

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136286.2 - phase: 0
(251 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
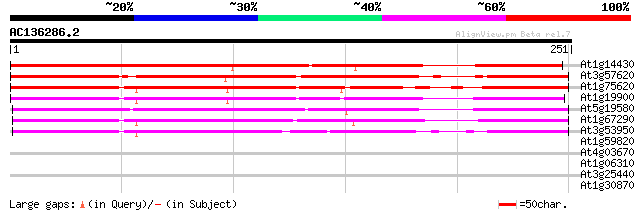
Score E
Sequences producing significant alignments: (bits) Value
At1g14430 hypothetical protein 236 9e-63
At3g57620 putative protein 231 2e-61
At1g75620 unknown protein 190 5e-49
At1g19900 unknown protein 186 8e-48
At5g19580 putative protein 184 3e-47
At1g67290 unknown protein 184 4e-47
At3g53950 unknown protein 181 3e-46
At1g59820 hypothetical protein 29 2.1
At4g03670 hypothetical protein 28 4.7
At1g06310 hypothetical protein 28 6.1
At3g25440 unknown protein 27 7.9
At1g30870 unknown protein 27 7.9
>At1g14430 hypothetical protein
Length = 849
Score = 236 bits (602), Expect = 9e-63
Identities = 128/249 (51%), Positives = 162/249 (64%), Gaps = 26/249 (10%)
Query: 1 MEVMPVPRVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPFMKFEL 60
ME MP PRVM DMLLLP G+V+I+NGAANGTAGWE+A N VL P+LY P + +FE+
Sbjct: 337 MEQMPSPRVMSDMLLLPNGDVLIINGAANGTAGWEDATNAVLNPILYLPEEPDQTRRFEI 396
Query: 61 LAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDFQAK-YPTELSLDAYYPDYLRPELDT 119
L P PRMYHS+++LL DGR+LVGGSNPHR Y+F A+ YPTELSL+AY P YL P+
Sbjct: 397 LTPTRIPRMYHSASLLLSDGRVLVGGSNPHRNYNFTARPYPTELSLEAYLPRYLDPQYAR 456
Query: 120 LRPVIVAVEVVNSTLSYESLFSVSFLLREVKDVN-RIRVSMVAPSFTTHSFAMNQRLLFL 178
+RP I+ VE+ + L Y F+V+F + + + V +VAPSF+THS AMNQRLL L
Sbjct: 457 VRPTIITVELAGNML-YGQAFAVTFAIPAFGMFDGGVSVRLVAPSFSTHSTAMNQRLLVL 515
Query: 179 EVTALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYYMLFVIH 238
V + + ++ YKA V GP + VAPPGYYM+FV+H
Sbjct: 516 RVRRVSQ-----------------------LSVFAYKADVDGPTNSYVAPPGYYMMFVVH 552
Query: 239 VGIPSVATW 247
GIPSVA W
Sbjct: 553 RGIPSVAVW 561
Score = 30.0 bits (66), Expect = 1.2
Identities = 14/35 (40%), Positives = 21/35 (60%)
Query: 68 RMYHSSAVLLPDGRILVGGSNPHRLYDFQAKYPTE 102
R ++S+ +LPDGRI++ G Y+F K P E
Sbjct: 169 RRWYSTNQILPDGRIIIVGGRRAFNYEFYPKDPGE 203
>At3g57620 putative protein
Length = 547
Score = 231 bits (590), Expect = 2e-61
Identities = 124/251 (49%), Positives = 168/251 (66%), Gaps = 29/251 (11%)
Query: 1 MEVMPVPRVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPFMKFEL 60
ME MP+PRVM DMLLLPTG+VII+NGA GTAGWE A +P++ PV+Y+P D+ F +
Sbjct: 324 METMPLPRVMGDMLLLPTGDVIIVNGAGAGTAGWEKARDPIIQPVIYQP-FDH---LFTV 379
Query: 61 LAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDF-QAKYPTELSLDAYYPDYLRPELDT 119
++ S PRMYHSSA+LLPDGR+LVGGSNPH Y+F +YPT+LSL+AY P YL D
Sbjct: 380 MSTPSRPRMYHSSAILLPDGRVLVGGSNPHVYYNFTNVEYPTDLSLEAYSPPYLFFTSDP 439
Query: 120 LRPVIVAVEVVNSTLSYESLFSVSFLLREVKDVNRIRVSMVAPSFTTHSFAMNQRLLFLE 179
+RP I+ + LSY+ LF+V F + + V+ + V +VAPSFTTHSFAMNQR++ L+
Sbjct: 440 IRPKILLTS--DKVLSYKRLFNVDFSIAQFLTVDLLSVRIVAPSFTTHSFAMNQRMVILK 497
Query: 180 VTALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYYMLFVIHV 239
+ ++ +DQ + NS Y+ + GP + +APPGYYM+F++H
Sbjct: 498 LLSV------TRDQ---------------LTNS-YRVSALGPSTAEIAPPGYYMIFLVHA 535
Query: 240 GIPSVATWVHV 250
GIPS A WV +
Sbjct: 536 GIPSSAAWVQI 546
>At1g75620 unknown protein
Length = 547
Score = 190 bits (483), Expect = 5e-49
Identities = 114/253 (45%), Positives = 156/253 (61%), Gaps = 30/253 (11%)
Query: 1 MEVMPVPRVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPF-MKFE 59
+E MP RVM DM+LLP GNV+++NG ++GTA WE PVL+P LY P D P +FE
Sbjct: 321 VEKMPRARVMGDMMLLPDGNVLLINGGSSGTAAWELGREPVLHPDLYHP--DKPVGSRFE 378
Query: 60 LLAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDFQ-AKYPTELSLDAYYPDYLRPELD 118
+ P++ PRMYHS A LL DGRILVGGSNPH Y+F +PTEL L+A+ P YL +
Sbjct: 379 VQNPSTIPRMYHSIATLLRDGRILVGGSNPHAFYNFTGVLFPTELRLEAFSPSYLDTKYS 438
Query: 119 TLRPVIVAVEVVNSTLSYESLFSVSFLLR-EVKDVNRIRVSMVAPSFTTHSFAMNQRLLF 177
+LRP IV +T++Y + + F++ VK + ++V+M+ PSFTTHSF+M+QRLL
Sbjct: 439 SLRPSIVDPR-PQTTVNYGRVLRLRFIVSGRVK--SPVKVTMLFPSFTTHSFSMHQRLL- 494
Query: 178 LEVTALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYYMLFVI 237
L+ V++ F G S +Y+ VR P S +APPGYYM+FV+
Sbjct: 495 ----VLDHVIS---------FKLGIS--------KIYEVRVRTPSSAILAPPGYYMVFVV 533
Query: 238 HVGIPSVATWVHV 250
+ IPS WV +
Sbjct: 534 NQDIPSEGLWVRL 546
>At1g19900 unknown protein
Length = 548
Score = 186 bits (473), Expect = 8e-48
Identities = 109/250 (43%), Positives = 147/250 (58%), Gaps = 28/250 (11%)
Query: 1 MEVMPVPRVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPF-MKFE 59
+E MP RVM DM+ LP G+V+++NG + GTA WE PVL P LY P +NP +FE
Sbjct: 322 VEKMPHARVMGDMIPLPNGDVLLINGGSFGTAAWELGRTPVLAPDLYHP--ENPVGSRFE 379
Query: 60 LLAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDFQ-AKYPTELSLDAYYPDYLRPELD 118
L P + PRMYHS+A+LL DGR+LVGGSNPH Y++ +PTELSL+A+ P YL+ E
Sbjct: 380 SLRPTTIPRMYHSAAILLRDGRVLVGGSNPHAFYNYTGVLFPTELSLEAFSPVYLQREFS 439
Query: 119 TLRPVIVAVEVVNSTLSYESLFSVSFLLREVKDVNRIRVSMVAPSFTTHSFAMNQRLLFL 178
LRP I++ E S + Y + + F + + +V+MV P+FTTHSFAMNQR+L L
Sbjct: 440 NLRPKIISPE-PQSMIKYGTNLKLKFSVTG-EVTTPAKVTMVFPTFTTHSFAMNQRVLVL 497
Query: 179 EVTALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYYMLFVIH 238
+ K + +Y+ VR P S N+A PGYYM+FV++
Sbjct: 498 DNVKFTR----------------------KGKSPMYEVQVRTPRSANIAWPGYYMIFVVN 535
Query: 239 VGIPSVATWV 248
IPS WV
Sbjct: 536 QDIPSEGVWV 545
>At5g19580 putative protein
Length = 594
Score = 184 bits (468), Expect = 3e-47
Identities = 96/250 (38%), Positives = 141/250 (56%), Gaps = 27/250 (10%)
Query: 2 EVMPVPRVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPFMKFELL 61
E+MP PR+M D ++LP G+++++NGA G +GW +P P+LYKP +F L
Sbjct: 370 EMMPTPRIMSDTVILPNGDILLVNGAKRGCSGWGYGKDPAFAPLLYKPHAARG-KRFRQL 428
Query: 62 APASTPRMYHSSAVLLPDGRILVGGSNPHRLYDFQAKYPTELSLDAYYPDYLRPELDTLR 121
P + PRMYHSSA++LPDG++LVGGSN + Y + ++PTEL ++ + P YL P L +R
Sbjct: 429 KPTTIPRMYHSSAIILPDGKVLVGGSNTNDGYKYNVEFPTELRVEKFSPPYLDPALANIR 488
Query: 122 PVIVAVEVVNSTLSYESLFSVSFLLREV-KDVNRIRVSMVAPSFTTHSFAMNQRLLFLEV 180
P IV + Y F+V L+E ++V+M+AP+FTTHS +MN R+L L V
Sbjct: 489 PKIVTTGTPKQ-VKYGQFFNVKVDLKEKGATKGNLKVTMLAPAFTTHSISMNMRMLILGV 547
Query: 181 TALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYYMLFVIHVG 240
+ K A + Y PP+ N+APPGYY++F I+ G
Sbjct: 548 NNV------------------------KPAGAGYDIQAVAPPNGNIAPPGYYLIFAIYKG 583
Query: 241 IPSVATWVHV 250
+PS W+ V
Sbjct: 584 VPSTGEWIQV 593
Score = 27.3 bits (59), Expect = 7.9
Identities = 19/74 (25%), Positives = 31/74 (41%), Gaps = 6/74 (8%)
Query: 22 IILNGAANGTAGWENAANPVLYPVLYKPGLDNPFMKFELLAPASTPRMYHSSAVLLPDGR 81
+ +NG T G+ AN Y +N K+E A + ++S+ LPDG+
Sbjct: 165 LTVNGTLVSTGGYGGGANTARY----LSSCEN--CKWEEYPQALAAKRWYSTQATLPDGK 218
Query: 82 ILVGGSNPHRLYDF 95
V G Y++
Sbjct: 219 FFVIGGRDALNYEY 232
>At1g67290 unknown protein
Length = 615
Score = 184 bits (467), Expect = 4e-47
Identities = 98/251 (39%), Positives = 141/251 (56%), Gaps = 28/251 (11%)
Query: 2 EVMPVPRVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPF-MKFEL 60
E MP RVM D ++LP G ++I+NGA G++GW A P P+LYKP + P +F+
Sbjct: 390 ETMPTSRVMSDTVILPNGEILIINGAKRGSSGWHLAKEPNFAPLLYKP--NKPLGQRFKE 447
Query: 61 LAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDFQAKYPTELSLDAYYPDYLRPELDTL 120
LAP++ PR+YHS A+ LPDG++LVGGSN + Y F +YPTEL ++ + P YL P L +
Sbjct: 448 LAPSTIPRVYHSIAIALPDGKVLVGGSNTNNGYQFNVEYPTELRIEKFSPPYLDPALANM 507
Query: 121 RPVIVAVEVVNSTLSYESLFSVSFLLREVKDV-NRIRVSMVAPSFTTHSFAMNQRLLFLE 179
RP IV + Y +F V L++ + V+M+APSFTTHS +MN RLL L
Sbjct: 508 RPRIVNT-ATPKQIKYGQMFDVKIELKQQNVAKENVMVTMLAPSFTTHSVSMNMRLLMLG 566
Query: 180 VTALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYYMLFVIHV 239
+ ++ V ++ PPS +APPGYY+LF ++
Sbjct: 567 INNVKNV-----------------------GGDNHQIQAVAPPSGKLAPPGYYLLFAVYN 603
Query: 240 GIPSVATWVHV 250
G+PSV W+ +
Sbjct: 604 GVPSVGEWIQI 614
>At3g53950 unknown protein
Length = 545
Score = 181 bits (460), Expect = 3e-46
Identities = 101/250 (40%), Positives = 148/250 (58%), Gaps = 30/250 (12%)
Query: 2 EVMPVPRVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPF-MKFEL 60
E MP R+M DM+ LPTG ++I+NGA G+ G+E ++P LYP+LY+P D P ++F
Sbjct: 323 EEMPFGRIMGDMVNLPTGEILIINGAQAGSQGFEMGSDPCLYPLLYRP--DQPIGLRFMT 380
Query: 61 LAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDFQAKYPTELSLDAYYPDYLRPELDTL 120
L P + PRMYHS+A LLPDGRIL+ GSNPH Y F A++PTEL ++A+ P+YL P+ L
Sbjct: 381 LNPGTVPRMYHSTANLLPDGRILLAGSNPHYFYKFNAEFPTELRIEAFSPEYLSPDRANL 440
Query: 121 RPVIVAVEVVNSTLSYESLFSVSFLLREVKDVNRIRVSMVAPSFTTHSFAMNQRLLFLEV 180
RP ++ + + Y +F V F+ + V I+++ + F THSF+ QRL+ L V
Sbjct: 441 RP---EIQEIPQIIRYGEVFDV-FVTVPLPVVGIIQMNWGSAPFATHSFSQGQRLVKLTV 496
Query: 181 TALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYYMLFVIHVG 240
S+ D G G+ Y+ PP+ V+PPGYYM F ++ G
Sbjct: 497 AP------SVPD------------GVGR-----YRIQCTAPPNGAVSPPGYYMAFAVNQG 533
Query: 241 IPSVATWVHV 250
+PS+A W+ +
Sbjct: 534 VPSIARWIRI 543
>At1g59820 hypothetical protein
Length = 1213
Score = 29.3 bits (64), Expect = 2.1
Identities = 12/30 (40%), Positives = 19/30 (63%)
Query: 43 YPVLYKPGLDNPFMKFELLAPASTPRMYHS 72
YP LY+ G+ N F K+ ++A +T +Y S
Sbjct: 967 YPELYREGIRNSFFKWRVVAVWATSAVYQS 996
>At4g03670 hypothetical protein
Length = 171
Score = 28.1 bits (61), Expect = 4.7
Identities = 14/38 (36%), Positives = 23/38 (59%), Gaps = 4/38 (10%)
Query: 149 VKDVNRIRVSMVAPSFTTHSFAMNQRLLFLEVTALEEV 186
++D++ I+ S+ +P FT H F ++ FLE T L V
Sbjct: 36 MEDIHVIKASVGSPPFTDHKF----KIKFLEYTKLSHV 69
>At1g06310 hypothetical protein
Length = 667
Score = 27.7 bits (60), Expect = 6.1
Identities = 34/135 (25%), Positives = 53/135 (39%), Gaps = 16/135 (11%)
Query: 11 PDMLLLPTGNVIILNGAANGTAGWENAANP----------VLYPVLYKPGLDNPFMKF-E 59
P++ + G+ I LNG NG ++N P VL Y + +P +F
Sbjct: 268 PNVRIADCGHKIGLNGVDNGRIWFDNLRIPRENLLNSVADVLADGKYVSSIKDPDQRFGA 327
Query: 60 LLAPASTPRMYHSSAVLLPDG---RILVGGSNPHRLYDFQAKYPTELSLDAYYPDYLRPE 116
LAP ++ R+ +S+ + + + S R + A P L LD YP + R
Sbjct: 328 FLAPLTSGRVTIASSAIYSAKLGLAVAIRYSLSRRAFSVAANGPEVLLLD--YPSHQRRL 385
Query: 117 LDTLRPVIVAVEVVN 131
L L VN
Sbjct: 386 LPLLAKTYAMSFAVN 400
>At3g25440 unknown protein
Length = 380
Score = 27.3 bits (59), Expect = 7.9
Identities = 13/36 (36%), Positives = 20/36 (55%)
Query: 170 AMNQRLLFLEVTALEEVVNSMQDQNFGEFGFGSSLG 205
A +R LFLE +N+ +D++FG+ G S G
Sbjct: 315 AKTERPLFLEEFEKFPAINNREDEDFGDLGKAKSEG 350
>At1g30870 unknown protein
Length = 349
Score = 27.3 bits (59), Expect = 7.9
Identities = 12/29 (41%), Positives = 19/29 (65%)
Query: 86 GSNPHRLYDFQAKYPTELSLDAYYPDYLR 114
G+ RLY++ A ++ S+DA Y DYL+
Sbjct: 221 GTIQSRLYNYNATSGSDPSIDAKYADYLQ 249
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.138 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,805,138
Number of Sequences: 26719
Number of extensions: 248075
Number of successful extensions: 549
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 506
Number of HSP's gapped (non-prelim): 21
length of query: 251
length of database: 11,318,596
effective HSP length: 97
effective length of query: 154
effective length of database: 8,726,853
effective search space: 1343935362
effective search space used: 1343935362
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC136286.2


