

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135566.1 + phase: 0 /pseudo
(437 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
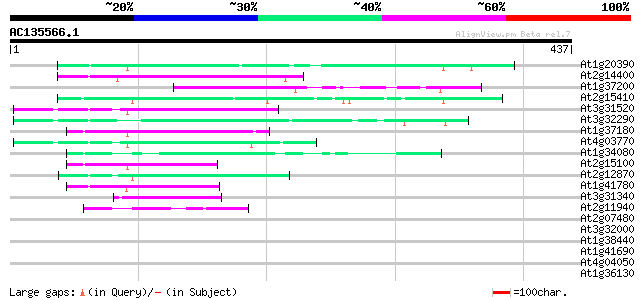
Score E
Sequences producing significant alignments: (bits) Value
At1g20390 hypothetical protein 79 6e-15
At2g14400 putative retroelement pol polyprotein 78 1e-14
At1g37200 hypothetical protein 70 3e-12
At2g15410 putative retroelement pol polyprotein 64 1e-10
At3g31520 hypothetical protein 64 2e-10
At3g32290 hypothetical protein 61 1e-09
At1g37180 hypothetical protein 60 3e-09
At4g03770 hypothetical protein 58 1e-08
At1g34080 putative protein 56 4e-08
At2g15100 putative retroelement pol polyprotein 49 4e-06
At2g12870 putative retroelement pol polyprotein 49 7e-06
At1g41780 hypothetical protein 44 1e-04
At3g31340 Athila ORF 1, putative 44 2e-04
At2g11940 putative retroelement gag/pol polyprotein 42 9e-04
At2g07480 F9A16.15 39 0.004
At3g32000 unknown protein 39 0.007
At1g38440 hypothetical protein 36 0.037
At1g41690 hypothetical protein 35 0.062
At4g04050 putative transposon protein 35 0.082
At1g36130 hypothetical protein 35 0.082
>At1g20390 hypothetical protein
Length = 1791
Score = 78.6 bits (192), Expect = 6e-15
Identities = 82/404 (20%), Positives = 153/404 (37%), Gaps = 61/404 (15%)
Query: 38 LDEPEPQPFSAKIWNAPVPDNFKPPHLPTFDGRGDPSEHVTVFNTRMSVYGVA----DSL 93
L+E + PF+ +I N + K L +++GR DP E +T FN ++ + D+
Sbjct: 149 LEETQQTPFTRRITNVSIRGAQKIK-LESYNGRNDPKEFLTSFNVAINRAELTIDNFDAG 207
Query: 94 KCKLLAGTFADAALRWYMSLPCFSIMGYQDMIKKFTQQFSGTRHRKVLSTSLFNVRQGPS 153
+C++ A W+ L SI + + F + ++ + + L+++ QG
Sbjct: 208 RCQIFIEHLTGPAHNWFSRLKPNSIDSFHQLTSSFLKHYAPLIENQTSNADLWSISQGAK 267
Query: 154 ESLREYLARFNDSTIKVSNPNQEVFVGAFQNGLW-AGQFNESIAQKPADSMEEIIARAEC 212
ESLR ++ RF ++ P++ V A +N +W +F + I ++E+ + RA
Sbjct: 268 ESLRSFVDRFKLVVTNITVPDEAAIV-ALRNAVWYDSRFRDDITLHAPSTLEDALHRASR 326
Query: 213 YVKGEESNAEKKAREAKERGNSGGERRNHYVPPNRDRGTFK-KPYERNQNRYVPEHFTPL 271
+++ EE EK K P +D K P + N+ R +
Sbjct: 327 FIELEE---EKLILARKHNSTK--------TPACKDAVVIKVGPDDSNEPRQHLDRNPSA 375
Query: 272 NTRPERILKEVFESKIIPPPPFSRARVMGQDKDAWCKYHLVQGHNTDDCVHLKREIEKLL 331
+P L P + R +C+YH + H+T++C L+ +
Sbjct: 376 GRKPTSFLVSTETPDAKPWNKYIRDADSPAAGPMYCEYHKSRAHSTENCRFLQGLLMAKY 435
Query: 332 LSGKL---------------RGYAKERRHNERQEDKPNPEQK------------------ 358
SG + R R++ Q P P ++
Sbjct: 436 KSGGITIECDRPPINNKNQRRNETTARQYLNDQTKPPTPAEQGIITSADDPAAKRQRNGK 495
Query: 359 ---------HTLHTISGGFAGGGESSNSMKKYARQVMLLGDGPS 393
+H I GG +S S+K+Y ++ ++ PS
Sbjct: 496 AIAAEPVVVRQVHVIMGGLQNCSDSVRSIKQYRKKAEMVVAWPS 539
>At2g14400 putative retroelement pol polyprotein
Length = 1466
Score = 77.8 bits (190), Expect = 1e-14
Identities = 50/199 (25%), Positives = 104/199 (52%), Gaps = 8/199 (4%)
Query: 38 LDEPEPQPFSAKIWNAPVPDNFKPPHLPTFDGRGDPSEHVTVFNT---RMSVYGVADSLK 94
+++ PF+ +I + + D+ K +L +++G DP ++ F R+ + + ++
Sbjct: 137 IEKARQTPFTPQITSLRIRDSRKL-NLESYNGLEDPKGYLAAFLIAAGRVDLNEADEDVR 195
Query: 95 -CKLLAGTFADAALRWYMSLPCFSIMGYQDMIKKFTQQFSGTRHRKVLSTSLFNVRQGPS 153
CKL + AL W+ L SI + ++ F +Q+S + + T L+N+ QGP+
Sbjct: 196 YCKLFSENLCGQALMWFTQLEPGSISNFNELSVVFLKQYSILMDKSISDTDLWNLSQGPN 255
Query: 154 ESLREYLARFNDSTIKVSNPNQEVFVGAFQNGLW-AGQFNESIAQKPADSMEEIIARAEC 212
E+LR ++ +F K+S +Q+ + A + GLW +F E + D++++ + RA
Sbjct: 256 ETLRAFITKFKYVLSKLSRISQQSALSALRKGLWYDSRFKEDLILHKLDTIQDALFRANN 315
Query: 213 Y--VKGEESNAEKKAREAK 229
+ V+ E+ + K+ ++AK
Sbjct: 316 WMEVEDEKESFAKRDKQAK 334
>At1g37200 hypothetical protein
Length = 1564
Score = 69.7 bits (169), Expect = 3e-12
Identities = 64/247 (25%), Positives = 110/247 (43%), Gaps = 45/247 (18%)
Query: 128 FTQQFSGTRHRKVLSTSLFNVRQGPSESLREYLARFNDSTIKVSNPNQEVFVGAFQNGLW 187
F +Q S ++ L+++ +ES R Y+ +F K++N N+EV + A +NGLW
Sbjct: 176 FLKQDSVLIEQRTSEADLWSLTLLQNESHRSYVEKFKAIKSKIANLNEEVAIAALRNGLW 235
Query: 188 -AGQFNESIAQKPADSMEEIIARAECYVKGEESNA--EKKAREAKERGNSGGERRNHYVP 244
+ +F E + + S+++ + +A + KGEE A K +E+K +
Sbjct: 236 FSSRFREELTVRQPISLDDALHKALHFAKGEEELAVLALKFKESKTQN-----------A 284
Query: 245 PNRDRGTFKKPYERNQNRYVPEHFTPLNTRPERILKEVFESKIIPPPPFSRARVMGQDKD 304
P + FKK NQ T+ + L + E+ P D
Sbjct: 285 PLAIKTPFKK---ENQ------------TQGQHTLFAIEEAAEDESPEL--------DLG 321
Query: 305 AWCKYHLVQGHNTDDCVHLKREIEKLLLSG----KLRGYAKERRHNERQEDKPNPEQKHT 360
+CKYH +GH+T++C R ++KL+ +G K E + QE++ P+QK
Sbjct: 322 KYCKYHNKRGHSTEEC----RAVKKLIAAGGKTKKGSNPKVETPPPDEQEEEQTPKQKKR 377
Query: 361 LHTISGG 367
T+ GG
Sbjct: 378 ERTLEGG 384
>At2g15410 putative retroelement pol polyprotein
Length = 1787
Score = 64.3 bits (155), Expect = 1e-10
Identities = 83/373 (22%), Positives = 148/373 (39%), Gaps = 35/373 (9%)
Query: 38 LDEPEPQPFSAKIWNAPVPDNFKPPHLPTFDGRGDPSEHVTVFNTRMSVYGVADSLK--- 94
L+E + PF++ I + + + L ++G DP +T + + +D +
Sbjct: 230 LEETQRSPFTSCISDVRIR-HISKIKLANYEGLVDPRPFLTSVSIAIGRAHFSDEDRDAG 288
Query: 95 -CKLLAGTFADAALRWYMSLPCFSIMGYQDMIKKFTQQFSGTRHRKVLSTSLFNVRQGPS 153
C+L + AAL W+ L SI + F + + + + L+ + Q
Sbjct: 289 SCQLFVKHLSGAALTWFSRLEANSIDSVHALTTSFLKNYGVFMEKGASNVDLWTMAQTAK 348
Query: 154 ESLREYLARFNDSTIKVSNPNQEVFVGAFQNGLWAGQFNESIAQ-KP---ADSMEEIIAR 209
ESLR ++ RF + V+ P+ + + A +N LW G + KP S + IA+
Sbjct: 349 ESLRSFIGRFKEIVTSVATPD-DAAIAALRNALWHGDSSRCTTLCKPFHRQGSAKAPIAK 407
Query: 210 AECYVKGEESNAEKKAREAKERGNSGGERRNHYVPPNRDRGTFKKPYER---------NQ 260
+ ES A AKE G ++YV R KP+ + +Q
Sbjct: 408 EKPKEGHHESRQHYDADYAKEEKLKKGT--SYYVGDTAPRSD--KPWNKWNRSADAKGDQ 463
Query: 261 NRY----VPEHFTPLNTRPERILKEVFESKIIPPPPFSRARVMGQDKDAWCKYHLVQGHN 316
Y +P H T + + +L F+ + P R R D+D + V+ +
Sbjct: 464 KYYEFHKIPGHSTDECRQLQTLLLTKFKKGDLDIEP-DRRRTGTNDRDN--THRRVENTD 520
Query: 317 TDDCVHLKREIEKLLLSGKL-----RGYAKERRHNERQEDKPNPEQKHTLHTISGGFAGG 371
D+ +R+ E + ++ R +R H+ ++ + P K + I GG
Sbjct: 521 RDNQDDKRRQDETDRRAERIHDDDRRAEPHKRNHDAPRQFEDEPASKRRRNMIMGGLTAC 580
Query: 372 GESSNSMKKYARQ 384
+S S+K Y RQ
Sbjct: 581 RDSVRSIKSYRRQ 593
>At3g31520 hypothetical protein
Length = 528
Score = 63.9 bits (154), Expect = 2e-10
Identities = 55/215 (25%), Positives = 88/215 (40%), Gaps = 18/215 (8%)
Query: 4 QIALLLEQNEALLASVETIQQVQQQEKTDSHHGDLDEPEPQPFSAKIWNAPVPDNFKPPH 63
+++L +NE L + V D L+E PF+ +I NA + D K
Sbjct: 188 KLSLAKAENEMKLVRSQMHNAVSSAPNIDRI---LEESHNTPFTHRISNAIISDPGKL-R 243
Query: 64 LPTFDGRGDPSEHVTVFNTRMSVYGVA---------DSLKCKLLAGTFADAALRWYMSLP 114
+ F+G DP H+ F + VA D+ C L AL W+ L
Sbjct: 244 IEYFNGSSDPKRHLKSF-----IISVARTKFRPEERDAGLCHLFVEHLKGPALCWFSRLE 298
Query: 115 CFSIMGYQDMIKKFTQQFSGTRHRKVLSTSLFNVRQGPSESLREYLARFNDSTIKVSNPN 174
S+ +Q++ F +Q+S L+++ Q P+E LR++LA+ + KV N
Sbjct: 299 GNSMDSFQELSTLFLKQYSVLIDLGTSDADLWSLSQQPNEPLRDFLAKLRSTLAKVEGIN 358
Query: 175 QEVFVGAFQNGLWAGQFNESIAQKPADSMEEIIAR 209
+ A + LW + S K S E II +
Sbjct: 359 DVAALSALKKALWVLKAGHSFTMKDEISPEPIITK 393
>At3g32290 hypothetical protein
Length = 612
Score = 60.8 bits (146), Expect = 1e-09
Identities = 80/374 (21%), Positives = 140/374 (37%), Gaps = 52/374 (13%)
Query: 4 QIALLLEQNEALLASVETIQQVQQQEKTDSHHGDLDEPEPQPFSAKIWNAPVPDNFKPPH 63
+++L +NE L + V D L+E PF+ I NA + D K
Sbjct: 126 KLSLAKAENEMKLVRSQMHNAVSSAPNIDRI---LEESHNTPFTHMISNAIISDPGKL-R 181
Query: 64 LPTFDGRGDPSEHVTVFNTRMSVYGVADSLKCKLLAGTFADAALRWYMSLPCFSIMGYQD 123
+ F+G DP H+ F + A A R S+ +Q+
Sbjct: 182 IEYFNGSSDPKGHLKSFIISV------------------ARAKFRQEERDAGNSVDSFQE 223
Query: 124 MIKKFTQQFSGTRHRKVLSTSLFNVRQGPSESLREYLARFNDSTIKVSNPNQEVFVGAFQ 183
+ F +Q+S L+++ Q P+E LR++LA+F + KV N + A +
Sbjct: 224 LSTLFLKQYSVLIDLGTSDADLWSLSQQPNEPLRDFLAKFRSTLAKVEGINDVAALSALK 283
Query: 184 NGLW-AGQFNESIAQKPADSMEEIIARAECYVKGEESNAEKKAREAKERGNSGGERRNHY 242
LW +F + + ++ + + RA YV EE E A+ + + ++ +
Sbjct: 284 KALWYKSEFRKELNLSKPLTIRDALHRASDYVSHEE-EMELVAKRHEPSKQTPRIDKSQH 342
Query: 243 VPPNRDRGTFKKPYERNQNR-YVPEHFTPLNTRPERILKEVFESKIIPPPPFSRARVMGQ 301
PN +G + ++ R + H N + + E + R R G+
Sbjct: 343 SAPNHKKGAQGWTFVHHEGRNFSGAH----NYKADTPRGEAARGR-----GRGRGRGRGR 393
Query: 302 DKDAW------------CKYHLVQGHNTDDCVHLKREIEKLLLSGKLRG------YAKER 343
+ W C+ H GH+T C L ++ L+G++ G E+
Sbjct: 394 ESYTWTKDQPAGNEQEYCELHKSYGHHTSRCRSLGAKLVAKFLAGEIGGGLTIEDLEAEK 453
Query: 344 RHNERQEDKPNPEQ 357
E+ NPEQ
Sbjct: 454 GKTEQVNAVANPEQ 467
>At1g37180 hypothetical protein
Length = 661
Score = 59.7 bits (143), Expect = 3e-09
Identities = 40/162 (24%), Positives = 76/162 (46%), Gaps = 6/162 (3%)
Query: 45 PFSAKIWNAPVPDNFKPPHLPTFDGRGDPSEHVTVFNTRMSVYGVA----DSLKCKLLAG 100
PF+ +I + + F+ LP ++G+GDP+EH+T F + + D+ CKL +
Sbjct: 298 PFTPRISKLRIRE-FRDFKLPVYNGKGDPNEHLTSFQVIVRRVPLEPYEEDAGLCKLFSE 356
Query: 101 TFADAALRWYMSLPCFSIMGYQDMIKKFTQQFSGTRHRKVLSTSLFNVRQGPSESLREYL 160
+ AL W+ L SI ++ + F +Q+ + L+N+ Q + LR Y+
Sbjct: 357 NLSGPALTWFTQLEEGSIDNFKQLSTAFIKQYGYFIKSDITEAHLWNLSQSADDPLRTYI 416
Query: 161 ARFNDSTIKVSNPNQEVFVGAFQNGLWAGQFNESIAQKPADS 202
F + +++ + + A +NG ++ + ES P S
Sbjct: 417 IVFKEIMVQIPSLSDSTAQFALKNGTFSVE-KESSKSSPKAS 457
Score = 32.3 bits (72), Expect = 0.53
Identities = 19/54 (35%), Positives = 25/54 (46%), Gaps = 8/54 (14%)
Query: 291 PPFSRARVMG--------QDKDAWCKYHLVQGHNTDDCVHLKREIEKLLLSGKL 336
PPFS G QD +A+C H + GH T DC L R + SG++
Sbjct: 461 PPFSPKGKNGAFPSNKWVQDLNAYCDIHKMNGHATKDCKALGRLLAAKYASGEI 514
>At4g03770 hypothetical protein
Length = 464
Score = 57.8 bits (138), Expect = 1e-08
Identities = 61/247 (24%), Positives = 100/247 (39%), Gaps = 21/247 (8%)
Query: 4 QIALLLEQNEALLASVETIQQVQQQEKTDSHHGDLDEPEPQPFSAKIWNAPVPDNFKPPH 63
+++L +NE L + V D L+E PF+ I NA + D K
Sbjct: 180 KLSLAKAENEMKLVRSQMHNAVSSAPNIDRI---LEESHNTPFTHMISNAIISDPGKL-R 235
Query: 64 LPTFDGRGDPSEHVTVFNTRMSVYGVA---------DSLKCKLLAGTFADAALRWYMSLP 114
+ F+G DP H+ +F + VA D+ C L AL W+ L
Sbjct: 236 IEYFNGSSDPKGHLKLF-----IISVARAKFRPEERDAGLCHLFVEHLKGPALNWFTRLK 290
Query: 115 CFSIMGYQDMIKKFTQQFSGTRHRKVLSTSLFNVRQGPSESLREYLARFNDSTIKVSNPN 174
S+ +Q++ F +Q+S L+++ Q P+E LR++LA+F + KV N
Sbjct: 291 GNSVDSFQELSTLFLKQYSVLIDPSTSDADLWSLSQQPNEPLRDFLAKFRSTLAKVEGIN 350
Query: 175 QEVFVGAFQNGLW--AGQFNESIAQKPADSMEEIIARAECYVKGEESNAEKKAREAKERG 232
+ + L A + +A++ + AE K E+ NA +A
Sbjct: 351 DLAALSTLKKALCLRARLSAKFLAREIGGGLTIKDLEAE-KGKTEQVNAVANPEQAAPAA 409
Query: 233 NSGGERR 239
NS G +R
Sbjct: 410 NSEGPKR 416
>At1g34080 putative protein
Length = 811
Score = 55.8 bits (133), Expect = 4e-08
Identities = 63/293 (21%), Positives = 108/293 (36%), Gaps = 90/293 (30%)
Query: 45 PFSAKIWNAPVPDNFKPPHLPTFDGRGDPSEHVTVFNTRMSVYGVADSLKCKLLAGTFAD 104
PF+ +I + + F+ LP ++G+GDP E++T F +++AG
Sbjct: 193 PFTLRISKLRIRE-FQDFKLPGYNGKGDPKEYLTSF---------------QVIAGREG- 235
Query: 105 AALRWYMSLPCFSIMGYQDMIKKFTQQFSGTRHRKVLSTSLFNVRQGPSESLREYLARFN 164
SI ++ + F +Q+ V L+N+ Q E LR Y F
Sbjct: 236 ------------SIDNFKQLSTAFIKQYEYFIKSDVTEAQLWNLSQAADEPLRTYNTAFK 283
Query: 165 DSTIKVSNPNQEVFVGAFQNGLW-AGQFNESIAQKPADSMEEIIARAECYVKGEESNAEK 223
+ ++V + A +NGLW +F ES+ +S+++ + GE
Sbjct: 284 EIMVQVPGLSDSAAQSALKNGLWHESRFRESLTVNRPESIQDAL-------HGELPKTSP 336
Query: 224 KAREAKERGNSGGERRNHYVPPNRDRGTFKKPYERNQNRYVPEHFTPLNTRPERILKEVF 283
KA +K PP +G K + N++V
Sbjct: 337 KASPSK--------------PPFSPKG---KNGAFSSNKWV------------------- 360
Query: 284 ESKIIPPPPFSRARVMGQDKDAWCKYHLVQGHNTDDCVHLKREIEKLLLSGKL 336
+D DA+C+ H + GH T DC L R + +SG++
Sbjct: 361 -----------------RDPDAYCEIHKMNGHATKDCKALGRLLAAKYVSGEI 396
>At2g15100 putative retroelement pol polyprotein
Length = 1329
Score = 49.3 bits (116), Expect = 4e-06
Identities = 33/122 (27%), Positives = 55/122 (45%), Gaps = 5/122 (4%)
Query: 45 PFSAKIWNAPVPDNFKPPHLPTFDGRGDPSEHVTVFNTRMSVYGVA----DSLKCKLLAG 100
PF+ +I + + F+ LP ++G+GD EH+T F + D+ CKL +
Sbjct: 157 PFTPRISKLRIRE-FRDFKLPVYNGKGDLKEHLTSFQVIAGRVPLEPHEEDAGLCKLFSE 215
Query: 101 TFADAALRWYMSLPCFSIMGYQDMIKKFTQQFSGTRHRKVLSTSLFNVRQGPSESLREYL 160
AL W+ L SI ++ + F +Q+ + + L+N Q E LR Y+
Sbjct: 216 NLFGLALTWFTQLEEGSIDNFKQLSTAFIKQYEYFINSDITEAHLWNFSQSADEPLRTYI 275
Query: 161 AR 162
R
Sbjct: 276 YR 277
>At2g12870 putative retroelement pol polyprotein
Length = 411
Score = 48.5 bits (114), Expect = 7e-06
Identities = 43/189 (22%), Positives = 76/189 (39%), Gaps = 14/189 (7%)
Query: 39 DEPEPQPFSAKIWNAPVPDNFKPPHLPTFDGRGDPSEHVTVFNTRMSVYGVADSLK---- 94
+E PF+ KI +A + D K + F+G P H+ F +Y D K
Sbjct: 55 EESHNTPFTRKISSAIISDPGKL-RIDYFNGSSYPKGHLKSF----IIYAARDKFKPEER 109
Query: 95 ----CKLLAGTFADAALRWYMSLPCFSIMGYQDMIKKFTQQFSGTRHRKVLSTSLFNVRQ 150
C L L ++ L + ++++ F +QFS L + L+++ Q
Sbjct: 110 DAGLCHLFVEHLKGPVLVLFLRLEGNFVDSFEELSTLFLKQFSVLIDPGTLDSDLWSLSQ 169
Query: 151 GPSESLREYLARFNDSTIKVSNPNQEVFVGAFQNGLW-AGQFNESIAQKPADSMEEIIAR 209
P+E L + L +F + V + A + LW +F + + ++ + + R
Sbjct: 170 QPNEPLMDLLTKFRSTLASVEGIIDVAALSALKEALWYKSEFRKELNLSKPQTICDALHR 229
Query: 210 AECYVKGEE 218
A YV EE
Sbjct: 230 ASDYVSHEE 238
>At1g41780 hypothetical protein
Length = 246
Score = 44.3 bits (103), Expect = 1e-04
Identities = 32/123 (26%), Positives = 54/123 (43%), Gaps = 5/123 (4%)
Query: 45 PFSAKIWNAPVPDNFKPPHLPTFDGRGDPSEHVTVFNTRMSVYGV----ADSLKCKLLAG 100
PF+ KI NA + D K + F G DP H+ F ++ +D+ C L
Sbjct: 125 PFTRKISNAIISDPEKL-RIEYFSGSSDPKGHLKSFKISVARAKFKPEESDAGLCHLFVE 183
Query: 101 TFADAALRWYMSLPCFSIMGYQDMIKKFTQQFSGTRHRKVLSTSLFNVRQGPSESLREYL 160
L W+ L S+ +Q++ F +Q+S L+++ Q P+E L ++L
Sbjct: 184 HLKGPVLDWFSRLEGNSVDSFQELSTLFLKQYSVLIDPGTSDVDLWSLSQQPNEPLSDFL 243
Query: 161 ARF 163
+F
Sbjct: 244 TKF 246
>At3g31340 Athila ORF 1, putative
Length = 781
Score = 43.9 bits (102), Expect = 2e-04
Identities = 28/85 (32%), Positives = 42/85 (48%), Gaps = 3/85 (3%)
Query: 82 TRMSVYGVADSL-KCKLLAGTFADAALRWYMSLPCFSIMGYQDMIKKFTQQFSGTRHRKV 140
TRM+ GV D + KC+L + AD A RW SL ++ ++D F Q+ +
Sbjct: 124 TRMN--GVPDDIIKCRLFIFSLADNAHRWLKSLDPINLRSWEDYKAAFLGQYFTQSRTAI 181
Query: 141 LSTSLFNVRQGPSESLREYLARFND 165
L + + +QG +ES E RF D
Sbjct: 182 LRNKISSFQQGGTESFPEAWERFKD 206
>At2g11940 putative retroelement gag/pol polyprotein
Length = 1212
Score = 41.6 bits (96), Expect = 9e-04
Identities = 31/129 (24%), Positives = 55/129 (42%), Gaps = 27/129 (20%)
Query: 58 NFKPPHLPTFDGRGDPSEHVTVFNTRMSVYGVADSLKCKLLAGTFADAALRWYMSLPCFS 117
+F+ LP+++ +GDP EH+T F +++AG + ++ ++ L
Sbjct: 116 DFRDFKLPSYNDKGDPKEHLTSF---------------QVIAGRVEEGSIDYFKQLSTTF 160
Query: 118 IMGYQDMIKKFTQQFSGTRHRKVLSTSLFNVRQGPSESLREYLARFNDSTIKVSNPNQEV 177
I Y IK V L+N+ Q E L+ Y+ F + ++V N +
Sbjct: 161 IKQYGYFIK-----------NDVSEAQLWNLSQS-DEPLQTYITAFKEIMVQVPNLSDSA 208
Query: 178 FVGAFQNGL 186
A +NGL
Sbjct: 209 AQSALKNGL 217
>At2g07480 F9A16.15
Length = 1012
Score = 39.3 bits (90), Expect = 0.004
Identities = 30/100 (30%), Positives = 44/100 (44%), Gaps = 4/100 (4%)
Query: 72 DPSEHVTVFNTRMS---VYGVAD-SLKCKLLAGTFADAALRWYMSLPCFSIMGYQDMIKK 127
DP +H+ F+ S + GV++ S K +L + D A W +LP SI + D K
Sbjct: 56 DPLDHLDNFDRLCSLTKINGVSEESFKLRLFPFSLGDKAHLWEKTLPVKSIDTWDDCKKA 115
Query: 128 FTQQFSGTRHRKVLSTSLFNVRQGPSESLREYLARFNDST 167
F +F + L + + Q SES E RF T
Sbjct: 116 FLAKFFSNSRKARLRSEISGFNQKNSESFSEAWERFKGYT 155
>At3g32000 unknown protein
Length = 839
Score = 38.5 bits (88), Expect = 0.007
Identities = 30/100 (30%), Positives = 43/100 (43%), Gaps = 4/100 (4%)
Query: 72 DPSEHVTVFNTRMS---VYGVADS-LKCKLLAGTFADAALRWYMSLPCFSIMGYQDMIKK 127
DP +H+ F+ + + GV+++ K +L + D AL W M+LP SI D K
Sbjct: 61 DPLDHLDEFDRLCNLTKINGVSENGFKLRLFPFSLGDKALIWEMNLPHDSITTRDDCKKA 120
Query: 128 FTQQFSGTRHRKVLSTSLFNVRQGPSESLREYLARFNDST 167
F +F L + Q ES E RF D T
Sbjct: 121 FLSKFFSNDRTARLRNEISGFSQKIGESFCEAWERFKDYT 160
>At1g38440 hypothetical protein
Length = 263
Score = 36.2 bits (82), Expect = 0.037
Identities = 29/100 (29%), Positives = 41/100 (41%), Gaps = 4/100 (4%)
Query: 72 DPSEHVTVFNTRMS---VYGVA-DSLKCKLLAGTFADAALRWYMSLPCFSIMGYQDMIKK 127
DP +H+ F+ + + GV+ D K +L + D A W +LP SI + D K
Sbjct: 14 DPLDHLDEFDRLYNLTKINGVSEDGFKLRLFPFSLGDKAHIWEKNLPHDSITTWDDCKKA 73
Query: 128 FTQQFSGTRHRKVLSTSLFNVRQGPSESLREYLARFNDST 167
F +F L + Q SES E RF T
Sbjct: 74 FLSKFFSNARTARLRNEISGFSQKTSESFCEAWERFKGYT 113
>At1g41690 hypothetical protein
Length = 371
Score = 35.4 bits (80), Expect = 0.062
Identities = 29/96 (30%), Positives = 39/96 (40%), Gaps = 4/96 (4%)
Query: 72 DPSEHVTVFNTRMS---VYGVA-DSLKCKLLAGTFADAALRWYMSLPCFSIMGYQDMIKK 127
DP +H+ F S + GV+ D K +L + D A W +LP SI + D K
Sbjct: 61 DPLDHLDEFERLCSLTKINGVSEDGFKLRLFPFSLGDKAHLWEKTLPQGSITTWDDCKKA 120
Query: 128 FTQQFSGTRHRKVLSTSLFNVRQGPSESLREYLARF 163
F +F L + Q SES E RF
Sbjct: 121 FLAKFFSYSRTARLRNEISGFTQKQSESFCEAWERF 156
>At4g04050 putative transposon protein
Length = 681
Score = 35.0 bits (79), Expect = 0.082
Identities = 21/69 (30%), Positives = 35/69 (50%), Gaps = 8/69 (11%)
Query: 302 DKDAWCKYHLVQGHNTDDCVHLKREIEKLLLSG----KLRGYAKERRHNERQEDKPNPEQ 357
D +CKYH +G++T++C R ++KL+ +G K E + QE++ P+Q
Sbjct: 381 DLGKYCKYHKKRGYSTEEC----RAVKKLIAAGGKTKKGSNPKVETPPPDEQEEEQTPKQ 436
Query: 358 KHTLHTISG 366
K T G
Sbjct: 437 KKRERTPEG 445
>At1g36130 hypothetical protein
Length = 914
Score = 35.0 bits (79), Expect = 0.082
Identities = 41/170 (24%), Positives = 64/170 (37%), Gaps = 13/170 (7%)
Query: 73 PSEHVTVFNT---RMSVYGVA-DSLKCKLLAGTFADAALRWYMSLPCFSIMGYQDMIKKF 128
P +H+ F + GV+ D L CKL + A A W L S+ ++ + F
Sbjct: 124 PMDHIERFEDLVLSIKANGVSEDYLLCKLFPYSLAGEADSWLKQLKAGSLKTWRSIKIAF 183
Query: 129 TQQFSGTRHRKVLSTSLFNVRQGPSESLREYLARFNDSTIKVSNPN-QEVFVGAFQNGLW 187
F + L L QGP+E+ + RF + + E+ + A N
Sbjct: 184 LNNFYDDAKSEELRNKLSTFTQGPAEAFKAAWVRFKEYQRDCPHHGFSEMALDAASN--- 240
Query: 188 AGQFNESIAQKPADSMEEIIARAECYVKGEESNAEKKAREAKERGNSGGE 237
G FN PAD+ +I C + ++ E+K GN E
Sbjct: 241 -GNFN---THYPADA-TTLIENLACINSTKNADFERKKIAGAISGNQMAE 285
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.134 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,857,063
Number of Sequences: 26719
Number of extensions: 524906
Number of successful extensions: 1536
Number of sequences better than 10.0: 39
Number of HSP's better than 10.0 without gapping: 30
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 1481
Number of HSP's gapped (non-prelim): 54
length of query: 437
length of database: 11,318,596
effective HSP length: 102
effective length of query: 335
effective length of database: 8,593,258
effective search space: 2878741430
effective search space used: 2878741430
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC135566.1


