

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135801.4 + phase: 0
(61 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
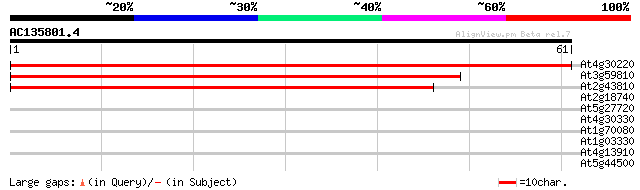
Sequences producing significant alignments: (bits) Value
At4g30220 snRNP Sm protein F - like 110 2e-25
At3g59810 U6 snRNA-associated Sm-like protein 50 3e-07
At2g43810 putative small nuclear ribonucleoprotein polypeptide F 49 4e-07
At2g18740 small nuclear ribonucleoprotein E like protein 30 0.23
At5g27720 glycine rich protein - like 29 0.39
At4g30330 small nuclear ribonucleoprotein homolog 28 1.1
At1g70080 unknown protein 25 5.6
At1g03330 unknown protein 25 5.6
At4g13910 putative disease resistance protein 25 7.3
At5g44500 unknown protein 25 9.5
>At4g30220 snRNP Sm protein F - like
Length = 88
Score = 110 bits (274), Expect = 2e-25
Identities = 51/61 (83%), Positives = 55/61 (89%)
Query: 1 MEYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVLYMRGVPEDEEIEDAAD 60
MEYKG+L SVDSYMNLQL NTEEYI+G TGNLGEILIRCNNVLY+RGVPEDEE+EDA
Sbjct: 28 MEYKGFLASVDSYMNLQLGNTEEYIDGQLTGNLGEILIRCNNVLYVRGVPEDEELEDADQ 87
Query: 61 D 61
D
Sbjct: 88 D 88
>At3g59810 U6 snRNA-associated Sm-like protein
Length = 91
Score = 49.7 bits (117), Expect = 3e-07
Identities = 21/49 (42%), Positives = 30/49 (60%)
Query: 1 MEYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVLYMRGV 49
++Y+G L +D YMN+ + TEEY+ G G+ IR NNVLY+ V
Sbjct: 35 VDYRGTLTCLDGYMNIAMEQTEEYVNGQLKNKYGDAFIRGNNVLYISTV 83
>At2g43810 putative small nuclear ribonucleoprotein polypeptide F
Length = 91
Score = 49.3 bits (116), Expect = 4e-07
Identities = 19/46 (41%), Positives = 29/46 (62%)
Query: 1 MEYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVLYM 46
++Y+G L +D YMN+ + TEEY+ G G+ +R NNVLY+
Sbjct: 35 VDYRGILTCLDGYMNIAMEQTEEYVNGQLKNTYGDAFVRGNNVLYI 80
>At2g18740 small nuclear ribonucleoprotein E like protein
Length = 88
Score = 30.0 bits (66), Expect = 0.23
Identities = 17/47 (36%), Positives = 25/47 (53%), Gaps = 1/47 (2%)
Query: 1 MEYKGYLVSVDSYMNLQLANTEEY-IEGNFTGNLGEILIRCNNVLYM 46
+ +G + D YMNL L EE I+ N LG IL++ +N+ M
Sbjct: 37 LRIEGRITGFDEYMNLVLDEAEEVSIKKNTRKPLGRILLKGDNITLM 83
>At5g27720 glycine rich protein - like
Length = 129
Score = 29.3 bits (64), Expect = 0.39
Identities = 22/63 (34%), Positives = 35/63 (54%), Gaps = 9/63 (14%)
Query: 3 YKGYLVSVDSYMNLQLANTEEYI----EGNFTGNLGEILIRCNNVLYMRGVPEDEEIEDA 58
Y G+LV+ D++MN+ L E I +G+ + E IR N + Y+R VP DE I+
Sbjct: 25 YNGHLVNCDTWMNIHL---REVICTSKDGDRFWRMPECYIRGNTIKYLR-VP-DEVIDKV 79
Query: 59 ADD 61
++
Sbjct: 80 QEE 82
>At4g30330 small nuclear ribonucleoprotein homolog
Length = 88
Score = 27.7 bits (60), Expect = 1.1
Identities = 16/47 (34%), Positives = 24/47 (51%), Gaps = 1/47 (2%)
Query: 1 MEYKGYLVSVDSYMNLQLANTEEY-IEGNFTGNLGEILIRCNNVLYM 46
+ +G + D YMNL L EE I+ LG IL++ +N+ M
Sbjct: 37 LRIEGRITGFDEYMNLVLDEAEEVSIKKKTRKPLGRILLKGDNITLM 83
>At1g70080 unknown protein
Length = 611
Score = 25.4 bits (54), Expect = 5.6
Identities = 13/35 (37%), Positives = 17/35 (48%)
Query: 4 KGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILI 38
KGYL + L + EEYIE GE+L+
Sbjct: 436 KGYLQEAEWSNRGHLPSHEEYIEVGVASTAGEVLL 470
>At1g03330 unknown protein
Length = 93
Score = 25.4 bits (54), Expect = 5.6
Identities = 17/51 (33%), Positives = 28/51 (54%), Gaps = 3/51 (5%)
Query: 4 KGYLVSVDSYMNLQLANTEEYIEGNFTGNLG--EILIRCNNVLYMRGVPED 52
+G L SVD Y+N++L NT + + L IR + V Y++ +P+D
Sbjct: 26 RGTLHSVDQYLNIKLENTRVVDQDKYPHMLSVRNCFIRGSVVRYVQ-LPKD 75
>At4g13910 putative disease resistance protein
Length = 509
Score = 25.0 bits (53), Expect = 7.3
Identities = 16/56 (28%), Positives = 29/56 (51%), Gaps = 4/56 (7%)
Query: 2 EYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVLYMRGVPEDEEIED 57
E+ +L + S L ++ + IEG +G L + LI+C ++ ++ ED I D
Sbjct: 166 EFPTFLQNQTSLEYLDISANQ--IEGQLSGQLPKSLIKCTDLEFLN--VEDNRIND 217
>At5g44500 unknown protein
Length = 254
Score = 24.6 bits (52), Expect = 9.5
Identities = 21/65 (32%), Positives = 30/65 (45%), Gaps = 15/65 (23%)
Query: 5 GYLVSVDSYMNLQLANTEEY-----IEGNFTGN--------LGEILIRCNNVLYM--RGV 49
G ++ D +MNL L + EE+ +GN N LG +L+R V+ M G
Sbjct: 29 GKFMAFDRHMNLVLGDCEEFRKLPPAKGNKKTNEEREERRTLGLVLLRGEEVISMTVEGP 88
Query: 50 PEDEE 54
P EE
Sbjct: 89 PPPEE 93
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.313 0.137 0.385
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,448,864
Number of Sequences: 26719
Number of extensions: 48985
Number of successful extensions: 66
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 61
Number of HSP's gapped (non-prelim): 10
length of query: 61
length of database: 11,318,596
effective HSP length: 37
effective length of query: 24
effective length of database: 10,329,993
effective search space: 247919832
effective search space used: 247919832
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 52 (24.6 bits)
Medicago: description of AC135801.4


