

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135796.7 - phase: 0
(277 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
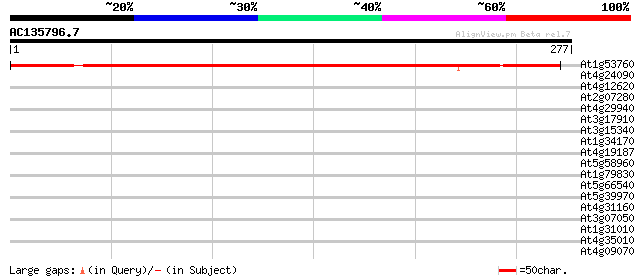
Score E
Sequences producing significant alignments: (bits) Value
At1g53760 hypothetical protein 305 2e-83
At4g24090 unknown protein 31 0.83
At4g12620 origin recognition complex subunit 1 -like protein 31 0.83
At2g07280 hypothetical protein 30 1.1
At4g29940 pathogenesis related homeodomain protein (PRHA) 29 2.4
At3g17910 putative surfeit 1 protein 29 2.4
At3g15340 hypothetical protein 29 3.2
At1g34170 putative protein 29 3.2
At4g19187 unknown protein 28 4.1
At5g58960 unknown protein 28 5.4
At1g79830 unknown protein (At1g79830) 28 5.4
At5g66540 unknown protein 28 7.0
At5g39970 predicted GPI-anchored protein 28 7.0
At4g31160 unknown protein 28 7.0
At3g07050 GTPase like protein 28 7.0
At1g31010 unknown protein 28 7.0
At4g35010 beta-galactosidase - like protein 27 9.2
At4g09070 putative protein 27 9.2
>At1g53760 hypothetical protein
Length = 272
Score = 305 bits (781), Expect = 2e-83
Identities = 151/273 (55%), Positives = 197/273 (71%), Gaps = 6/273 (2%)
Query: 1 MKAKLIVFPIRGRNWCFTRSIDHTLPASSSTADFSQSPSTLKQLWKSINTGDKPFNAKAE 60
M+A+L+VFPI+G+ WCF+RS+D S S +P+T++ LWK I++ KP NA AE
Sbjct: 1 MRARLVVFPIKGKKWCFSRSVDPFAAQSPSGV----TPTTVRGLWKKISSESKPINANAE 56
Query: 61 LFTDYVANKMNNGWVTLENAPDGSFKKKIHGLGLWLLSRVKPSEIFLKSISKDVTGVEVV 120
L D++++KMN WV LE APDGS K KIHG GL LL+RVKPSEIFLKSISK+VT V+V
Sbjct: 57 LLVDFISDKMNKAWVGLEKAPDGSIKNKIHGFGLKLLARVKPSEIFLKSISKEVTSVQVT 116
Query: 121 YPSSMNARLVRRRLRHIAMRGTIIHRKFFYGSVSLIPLSSAFSILPLPNVPFFWILFRSY 180
YP S++ RLVRRRLRHIAM GTI+H+K+ GSV+L+PL+SAF +LPLPN+PFFW+LFR+Y
Sbjct: 117 YPPSLDPRLVRRRLRHIAMSGTILHKKYLVGSVTLLPLTSAFMVLPLPNIPFFWVLFRTY 176
Query: 181 SHWRALQGSEKLFQLVSDGSQSSNTYSGKKETEHEDSENE-SLGLDEPHWVLTPSKELEN 239
SHWRALQGSEKL +L+S+ + S E ++S + P +L PS+EL
Sbjct: 177 SHWRALQGSEKLLKLISNEANPDKPDSTDDADESKNSNTKPEQKSQSPTCILLPSEELYQ 236
Query: 240 IVRQEDGNDGLSRGTIEEICKIYDLNTQDVVKY 272
++R E +GL TI EICK +DLN DV+KY
Sbjct: 237 LIR-EASEEGLDEATIIEICKSFDLNKNDVLKY 268
>At4g24090 unknown protein
Length = 308
Score = 30.8 bits (68), Expect = 0.83
Identities = 25/75 (33%), Positives = 36/75 (47%), Gaps = 7/75 (9%)
Query: 177 FRSYSHWRALQGSEKLFQLVSDGSQSSNTYSGKKETEHED----SENESL--GLDEPHW- 229
+R +WR SE + SDG N+ SG + E+ EN+ L GL++
Sbjct: 36 YRRTRNWRIRSSSEDNVAISSDGGDLKNSLSGIVGNQVEELLSREENKGLLDGLEKASLR 95
Query: 230 VLTPSKELENIVRQE 244
V +ELE+I RQE
Sbjct: 96 VEIAKRELEDIERQE 110
>At4g12620 origin recognition complex subunit 1 -like protein
Length = 813
Score = 30.8 bits (68), Expect = 0.83
Identities = 14/39 (35%), Positives = 23/39 (58%), Gaps = 3/39 (7%)
Query: 182 HWRALQGSEKLFQLVSDGSQSSNTYSGKKETEHEDSENE 220
HWR+ + +L +L S S ++G+KE E +DS+ E
Sbjct: 336 HWRSFK---RLAELADGDSDSDQEWNGRKEEEVDDSDEE 371
>At2g07280 hypothetical protein
Length = 401
Score = 30.4 bits (67), Expect = 1.1
Identities = 17/50 (34%), Positives = 26/50 (52%)
Query: 201 QSSNTYSGKKETEHEDSENESLGLDEPHWVLTPSKELENIVRQEDGNDGL 250
+S NT + ++E+ D E+E +GLD P + P K+ D N GL
Sbjct: 256 ESFNTALTEHQSENTDKESEEVGLDVPTVISQPLKDKPQEFPSVDINIGL 305
>At4g29940 pathogenesis related homeodomain protein (PRHA)
Length = 796
Score = 29.3 bits (64), Expect = 2.4
Identities = 21/81 (25%), Positives = 36/81 (43%), Gaps = 3/81 (3%)
Query: 186 LQGSEKLFQLVSDGSQSSNTYSGKKETEHEDSENESLGLDEPHWVLTPSKELENIVRQED 245
L+ ++ F LV QSS +T E++E ES + EPH L+ L+ +++
Sbjct: 594 LEQNDSSFVLVPHEKQSSEI---SLKTAVEENETESKMMKEPHEELSSEMSLKTAAEEKE 650
Query: 246 GNDGLSRGTIEEICKIYDLNT 266
+ EE+ + L T
Sbjct: 651 TESKMIEEPHEELSREMSLKT 671
>At3g17910 putative surfeit 1 protein
Length = 354
Score = 29.3 bits (64), Expect = 2.4
Identities = 35/160 (21%), Positives = 60/160 (36%), Gaps = 22/160 (13%)
Query: 29 SSTADFSQSPSTLKQLWK--------SINTGDKPFNAKAELFTDYVANKMNNGWVTLENA 80
S+T S SPS KQ W S ++ +++ NK + W L
Sbjct: 20 STTTSISASPSLPKQFWSRHFSAVADSSSSSSAALGSQSSSSAPPQENKRGSKWSQLLLF 79
Query: 81 PDGSFKKKIHGLGLWLLSR----VKPSEIFLKSISKDVTGVEVVYPSSMNARLVRRRLRH 136
G+ GLG W + R K E + ++ + + + +P N + R
Sbjct: 80 LPGAI---TFGLGSWQIVRREEKFKTLEYQQQRLNMEPIKLNIDHPLDKNLNAL--EFRR 134
Query: 137 IAMRGTIIHRKFFY-----GSVSLIPLSSAFSILPLPNVP 171
++ +G ++ Y S+S I + F I PL +P
Sbjct: 135 VSCKGVFDEQRSIYLGPRSRSISGITENGFFVITPLMPIP 174
>At3g15340 hypothetical protein
Length = 478
Score = 28.9 bits (63), Expect = 3.2
Identities = 12/32 (37%), Positives = 19/32 (58%)
Query: 214 HEDSENESLGLDEPHWVLTPSKELENIVRQED 245
H E+ S+G +E WVL +++ + IV ED
Sbjct: 166 HYQIEHGSIGFEEEDWVLKETEKADGIVLSED 197
>At1g34170 putative protein
Length = 505
Score = 28.9 bits (63), Expect = 3.2
Identities = 14/37 (37%), Positives = 20/37 (53%), Gaps = 4/37 (10%)
Query: 21 IDHTLPASSSTADFSQSPSTLKQLWKSINTGDKPFNA 57
I+H +P+S D SQS +TL LW G + N+
Sbjct: 298 IEHLIPSS----DISQSSATLSNLWTCQEIGQRSMNS 330
>At4g19187 unknown protein
Length = 595
Score = 28.5 bits (62), Expect = 4.1
Identities = 18/62 (29%), Positives = 33/62 (53%), Gaps = 2/62 (3%)
Query: 179 SYSHWRALQGSEK-LFQLVSDGSQSSNTYSGKKETEHEDSENESLGLDEPHWVLTPSKEL 237
S H R+++G EK +++ D + S+ Y K+TE DSE+ + H + + +E
Sbjct: 523 SEKHHRSVKGKEKHVYEASDDPEEFSDRYRSTKKTE-SDSESNRRSRKKKHELSSEEEEG 581
Query: 238 EN 239
E+
Sbjct: 582 ES 583
>At5g58960 unknown protein
Length = 484
Score = 28.1 bits (61), Expect = 5.4
Identities = 14/55 (25%), Positives = 29/55 (52%)
Query: 209 KKETEHEDSENESLGLDEPHWVLTPSKELENIVRQEDGNDGLSRGTIEEICKIYD 263
++E + E+ ENE G + L + +R ++G +G+S T++E+ + D
Sbjct: 36 REEYDTEEEENEEEGEIQYEDALEKENGKDETIRAKNGRNGVSVETVQEMEMVMD 90
>At1g79830 unknown protein (At1g79830)
Length = 918
Score = 28.1 bits (61), Expect = 5.4
Identities = 16/49 (32%), Positives = 26/49 (52%)
Query: 185 ALQGSEKLFQLVSDGSQSSNTYSGKKETEHEDSENESLGLDEPHWVLTP 233
+LQ +EKL S SQ S +E++ EDSE + + ++ V +P
Sbjct: 174 SLQPNEKLEMTASQDSQPEQPKSEAEESQPEDSEAKEVTVENKDTVHSP 222
>At5g66540 unknown protein
Length = 524
Score = 27.7 bits (60), Expect = 7.0
Identities = 21/67 (31%), Positives = 29/67 (42%), Gaps = 4/67 (5%)
Query: 198 DGSQSSNTYSGKKETEHEDSENESLGLDEPHWVLTPSKELENIVRQEDGNDGLSRGTIEE 257
DG S + KE E DSE G DE +E E +E+ DG + G ++
Sbjct: 117 DGFDSDDVDDEDKEIESNDSE----GEDEEEEEEDEEEEEEEEEEEEEEKDGDNEGIEDK 172
Query: 258 ICKIYDL 264
KI +L
Sbjct: 173 FFKIKEL 179
>At5g39970 predicted GPI-anchored protein
Length = 677
Score = 27.7 bits (60), Expect = 7.0
Identities = 15/47 (31%), Positives = 24/47 (50%), Gaps = 1/47 (2%)
Query: 14 NWCFTRSIDHTLPASSSTADFSQSPSTLKQLWKSINTGDKPFNAKAE 60
N C + S+ +T P +S D ST+ ++WKS N K F ++
Sbjct: 117 NECQSLSVTNT-PFASQAGDGGNITSTISEIWKSSNDFCKIFGGASD 162
>At4g31160 unknown protein
Length = 1846
Score = 27.7 bits (60), Expect = 7.0
Identities = 14/51 (27%), Positives = 24/51 (46%)
Query: 199 GSQSSNTYSGKKETEHEDSENESLGLDEPHWVLTPSKELENIVRQEDGNDG 249
G N+ SG + ED+E+ DE +L +E ++ E+ +DG
Sbjct: 1764 GLAGDNSDSGDDDLSSEDNEDSVSDFDEEADILIDGDFMEELIEGENEDDG 1814
>At3g07050 GTPase like protein
Length = 582
Score = 27.7 bits (60), Expect = 7.0
Identities = 14/57 (24%), Positives = 28/57 (48%)
Query: 209 KKETEHEDSENESLGLDEPHWVLTPSKELENIVRQEDGNDGLSRGTIEEICKIYDLN 265
KKE E ++ LGL + T + +E++ + + D R +E+ K+ +L+
Sbjct: 81 KKEARKERAKKRKLGLVDDEDTKTEGETIEDLPKVVNVRDNSERAFYKELVKVIELS 137
>At1g31010 unknown protein
Length = 360
Score = 27.7 bits (60), Expect = 7.0
Identities = 29/93 (31%), Positives = 39/93 (41%), Gaps = 15/93 (16%)
Query: 39 STLKQLWKSINTGDKPFNAKAELFTDYVANKMNNGW---VTLENAPDGSFKKKIHGLGLW 95
S LK + I K A E + D V N MN W V FK K G+GLW
Sbjct: 273 SELKDVKFDIPKYAKQPKAGEESWKDLVDN-MNKWWDNRVDKRTPKSPDFKHKETGVGLW 331
Query: 96 L-------LSRVKPSEIFLKSISKDVTGVEVVY 121
L L ++ P KS + D+ GV+ ++
Sbjct: 332 LSDSPSWVLEKLPPP----KSKTSDIYGVQEMF 360
>At4g35010 beta-galactosidase - like protein
Length = 831
Score = 27.3 bits (59), Expect = 9.2
Identities = 16/64 (25%), Positives = 32/64 (50%), Gaps = 4/64 (6%)
Query: 174 WILFRSYSHWRALQGSEKLFQLVSDGSQSSNTYSGKKETEHEDSENESLGLDEPHWVLTP 233
WILF++ R + L L+S SS++++ KK+ + + N+ + D ++
Sbjct: 10 WILFQTRKFLRKPENLTVLVVLLS----SSSSFAAKKDAKKKKKSNKEVTYDGTSLIIDG 65
Query: 234 SKEL 237
+EL
Sbjct: 66 KREL 69
>At4g09070 putative protein
Length = 219
Score = 27.3 bits (59), Expect = 9.2
Identities = 14/42 (33%), Positives = 24/42 (56%), Gaps = 5/42 (11%)
Query: 170 VPFFWILFRSYSHWRALQGSEKLFQLVSDGSQSSNTYSGKKE 211
+P W+L HW+ QGS Q++++ +QS + +G KE
Sbjct: 1 MPVKWLL-----HWQPNQGSTFSSQILNEVTQSIESLNGVKE 37
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.134 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,480,365
Number of Sequences: 26719
Number of extensions: 283570
Number of successful extensions: 953
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 941
Number of HSP's gapped (non-prelim): 21
length of query: 277
length of database: 11,318,596
effective HSP length: 98
effective length of query: 179
effective length of database: 8,700,134
effective search space: 1557323986
effective search space used: 1557323986
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC135796.7


