

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135799.3 + phase: 0
(242 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
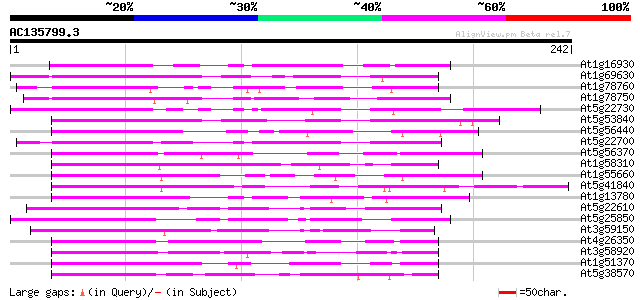
Score E
Sequences producing significant alignments: (bits) Value
At1g16930 hypothetical protein 86 2e-17
At1g69630 hypothetical protein 78 5e-15
At1g78760 hypothetical protein 75 4e-14
At1g78750 hypothetical protein 74 7e-14
At5g22730 unknown protein 73 1e-13
At5g53840 putative protein 71 5e-13
At5g56440 putative protein 70 1e-12
At5g22700 unknown protein 70 1e-12
At5g56370 unknown protein 68 4e-12
At1g58310 hypothetical protein 68 5e-12
At1g55660 hypothetical protein 68 5e-12
At5g41840 putative protein 67 9e-12
At1g13780 hypothetical protein 67 1e-11
At5g22610 unknown protein 66 1e-11
At5g25850 putative protein 66 2e-11
At3g59150 unknown protein 66 2e-11
At4g26350 putative protein 65 3e-11
At3g58920 putative protein 65 3e-11
At1g51370 unknown protein 65 3e-11
At5g38570 putative protein 65 4e-11
>At1g16930 hypothetical protein
Length = 449
Score = 85.5 bits (210), Expect = 2e-17
Identities = 56/173 (32%), Positives = 93/173 (53%), Gaps = 33/173 (19%)
Query: 18 ADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKF 77
+D+IS LPD +L ILS +STKE+V TS+LSKRW NLWL +P +D ++SN+
Sbjct: 14 SDRISNLPDSLLCQILSDLSTKESVCTSVLSKRWRNLWLHVPVLD--------LDSNNFP 65
Query: 78 IDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWVNLV 137
D V+ F+NRF + N L + + + + W+N V
Sbjct: 66 DDDVF------------VSFVNRF---LGSENEQHLERFKLIYEVNEHDASRFKSWINAV 110
Query: 138 VQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRTLVSLDLHHFSV 190
++RR+ + +++H +VDD D L K+P+++++C LV+L L+ ++
Sbjct: 111 IKRRVCH--------FNVHNEVDDDDE--LVKMPLSLYSCERLVNLQLYRVAL 153
>At1g69630 hypothetical protein
Length = 451
Score = 77.8 bits (190), Expect = 5e-15
Identities = 56/186 (30%), Positives = 87/186 (46%), Gaps = 34/186 (18%)
Query: 1 MQRQKRRRRSSKASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPN 60
M + + +K S V D IS+LPD +L +L + TK+ V TS+LS+RW NLW +P
Sbjct: 1 MDEDREKHVRAKGSDEV-DWISKLPDCLLCEVLLNLPTKDVVKTSVLSRRWRNLWKHVPG 59
Query: 61 IDFNNIKIDSIESNSKFIDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPH 120
+D +N + F+DS F+ +F L Y D+
Sbjct: 60 LDLDNTDFQEFNTFLSFVDSFLD--------FNSESFLQKFILK---------YDCDDE- 101
Query: 121 LLYKRSCHNVVKWVNLVVQRRLKYLRLNLRLGYDLHLDV-DDYDNSYLPKLPITIFTCRT 179
Y + +W+N +V R+++ H+DV DD S+ +LP +I+TC +
Sbjct: 102 --YDPDIFLIGRWINTIVTRKVQ------------HIDVLDDSYGSWEVQLPSSIYTCES 147
Query: 180 LVSLDL 185
LVSL L
Sbjct: 148 LVSLKL 153
>At1g78760 hypothetical protein
Length = 452
Score = 74.7 bits (182), Expect = 4e-14
Identities = 60/188 (31%), Positives = 98/188 (51%), Gaps = 50/188 (26%)
Query: 4 QKRRRRSSKASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLP--NI 61
+KR RRS + D++S LP+ ++ I+ + TK+ V +S+LSKRW NLW +P N+
Sbjct: 7 EKRARRSEE------DRLSNLPESLICQIMLNIPTKDVVKSSVLSKRWRNLWRYVPGLNV 60
Query: 62 DFNNIKIDSIESNSKFIDSVYSVLVSPDTAIGGSHFINRF-CLDVQ--FRNGNPLYGYDN 118
++N +F+D Y+ VS F++RF LD + F++ Y D
Sbjct: 61 EYN-----------QFLD--YNAFVS---------FVDRFLALDRESCFQSFRLRYDCDE 98
Query: 119 PHLLYKRSCHNVVKWVNLVVQRRLKYLRLNLRLGYDLHLDVDDYD-NSYLPKLPITIFTC 177
+R+ NV +W+N+VV ++LK LDV DY + ++P +++TC
Sbjct: 99 E----ERTISNVKRWINIVVDQKLKV------------LDVLDYTWGNEEVQIPPSVYTC 142
Query: 178 RTLVSLDL 185
+LVSL L
Sbjct: 143 ESLVSLKL 150
>At1g78750 hypothetical protein
Length = 458
Score = 73.9 bits (180), Expect = 7e-14
Identities = 61/191 (31%), Positives = 91/191 (46%), Gaps = 39/191 (20%)
Query: 7 RRRSSKASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNI----- 61
+R +K S V D IS LP+ +L +L + TK+ V +S+LS RW NLW +P
Sbjct: 7 KRVRAKGSREV-DWISNLPETLLCQVLFYLPTKDVVKSSVLSSRWRNLWKYVPGFNLSYC 65
Query: 62 DFNNIKIDSIESNS--KFIDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNP 119
DF+ S + N+ +F+DS F ++ CL FR GY P
Sbjct: 66 DFHVRNTFSYDHNTFLRFVDSFMG-------------FNSQSCLQ-SFRLEYDSSGYGEP 111
Query: 120 HLLYKRSCHNVVKWVNLVVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRT 179
L R +W+N VV R++KYL + +DD ++Y ++P T++TC T
Sbjct: 112 KLALIR------RWINSVVSRKVKYLGV-----------LDDSCDNYEFEMPPTLYTCET 154
Query: 180 LVSLDLHHFSV 190
LV L L S+
Sbjct: 155 LVYLTLDGLSL 165
>At5g22730 unknown protein
Length = 457
Score = 73.2 bits (178), Expect = 1e-13
Identities = 64/233 (27%), Positives = 108/233 (45%), Gaps = 49/233 (21%)
Query: 1 MQRQKRRRRSSKASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPN 60
M+ + R +A D IS+LPD ++ IL + K+ V TS LS RW +LWL +P
Sbjct: 1 MKAKYGERSQGTYFMAGEDLISKLPDSLITQILLYLPIKDIVRTSSLSSRWKSLWLLIPR 60
Query: 61 IDFNNIKIDSIESNSKFIDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPH 120
+D +DS E Y+ V F+N+F + F +G D
Sbjct: 61 LD-----LDSEEFQD------YNAFVG---------FMNKF---IDF-SGEEKICLDKLK 96
Query: 121 LLYKRSCHN---VVKWVNLVVQRRLKYLRLNLRLGYDLHLDVDDYDN-SYLPKLPITIFT 176
L +++ ++ V +W++ VV+R+LK HLDV+ N +L ++P++++
Sbjct: 97 LSSRKTVNDLPCVTRWIDFVVRRKLK------------HLDVECLVNRKFLEEMPLSLYV 144
Query: 177 CRTLVSLDLHHFSVKGVCYRFLLKALSTSNSLRLDTFKLYRSIYQVRFIMNTM 229
C TLV+L LH R LL + L T +L ++Y ++ ++
Sbjct: 145 CDTLVNLRLH---------RVLLGKFEAVSLPCLKTMRLEENVYANDVVLESL 188
>At5g53840 putative protein
Length = 444
Score = 71.2 bits (173), Expect = 5e-13
Identities = 53/200 (26%), Positives = 94/200 (46%), Gaps = 43/200 (21%)
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKFI 78
+++S+LPD ++ ILS +STK+AV TSILS RW NLW +P +DF++ ++ S F
Sbjct: 18 ERLSQLPDHLICVILSHLSTKDAVRTSILSTRWRNLWQLVPVLDFDSRELRSFSEFVSFA 77
Query: 79 DSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWVNLVV 138
S + + +I + + + GN + + W++LV
Sbjct: 78 GSFFY--------LHKDSYIQKLRVCIYDLAGN----------------YYLTSWIDLVT 113
Query: 139 QRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRTLVSLDLHHFSVKGV----- 193
+ R++ H+D+ + S +P++++TC TLV L L ++ V
Sbjct: 114 RHRIQ------------HIDISVFTCSGFGVIPLSLYTCDTLVHLKLSRVTMVNVEFVSL 161
Query: 194 -CYRFL-LKALSTSNSLRLD 211
C + L L ++ +N LD
Sbjct: 162 PCLKILDLDFVNFTNETTLD 181
>At5g56440 putative protein
Length = 430
Score = 70.1 bits (170), Expect = 1e-12
Identities = 55/192 (28%), Positives = 90/192 (46%), Gaps = 48/192 (25%)
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKFI 78
D+IS LPDDV+ ILS V TK V+T++LSKRW LW +P +D+ + + E
Sbjct: 2 DRISLLPDDVVFKILSFVPTKVVVSTNLLSKRWRYLWKHVPKLDYRDPSLVDTEHWR--- 58
Query: 79 DSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSC--HNVVKWVNL 136
S F+++F L + P+ + HL R+C ++ W+++
Sbjct: 59 ---------------ASRFVDKFLL----LHEAPVL--ETLHLSLSRNCPPSDIETWISV 97
Query: 137 VVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLP-----KLPITIFTCRTLVSLD-LHHFSV 190
+ RR++ L + Y+P +LP +++TC TLVSL L F+V
Sbjct: 98 AISRRVRNLHIY----------------RYIPSTGPIRLPRSLYTCETLVSLYLLLDFTV 141
Query: 191 KGVCYRFLLKAL 202
+ F ++L
Sbjct: 142 DDAPFMFCFRSL 153
>At5g22700 unknown protein
Length = 452
Score = 70.1 bits (170), Expect = 1e-12
Identities = 55/183 (30%), Positives = 87/183 (47%), Gaps = 40/183 (21%)
Query: 4 QKRRRRSSKASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDF 63
QKR + S++ D+IS LPD++L ILS + TK AV TSILS RW ++WLS P +D
Sbjct: 11 QKRNQMSNRG-----DRISSLPDELLCQILSNLPTKNAVTTSILSTRWRSIWLSTPVLD- 64
Query: 64 NNIKIDSIESNSKFIDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLY 123
I ID+ + + F+ F +RF ++F + L+ +
Sbjct: 65 --IDIDAFDDATTFVS-----------------FASRF---LEFSKDSCLHKFKLSVERD 102
Query: 124 KRSCHNVVKWVNLVVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRTLVSL 183
++ W+ V RR++ HL+VD + + + +T++ TLVSL
Sbjct: 103 DVDMCTIMPWIQDAVNRRIQ------------HLEVDCRFSFHFEAVYLTLYLSETLVSL 150
Query: 184 DLH 186
LH
Sbjct: 151 RLH 153
>At5g56370 unknown protein
Length = 421
Score = 68.2 bits (165), Expect = 4e-12
Identities = 57/188 (30%), Positives = 85/188 (44%), Gaps = 38/188 (20%)
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKFI 78
D IS LPDD L ILSL+ TK+ + TS+LSKRW LW +P + ++ ID + F+
Sbjct: 2 DSISLLPDDFLLRILSLLPTKDVLNTSVLSKRWRYLWKLVPKLQYS--LIDKNADHGTFV 59
Query: 79 DSV-YSVLVSPDTAIGGSHF-INRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWVNL 136
V S+L+S + H + R C +V ++ WV +
Sbjct: 60 RFVDRSLLLSMAPVLESLHLKLGRQCSEV-----------------------DIGFWVRI 96
Query: 137 VVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRTLVSLDLHHFSVKGVCYR 196
V++ L +L D + Y +LP ++FTC TL L L + S+K V +
Sbjct: 97 AVEKGL----------CELDFDYEHYKTEPC-RLPQSLFTCGTLTVLKLKNVSLKDVQFP 145
Query: 197 FLLKALST 204
K L T
Sbjct: 146 VCFKLLKT 153
>At1g58310 hypothetical protein
Length = 505
Score = 67.8 bits (164), Expect = 5e-12
Identities = 51/171 (29%), Positives = 90/171 (51%), Gaps = 21/171 (12%)
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDF-NNIKIDSIESNSKF 77
D IS LPD +L +ILS ++TKEA +TS+L+K+W L+ S+PN+DF +++ + + N
Sbjct: 8 DIISGLPDSLLCHILSFLNTKEAASTSVLAKKWRYLFASVPNLDFDDSVHLRLGKRNPAV 67
Query: 78 IDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVK---WV 134
Y +++ + + F++ ++ ++ +PL+ + L R C ++V+ W+
Sbjct: 68 SGEDYLKMINERSDQLSTSFMDFVDQVLRLQDNSPLHKFS----LKIRDCVDIVRIICWI 123
Query: 135 NLVVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRTLVSLDL 185
V++R + L L D+HL + LP IF TLV L L
Sbjct: 124 LKVLERGVSDLEL------DMHL-------KWKSSLPSKIFLSETLVRLKL 161
>At1g55660 hypothetical protein
Length = 492
Score = 67.8 bits (164), Expect = 5e-12
Identities = 59/190 (31%), Positives = 94/190 (49%), Gaps = 36/190 (18%)
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFN--NIKIDSIESNSK 76
DKIS+LPD++L +LS +STK+AV+TSILS RW +LW+ LP +++N + + + ++
Sbjct: 53 DKISQLPDELLVKVLSFLSTKDAVSTSILSMRWKSLWMWLPKLEYNFRHYSVSEGQGLAR 112
Query: 77 FIDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWVNL 136
FI S V +P L ++FR G G P +Y WV+L
Sbjct: 113 FITSSLRVHKAPAIE----------SLSLKFRYG--AIGSIKPKDIY--------LWVSL 152
Query: 137 VVQ-RRLKYLRLNLRLGYDLHLDVDDYDNSYLP-KLPITIFTCRTLVSLDLHHFSVKGVC 194
V ++ L L L ++ + LP KLP ++ C+++V L L + V
Sbjct: 153 AVHVSNVRELSLKL------------FNFAELPTKLPKSLCKCKSIVILKLKDEILVDVP 200
Query: 195 YRFLLKALST 204
+ L +L T
Sbjct: 201 RKVCLPSLKT 210
>At5g41840 putative protein
Length = 540
Score = 67.0 bits (162), Expect = 9e-12
Identities = 67/241 (27%), Positives = 110/241 (44%), Gaps = 56/241 (23%)
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFN-----------NIK 67
D+IS LPD ++ +ILS + TKEA +T++L+KRW L +PN++F+ N
Sbjct: 14 DRISGLPDALICHILSFLPTKEAASTTVLAKRWKPLLAFVPNLNFDDSIYFHPRARRNKY 73
Query: 68 IDSIESNSKFIDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSC 127
S ES F+DSV ++ T + H C DV ++
Sbjct: 74 SKSYESFMSFVDSVLALQAKTKTPLKRFHV---KCEDVVDQSW----------------- 113
Query: 128 HNVVKWVNLVVQRRLKYLRLNLRLGYDLHLDVD-DY-DNSYLPKLPITIFTCRTLVSLDL 185
V++W+ V++R + L DLH+ +Y +NS LP IF +TLV L +
Sbjct: 114 --VLEWIPKVLKRGV--------LDIDLHITSSRNYCENSSFYSLPSKIFVSKTLVRLKI 163
Query: 186 H-----HFSVKGVCYRFLLKALSTSNSLRLDTFKLYRSIYQVRFIMNTMHVIYFLFLSEI 240
H V+G +L +L LD FK+ S+ + +++ H + L L+ +
Sbjct: 164 QFQDGVHIDVEGGV------SLPKLKTLHLDYFKIETSM--LNKLLSGCHALEELVLANL 215
Query: 241 L 241
+
Sbjct: 216 M 216
>At1g13780 hypothetical protein
Length = 451
Score = 66.6 bits (161), Expect = 1e-11
Identities = 51/184 (27%), Positives = 89/184 (47%), Gaps = 31/184 (16%)
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKFI 78
D+IS LP+ ++ IL + TK +V TS+LS RW NLWL++P +D N +N K +
Sbjct: 5 DRISELPESLISQILLHLPTKASVKTSVLSTRWKNLWLNVPGLDLNCRDFPFQNNNEKLL 64
Query: 79 DSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWVNLV- 137
FI+RF +QF N + L + + S ++K+ + +
Sbjct: 65 ----------------IDFIDRF---LQFNNESRLQKFKVDY-----SRDKIIKFSDRIG 100
Query: 138 --VQRRLKYLRLNLRLGYDLHLDVDD-YDNSYLPKLPITIFTCRTLVSLDLHHFSVKGVC 194
+ R ++ L + Y D DD D + +P+ +++C+TLVSL L + ++
Sbjct: 101 DAISRGIRVLDVESNTYY---RDADDCIDYPCIEFMPLNLYSCKTLVSLKLSYSGLEDPG 157
Query: 195 YRFL 198
+ +L
Sbjct: 158 FVYL 161
>At5g22610 unknown protein
Length = 502
Score = 66.2 bits (160), Expect = 1e-11
Identities = 46/179 (25%), Positives = 87/179 (47%), Gaps = 29/179 (16%)
Query: 8 RRSSKASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIK 67
+R LA+ D IS+LP+ +L ILS + TK+ V TS+LSKRW ++WL +P +D ++ +
Sbjct: 7 KRICVEQLAIEDLISKLPEVLLSQILSYLPTKDIVRTSVLSKRWKSVWLLIPGLDLDSSE 66
Query: 68 IDSIESNSKFIDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSC 127
++ F+D + L + +++ L +Q +P SC
Sbjct: 67 FPHYDT---FVDFMNEFLFFSREE---NPCLHKLKLSIQKNENDP-------------SC 107
Query: 128 HNVVKWVNLVVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRTLVSLDLH 186
V W + V + +L++L D+ + + +P++++ C+TL+ L L+
Sbjct: 108 --VTLWTDCVARGKLQHL--------DVEFGGRVMEREFWEMMPLSLYICKTLLHLRLY 156
>At5g25850 putative protein
Length = 481
Score = 65.9 bits (159), Expect = 2e-11
Identities = 50/190 (26%), Positives = 90/190 (47%), Gaps = 37/190 (19%)
Query: 1 MQRQKRRRRSSKASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPN 60
M R+ ++ L + +++S+L D ++ ILS + KE V TS+LS RW N+WLS+P+
Sbjct: 2 MGRKNSKQVRYSNRLILINRLSQLSDPLICQILSHLPIKEVVTTSVLSTRWKNIWLSVPS 61
Query: 61 IDFNNIKIDSIESNSKFIDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPH 120
++ +YS+ + +T F +RF + + L Y + H
Sbjct: 62 LEL-----------------IYSIFPNFNTF---RSFGDRFFDSTRVSCIHNLKLYFDEH 101
Query: 121 LLYKRSCHNVVKWVNLVVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRTL 180
+C+ + W+N V R ++ L + G +Y +LP++++ C TL
Sbjct: 102 ----DACY-LTSWINAFVTRNIRRLYVRRVRG------------NYFHELPLSLYVCETL 144
Query: 181 VSLDLHHFSV 190
VSL L H ++
Sbjct: 145 VSLKLFHLTL 154
>At3g59150 unknown protein
Length = 475
Score = 65.9 bits (159), Expect = 2e-11
Identities = 57/177 (32%), Positives = 83/177 (46%), Gaps = 33/177 (18%)
Query: 10 SSKASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNN---I 66
+ KA + D IS LP+D+L ILS +STKEA TSILSKRW+NL LS+P +DF++ +
Sbjct: 4 NKKAKPSYGDVISNLPNDLLCRILSYLSTKEAALTSILSKRWSNLLLSIPILDFDDSVLL 63
Query: 67 KIDSIESNSKFIDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRS 126
K + + F + L+S + S + R L + P
Sbjct: 64 KPQKGQRKNVFFKAFVDRLLS--QRVETSSHVQRVSLKCRQGGVEP-------------D 108
Query: 127 CHNVVKWVNLVVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRTLVSL 183
C V+KW+ L R L L L+L + + + LP +F +TLV L
Sbjct: 109 C--VIKWI-LTTVRDLGVLDLSLCIDFGIF------------HLPFNVFRSKTLVKL 150
>At4g26350 putative protein
Length = 431
Score = 65.1 bits (157), Expect = 3e-11
Identities = 44/167 (26%), Positives = 78/167 (46%), Gaps = 35/167 (20%)
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKFI 78
D IS+ PD +L ILS + TK+ + TS+LSKRW +LW +P +++ D N +F+
Sbjct: 2 DIISQCPDHLLLRILSFIPTKDVIVTSLLSKRWGSLWRWVPKLEY-----DFTRQNMRFV 56
Query: 79 DSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWVNLVV 138
VY L+ + + S + L + R ++ W+++ V
Sbjct: 57 KFVYRSLLQNNAPVLESLHLKNIILYAECRT------------------VDIGGWIDIAV 98
Query: 139 QRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRTLVSLDL 185
RR++ +L + ++ D + +LP +++TC TL S L
Sbjct: 99 SRRVR----------ELEISINCSDEKF--RLPSSLYTCGTLESFIL 133
>At3g58920 putative protein
Length = 470
Score = 65.1 bits (157), Expect = 3e-11
Identities = 56/169 (33%), Positives = 85/169 (50%), Gaps = 32/169 (18%)
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKFI 78
D+IS LP++++ +I+S +S KEA S+LSKRW NL+ + ++F+ DS+++ +
Sbjct: 2 DRISNLPNEIICHIVSFLSAKEAAFASVLSKRWQNLFTIVQKLEFD----DSVKNQGSLM 57
Query: 79 DSVYSVLVSPDTAIGGSHFINRF--CLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWVNL 136
D V VL P T S F RF L + G + PH++ + C NV+K
Sbjct: 58 DFVNGVLALPVTT-RISKFSLRFDSFLKRKLETGLVI----GPHVVNRCLC-NVLK---- 107
Query: 137 VVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRTLVSLDL 185
R DL L++ D YL LP +FTC+T+V L L
Sbjct: 108 -------------RGVLDLKLEIYGED-GYL--LPSEVFTCKTIVDLKL 140
>At1g51370 unknown protein
Length = 435
Score = 65.1 bits (157), Expect = 3e-11
Identities = 47/170 (27%), Positives = 81/170 (47%), Gaps = 45/170 (26%)
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKFI 78
D+IS+LP+ ++ IL +STK++V TS LS +W LW S+P +D + + + F+
Sbjct: 19 DRISQLPEPLISEILFHLSTKDSVRTSALSTKWRYLWQSVPGLDLDPYASSNTNTIVSFV 78
Query: 79 DSVYSVLVSPDTAIGGSH---FINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWVN 135
+S + SH +I + LD L Y ++++ W++
Sbjct: 79 ESFFD-----------SHRDSWIRKLRLD----------------LGYHHDKYDLMSWID 111
Query: 136 LVVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRTLVSLDL 185
RR++ HLDV + ++ K+P++I+TC TLV L L
Sbjct: 112 AATTRRIQ------------HLDVHCFHDN---KIPLSIYTCTTLVHLRL 146
>At5g38570 putative protein
Length = 411
Score = 64.7 bits (156), Expect = 4e-11
Identities = 50/171 (29%), Positives = 79/171 (45%), Gaps = 30/171 (17%)
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKFI 78
D I+ LPDD+L ILS V T AV+T +LSKRW LW+ LPN++F + +S +
Sbjct: 2 DNINGLPDDLLVKILSFVPTYVAVSTCVLSKRWEFLWMWLPNLEF----VSPWDSRPGIV 57
Query: 79 DSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWVNLVV 138
D ++ I + I R C+ + NPH+ + +W+ + V
Sbjct: 58 D-----FINKKLPIHRAPVIERLCIHIN----------SNPHI----KPEYIKRWIEIAV 98
Query: 139 QRRLKYLRLNL--RLGYDLHLDVDDY--DNSYLPKLPITIFTCRTLVSLDL 185
++ L+++ + L L+ + Y D YL +L I C L L L
Sbjct: 99 SHYVRELQIDYHSKTKITLQLEAESYFIDGKYLQQL---ISGCPVLEDLSL 146
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.326 0.141 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,269,064
Number of Sequences: 26719
Number of extensions: 221702
Number of successful extensions: 1255
Number of sequences better than 10.0: 227
Number of HSP's better than 10.0 without gapping: 199
Number of HSP's successfully gapped in prelim test: 28
Number of HSP's that attempted gapping in prelim test: 967
Number of HSP's gapped (non-prelim): 266
length of query: 242
length of database: 11,318,596
effective HSP length: 96
effective length of query: 146
effective length of database: 8,753,572
effective search space: 1278021512
effective search space used: 1278021512
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC135799.3


