

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136142.6 + phase: 0
(147 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
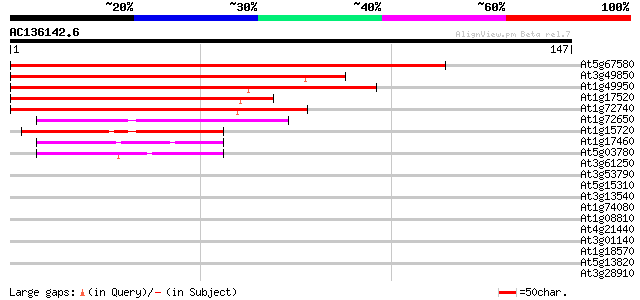
Sequences producing significant alignments: (bits) Value
At5g67580 telomere repeat binding factor 2 (TRB2) 152 8e-38
At3g49850 MYB -like protein 132 5e-32
At1g49950 telomere repeat binding factor 1 (TRB1)/DNA-binding pr... 130 3e-31
At1g17520 myb-related DNA-binding like protein 88 1e-18
At1g72740 DNA-binding like protein 86 9e-18
At1g72650 unknown protein 48 2e-06
At1g15720 unknown protein 47 4e-06
At1g17460 unknown protein (At1g17460) 44 3e-05
At5g03780 myb -like protein 39 8e-04
At3g61250 putative transcription factor (MYB17) 38 0.002
At3g53790 putative protein 38 0.002
At5g15310 myb-related protein - like 35 0.014
At3g13540 myb-related protein 5 35 0.014
At1g74080 putative transcription factor 35 0.014
At1g08810 transcription factor like protein 35 0.014
At4g21440 myb-related protein M4 35 0.019
At3g01140 putative Myb-related transcription factor 35 0.019
At1g18570 unknown protein 35 0.019
At5g13820 telomeric DNA-binding protein 1 (TBP1) 34 0.025
At3g28910 MYB family transcription factor (hsr1), putative 34 0.025
>At5g67580 telomere repeat binding factor 2 (TRB2)
Length = 299
Score = 152 bits (383), Expect = 8e-38
Identities = 76/114 (66%), Positives = 86/114 (74%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTA 60
MGAPKQKWT EEEAALKAGV+KHG GKWRTIL D EFS IL++RSNVDLKDKWRNI+VTA
Sbjct: 1 MGAPKQKWTPEEEAALKAGVLKHGTGKWRTILSDTEFSLILKSRSNVDLKDKWRNISVTA 60
Query: 61 IWGSRQKAKLALKNSPPAPKTDNNQLALGKVVQREDFLDIKPLTISGGTFQSPK 114
+WGSR+KAKLALK +PP K D+N AL V D KP + G SP+
Sbjct: 61 LWGSRKKAKLALKRTPPGTKQDDNNTALTIVALTNDDERAKPTSPGGSGGGSPR 114
>At3g49850 MYB -like protein
Length = 295
Score = 132 bits (333), Expect = 5e-32
Identities = 63/89 (70%), Positives = 75/89 (83%), Gaps = 1/89 (1%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTA 60
MGAPK KWT EEE ALKAGV+KHG GKWRTIL DP +S+IL++RSNVDLKDKWRNI+VTA
Sbjct: 1 MGAPKLKWTPEEETALKAGVLKHGTGKWRTILSDPVYSTILKSRSNVDLKDKWRNISVTA 60
Query: 61 IWGSRQKAKLALKNSP-PAPKTDNNQLAL 88
+WGSR+KAKLALK +P + D+N A+
Sbjct: 61 LWGSRKKAKLALKRTPLSGSRQDDNATAI 89
>At1g49950 telomere repeat binding factor 1 (TRB1)/DNA-binding
protein PcMYB1, putative
Length = 300
Score = 130 bits (326), Expect = 3e-31
Identities = 62/97 (63%), Positives = 77/97 (78%), Gaps = 1/97 (1%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTA 60
MGAPKQKWT EEE+ALK+GV+KHG GKWRTIL DPEFS +L RSNVDLKDKWRN++V A
Sbjct: 1 MGAPKQKWTQEEESALKSGVIKHGPGKWRTILKDPEFSGVLYLRSNVDLKDKWRNMSVMA 60
Query: 61 I-WGSRQKAKLALKNSPPAPKTDNNQLALGKVVQRED 96
WGSR+K++LA+K + PK + N LAL +Q ++
Sbjct: 61 NGWGSREKSRLAVKRTFSLPKQEENSLALTNSLQSDE 97
>At1g17520 myb-related DNA-binding like protein
Length = 296
Score = 88.2 bits (217), Expect = 1e-18
Identities = 43/70 (61%), Positives = 51/70 (72%), Gaps = 1/70 (1%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVT- 59
MG K KWTAEEE AL AGV KHG GKW+ IL DPE + L +RSN+DLKDKWRN++V
Sbjct: 1 MGNQKLKWTAEEEEALLAGVRKHGPGKWKNILRDPELAEQLSSRSNIDLKDKWRNLSVAP 60
Query: 60 AIWGSRQKAK 69
I GS+ K +
Sbjct: 61 GIQGSKDKIR 70
>At1g72740 DNA-binding like protein
Length = 291
Score = 85.5 bits (210), Expect = 9e-18
Identities = 43/82 (52%), Positives = 54/82 (65%), Gaps = 4/82 (4%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINV-- 58
MG K KWTAEEE AL AG+ KHG GKW+ IL DPEF+ L RSN+DLKDKWRN++V
Sbjct: 1 MGNQKLKWTAEEEEALLAGIRKHGPGKWKNILRDPEFADQLIHRSNIDLKDKWRNLSVPP 60
Query: 59 --TAIWGSRQKAKLALKNSPPA 78
++ + AK+ + PA
Sbjct: 61 GTQSLTNKARPAKVKEEGDTPA 82
>At1g72650 unknown protein
Length = 624
Score = 48.1 bits (113), Expect = 2e-06
Identities = 29/66 (43%), Positives = 36/66 (53%), Gaps = 2/66 (3%)
Query: 8 WTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTAIWGSRQK 67
WT E A L GV K+GAGKW I S R++VDLKDKWRN+ T+ S
Sbjct: 537 WTLSEIAKLVEGVSKYGAGKWSEI--KKHLFSSHSYRTSVDLKDKWRNLLKTSFAQSPSN 594
Query: 68 AKLALK 73
+ +LK
Sbjct: 595 SVGSLK 600
>At1g15720 unknown protein
Length = 390
Score = 47.0 bits (110), Expect = 4e-06
Identities = 24/53 (45%), Positives = 33/53 (61%), Gaps = 3/53 (5%)
Query: 4 PKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNI 56
P + WT+EE AAL+ GV ++G W+ I + + RS VDLKDKWRN+
Sbjct: 337 PMKFWTSEEVAALREGVKEYGKS-WKDI--KNSYPVVFADRSEVDLKDKWRNL 386
>At1g17460 unknown protein (At1g17460)
Length = 604
Score = 43.9 bits (102), Expect = 3e-05
Identities = 24/49 (48%), Positives = 28/49 (56%), Gaps = 2/49 (4%)
Query: 8 WTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNI 56
WT E L GV K+G GKW T + FS R+ VDLKDKWRN+
Sbjct: 499 WTISEVEKLVEGVSKYGVGKW-TEIKKLSFSPYTH-RTTVDLKDKWRNL 545
>At5g03780 myb -like protein
Length = 420
Score = 39.3 bits (90), Expect = 8e-04
Identities = 25/54 (46%), Positives = 28/54 (51%), Gaps = 6/54 (11%)
Query: 8 WTAEEEAALKAGVVKHGAGK-----WRTILMDPEFSSILRTRSNVDLKDKWRNI 56
WT EEE LK GV K A WR IL E TR+ DLKDKWR++
Sbjct: 350 WTYEEEEMLKVGVEKFAAEANKNMPWRKILEMGE-KVFHETRTPADLKDKWRSM 402
>At3g61250 putative transcription factor (MYB17)
Length = 299
Score = 38.1 bits (87), Expect = 0.002
Identities = 20/55 (36%), Positives = 30/55 (54%), Gaps = 5/55 (9%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRN 55
+G K WT EE+ L A + K+G G WRT+ P+ + +LR + L +W N
Sbjct: 10 IGLKKGPWTPEEDEVLVAHIKKNGHGSWRTL---PKLAGLLRCGKSCRL--RWTN 59
>At3g53790 putative protein
Length = 400
Score = 37.7 bits (86), Expect = 0.002
Identities = 22/63 (34%), Positives = 35/63 (54%), Gaps = 2/63 (3%)
Query: 5 KQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTAIWGS 64
++ +T E AL V + G G+WR + F+ + R+ VDLKDKW+ + TA +
Sbjct: 315 RRPFTVSEVEALVQAVERLGTGRWRDV-KSHAFNHV-NHRTYVDLKDKWKTLVHTAKISA 372
Query: 65 RQK 67
RQ+
Sbjct: 373 RQR 375
>At5g15310 myb-related protein - like
Length = 326
Score = 35.0 bits (79), Expect = 0.014
Identities = 19/55 (34%), Positives = 29/55 (52%), Gaps = 5/55 (9%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRN 55
+G K WT EE+ L A + +HG G WR++ PE + + R + L +W N
Sbjct: 10 LGLKKGPWTPEEDQKLLAYIEEHGHGSWRSL---PEKAGLHRCGKSCRL--RWTN 59
>At3g13540 myb-related protein 5
Length = 249
Score = 35.0 bits (79), Expect = 0.014
Identities = 18/55 (32%), Positives = 30/55 (53%), Gaps = 5/55 (9%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRN 55
MG + WT EE+ L + + K G G+WR++ P+ + +LR + L +W N
Sbjct: 21 MGMKRGPWTVEEDEILVSFIKKEGEGRWRSL---PKRAGLLRCGKSCRL--RWMN 70
>At1g74080 putative transcription factor
Length = 333
Score = 35.0 bits (79), Expect = 0.014
Identities = 20/54 (37%), Positives = 28/54 (51%), Gaps = 5/54 (9%)
Query: 2 GAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRN 55
G K WT EE+ L A V +HG G WRT+ P+ + + R + L +W N
Sbjct: 11 GLKKGAWTQEEDQKLIAYVQRHGEGGWRTL---PDKAGLKRCGKSCRL--RWAN 59
>At1g08810 transcription factor like protein
Length = 280
Score = 35.0 bits (79), Expect = 0.014
Identities = 18/55 (32%), Positives = 29/55 (52%), Gaps = 5/55 (9%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRN 55
+G K WT EE+ L + + +HG G WR++ P + +LR + L +W N
Sbjct: 10 IGIKKGPWTPEEDIILVSYIQEHGPGNWRSV---PTNTGLLRCSKSCRL--RWTN 59
>At4g21440 myb-related protein M4
Length = 350
Score = 34.7 bits (78), Expect = 0.019
Identities = 19/54 (35%), Positives = 28/54 (51%), Gaps = 5/54 (9%)
Query: 2 GAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRN 55
G K WT+EE+ L + KHG G WRT+ P+ + + R + L +W N
Sbjct: 11 GLKKGPWTSEEDQKLVDYIQKHGYGNWRTL---PKNAGLQRCGKSCRL--RWTN 59
>At3g01140 putative Myb-related transcription factor
Length = 345
Score = 34.7 bits (78), Expect = 0.019
Identities = 19/54 (35%), Positives = 28/54 (51%), Gaps = 5/54 (9%)
Query: 2 GAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRN 55
G K WT EE+ L A + +HG G WR++ PE + + R + L +W N
Sbjct: 11 GLKKGPWTPEEDQKLLAYIEEHGHGSWRSL---PEKAGLQRCGKSCRL--RWTN 59
>At1g18570 unknown protein
Length = 352
Score = 34.7 bits (78), Expect = 0.019
Identities = 19/55 (34%), Positives = 29/55 (52%), Gaps = 5/55 (9%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRN 55
+G K WT EE+ L + + +HG G WRT+ PE + + R + L +W N
Sbjct: 11 LGLKKGAWTPEEDQKLLSYLNRHGEGGWRTL---PEKAGLKRCGKSCRL--RWAN 60
>At5g13820 telomeric DNA-binding protein 1 (TBP1)
Length = 640
Score = 34.3 bits (77), Expect = 0.025
Identities = 20/63 (31%), Positives = 35/63 (54%), Gaps = 2/63 (3%)
Query: 5 KQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTAIWGS 64
++ ++ E AL + V + G G+WR + + ++ RT VDLKDKW+ + TA
Sbjct: 535 RRPFSVTEVEALVSAVEEVGTGRWRDVKLRSFENASHRTY--VDLKDKWKTLVHTASISP 592
Query: 65 RQK 67
+Q+
Sbjct: 593 QQR 595
>At3g28910 MYB family transcription factor (hsr1), putative
Length = 323
Score = 34.3 bits (77), Expect = 0.025
Identities = 18/54 (33%), Positives = 26/54 (47%), Gaps = 5/54 (9%)
Query: 2 GAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRN 55
G K WT EE+ L + +HG G WR + P + +LR + L +W N
Sbjct: 11 GVKKGPWTPEEDIILVTYIQEHGPGNWRAV---PTNTGLLRCSKSCRL--RWTN 59
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.314 0.130 0.381
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,476,326
Number of Sequences: 26719
Number of extensions: 136468
Number of successful extensions: 461
Number of sequences better than 10.0: 143
Number of HSP's better than 10.0 without gapping: 114
Number of HSP's successfully gapped in prelim test: 29
Number of HSP's that attempted gapping in prelim test: 324
Number of HSP's gapped (non-prelim): 154
length of query: 147
length of database: 11,318,596
effective HSP length: 90
effective length of query: 57
effective length of database: 8,913,886
effective search space: 508091502
effective search space used: 508091502
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 55 (25.8 bits)
Medicago: description of AC136142.6


