGGETLKMECSCK 278
Query: 90 GSIKFVHQDCLLQWLNHSNARQCEVCKHPFSFSPV 124
G + H+DC L+W + CEVCK PV
Sbjct: 279 GELALAHKDCALKWFTIKGNKTCEVCKQEVKNLPV 313
>At5g38070 putative protein
Length = 259
Score = 56.2 bits (134), Expect = 9e-08
Identities = 31/101 (30%), Positives = 48/101 (46%), Gaps = 18/101 (17%)
Query: 46 VATASTAPPSAKYDDDDEDEEDV-------------CRICRNPGDADNPLRYPCACSGSI 92
+ T + A P A ++D E + CRIC + D D + PC+CSG++
Sbjct: 18 LGTVNRADPKADSVNEDGVSESISAGADLCESKFVQCRICHDE-DEDTNMDTPCSCSGTL 76
Query: 93 KFVHQDCLLQWLNHSNARQCEVCKHPFSFSPVYAENAPARL 133
KF H +C+ +W N CE+C+ + P Y AP +L
Sbjct: 77 KFAHHNCVQRWCNEKGDTVCEICRQ--QYKPGY--TAPRQL 113
>At1g02610 hypothetical protein
Length = 146
Score = 53.1 bits (126), Expect = 7e-07
Identities = 20/47 (42%), Positives = 30/47 (63%), Gaps = 1/47 (2%)
Query: 69 CRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVC 115
CRIC +A++ PC+CSG+IKF H+DC+ +W + CE+C
Sbjct: 20 CRICHEE-EAESYFEAPCSCSGTIKFAHRDCIQRWCDEKGNTICEIC 65
>At4g02075 PIT1 (Pit1)
Length = 218
Score = 52.8 bits (125), Expect = 1e-06
Identities = 19/47 (40%), Positives = 29/47 (61%), Gaps = 1/47 (2%)
Query: 69 CRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVC 115
CRIC + ++ PCACSG++KF H++C+ +W N CE+C
Sbjct: 20 CRICHEE-EEESFFEVPCACSGTVKFAHRNCIQRWCNEKGNTTCEIC 65
>At3g09760 unknown protein
Length = 491
Score = 51.2 bits (121), Expect = 3e-06
Identities = 22/66 (33%), Positives = 32/66 (48%), Gaps = 1/66 (1%)
Query: 60 DDDEDEEDVCRICR-NPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHP 118
+D +EE VCRIC G+ + C C G + H++C ++W R C+VCK
Sbjct: 257 EDVPEEEAVCRICLVELGEDSEAFKMECLCRGELALAHKECTIKWFTIKGNRTCDVCKQE 316
Query: 119 FSFSPV 124
PV
Sbjct: 317 VQNLPV 322
>At5g03180 unknown protein
Length = 462
Score = 47.8 bits (112), Expect = 3e-05
Identities = 28/93 (30%), Positives = 40/93 (42%), Gaps = 2/93 (2%)
Query: 33 PRGSKGKEIDAEAVATASTAPPSAKYDDDDEDEEDVCRICRNPGDADNP-LRYPCACSGS 91
P S+G E E + AS + +D +EE VCRIC + D + C C G
Sbjct: 212 PTPSRGDEKRLE-MTQASKLNENDDGGEDVPEEEAVCRICMVEMEEDEEAFKMECMCKGE 270
Query: 92 IKFVHQDCLLQWLNHSNARQCEVCKHPFSFSPV 124
+ H+ C ++W C+VCK PV
Sbjct: 271 LALAHKTCTIKWFTIKGNITCDVCKQEVRNLPV 303
>At1g11020 unknown protein
Length = 321
Score = 47.4 bits (111), Expect = 4e-05
Identities = 36/126 (28%), Positives = 49/126 (38%), Gaps = 20/126 (15%)
Query: 2 EIANEPPPSL---DGTPIAATTPSSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSAKY 58
E+ +PP S D P+ +SSS S S++S EI AE
Sbjct: 5 EVQLQPPDSQKLSDSAPLLGDHTNSSSASPSAASVVAGNSDEIKAEEDL----------- 53
Query: 59 DDDDEDEEDVCRICRNPGDA--DNPLRYPCACSGSIKFVHQDCLLQWLNHSNA---RQCE 113
++D CRIC + L PC C G+ +FVH+ CL W + C
Sbjct: 54 -ENDASSAPCCRICLEDDSELLGDELISPCMCKGTQQFVHRSCLDHWRSVKEGFAFSHCT 112
Query: 114 VCKHPF 119
CK F
Sbjct: 113 TCKAQF 118
>At2g22120 unknown protein
Length = 324
Score = 46.2 bits (108), Expect = 9e-05
Identities = 42/169 (24%), Positives = 72/169 (41%), Gaps = 22/169 (13%)
Query: 43 AEAVATASTAPPSAKYDDDDED-------EEDVCRICRNPGDADNPLRYPCACSGSIKFV 95
A+ + + PPS + + D E+ CRIC D PC C G+ K+V
Sbjct: 2 ADELELSPLVPPSPMVEPSEIDLEAGGPGEQIQCRICLETDGRD--FIAPCKCKGTSKYV 59
Query: 96 HQDCLLQWLNHSNA---RQCEVCKHPFSFSPVYAENAPAR-LPFQEFVVGMAMKACHVLQ 151
H+DCL W C CK P+ A + R L F+ FV +L
Sbjct: 60 HRDCLDHWRAIKEGFAFAHCTTCKAPYYLRVHSAGDRKWRTLKFRFFVTR------DILS 113
Query: 152 FFVRLSFVLSVWLLIIPFITFWIWRLAFVRS-FGEAQRLFFYHLFTAVI 199
F+ + V++ ++ FI ++ +++R +G + FY++ A++
Sbjct: 114 IFLAVQLVIAALAYMVYFID--SYQQSWLRHIWGFDSEVTFYYMCGALL 160
>At3g60080 unknown protein
Length = 306
Score = 45.8 bits (107), Expect = 1e-04
Identities = 31/102 (30%), Positives = 49/102 (47%), Gaps = 9/102 (8%)
Query: 16 IAATTPSSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSAKYDDDDEDEEDVCRICRNP 75
+A+ SSS SSSSSS K +ID+ S++ DD D D +C +C+
Sbjct: 117 LASDNSGSSSSSSSSSSSSLLKSSDIDSIPTIQISSS-LLCSTDDSDPDSVLLCAVCKED 175
Query: 76 G-DADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCK 116
++ R PC+ H DC++ WL+ N+ C +C+
Sbjct: 176 FIIGESARRLPCS-----HIYHSDCIVPWLSDHNS--CPLCR 210
>At1g50440 unknown protein
Length = 250
Score = 45.4 bits (106), Expect = 2e-04
Identities = 45/165 (27%), Positives = 68/165 (40%), Gaps = 21/165 (12%)
Query: 56 AKYDDDD-----EDEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNAR 110
A+ DDD+ ++ CRIC + G D L PC C G+ K VH+ CL W +
Sbjct: 46 AETDDDETTLLVSGDQPQCRICLDVGGED--LIAPCNCKGTQKHVHRSCLDNWRSTKEGF 103
Query: 111 QCEVCKHPFSFSPVYAENAPA-----RLPFQEFVVGMAMKACHVLQFFVRLSFVLSVWLL 165
C +F + A N PA RL FQ V F+ +S + V L
Sbjct: 104 AFSHCTECRAFFKLRA-NVPADRWWLRLRFQLLVARD--------HAFIFISVQMIVAFL 154
Query: 166 IIPFITFWIWRLAFVRSFGEAQRLFFYHLFTAVILTDCLHGFLLS 210
+ F+ L + + E F+ A++L L+GF ++
Sbjct: 155 GLLVYKFYGEELREMFGYEEHPYGFYTLAVLAIVLVGLLYGFFIA 199
>At4g11370 RING-H2 finger protein RHA1a -like protein
Length = 159
Score = 44.3 bits (103), Expect = 3e-04
Identities = 22/84 (26%), Positives = 39/84 (46%), Gaps = 3/84 (3%)
Query: 45 AVATASTAPPSAKYDDDDEDEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWL 104
+ + A+ P ++ D D ED C +C + ++D+ +R C H CL +W+
Sbjct: 62 SASLANELIPVVRFSDLPTDPEDCCTVCLSDFESDDKVRQLPKCG---HVFHHYCLDRWI 118
Query: 105 NHSNARQCEVCKHPFSFSPVYAEN 128
N +C VC+H F Y ++
Sbjct: 119 VDYNKMKCPVCRHRFLPKEKYTQS 142
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.328 0.141 0.441
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,836,833
Number of Sequences: 26719
Number of extensions: 882635
Number of successful extensions: 5866
Number of sequences better than 10.0: 113
Number of HSP's better than 10.0 without gapping: 39
Number of HSP's successfully gapped in prelim test: 76
Number of HSP's that attempted gapping in prelim test: 5649
Number of HSP's gapped (non-prelim): 219
length of query: 982
length of database: 11,318,596
effective HSP length: 109
effective length of query: 873
effective length of database: 8,406,225
effective search space: 7338634425
effective search space used: 7338634425
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 65 (29.6 bits)
Medicago: description of AC136449.11


