

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135801.5 - phase: 0
(736 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
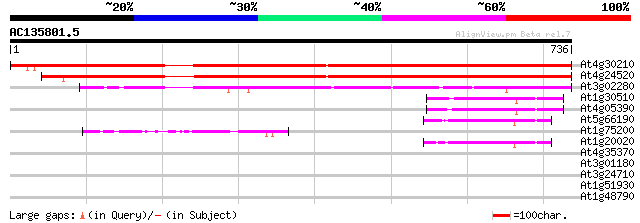
Score E
Sequences producing significant alignments: (bits) Value
At4g30210 NADPH-ferrihemoprotein reductase (ATR2) 1055 0.0
At4g24520 NADPH-ferrihemoprotein reductase ATR1 886 0.0
At3g02280 NADPH-ferrihemoprotein reductase like protein 253 3e-67
At1g30510 ferrodoxin NADP oxidoreductase - like protein 68 2e-11
At4g05390 ferredoxin-NADP+ reductase like protein 67 5e-11
At5g66190 ferredoxin-NADP+ reductase 63 5e-10
At1g75200 unknown protein 61 2e-09
At1g20020 ferredoxin--NADP reductase precursor, putative 57 3e-08
At4g35370 putative protein 35 0.20
At3g01180 putative glycogen synthase 35 0.20
At3g24710 hypothetical protein 31 2.2
At1g51930 hypothetical protein 30 4.8
At1g48790 unknown protein 30 6.3
>At4g30210 NADPH-ferrihemoprotein reductase (ATR2)
Length = 711
Score = 1055 bits (2729), Expect = 0.0
Identities = 520/747 (69%), Positives = 607/747 (80%), Gaps = 47/747 (6%)
Query: 1 MQDSSSMKFSPLDLMSALIKGK----IDPSNGTVP-------ASLILENREFVMILTTSI 49
M SSS S +DLM+A+IKG+ DP+N + +S+++ENR+F MI+TTSI
Sbjct: 1 MSSSSSSSTSMIDLMAAIIKGEPVIVSDPANASAYESVAAELSSMLIENRQFAMIVTTSI 60
Query: 50 AVLIGCVVVLIWRRSNSQKPKPIEVPKRVIEKLPELEIDDGTKKVTVFFGTQTGTAEGFA 109
AVLIGC+V+L+WRRS S K +E K ++ K E EIDDG KKVT+FFGTQTGTAEGFA
Sbjct: 61 AVLIGCIVMLVWRRSGSGNSKRVEPLKPLVIKPREEEIDDGRKKVTIFFGTQTGTAEGFA 120
Query: 110 KAIAEEAKARYEKAKFKVVDMDDYAADDDEYEEKLKKETMALFFLATYGDGEPTDNAARF 169
KA+ EEAKARYEK +FK+VD+DDYAADDDEYEEKLKKE +A FFLATYGDGEPTDNAARF
Sbjct: 121 KALGEEAKARYEKTRFKIVDLDDYAADDDEYEEKLKKEDVAFFFLATYGDGEPTDNAARF 180
Query: 170 YKWFEEFEGEEDSFKNLQYGVFGLGNRQYEHFNKVCTQALMHSCGFLSFDYSDGVASFRA 229
YKWF E + KNL+YGVFGLGNRQYEHFNKV
Sbjct: 181 YKWFTEGNDRGEWLKNLKYGVFGLGNRQYEHFNKV------------------------- 215
Query: 230 FSDIYHSTQVAKIVDDKLLEKGGNRLVPVGLGDDDQCIEDDFTAWKEELWPALDQLLRDE 289
AK+VDD L+E+G RLV VGLGDDDQCIEDDFTAW+E LWP LD +LR+E
Sbjct: 216 ----------AKVVDDILVEQGAQRLVQVGLGDDDQCIEDDFTAWREALWPELDTILREE 265
Query: 290 DDATVATPYTASVLEYRVVIRDQLDATVDEKKQLNGNGHAVVDAHHPVRANVAVRKELHT 349
D VATPYTA+VLEYRV I D DA ++ NGNG+ V DA HP +ANVAV++ELHT
Sbjct: 266 GDTAVATPYTAAVLEYRVSIHDSEDAKFNDINMANGNGYTVFDAQHPYKANVAVKRELHT 325
Query: 350 PASDRSCTHLEFDISGTGVVYETGDHVGVYCENLSDTVEEAERILGLSPDTYFSVHTDDE 409
P SDRSC HLEFDI+G+G+ YETGDHVGV C+NLS+TV+EA R+L +SPDTYFS+H + E
Sbjct: 326 PESDRSCIHLEFDIAGSGLTYETGDHVGVLCDNLSETVDEALRLLDMSPDTYFSLHAEKE 385
Query: 410 DGKPLGGSSLPPPFPPCTLRTALAKYADVLSSPKKSALLALAAHASDPSEADRLRHLASP 469
DG P+ SSLPPPFPPC LRTAL +YA +LSSPKKSAL+ALAAHASDP+EA+RL+HLASP
Sbjct: 386 DGTPIS-SSLPPPFPPCNLRTALTRYACLLSSPKKSALVALAAHASDPTEAERLKHLASP 444
Query: 470 AGKDEYAEWVIASQRSLLEVMAEFSSAKPPIGVFFASVAPRLQPRYYSISSSPRVAPSRI 529
AGKDEY++WV+ SQRSLLEVMAEF SAKPP+GVFFA VAPRLQPR+YSISSSP++A +RI
Sbjct: 445 AGKDEYSKWVVESQRSLLEVMAEFPSAKPPLGVFFAGVAPRLQPRFYSISSSPKIAETRI 504
Query: 530 HVTCALVHDKMPTGRIHQGVCSTWMKNSAPLEKSQDCSWAPIFVRQSNFRLPADNKVPII 589
HVTCALV++KMPTGRIH+GVCSTWMKN+ P EKS++CS APIFVRQSNF+LP+D+KVPII
Sbjct: 505 HVTCALVYEKMPTGRIHKGVCSTWMKNAVPYEKSENCSSAPIFVRQSNFKLPSDSKVPII 564
Query: 590 MIGPGTGLAPFRGFLQERLALKEEGAELGPSVLFFGCRNRQVDYIYEDELNHFVHGGALS 649
MIGPGTGLAPFRGFLQERLAL E G ELGPSVLFFGCRNR++D+IYE+EL FV GAL+
Sbjct: 565 MIGPGTGLAPFRGFLQERLALVESGVELGPSVLFFGCRNRRMDFIYEEELQRFVESGALA 624
Query: 650 ELIVAFSREGPTKEYVQHKMIEKASDIWNMISQGAYIYVCGDAKGMAKDVHRTLHTILQE 709
EL VAFSREGPTKEYVQHKM++KASDIWNMISQGAY+YVCGDAKGMA+DVHR+LHTI QE
Sbjct: 625 ELSVAFSREGPTKEYVQHKMMDKASDIWNMISQGAYLYVCGDAKGMARDVHRSLHTIAQE 684
Query: 710 QGSLDNSKTESMVKNLQMTGRYLRDVW 736
QGS+D++K E VKNLQ +GRYLRDVW
Sbjct: 685 QGSMDSTKAEGFVKNLQTSGRYLRDVW 711
>At4g24520 NADPH-ferrihemoprotein reductase ATR1
Length = 692
Score = 886 bits (2290), Expect = 0.0
Identities = 435/701 (62%), Positives = 541/701 (77%), Gaps = 43/701 (6%)
Query: 42 VMILTTSIAVLIGCVVVLIWRRSNSQKP---KPIEVPKRVIEKLPE--LEIDDGTKKVTV 96
++I TTS+A++ G VVL+W+++ + + KP+ +PK ++ K + L++ G +V++
Sbjct: 29 LVIATTSLALVAG-FVVLLWKKTTADRSGELKPLMIPKSLMAKDEDDDLDLGSGKTRVSI 87
Query: 97 FFGTQTGTAEGFAKAIAEEAKARYEKAKFKVVDMDDYAADDDEYEEKLKKETMALFFLAT 156
FFGTQTGTAEGFAKA++EE KARYEKA KV+D+DDYAADDD+YEEKLKKET+A F +AT
Sbjct: 88 FFGTQTGTAEGFAKALSEEIKARYEKAAVKVIDLDDYAADDDQYEEKLKKETLAFFCVAT 147
Query: 157 YGDGEPTDNAARFYKWFEEFEGEEDSFKNLQYGVFGLGNRQYEHFNKVCTQALMHSCGFL 216
YGDGEPTDNAARFYKWF E + + L YGVF LGNRQYEHFNK+
Sbjct: 148 YGDGEPTDNAARFYKWFTEENERDIKLQQLAYGVFALGNRQYEHFNKI------------ 195
Query: 217 SFDYSDGVASFRAFSDIYHSTQVAKIVDDKLLEKGGNRLVPVGLGDDDQCIEDDFTAWKE 276
G+ ++D++L +KG RL+ VGLGDDDQ IEDDF AWKE
Sbjct: 196 ------GI-----------------VLDEELCKKGAKRLIEVGLGDDDQSIEDDFNAWKE 232
Query: 277 ELWPALDQLLRDEDDATVATPYTASVLEYRVVIRDQLDATVDEKKQLNGNGHAVVDAHHP 336
LW LD+LL+DEDD +VATPYTA + EYRVV D T + NG+ +D HHP
Sbjct: 233 SLWSELDKLLKDEDDKSVATPYTAVIPEYRVVTHDPRFTTQKSMESNVANGNTTIDIHHP 292
Query: 337 VRANVAVRKELHTPASDRSCTHLEFDISGTGVVYETGDHVGVYCENLSDTVEEAERILGL 396
R +VAV+KELHT SDRSC HLEFDIS TG+ YETGDHVGVY EN + VEEA ++LG
Sbjct: 293 CRVDVAVQKELHTHESDRSCIHLEFDISRTGITYETGDHVGVYAENHVEIVEEAGKLLGH 352
Query: 397 SPDTYFSVHTDDEDGKPLGGSSLPPPFP-PCTLRTALAKYADVLSSPKKSALLALAAHAS 455
S D FS+H D EDG PL S++PPPFP PCTL T LA+YAD+L+ P+KSAL+ALAA+A+
Sbjct: 353 SLDLVFSIHADKEDGSPLE-SAVPPPFPGPCTLGTGLARYADLLNPPRKSALVALAAYAT 411
Query: 456 DPSEADRLRHLASPAGKDEYAEWVIASQRSLLEVMAEFSSAKPPIGVFFASVAPRLQPRY 515
+PSEA++L+HL SP GKDEY++W++ASQRSLLEVMA F SAKPP+GVFFA++APRLQPRY
Sbjct: 412 EPSEAEKLKHLTSPDGKDEYSQWIVASQRSLLEVMAAFPSAKPPLGVFFAAIAPRLQPRY 471
Query: 516 YSISSSPRVAPSRIHVTCALVHDKMPTGRIHQGVCSTWMKNSAPLEKSQDCSWAPIFVRQ 575
YSISSSPR+APSR+HVT ALV+ PTGRIH+GVCSTWMKN+ P EKS +CS APIF+R
Sbjct: 472 YSISSSPRLAPSRVHVTSALVYGPTPTGRIHKGVCSTWMKNAVPAEKSHECSGAPIFIRA 531
Query: 576 SNFRLPADNKVPIIMIGPGTGLAPFRGFLQERLALKEEGAELGPSVLFFGCRNRQVDYIY 635
SNF+LP++ PI+M+GPGTGLAPFRGFLQER+ALKE+G ELG S+LFFGCRNRQ+D+IY
Sbjct: 532 SNFKLPSNPSTPIVMVGPGTGLAPFRGFLQERMALKEDGEELGSSLLFFGCRNRQMDFIY 591
Query: 636 EDELNHFVHGGALSELIVAFSREGPTKEYVQHKMIEKASDIWNMISQGAYIYVCGDAKGM 695
EDELN+FV G +SELI+AFSREG KEYVQHKM+EKA+ +W++I + Y+YVCGDAKGM
Sbjct: 592 EDELNNFVDQGVISELIMAFSREGAQKEYVQHKMMEKAAQVWDLIKEEGYLYVCGDAKGM 651
Query: 696 AKDVHRTLHTILQEQGSLDNSKTESMVKNLQMTGRYLRDVW 736
A+DVHRTLHTI+QEQ + +S+ E++VK LQ GRYLRDVW
Sbjct: 652 ARDVHRTLHTIVQEQEGVSSSEAEAIVKKLQTEGRYLRDVW 692
>At3g02280 NADPH-ferrihemoprotein reductase like protein
Length = 623
Score = 253 bits (646), Expect = 3e-67
Identities = 186/668 (27%), Positives = 309/668 (45%), Gaps = 74/668 (11%)
Query: 92 KKVTVFFGTQTGTAEGFAKAIAEEAKARYEKAKFKVVDMDDYAADDDEYEEKLKKETMAL 151
+K+ V + +QTG A A+ I EA+ R A VV D++ + E+ +
Sbjct: 6 RKLLVLYASQTGNALDAAERIGREAERRGLPAS--VVSTDEFDTSSLPHHEEA-----VV 58
Query: 152 FFLATYGDGEPTDNAARFYKWFEEFEGEEDSFKNLQYGVFGLGNRQYEHFNKVCTQALMH 211
F ++T G G+ D+ F+++ + + ++Y VFGLG+ Y+ +N V
Sbjct: 59 FVVSTTGQGDSPDSFKAFWRFLLQRNLGNYWLQQVRYAVFGLGDSGYQKYNFV------- 111
Query: 212 SCGFLSFDYSDGVASFRAFSDIYHSTQVAKIVDDKLLEKGGNRLVPVGLGDDDQC--IED 269
AK +D +L + G ++ GLGDD E
Sbjct: 112 ----------------------------AKKLDKRLSDLGATTIIEKGLGDDQHPSGYEG 143
Query: 270 DFTAWKEELWPALDQL----LRDEDDATVATPYTASVLEYRVVIRDQ--------LDATV 317
W LW L Q+ D + +YR++ Q D+ +
Sbjct: 144 TLDPWMLSLWRTLYQINPKYFPKGPDVKIPQDEVIDKPKYRILFHKQEKLEPKLLSDSDI 203
Query: 318 DEKKQLNGNGHAVVDAHHPVRANVAVRKELHTPA-SDRSCTHLEFDISGTGVVYETGDHV 376
++ + G D P R E+ T A S + H EF + + YE GD V
Sbjct: 204 IQRARGMSPGKLFKDKSKPDCFLKMTRNEVLTKAESTKDVRHFEFQFVSSTIEYEVGDVV 263
Query: 377 GVYCENLSDTVEEAERILGLSPDTYFSVHTDDEDGKPLGGSSLPPPFPPCTLRTALAKYA 436
+ S V+ GL P+++ +V + + + P L+T +
Sbjct: 264 ELLPSQNSSVVDAFIERCGLDPESFITVGPRETENSSFSEEMITQI--PIKLKTFVELTM 321
Query: 437 DVLS-SPKKSALLALAAHASDPSEADRLRHLASPAGKDEYAEWVIASQRSLLEVMAEFSS 495
DV S SP++ ++ +A+ E +RL++ ASP G+D+ + +RS+LEV+ +F S
Sbjct: 322 DVTSASPRRYFFEIMSFYATAEHEKERLQYFASPEGRDDLYNYNQKERRSILEVLEDFPS 381
Query: 496 AKPPIGVFFASVAPRLQPRYYSISSSPRVAPSRIHVTCALVHDKMPTGRIHQGVCSTWMK 555
+ P + + P L+PR +SISSSP P+ +H+T ++V P R +G+CS+W+
Sbjct: 382 VQIPFD-WLVQLVPPLKPRAFSISSSPLAHPAAVHLTVSIVSWITPYKRTRKGLCSSWLA 440
Query: 556 NSAPLEKSQDCSWAPIFVRQSNFRLPADNKVPIIMIGPGTGLAPFRGFLQERLALKEEGA 615
+ AP ++ P++ + + P+ + +P+I++GPGTG APFRGF+ ER A++ + +
Sbjct: 441 SLAPEQEVN----IPVWFHKGSLPAPSQS-LPLILVGPGTGCAPFRGFIAER-AVQAQSS 494
Query: 616 ELGPSVLFFGCRNRQVDYIYEDEL-NHFVHGGALSE-----LIVAFSREGPTKEYVQHKM 669
+ P + FFGCRN+ D++Y D +H GG LSE AFSR+ P K YVQHK+
Sbjct: 495 PVAPVMFFFGCRNKDTDFLYRDFWESHAREGGMLSEGKGGGFYTAFSRDQPKKVYVQHKI 554
Query: 670 IEKASDIWNMISQGAYIYVCGDAKGMAKDVHRTLHTILQEQ-GSLDNSKTESMVKNLQMT 728
E + +W+++ GA +YV G + M DV I+ E+ G +K L+ T
Sbjct: 555 REMSKRVWDLLCDGAAVYVAGSSTKMPCDVMSAFEDIVSEETGGGSKEVASRWLKALEKT 614
Query: 729 GRYLRDVW 736
GRY + W
Sbjct: 615 GRYNVEAW 622
>At1g30510 ferrodoxin NADP oxidoreductase - like protein
Length = 381
Score = 68.2 bits (165), Expect = 2e-11
Identities = 57/187 (30%), Positives = 89/187 (47%), Gaps = 16/187 (8%)
Query: 548 GVCSTWMKNSAPLEKSQDC--SWAPIFVRQSNFRLPADNKVPIIMIGPGTGLAPFRGFLQ 605
GVCS ++ +S P +K Q S + + +S D IMI GTG+AP+RG+L+
Sbjct: 197 GVCSNFLCDSKPGDKIQITGPSGKVMLLPES------DPNATHIMIATGTGVAPYRGYLR 250
Query: 606 ERLALKEEGAEL-GPSVLFFGCRNRQVDYIYEDELNHFVHGGALS-ELIVAFSREGPTKE 663
G + LF G N +Y++E ++ + A SRE K+
Sbjct: 251 RMFMENVPNKTFSGLAWLFLGVANTD-SLLYDEEFTKYLKDHPDNFRFDKALSREEKNKK 309
Query: 664 ----YVQHKMIEKASDIWNMISQGAYIYVCGDAKGMAKDVHRTLHTILQEQGSLDNSKTE 719
YVQ K+ E + +I+ ++ GA+IY CG KGM + TL + +E+G + K
Sbjct: 310 GGKMYVQDKIEEYSDEIFKLLDNGAHIYFCG-LKGMMPGIQDTLKRVAEERGESWDLKLS 368
Query: 720 SMVKNLQ 726
+ KN Q
Sbjct: 369 QLRKNKQ 375
>At4g05390 ferredoxin-NADP+ reductase like protein
Length = 378
Score = 66.6 bits (161), Expect = 5e-11
Identities = 57/186 (30%), Positives = 89/186 (47%), Gaps = 14/186 (7%)
Query: 548 GVCSTWMKNSAPLEKSQDCSWAPIFVRQSNFRLPADN-KVPIIMIGPGTGLAPFRGFLQE 606
GVCS ++ N+ P +K + + + LP D+ K IMI GTG+AP+RG+L+
Sbjct: 194 GVCSNFLCNAKPGDKVKITGPSGKVML-----LPEDDPKATHIMIATGTGVAPYRGYLRR 248
Query: 607 RLALKEEGAEL-GPSVLFFGCRNRQVDYIYEDELNHFVHGGALS-ELIVAFSREGPTKE- 663
+ G + LF G N +Y++E + + A SRE K+
Sbjct: 249 MFMENVPNFKFDGLAWLFLGVANSD-SLLYDEEFAGYRKDYPENFRYDKALSREEKNKKG 307
Query: 664 ---YVQHKMIEKASDIWNMISQGAYIYVCGDAKGMAKDVHRTLHTILQEQGSLDNSKTES 720
YVQ K+ E + +I+ ++ GA+IY CG KGM + TL + +E+G K
Sbjct: 308 GKMYVQDKIEEYSDEIFKLLDNGAHIYFCG-LKGMMPGIQDTLKRVAEERGESWEQKLTQ 366
Query: 721 MVKNLQ 726
+ KN Q
Sbjct: 367 LRKNKQ 372
>At5g66190 ferredoxin-NADP+ reductase
Length = 360
Score = 63.2 bits (152), Expect = 5e-10
Identities = 50/176 (28%), Positives = 85/176 (47%), Gaps = 14/176 (7%)
Query: 543 GRIHQGVCSTWMKNSAPLEKSQDCSWAPIFVRQSNFRLPADNKVPIIMIGPGTGLAPFRG 602
G I +GVCS ++ + P ++++ P+ +P D IIM+G GTG+APFR
Sbjct: 171 GEIVKGVCSNFLCDLKPGDEAKITG--PV---GKEMLMPKDPNATIIMLGTGTGIAPFRS 225
Query: 603 FLQERLALKEEGAEL-GPSVLFFGCRNRQVDYIYEDELNHFVHGGALS-ELIVAFSREGP 660
FL + + E + G + LF G +Y++E + L A SRE
Sbjct: 226 FLWKMFFEEHEDYKFNGLAWLFLGVPTSS-SLLYKEEFEKMKEKNPDNFRLDFAVSREQT 284
Query: 661 T----KEYVQHKMIEKASDIWNMISQ-GAYIYVCGDAKGMAKDVHRTLHTILQEQG 711
K Y+Q +M E A ++W ++ + ++Y+CG KGM K + + ++ + G
Sbjct: 285 NEKGEKMYIQTRMAEYAEELWELLKKDNTFVYMCG-LKGMEKGIDDIMVSLAAKDG 339
>At1g75200 unknown protein
Length = 647
Score = 61.2 bits (147), Expect = 2e-09
Identities = 76/286 (26%), Positives = 117/286 (40%), Gaps = 62/286 (21%)
Query: 96 VFFGTQTGTAEGFAKAIAEEAKARYEKAKFKVVDMDDYAADDDEYEEKLKKETMALFFLA 155
+FF +QTGTA+ A+ + E + F +VD Y +D L KET+ LF +
Sbjct: 52 IFFISQTGTAKALAQRLHELCASN--DIAFDIVDPHSYEPED------LPKETLVLFIAS 103
Query: 156 TYGDGEPTDNAARFYKWFEEFEGEEDSFKNLQYGVFGLGNRQYEHFNKVCTQALMHSCGF 215
T+ G+P N W E ED F +G+ L+ C F
Sbjct: 104 TWDGGKPPKNGEFLVNWLG--ESAED---------FRVGS------------LLLSDCKF 140
Query: 216 LSFDYSDGVASFRAFSDIYHSTQVAKIVDDKLLEKGGNRLVPVGLGD-DDQCIEDDFTAW 274
F GV S RA+ + Y++ VAK + +++ GG ++PVG GD DD ++ F W
Sbjct: 141 AVF----GVGS-RAYGESYNA--VAKELSSRMIGLGGLEMIPVGEGDVDDGELDRAFQDW 193
Query: 275 KEELWPALDQLLRDEDDATVATPYTASVLEYRVVIRDQLD--ATVDEKKQLNGNGHAVVD 332
+ + L E T V + + D L+ + DE+ + N +VD
Sbjct: 194 CDGVIRVLKGGSAQE---------TNGVSQQIGAVEDDLEYYDSTDEEDEDNDADGGIVD 244
Query: 333 AHH-----PVRANVAV-------RKELHTPASDRSCTHLEFDISGT 366
P + N V +KE+ TP S T + I G+
Sbjct: 245 LEDIAGKAPSKRNGVVKVTKVDGKKEMVTPVIRASLTKQGYKIIGS 290
>At1g20020 ferredoxin--NADP reductase precursor, putative
Length = 369
Score = 57.4 bits (137), Expect = 3e-08
Identities = 47/176 (26%), Positives = 81/176 (45%), Gaps = 14/176 (7%)
Query: 543 GRIHQGVCSTWMKNSAPLEKSQDCSWAPIFVRQSNFRLPADNKVPIIMIGPGTGLAPFRG 602
G +GVCS ++ + AP S P+ +P D +IM+ GTG+APFR
Sbjct: 180 GETVKGVCSNFLCDLAP--GSDVKLTGPV---GKEMLMPKDPNATVIMLATGTGIAPFRS 234
Query: 603 FLQERLALKEEGAEL-GPSVLFFGCRNRQVDYIYEDELNHF-VHGGALSELIVAFSREGP 660
FL + K + + G + LF G +Y++E + + A SRE
Sbjct: 235 FLWKMFFEKHDDYKFNGLAWLFLGVPTTS-SLLYQEEFDKMKAKAPENFRVDYAISREQA 293
Query: 661 T----KEYVQHKMIEKASDIWNMISQ-GAYIYVCGDAKGMAKDVHRTLHTILQEQG 711
K Y+Q +M + A+++W ++ + ++Y+CG KGM K + + ++ G
Sbjct: 294 NDKGEKMYIQTRMAQYAAELWELLKKDNTFVYMCG-LKGMEKGIDDIMVSLAANDG 348
>At4g35370 putative protein
Length = 414
Score = 34.7 bits (78), Expect = 0.20
Identities = 23/87 (26%), Positives = 38/87 (43%), Gaps = 5/87 (5%)
Query: 56 VVVLIWRRSNSQKPKPIEVPKRVIEKLPELEIDDGTKKVTVFFGTQTGTAEGFAKAIAEE 115
+V L W + K P+ +E+ ++DD + V + G +K +
Sbjct: 3 IVALSWIPKEASKAMPVAAESPCMEE----KMDDEIQNAQVTHAKSVAKSFGKSKVASSS 58
Query: 116 AKARYEKAKF-KVVDMDDYAADDDEYE 141
+ E KF K +DMD+Y +DDE E
Sbjct: 59 STDADEVVKFLKELDMDNYDEEDDEIE 85
>At3g01180 putative glycogen synthase
Length = 792
Score = 34.7 bits (78), Expect = 0.20
Identities = 24/81 (29%), Positives = 41/81 (49%), Gaps = 4/81 (4%)
Query: 381 ENLSDTVEEAERILGLSPDTYFSVHTD-DEDGKPLGG---SSLPPPFPPCTLRTALAKYA 436
E+L + + + ++ SP T +D +GKP SS+ PP+ P ++ T+ K +
Sbjct: 171 ESLKNRKQSSASVISSSPVTSPQKPSDVATNGKPWSSVVASSVDPPYKPSSVMTSPEKTS 230
Query: 437 DVLSSPKKSALLALAAHASDP 457
D ++SP K + A SDP
Sbjct: 231 DPVTSPGKPSKSRAGAFWSDP 251
>At3g24710 hypothetical protein
Length = 126
Score = 31.2 bits (69), Expect = 2.2
Identities = 12/35 (34%), Positives = 20/35 (56%)
Query: 676 IWNMISQGAYIYVCGDAKGMAKDVHRTLHTILQEQ 710
+W+++ GA +YV G + M DV L I+ E+
Sbjct: 5 VWDLLCDGAVVYVAGSSTEMPSDVMSALGEIVSEE 39
>At1g51930 hypothetical protein
Length = 132
Score = 30.0 bits (66), Expect = 4.8
Identities = 25/99 (25%), Positives = 45/99 (45%), Gaps = 19/99 (19%)
Query: 52 LIGCVVVLIWRRSNSQKPKPIEVPKRVIEKL--PELEIDDGTKKVTVFFGTQTGTAEGFA 109
+IG +V++I ++ P P +P I + P+ +I+ G K VF
Sbjct: 13 IIGLIVIVIANCCDNIDPPPTRLPPETIHQTVQPQQDIETGQSKALVF------------ 60
Query: 110 KAIAEEAKARYEKA---KFKVVDMDDYAADDDEYEEKLK 145
K I EE R E+ +F + +++Y +DD +L+
Sbjct: 61 KDIKEEEGGREEEGGGKRFCPICLEEY--EDDHQIRRLR 97
>At1g48790 unknown protein
Length = 507
Score = 29.6 bits (65), Expect = 6.3
Identities = 32/118 (27%), Positives = 47/118 (39%), Gaps = 17/118 (14%)
Query: 486 LLEVMAEFSSAKPPIGVFFASVAPRLQPRYY--------SISSSPRVAPSRIHVTCALVH 537
LL+V+ E KP + + P+L+PRY S+ S V PS A V
Sbjct: 87 LLDVLTELEKLKPVVQQRIDELYPKLKPRYNVQAHPANGSLGWSSAVKPSFNSYDHAKVR 146
Query: 538 DKMPTGRIHQGVCSTWMKNSAPLEKSQDCSWAPIFVRQS-NFRLPADNKVPIIMIGPG 594
+ + G N+APLE+ F + S NFR + ++GPG
Sbjct: 147 NPPGHNSGYMGSRGQQFLNAAPLEER--------FRKMSVNFRPNEETLSKHSILGPG 196
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.135 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,776,881
Number of Sequences: 26719
Number of extensions: 828509
Number of successful extensions: 2331
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 2298
Number of HSP's gapped (non-prelim): 17
length of query: 736
length of database: 11,318,596
effective HSP length: 107
effective length of query: 629
effective length of database: 8,459,663
effective search space: 5321128027
effective search space used: 5321128027
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 64 (29.3 bits)
Medicago: description of AC135801.5


