

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
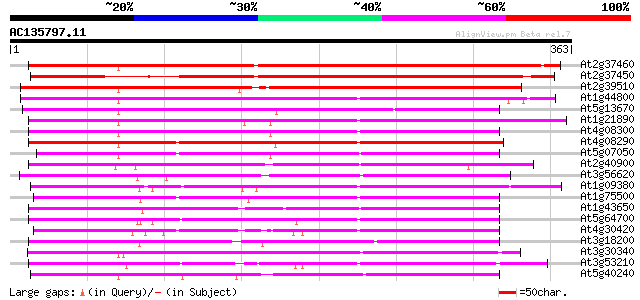
Query= AC135797.11 + phase: 0 /pseudo
(363 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At2g37460 nodulin-like protein 353 1e-97
At2g37450 nodulin-like protein 334 4e-92
At2g39510 nodulin-like protein 301 5e-82
At1g44800 nodulin like protein 255 3e-68
At5g13670 unknown protein 250 1e-66
At1g21890 nodulin-like protein 250 1e-66
At4g08300 nodulin-like protein 249 2e-66
At4g08290 nodulin-like protein 248 5e-66
At5g07050 MtN21 nodulin protein-like 246 1e-65
At2g40900 putative integral membrane protein nodulin 226 2e-59
At3g56620 nodulin-like protein 202 2e-52
At1g09380 nodulin N21 like protein 172 2e-43
At1g75500 nodulin-like protein 169 2e-42
At1g43650 nodulin-like protein 167 1e-41
At5g64700 nodulin-like protein 165 3e-41
At4g30420 nodulin-like protein 159 2e-39
At3g18200 integral membrane protein, putative 159 3e-39
At3g30340 unknown protein 155 3e-38
At3g53210 unknown protein 148 5e-36
At5g40240 unknown protein 133 2e-31
>At2g37460 nodulin-like protein
Length = 380
Score = 353 bits (905), Expect = 1e-97
Identities = 185/360 (51%), Positives = 237/360 (65%), Gaps = 19/360 (5%)
Query: 13 KPFIAVVLLQFGYAGMDILSKSALNKGMSCYVLVVYRHAVAFVVIVPFAVILENGSQ--- 69
+PFI++V+LQ G AGMDILSK+ LNKGMS YVLVVYRHAVA +V+ PFA + +
Sbjct: 14 RPFISMVVLQVGLAGMDILSKAVLNKGMSNYVLVVYRHAVATIVMAPFAFYFDKKVRPKM 73
Query: 70 ------------CARACY*SKFIFFGNEVYNSYFCSCHVQCPSCYYLCRGLDSQMEKIKM 117
+ G + + F + +E++K+
Sbjct: 74 TLMIFFKISLLGLLEPVIDQNLYYLGMKYTTATFATAMYNVLPAITFVLAYIFGLERVKL 133
Query: 118 RSVHSQAKIVGTIATVAGAMVMTLIKGPILNLFGIHESSAQIQHNGGVNLQHAVKGSIMI 177
R + S K+VGT+ATV GAM+MTL+KGP+L+LF SA + G ++ A+KG++++
Sbjct: 134 RCIRSTGKVVGTLATVGGAMIMTLVKGPVLDLFWTKGVSAH--NTAGTDIHSAIKGAVLV 191
Query: 178 TIGCFSCACFTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEPSVWSLSWDT 237
TIGCFS ACF ILQA+TL TYPAELSLTAWICL+GT+EG VAL+ME+G PS W++ WDT
Sbjct: 192 TIGCFSYACFMILQAITLRTYPAELSLTAWICLMGTIEGTAVALVMEKGNPSAWAIGWDT 251
Query: 238 KLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEKIYLGRV 297
KLL A YSGIVCS +AYY+ GVVM+ RGPVFVT F+PLCM+IVAIMS I AE++YLGRV
Sbjct: 252 KLLTATYSGIVCSALAYYVGGVVMKTRGPVFVTAFSPLCMIIVAIMSTIIFAEQMYLGRV 311
Query: 298 IGAVVIILGLYLVVWGKSKDYDRPSPI-IKDEILPAKQTIENNDKEKFHSHEVITSSNFG 356
+GAVVI GLYLV+WGK KDY S + + DE K + N K+ HEVIT S G
Sbjct: 312 LGAVVICAGLYLVIWGKGKDYKYNSTLQLDDESAQPKLELSGNGKDNV-DHEVITISKQG 370
>At2g37450 nodulin-like protein
Length = 297
Score = 334 bits (857), Expect = 4e-92
Identities = 177/340 (52%), Positives = 221/340 (64%), Gaps = 53/340 (15%)
Query: 14 PFIAVVLLQFGYAGMDILSKSALNKGMSCYVLVVYRHAVAFVVIVPFAVILENGSQCARA 73
PFI +VLLQ GYAGMDIL+K LNKGMS YVL VYRH VA VV+ PFA
Sbjct: 9 PFILMVLLQIGYAGMDILTKDVLNKGMSIYVLSVYRHGVATVVMAPFAF----------- 57
Query: 74 CY*SKFIFFGNEVYNSYFCSCHVQCPSCYYLCRGLDSQMEKIKMRSVHSQAKIVGTIATV 133
YF ++E +K +S+ S AK+VGT+ TV
Sbjct: 58 ----------------YF------------------DKLESVKFQSIRSAAKVVGTVTTV 83
Query: 134 AGAMVMTLIKGPILNLFGIHESSAQIQHNGGVNLQHAVKGSIMITIGCFSCACFTILQAV 193
G MVMTL+KGP L+LF SAQ + G ++ ++KG++++TIGCFS ACF ILQA+
Sbjct: 84 GGIMVMTLVKGPALDLFWTKGPSAQ--NTVGTDIHSSIKGAVLVTIGCFSYACFMILQAI 141
Query: 194 TLETYPAELSLTAWICLLGTVEGGIVALIMERGEPSVWSLSWDTKLLAAVYSGIVCSGMA 253
TL+TYPAELSL WICL+GT+EG +VAL+ME+G PSVW++ WDTKLL YSGIVCS +
Sbjct: 142 TLKTYPAELSLATWICLIGTIEGVVVALVMEKGNPSVWAIGWDTKLLTITYSGIVCSALG 201
Query: 254 YYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEKIYLGRVIGAVVIILGLYLVVWG 313
YYI GVVM+ RGPVFVT F PLCM++VAIMS I E++YLGR +GA VI +GLYLV+WG
Sbjct: 202 YYIGGVVMKTRGPVFVTAFKPLCMIVVAIMSSIIFDEQMYLGRALGATVICVGLYLVIWG 261
Query: 314 KSKDYDRPS-PIIKDEILPAKQTIENNDKEKFHSHEVITS 352
K+KDY+ PS P I D++ A K+K VI S
Sbjct: 262 KAKDYEYPSTPQIDDDLAQA-----TTSKQKEQRRTVIES 296
>At2g39510 nodulin-like protein
Length = 374
Score = 301 bits (770), Expect = 5e-82
Identities = 162/344 (47%), Positives = 219/344 (63%), Gaps = 25/344 (7%)
Query: 8 SFQNWKPFIAVVLLQFGYAGMDILSKSALNKGMSCYVLVVYRHAVAFVVIVPFAVILENG 67
+ + WKPFI VV LQFGYAG+ I++K ALN+GMS +VL YRH VA + I PFA L+
Sbjct: 2 ALKTWKPFITVVSLQFGYAGLSIIAKFALNQGMSPHVLASYRHIVATIFIAPFAYFLDRK 61
Query: 68 SQ---------------CARACY*SKFIFFGNEVYNSYFCSCHVQCPSCYYLCRGLDSQM 112
+ + G + ++ F + + ++
Sbjct: 62 IRPKMTLSIFFKILLLGLLEPTIDQNLYYTGMKYTSATFTAAMTNVLPAFAFIMAWIFRL 121
Query: 113 EKIKMRSVHSQAKIVGTIATVAGAMVMTLIKGPIL-----NLFGIHESSAQIQHNGGVNL 167
EK+ ++ +HSQAKI+GTI TV GAM+MT++KGP++ N IH+ S+ N GV
Sbjct: 122 EKVNVKKIHSQAKILGTIVTVGGAMLMTVVKGPLIPLPWANPHDIHQDSS----NTGVK- 176
Query: 168 QHAVKGSIMITIGCFSCACFTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGE 227
Q KG+ +I IGC A F LQA+TL++YP ELSLTA+IC LG++E IVAL +ERG
Sbjct: 177 QDLTKGASLIAIGCICWAGFINLQAITLKSYPVELSLTAYICFLGSIESTIVALFIERGN 236
Query: 228 PSVWSLSWDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFI 287
PS W++ D+KLLAAVY G++CSG+ YY+QGV+M+ RGPVFVT FNPL MVIVAI+ I
Sbjct: 237 PSAWAIHLDSKLLAAVYGGVICSGIGYYVQGVIMKTRGPVFVTAFNPLSMVIVAILGSII 296
Query: 288 LAEKIYLGRVIGAVVIILGLYLVVWGKSKDYDRPSPIIKDEILP 331
LAE ++LGR++GA+VI+LGLY V+WGKSKD S D+ LP
Sbjct: 297 LAEVMFLGRILGAIVIVLGLYSVLWGKSKDEPSSSFSDMDKELP 340
>At1g44800 nodulin like protein
Length = 370
Score = 255 bits (651), Expect = 3e-68
Identities = 150/368 (40%), Positives = 208/368 (55%), Gaps = 24/368 (6%)
Query: 8 SFQNWKPFIAVVLLQFGYAGMDILSKSALNKGMSCYVLVVYRHAVAFVVIVPFAVILENG 67
S + KP +A++ LQFGYAGM I++ + GM +VL YRH VA VV+ PFA++ E
Sbjct: 5 SMEKIKPILAIISLQFGYAGMYIITMVSFKHGMDHWVLATYRHVVATVVMAPFALMFERK 64
Query: 68 SQ---------------CARACY*SKFIFFGNEVYNSYFCSCHVQCPSCYYLCRGLDSQM 112
+ + G + ++ + S L ++
Sbjct: 65 IRPKMTLAIFWRLLALGILEPLMDQNLYYIGLKNTSASYTSAFTNALPAVTFILALIFRL 124
Query: 113 EKIKMRSVHSQAKIVGTIATVAGAMVMTLIKGPILNLFGIHESSAQIQHNGGVNLQHAVK 172
E + R VHS AK+VGT+ TV GAM+MTL KGP + + +S + QH V
Sbjct: 125 ETVNFRKVHSVAKVVGTVITVGGAMIMTLYKGPAIEIVKAAHNSFHGGSSSTPTGQHWVL 184
Query: 173 GSIMITIGCFSCACFTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEPSVWS 232
G+I I + A F ILQ+ TL+ YPAELSL IC +GT+ I +LIM R +PS W
Sbjct: 185 GTIAIMGSISTWAAFFILQSYTLKVYPAELSLVTLICGIGTILNAIASLIMVR-DPSAWK 243
Query: 233 LSWDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEKI 292
+ D+ LAAVYSG+VCSG+AYYIQ +V++ RGPVF T+F+P+CM+I A + +LAEKI
Sbjct: 244 IGMDSGTLAAVYSGVVCSGIAYYIQSIVIKQRGPVFTTSFSPMCMIITAFLGALVLAEKI 303
Query: 293 YLGRVIGAVVIILGLYLVVWGKSKDYDRP---SPIIKDEILP----AKQTIENNDKEKFH 345
+LG +IGAV I+LGLY VVWGKSKD P + K + LP KQT +D
Sbjct: 304 HLGSIIGAVFIVLGLYSVVWGKSKDEVNPLDEKIVAKSQELPITNVVKQT-NGHDVSGAP 362
Query: 346 SHEVITSS 353
++ V+TS+
Sbjct: 363 TNGVVTST 370
>At5g13670 unknown protein
Length = 377
Score = 250 bits (638), Expect = 1e-66
Identities = 139/327 (42%), Positives = 196/327 (59%), Gaps = 19/327 (5%)
Query: 9 FQNWKPFIAVVLLQFGYAGMDILSKSALNKGMSCYVLVVYRHAVAFVVIVPFAVILENGS 68
F+ +PFIA+V +Q YA M I++K ALNKGMS +VLV YR AVA +I PFA+ILE +
Sbjct: 3 FERARPFIAIVFIQCLYALMSIVAKLALNKGMSPHVLVAYRMAVASALITPFALILERNT 62
Query: 69 Q---------------CARACY*SKFIFFGNEVYNSYFCSCHVQCPSCYYLCRGLDSQME 113
+ + G ++ + F S ++E
Sbjct: 63 RPKLTFKILLQIAILSLFEPVVEQNLYYSGMKLTTATFTSALCNALPAMTFIMACVFKLE 122
Query: 114 KIKMRSVHSQAKIVGTIATVAGAMVMTLIKGPILNLFGIHESSAQIQHNGGVNLQHAV-- 171
K+ + HSQAK+VGT+ + GAM+MT +KG ++ L S H + +
Sbjct: 123 KVTIERRHSQAKLVGTMVAIGGAMLMTFVKGNVIELPWTSNSRGLNGHTHAMRIPKQADI 182
Query: 172 -KGSIMITIGCFSCACFTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEPSV 230
+GSIM+ CFS +C+ ILQA L Y AELSLTA +C++G +E ++ LI ER SV
Sbjct: 183 ARGSIMLVASCFSWSCYIILQAKILAQYKAELSLTALMCIMGMLEATVMGLIWERKNMSV 242
Query: 231 WSLSWDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAE 290
W ++ D LLA++Y G+V SG+AYY+ G + RGPVFV+ FNPL MV+VAI+S F+ E
Sbjct: 243 WKINPDVTLLASIYGGLV-SGLAYYVIGWASKERGPVFVSAFNPLSMVLVAILSTFVFLE 301
Query: 291 KIYLGRVIGAVVIILGLYLVVWGKSKD 317
K+Y+GRVIG+VVI++G+YLV+WGKSKD
Sbjct: 302 KVYVGRVIGSVVIVIGIYLVLWGKSKD 328
>At1g21890 nodulin-like protein
Length = 389
Score = 250 bits (638), Expect = 1e-66
Identities = 149/372 (40%), Positives = 208/372 (55%), Gaps = 25/372 (6%)
Query: 13 KPFIAVVLLQFGYAGMDILSKSALNKGMSCYVLVVYRHAVAFVVIVPFAVILENGSQ--- 69
KP++A++ +QFGYAGM I++ +L GM+ YVL VYRHA+A VI PFA+ E +
Sbjct: 10 KPYLAMISMQFGYAGMYIITMVSLKHGMNHYVLAVYRHAIATAVIAPFALFHERKIRPKM 69
Query: 70 ------------CARACY*SKFIFFGNEVYNSYFCSCHVQCPSCYYLCRGLDSQMEKIKM 117
+ G ++ F S + ++E +
Sbjct: 70 TFRIFLQIALLGFIEPVLDQNLYYVGMTYTSATFASATANVLPAITFVLAIIFRLESVNF 129
Query: 118 RSVHSQAKIVGTIATVAGAMVMTLIKGPILNLF------GIHESSAQIQHNGGVNL---Q 168
+ V S AK+VGT+ TV+GA++MTL KGPI++ G A H G +
Sbjct: 130 KKVRSIAKVVGTVITVSGALLMTLYKGPIVDFIRFGGGGGGGSDGAGGSHGGAGAAAMDK 189
Query: 169 HAVKGSIMITIGCFSCACFTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEP 228
H + G++M+ F A F ILQ+ TL+ YPAELSLT ICL+GT+EG V+L+ R +
Sbjct: 190 HWIPGTLMLLGRTFGWAGFFILQSFTLKQYPAELSLTTLICLMGTLEGTAVSLVTVR-DL 248
Query: 229 SVWSLSWDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFIL 288
S W + +D+ L AA YSG++CSG+AYY+QGVVMR RGPVFV TFNPLC+VI A + +L
Sbjct: 249 SAWKIGFDSNLFAAAYSGVICSGVAYYVQGVVMRERGPVFVATFNPLCVVITAALGVVVL 308
Query: 289 AEKIYLGRVIGAVVIILGLYLVVWGKSKDYDRPSPIIKDEILPAKQTIENNDKEKFHSHE 348
+E I+LG VIG + II+GLY VVWGK KD + LP K ++ D K + E
Sbjct: 309 SESIHLGSVIGTLFIIVGLYTVVWGKGKDKRMTDDDEDCKGLPIKSPVKPVDTGKGLAAE 368
Query: 349 VITSSNFGAIAR 360
+ S G A+
Sbjct: 369 LEMKSKEGQEAK 380
>At4g08300 nodulin-like protein
Length = 373
Score = 249 bits (635), Expect = 2e-66
Identities = 138/323 (42%), Positives = 191/323 (58%), Gaps = 19/323 (5%)
Query: 13 KPFIAVVLLQFGYAGMDILSKSALNKGMSCYVLVVYRHAVAFVVIVPFAVILENGSQ--- 69
KP IA++ LQFGYAGM I++ + GM+ ++L YRH VA +VI PFA+ILE +
Sbjct: 10 KPIIAIISLQFGYAGMYIITMVSFKHGMNHWILATYRHVVATIVIAPFALILERKIRPKM 69
Query: 70 ------------CARACY*SKFIFFGNEVYNSYFCSCHVQCPSCYYLCRGLDSQMEKIKM 117
+ G + ++ + S V + ++E + +
Sbjct: 70 TWPLFLRILALGFLEPLLDQNLYYIGMKATSATYSSAFVNALPAITFIMAVIFRIETVNL 129
Query: 118 RSVHSQAKIVGTIATVAGAMVMTLIKGPILNLFGIHESSAQIQHNGGVNL---QHAVKGS 174
+ S AK++GT TV GAMVMTL KGP + LF SS +G + Q+ V G+
Sbjct: 130 KKTRSLAKVIGTAITVGGAMVMTLYKGPAIELFKTAHSSLHGGSSGTSSETTDQNWVTGT 189
Query: 175 IMITIGCFSCACFTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEPSVWSLS 234
+ + + A F ILQ+ TL+ YPAELSL WIC +GTV I +LIM R + S W +
Sbjct: 190 LAVMGSITTWAGFFILQSFTLKKYPAELSLVMWICAMGTVLNTIASLIMVR-DVSAWKVG 248
Query: 235 WDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEKIYL 294
D+ LAAVYSG+VCSGMAYYIQ +V+R RGPVF T+F+P+CM+I A + +LAEKI+L
Sbjct: 249 MDSGTLAAVYSGVVCSGMAYYIQSIVIRERGPVFTTSFSPMCMIITAFLGVLVLAEKIHL 308
Query: 295 GRVIGAVVIILGLYLVVWGKSKD 317
G +IGA+ I+ GLY VVWGK+KD
Sbjct: 309 GSIIGAIFIVFGLYSVVWGKAKD 331
>At4g08290 nodulin-like protein
Length = 384
Score = 248 bits (632), Expect = 5e-66
Identities = 134/325 (41%), Positives = 207/325 (63%), Gaps = 20/325 (6%)
Query: 13 KPFIAVVLLQFGYAGMDILSKSALNKGMSCYVLVVYRHAVAFVVIVPFAVILENGSQ--- 69
+P++ ++ LQFG AG I+ + LN+G + YV++VYR+ VA +V+ PFA+I E +
Sbjct: 12 RPYLLMIFLQFGAAGTYIVIMATLNQGQNRYVVIVYRNLVAALVLAPFALIFERKVRPKM 71
Query: 70 ------------CARACY*SKFIFFGNEVYNSYFCSCHVQC-PSCYYLCRGLDSQMEKIK 116
F + G + ++ + S + PS ++ + +MEK+
Sbjct: 72 TLSVLWKIMALGFLEPVLDQGFGYLGMNMTSATYTSAIMNILPSVTFIIAWI-LRMEKVN 130
Query: 117 MRSVHSQAKIVGTIATVAGAMVMTLIKGPILNLFGIHESSAQIQHNGGVNLQHA--VKGS 174
+ V S+AKI+GT+ + GA+VMTL KGP++ L + + Q + + H V G+
Sbjct: 131 IAEVRSKAKIIGTLVGLGGALVMTLYKGPLIPLPWSNPNMDQQNGHTNNSQDHNNWVVGT 190
Query: 175 IMITIGCFSCACFTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEPSVWSLS 234
++I +GC + + F +LQ++T++TYPA+LSL+A ICL G V+ VAL++ER PS W++
Sbjct: 191 LLILLGCVAWSGFYVLQSITIKTYPADLSLSALICLAGAVQSFAVALVVER-HPSGWAVG 249
Query: 235 WDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEKIYL 294
WD +L A +Y+GIV SG+ YY+QG+VM+ RGPVFVT FNPLCM++VA+++ FIL E+I+
Sbjct: 250 WDARLFAPLYTGIVSSGITYYVQGMVMKTRGPVFVTAFNPLCMILVALIASFILHEQIHF 309
Query: 295 GRVIGAVVIILGLYLVVWGKSKDYD 319
G VIG VI GLY+VVWGK KDY+
Sbjct: 310 GCVIGGAVIAAGLYMVVWGKGKDYE 334
>At5g07050 MtN21 nodulin protein-like
Length = 381
Score = 246 bits (629), Expect = 1e-65
Identities = 141/324 (43%), Positives = 193/324 (59%), Gaps = 26/324 (8%)
Query: 18 VVLLQFGYAGMDILSKSALNKGMSCYVLVVYRHAVAFVVIVPFAVILENGSQ-------- 69
++ LQFGYAGM+I++K +LN GMS YVLVVYRHA+A VI PFA E +Q
Sbjct: 1 MISLQFGYAGMNIITKISLNTGMSHYVLVVYRHAIATAVIAPFAFFFERKAQPKITFSIF 60
Query: 70 -------CARACY*SKFIFFGNEVYNSYF-CSCHVQCPSCYYLCRGLDSQMEKIKMRSVH 121
F + G + + F C+ P+ ++ L +ME + ++ +
Sbjct: 61 MQLFILGLLGPVIDQNFYYMGLKYTSPTFSCAMSNMLPAMTFILAVL-FRMEMLDLKKLW 119
Query: 122 SQAKIVGTIATVAGAMVMTLIKGPILNLFGIHESSAQIQHNGGVNL-------QHAVKGS 174
QAKI GT+ TVAGAM+MT+ KGPI+ LF Q + + +KGS
Sbjct: 120 CQAKIAGTVVTVAGAMLMTIYKGPIVELFWTKYMHIQDSSHANTTSSKNSSSDKEFLKGS 179
Query: 175 IMITIGCFSCACFTILQAVTLETYPA-ELSLTAWICLLGTVEGGIVALIMERGEPSVWSL 233
I++ + A +LQA L+TY +LSLT IC +GT++ V +ME PS W +
Sbjct: 180 ILLIFATLAWASLFVLQAKILKTYAKHQLSLTTLICFIGTLQAVAVTFVMEHN-PSAWRI 238
Query: 234 SWDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEKIY 293
WD LLAA YSGIV S ++YY+QG+VM+ RGPVF T F+PL MVIVA+M F+LAEKI+
Sbjct: 239 GWDMNLLAAAYSGIVASSISYYVQGIVMKKRGPVFATAFSPLMMVIVAVMGSFVLAEKIF 298
Query: 294 LGRVIGAVVIILGLYLVVWGKSKD 317
LG VIGAV+I++GLY V+WGK K+
Sbjct: 299 LGGVIGAVLIVIGLYAVLWGKQKE 322
>At2g40900 putative integral membrane protein nodulin
Length = 400
Score = 226 bits (575), Expect = 2e-59
Identities = 130/348 (37%), Positives = 201/348 (57%), Gaps = 27/348 (7%)
Query: 13 KPFIAVVLLQFGYAGMDILSKSALNKGMSCYVLVVYRHAVAFVVIVPFAVILENG--SQC 70
KP+ A+V LQFGYAGM++++K+ L++GMS YVLV YR+A A I PFA++ E S+
Sbjct: 10 KPYFAMVCLQFGYAGMNLVTKTVLDRGMSHYVLVAYRNAFATAAIAPFALLSERKVRSKM 69
Query: 71 ARACY*SKFI-------------FFGNEVYNSYFCSCHVQCPSCYYLCRGLDSQMEKIKM 117
+ F+ + G ++ + F S + +MEK++M
Sbjct: 70 TFPIFMRIFLLALLGPVIDQNLYYIGLKLTSPTFSSAVSNIVPAITIILATLFRMEKVEM 129
Query: 118 RSVHSQAKIVGTIATVAGAMVMTLIKGPILNLFGIHESSAQIQHNGGVNLQHAVKGSIMI 177
R V K++GT+ TV G+++M KGP +N F H ++A +K ++ +
Sbjct: 130 RKVRCLVKVMGTLVTVVGSILMIFYKGPFINFFRSHLTAASSPPTADY-----LKAAVFL 184
Query: 178 TIGCFSCACFTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEPSVWSLSWDT 237
+ S A F +LQA TL+ Y A LS++ +C +GT++ +A +ME PS ++ +D
Sbjct: 185 LLASLSWASFFVLQAATLKKYSAHLSMSTMVCFMGTLQSLALAFVMEHN-PSALNIGFDM 243
Query: 238 KLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEKIYLG-- 295
LLA+ Y+GI+ S +AYY+QG++M+ +GPVFVT FNPL +VIV+IMS F+L + IYLG
Sbjct: 244 NLLASAYAGIMSSSIAYYVQGLMMQRKGPVFVTAFNPLIVVIVSIMSFFVLGQGIYLGGY 303
Query: 296 ----RVIGAVVIILGLYLVVWGKSKDYDRPSPIIKDEILPAKQTIENN 339
RVIG VV+++G+Y V+WGK D D +D ++ K NN
Sbjct: 304 VNNNRVIGVVVLMVGVYAVLWGKHVDDDGEETRHEDNVVAVKCCSGNN 351
>At3g56620 nodulin-like protein
Length = 377
Score = 202 bits (515), Expect = 2e-52
Identities = 117/333 (35%), Positives = 187/333 (56%), Gaps = 20/333 (6%)
Query: 7 KSFQNWKPFIAVVLLQFGYAGMDILSKSALNKGMSCYVLVVYRHAVAFVVIVPFAVILEN 66
K ++ KP+ A+V LQFGYAGM++++K L++GMS YVLV YR+A A I PFA++ E
Sbjct: 4 KMSESAKPYFAMVCLQFGYAGMNLVTKVVLDRGMSHYVLVAYRNAFATAAIAPFALLSER 63
Query: 67 GSQCARACY*SKFIF----FGNEV-YNSYFCSCHVQCPS----------CYYLCRGLDSQ 111
+ IF G + N Y+ + P+ + +
Sbjct: 64 KVRPKMTFPIFMQIFVLALLGPLIDQNLYYAGLKLTSPTFAGAVTNIVPALTFIISIICR 123
Query: 112 MEKIKMRSVHSQAKIVGTIATVAGAMVMTLIKGPILNLFGIHESSAQIQHNGGVNLQHAV 171
MEK++MR V QAK+VGT+ V GAM+M L K P++ H + + G + +
Sbjct: 124 MEKVEMRKVRFQAKVVGTLVIVVGAMLMILFKIPLITFLRSHLTGHALSPAG----EDYL 179
Query: 172 KGSIMITIGCFSCACFTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEPSVW 231
K ++ + I FS A F +LQA TL+ Y + LSL+ +C +GT++ + +ME S W
Sbjct: 180 KATVFLLIASFSWASFFVLQAATLKRYSSHLSLSTMVCFMGTLQSTALTFVMEPNL-SAW 238
Query: 232 SLSWDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEK 291
++ +D LLA+ Y+GI+ S +AYY+QG++ + + +FVT FNPL ++I +I+ IL +
Sbjct: 239 NIGFDMNLLASAYAGIMSSSIAYYVQGMMTKQKSVIFVTAFNPLVVIIGSIIGFLILNQT 298
Query: 292 IYLGRVIGAVVIILGLYLVVWGKSKDYDRPSPI 324
+ LG V+G ++++G+ V+WGK D D I
Sbjct: 299 LNLGGVLGMAILVVGVCTVLWGKEGDIDEEENI 331
>At1g09380 nodulin N21 like protein
Length = 374
Score = 172 bits (437), Expect = 2e-43
Identities = 111/366 (30%), Positives = 189/366 (51%), Gaps = 27/366 (7%)
Query: 14 PFIAVVLLQFGYAGMDILSKSALNKGMSCYVLVVYRHAVAFVVIVPFAVILENGSQCARA 73
PF+A+VL+Q GYAGM+I SK A+ GM +LV YR A + P A LE ++
Sbjct: 8 PFLAMVLVQIGYAGMNITSKMAMEAGMKPLILVAYRQIFATIATFPVAFFLERKTRPKIT 67
Query: 74 CY*SKFIFF-------GNEVYNSYF-----------CSCHVQCPSCYYLCRGLDSQMEKI 115
+FF GN+V YF C+ P+ +L + Q E +
Sbjct: 68 LRILVQVFFCSITGATGNQVL--YFVGLQNSSPTIACALTNLLPAVTFLLAAIFRQ-ETV 124
Query: 116 KMRSVHSQAKIVGTIATVAGAMVMTLIKGPILNL--FGIHESSAQ--IQHNGGVNLQHAV 171
++ QAK++GT+ V GAMV++ G + + IH + A+ +H +
Sbjct: 125 GIKKASGQAKVIGTLVCVIGAMVLSFYHGHTIGIGESKIHWAYAENITKHGSSSGHSNFF 184
Query: 172 KGSIMITIGCFSCACFTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEPSVW 231
G +I S A + I+Q ET+ A + T +CL+G+++ G +ALI + S W
Sbjct: 185 LGPFLIMAAAVSWAAWFIIQTKMSETFAAPYTSTLLMCLMGSIQCGAIALISDH-TISDW 243
Query: 232 SLSWDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEK 291
SLS + ++A+Y+G+V S +A+ + M+ +GP++V+ F+PL +V+VAI S +L EK
Sbjct: 244 SLSSPLRFISALYAGVVASALAFCLMSWAMQRKGPLYVSVFSPLLLVVVAIFSWALLEEK 303
Query: 292 IYLGRVIGAVVIILGLYLVVWGKSKDYDRPSPIIKDEILPAKQTIENNDKEKFHSHEVIT 351
+Y G +G+ ++++GLY V+WGK ++ ++++ +++ E S +
Sbjct: 304 LYTGTFMGSALVVIGLYGVLWGKDREVSEKEE-EREKVKQQNHKVKSESNEDIESRLPVA 362
Query: 352 SSNFGA 357
SS G+
Sbjct: 363 SSGNGS 368
>At1g75500 nodulin-like protein
Length = 389
Score = 169 bits (428), Expect = 2e-42
Identities = 109/325 (33%), Positives = 175/325 (53%), Gaps = 25/325 (7%)
Query: 16 IAVVLLQFGYAGMDILSKSALNKGMSCYVLVVYRHAVAFVVIVPFAVILENGSQCARAC- 74
IA++ LQFGYAG ++S++ALN G+S V VYR+ +A ++++PFA LE + A
Sbjct: 22 IAMLTLQFGYAGFHVVSRAALNMGISKLVFPVYRNIIALLLLLPFAYFLEKKERPAITLN 81
Query: 75 Y*SKFIFFG---------------NEVYNSYFCSCHVQCPSCYYLCRGLDSQMEKIKMRS 119
+ +F F + ++ S P+ +L L ++EK+++
Sbjct: 82 FLIQFFFLALIGITANQGFYLLGLDNTSPTFASSMQNSVPAITFLMAAL-LRIEKVRINR 140
Query: 120 VHSQAKIVGTIATVAGAMVMTLIKGPILNLFGIH-------ESSAQIQHNGGVNLQHAVK 172
+KI+GT VAGA V+TL KGP + H +SA + G ++
Sbjct: 141 RDGISKILGTALCVAGASVITLYKGPTIYTPASHLHAHLLTTNSAVLAPLGNAAPKNWTL 200
Query: 173 GSIMITIGCFSCACFTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEPSVWS 232
G I + C S + + + QA L++YPA LS+T++ C G ++ I+A ER + W
Sbjct: 201 GCIYLIGHCLSWSGWLVFQAPVLKSYPARLSVTSYTCFFGIIQFLIIAAFCER-DSQAWV 259
Query: 233 LSWDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEKI 292
+L +Y+GIV SG+A+ +Q + GPVFV + P+ ++VAIM+ L E+
Sbjct: 260 FHSGWELFTILYAGIVASGIAFAVQIWCIDRGGPVFVAVYQPVQTLVVAIMASIALGEEF 319
Query: 293 YLGRVIGAVVIILGLYLVVWGKSKD 317
YLG +IGAV+II GLY V++GKS++
Sbjct: 320 YLGGIIGAVLIIAGLYFVLYGKSEE 344
>At1g43650 nodulin-like protein
Length = 341
Score = 167 bits (422), Expect = 1e-41
Identities = 101/322 (31%), Positives = 173/322 (53%), Gaps = 23/322 (7%)
Query: 13 KPFIAVVLLQFGYAGMDILSKSALNKGMSCYVLVVYRHAVAFVVIVPFAVILENGSQCAR 72
K +A+V +Q YAGM +LSK A+++G + +V V YR A A + + PFA LE+
Sbjct: 4 KANMAMVFVQIVYAGMPLLSKVAISQGTNPFVFVFYRQAFAALALSPFAFFLESSKSSPL 63
Query: 73 ACY*SKFIFFGN---------------EVYNSYFCSCHVQCPSCYYLCRGLDSQMEKIKM 117
+ IFF + E + F + L ++E + +
Sbjct: 64 SFILLLKIFFISLCGLTLSLNLYYVAIENTTATFAAATTNAIPSITFVLALLFRLETVTL 123
Query: 118 RSVHSQAKIVGTIATVAGAMVMTLIKGP-ILNLFGIHESSAQIQHNGGVNLQHAVKGSIM 176
+ H AK+ G++ + GA+V +KGP ++N H +S+ I + + +++VKGSI
Sbjct: 124 KKSHGVAKVTGSMVGMLGALVFAFVKGPSLIN----HYNSSTIPNGTVPSTKNSVKGSIT 179
Query: 177 ITIGCFSCAC-FTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEPSVWSLSW 235
+ + +C C + I+Q+ ++ YPA+L L A CL ++ + A+ + R PSVW + +
Sbjct: 180 M-LAANTCWCLWIIMQSKVMKEYPAKLRLVALQCLFSCIQSAVWAVAVNRN-PSVWKIEF 237
Query: 236 DTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEKIYLG 295
LL+ Y GI+ +G+ Y++Q + +GPVF + PL +++ I+S F+ E YLG
Sbjct: 238 GLPLLSMAYCGIMVTGLTYWLQVWAIEKKGPVFTALYTPLALILTCIVSSFLFKETFYLG 297
Query: 296 RVIGAVVIILGLYLVVWGKSKD 317
V GAV+++ GLYL +WGK+K+
Sbjct: 298 SVGGAVLLVCGLYLGLWGKTKE 319
>At5g64700 nodulin-like protein
Length = 359
Score = 165 bits (418), Expect = 3e-41
Identities = 102/328 (31%), Positives = 168/328 (51%), Gaps = 25/328 (7%)
Query: 13 KPFIAVVLLQFGYAGMDILSKSALNKGMSCYVLVVYRHAVAFVVIVPFAVILENGSQCAR 72
KP++ V ++Q Y M ++SK+ N GM+ +V V YR A A + + P A E S
Sbjct: 7 KPYLMVTIIQVIYTIMFLISKAVFNGGMNTFVFVFYRQAFATIFLAPLAFFFERKSAPPL 66
Query: 73 ACY*SKFIF----FG-------NEVYNSYF-----CSCHVQCPSCYYLCRGLDSQMEKIK 116
+ IF FG N + SY + P+ + L ME++K
Sbjct: 67 SFVTFIKIFMLSLFGVTLSLDLNGIALSYTSATLAAATTASLPAITFFLALLFG-MERLK 125
Query: 117 MRSVHSQAKIVGTIATVAGAMVMTLIKGPILNLFGIHESSAQIQHNGGVNLQHAVKGSIM 176
++S+ AK+VG + G +++ + KGP+L L +H N H GS
Sbjct: 126 VKSIQGTAKLVGITVCMGGVIILAIYKGPLLKLPLCPHFYHGQEHPHRNNPGHVSGGSTS 185
Query: 177 ITIGCFSC-------ACFTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEPS 229
GC + +LQ L+ YP++L T CLL +++ ++A+ +ER + S
Sbjct: 186 WLKGCVLMITSNILWGLWLVLQGRVLKVYPSKLYFTTLHCLLSSIQSFVIAIALER-DIS 244
Query: 230 VWSLSWDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILA 289
W L W+ +L+A +Y G + +G+AYY+Q V+ RGPVF++ F PL ++ + S +L
Sbjct: 245 AWKLGWNLRLVAVIYCGFIVTGVAYYLQSWVIEKRGPVFLSMFTPLSLLFTLLSSAILLC 304
Query: 290 EKIYLGRVIGAVVIILGLYLVVWGKSKD 317
E I LG ++G +++I+GLY V+WGKS++
Sbjct: 305 EIISLGSIVGGLLLIIGLYCVLWGKSRE 332
>At4g30420 nodulin-like protein
Length = 373
Score = 159 bits (403), Expect = 2e-39
Identities = 113/329 (34%), Positives = 173/329 (52%), Gaps = 37/329 (11%)
Query: 16 IAVVLLQFGYAGMDILSKSALNKGMSCYVLVVYRHAVAFVVIVPFAVILENGSQCARACY 75
+A+ ++Q YAG+ + +++ L G+S V ++YR A A + I PF + S+ A +
Sbjct: 1 MAMTMIQLCYAGVTLFARATLVHGLSPRVFILYRQAFATIFIFPFLYLSRRKSKIAISSL 60
Query: 76 *SK---FIFFGNEV-----YNSYFCSCHVQC-----------PSCYYLCRGLDSQMEKIK 116
K IF + + N Y ++ P+ +L L + EK+
Sbjct: 61 DLKSFSLIFLVSLIGITINQNLYLEGLYLTSSSMGSAVGNIIPAITFLISFL-AGYEKLN 119
Query: 117 MRSVHSQAKIVGTIATVAGAMVMTLIKGP-ILNLFGIHESSAQIQHNGGVNLQHAVKGSI 175
+R + AKI GTI VAGA+ MTL++GP ILN ES+ I + L H +K
Sbjct: 120 LRDIRGLAKIAGTILCVAGAISMTLLRGPKILN----SESALPIAKSV---LGH-LKDQN 171
Query: 176 MITIGCF----SCACFT---ILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEP 228
IGC S C++ ILQ YP LSL+AW+CL GT++ +V +E+ +P
Sbjct: 172 TWLIGCLFLFSSTLCWSFWLILQVPISAYYPDNLSLSAWMCLFGTIQCAVVTFFLEK-DP 230
Query: 229 SVWSLSWDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFIL 288
+ W L ++ +Y+GI S +++ +Q + RGPVF FNPLC VIV I++
Sbjct: 231 NAWILHSYSEFATCLYAGIGASALSFTVQAWAIAKRGPVFSALFNPLCTVIVTILAALFF 290
Query: 289 AEKIYLGRVIGAVVIILGLYLVVWGKSKD 317
E+IY G +IG + +ILGLY V+WGK+KD
Sbjct: 291 HEEIYTGSLIGGLGVILGLYTVLWGKAKD 319
>At3g18200 integral membrane protein, putative
Length = 360
Score = 159 bits (401), Expect = 3e-39
Identities = 104/323 (32%), Positives = 165/323 (50%), Gaps = 24/323 (7%)
Query: 13 KPFIAVVLLQFGYAGMDILSKSALNKGMSCYVLVVYRHAVAFVVIVPFAVILENGSQCAR 72
K +A++ LQF +AG I+S+ ALN G+S V VYR+ +A ++I PFA E +
Sbjct: 11 KLVVALITLQFCFAGFHIVSRVALNIGVSKVVYPVYRNLLALLLIGPFAYFFEKKERPPL 70
Query: 73 ACY*SKFIFF---------------GNEVYNSYFCSCHVQCPSCYYLCRGLDSQMEKIKM 117
FF G F S ++E I +
Sbjct: 71 TISLLAQFFFLALIGITANQGFYLLGLYYATPTFASAMQNSVPAITFIMACALRLEHIDL 130
Query: 118 RSVHSQAKIVGTIATVAGAMVMTLIKGPILNLFGIHESSAQIQHN---GGVNLQHAVKGS 174
H AK++GT+ ++ GA V+TL +G F I + +Q G N G
Sbjct: 131 VRKHGVAKVLGTLVSIGGATVITLYRG-----FPIFDQGLNMQKEEVVGSDNSHSLTLGW 185
Query: 175 IMITIGCFSCACFTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEPSVWSLS 234
+ + C S A + +LQA L+ YPA+L+LT++ C G ++ ++AL +E + +S
Sbjct: 186 LYLMGHCLSWAGWMVLQAPVLKQYPAKLTLTSFTCFFGLIQFLVIALFVETDLNNWIIVS 245
Query: 235 WDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEKIYL 294
W+ +L +Y+GI+ SG+ Y+Q + GPVFV F PL ++VA M+ IL +++Y
Sbjct: 246 WE-ELFTILYAGIIASGLVVYLQTWCIYKSGPVFVAVFQPLQTLLVAAMAFLILGDQLYS 304
Query: 295 GRVIGAVVIILGLYLVVWGKSKD 317
G ++GAV I+LGLYLV+WGK+++
Sbjct: 305 GGIVGAVFIMLGLYLVLWGKNEE 327
>At3g30340 unknown protein
Length = 364
Score = 155 bits (392), Expect = 3e-38
Identities = 102/335 (30%), Positives = 169/335 (50%), Gaps = 19/335 (5%)
Query: 12 WKPFIAVVLLQFGYAGMDILSKSALNKGMSCYVLVVYRHAVAFVVIVPFAVILENGSQ-- 69
WK + + ++ G + ++++ K +++G++ V YR AV + ++PFA+ LE ++
Sbjct: 9 WKAVLMMSMINIGLSVVNVMFKKMIDEGLNRMVATTYRLAVGTLFLIPFAIFLERHNRPK 68
Query: 70 ------CAR-------ACY*SKFIFFGNEVYNSYFCSCHVQCPSCYYLCRGLDSQMEKIK 116
C+ F G E +S F L + E +
Sbjct: 69 LTGRILCSLFFSALLGTSLVQYFFLIGLEYTSSTFSLAFSNMVPSVTFALALVFRQETLN 128
Query: 117 MRSVHSQAKIVGTIATVAGAMVMTLIKGPILNL-FGIHESSAQIQHNGGVNLQHAVKGSI 175
++S +AK++GT+ + GA+V+TL KG L+ H + + G Q GSI
Sbjct: 129 IKSNVGRAKLLGTMICICGALVLTLYKGTALSREHSTHMETHTRTDSTGAMTQKWAMGSI 188
Query: 176 MITIGCFSCACFTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEPSVWSLSW 235
M+ I + + I+QA YP + + T + G ++ +++LI ER S+W +
Sbjct: 189 MLVISIIIWSSWFIVQAKISRVYPCQYTSTTILSFFGVIQSALLSLISERST-SMWVVKD 247
Query: 236 DTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEKIYLG 295
++LA +YSGIV SG+ Y +R RG VF ++F PL V AI S L E+IY G
Sbjct: 248 KFQVLALLYSGIVGSGLCYVGMSWCLRQRGAVFTSSFIPLIQVFAAIFSFSFLHEQIYCG 307
Query: 296 RVIGAVVIILGLYLVVWGKSKDYDRPSPIIKDEIL 330
VIG++VII+GLY+++WGKSK D+ + + K E L
Sbjct: 308 SVIGSMVIIVGLYILLWGKSK--DKSASVTKQEPL 340
>At3g53210 unknown protein
Length = 369
Score = 148 bits (373), Expect = 5e-36
Identities = 111/359 (30%), Positives = 183/359 (50%), Gaps = 33/359 (9%)
Query: 13 KPFIAVVLLQFGYAGMDILSKSALNKGMSCYVLVVYRHAVAFVVIVPFAVILENGSQCAR 72
K IA+V+ Q GYAG ++ + ALN G+S V +YR VAF V+ P A LE + A
Sbjct: 9 KLHIAMVVFQTGYAGNHVIMRYALNLGVSKLVFPLYRTIVAFSVLAPSAYFLEKKERPAM 68
Query: 73 AC---------------Y*SKFIFFGNEVYNSYFCSCHVQC-PSCYYLCRGLDSQMEKIK 116
F FG + + F S P+ +L L +EK++
Sbjct: 69 KISFLIQFFLLGLVGITLNQGFYIFGLDNTSPTFASATENVVPAVSFLMAALLG-IEKVE 127
Query: 117 MRSVHSQAKIVGTIATVAGAMVMTLIKGPILNLFGIHESSAQIQHNGGVNLQHAVKGSIM 176
+ AK+VGTI +VAG++V+TL KGP I++ S I N + + A + +
Sbjct: 128 WKRKDGIAKVVGTIVSVAGSLVITLYKGPT-----IYQPSLNIV-NQTIKPEEAEEENKN 181
Query: 177 ITIGCFS----CACFT---ILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEPS 229
T+GC C C++ +LQ+ L+ YPA S ++ C ++ ++ ER +
Sbjct: 182 WTLGCLCLMGHCLCWSSWIVLQSPLLKKYPARFSFVSYSCFFAVIQFFGISAYFER-DLE 240
Query: 230 VWSLSWDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILA 289
W + +L A +Y+G+V S M + IQ V+ GP+FV+ + PL +I A+++ L
Sbjct: 241 RWKIISGGELYALLYTGLVGSAMVFAIQIYVVERGGPLFVSAYLPLQTLIAAVLATLALG 300
Query: 290 EKIYLGRVIGAVVIILGLYLVVWGKSKDYDRPSPIIKDEILPAKQTIENNDKEKFHSHE 348
E YLG +IGA++I+ GLYLVV GKS ++ + + + + + + D+E +H+++
Sbjct: 301 EHFYLGGLIGAILIMSGLYLVVMGKS--WENQALCQQQQHMISSAASDFGDEEDYHNNK 357
>At5g40240 unknown protein
Length = 368
Score = 133 bits (334), Expect = 2e-31
Identities = 88/329 (26%), Positives = 158/329 (47%), Gaps = 33/329 (10%)
Query: 14 PFIAVVLLQFGYAGMDILSKSALNKGMSCYVLVVYRHAVAFVVIVPFAVILENGSQ--CA 71
PF A+ ++ G + L K+A +G+S YV V Y + V+ ++++P +VI + A
Sbjct: 20 PFAAMFAVECATVGSNTLFKAATLRGLSFYVFVFYSYIVSTLLLLPLSVIFGRSRRLPAA 79
Query: 72 RACY*SKFIFFGNEVYNSYFCSCHVQCPSCYYLCRGLDS-------------QMEKIKMR 118
++ K G + S C S L + + +ME++++R
Sbjct: 80 KSPLFFKIFLLGLVGFMSQIAGCKGIAYSSPTLASAISNLTPAFTFTLAVIFRMEQVRLR 139
Query: 119 SVHSQAKIVGTIATVAGAMVMTLIKGP----------ILNLFGIHESSAQIQHNGGVNLQ 168
S +QAKI+G I +++GA+V+ L KGP +L +H+ I+ +
Sbjct: 140 SSATQAKIIGAILSISGALVVVLYKGPQVLASASFTTVLPTVTLHQQLTSIESSW----- 194
Query: 169 HAVKGSIMITIGCFSCACFTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEP 228
+ G +++ F + + ILQ +E YP E+++ + L T+ V L E
Sbjct: 195 --IIGGLLLASQYFLISVWYILQTRVMEVYPEEITVVFFYNLFATLISVPVCLFAESNLT 252
Query: 229 SVWSLSWDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFIL 288
S W L D L A +YSG+ S + + +GPV+++ F PL + I M L
Sbjct: 253 S-WVLKPDISLAAIIYSGVFVSLFSALTHTWGLHLKGPVYISLFRPLSIAIAVAMGAIFL 311
Query: 289 AEKIYLGRVIGAVVIILGLYLVVWGKSKD 317
+ ++LG VIG++++ +G Y V+WGK+++
Sbjct: 312 GDALHLGSVIGSMILCIGFYTVIWGKARE 340
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.328 0.140 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,640,327
Number of Sequences: 26719
Number of extensions: 302196
Number of successful extensions: 1022
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 52
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 841
Number of HSP's gapped (non-prelim): 85
length of query: 363
length of database: 11,318,596
effective HSP length: 101
effective length of query: 262
effective length of database: 8,619,977
effective search space: 2258433974
effective search space used: 2258433974
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC135797.11


